Abstract
The 26S proteasome typically degrades proteins marked by ubiquitin chains. However, a distinct, ubiquitin-independent degradation pathway for nuclear proteins exists, mediated by the adaptor protein midnolin, yet its molecular mechanism remains poorly understood. Here, we present nine cryo-electron microscopy structures of the human 26S proteasome in complex with midnolin, which collectively delineate a near-complete catalytic cycle. Our structures reveal that midnolin binds to the proteasome via the RPN1 subunit by its C-terminal helix. Unexpectedly, its ubiquitin-like domain interacts with the RPN11 deubiquitinase in a non-catalytic role. This interaction positions the adjacent Catch domain, which is responsible for substrate binding, directly above the proteasomal entrance, potentially facilitating substrate entry into the proteasome. Furthermore, we observe four consecutive spiral staircase conformations of the AAA+ ATPase hexamer during substrate translocation. These findings provide insights into the mechanisms underlying ubiquitin-independent nuclear protein degradation and may help develop strategies for targeting nuclear proteins via direct proteasomal degradation.
Similar content being viewed by others
Introduction
The 26S proteasome, consisting of a 19S regulatory particle and a 20S catalytic core, has been established as the principal route for regulated protein degradation, traditionally relying on ubiquitin-mediated substrate recognition1,2. This 19S regulatory particle typically engages ubiquitinated proteins, facilitating their unfolding for degradation by the 20S core. Nevertheless, the identification of the midnolin (MIDN)-proteasome pathway expands this established view by revealing an alternative, ubiquitin-independent mechanism for the degradation of numerous nuclear proteins3,4. This alternative degradation system proves particularly useful for maintaining cellular homeostasis through rapid clearance of nuclear immediate early gene (IEG) products, including key transcriptional regulators such as EGR1, FosB, and IRF43,5. Functioning as molecular switches, these IEG proteins orchestrate cellular responses to stimuli by regulating growth, differentiation, stress adaptation, and inflammatory processes6,7,8,9,10.
Midnolin, initially characterized as a developmentally regulated gene in mouse midbrain11, employs three folded domains to execute its degradation functions: an N-terminal ubiquitin-like (Ubl) domain, a middle substrate-capturing Catch domain, and a C-terminal α-helix (hereafter referred to as αHelix-C) proteasome-binding module3. The Catch domain specifically recognizes a β strand degron of IEG proteins such as EGR1, FosB, and IRF4. Notably, αHelix-C of midnolin has been proven to be both necessary and sufficient for proteasome binding, but how the midnolin αHelix-C functions is unclear. Paradoxically, the Ubl domain, which is essential for degradation activity, is dispensable for substrate and proteasome binding. This is unexpected because shuttle factors, such as Rad23 and ubiquilins, use Ubl domains for binding to the proteasome to deliver certain ubiquitinated substrates1. Despite recent structural insights into the midnolin-proteasome12,13, key mechanistic questions remain unanswered. In particular, the binding kinetics of midnolin to the proteasome and the functional role of the Catch domain beyond substrate capture are still poorly understood. It remains unclear whether the Catch domain solely functions in substrate binding or also plays an active role in facilitating efficient degradation, such as positioning the substrate optimally toward the proteolytic entry site. Meanwhile, a key unresolved question is whether midnolin binds its substrates prior to proteasome association, or instead constitutively engages the proteasome before recruiting substrates. These knowledge gaps fundamentally limit our understanding of how midnolin achieves its substrate delivery and degradation efficiency.
Here, we show the structural and molecular basis of midnolin-mediated substrate degradation through single-particle cryo-electron microscopy (cryo-EM) and biochemical analysis. Our findings demonstrate that midnolin’s αHelix-C anchors to the 19S regulatory particle by binding RPN1 (PSMD2), while the coordinated Ubl-Catch domains position substrates at the substrate entrance oligonucleotide- or oligosaccharide-binding (OB) ring of the AAA+ (ATPase associated with diverse cellular activities) ATPase motor. ATP hydrolysis-driven and potential Mg²+-release-driven mechanical forces generated by the AAA+ ATPase ring then mediate substrate unfolding and translocation into the 20S catalytic core. Furthermore, our work demonstrates that midnolin achieves optimal activity by co-expressing with substrates. This work advances our perspective on proteasome substrate processing by revealing an evolutionarily optimized degradation pathway that bypasses ubiquitination constraints to meet the urgent temporal demands of nuclear signaling.
Results
Cryo-EM structures of substrate-engaged human midnolin-26S proteasome
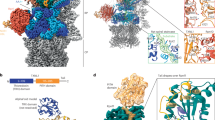
To understand how the midnolin-proteasome degrade substrates without ubiquitylation, we expressed midnolin in HEK293F and treated cells with proteasome inhibitor MG132 to prevent substrate or midnolin degradation and with phorbol 12-myristate 13-acetate (PMA), a protein kinase C agonist, to increase the level of the EGR1 protein as previously reported3. We purified midnolin by two-step tandem affinity chromatography followed by size-exclusion chromatography (Supplementary Fig. 1a). Endogenous 26S proteasome was co-purified with midnolin (Supplementary Fig. 1b, c), and proteasome-like particles were revealed by negative staining (Supplementary Fig. 1d). Using single-particle cryo-EM, we determined structures of midnolin-26S proteasome. As previously reported, 20S core particle is relatively stable and 19S regulatory particle is highly dynamic14,15,16. After 3D classification, we determined four distinct structural states (designated MIDN-PSRPT1, -PSRPT5, -PSRPT2, and -PSRPT6 for subsequent discussion). Among these states, the RPN1 subunit exhibited varying degrees of resolution and occupied distinct positions relative to the AAA+ motor (Supplementary Figs. 2, 3, and Supplementary Table 1). The 3D classification yielded structural states that all represent substrate-processing conformations. This includes classes reconstructed from particles that were not assigned to a dominant state but still contributed to the overall conformational landscape. The highest resolution (2.87 Å) was achieved for the MIDN-PSRPT1 state (Fig. 1a). RPN1 shows extra density when compared with previous structures15. However, its high dynamic precludes its resolution. We performed local refinement on RPN1 and refined it to high resolution of 3.07 Å, which indicated the αHelix-C of midnolin interacts with RPN1 (Fig. 1b, c, and Supplementary Table 1). When the map is low pass filtered to 15 Å, a more distinct yet low-resolution density at the entrance of the ATPase N-ring potentially represents the Catch domain of midnolin or substrate has been translocated into the central channel (Fig. 1d). Therefore, we solved actively processing conformations of the midnolin-bound 26S proteasome, which showed substrate density in the central channel, reaching through the N-ring, the ATPase ring, and into the degradation chamber of the 20S core peptidase (Fig. 1e). Here, we have captured the structures of the natural endogenous substrate-bound 26S proteasome in action.
a Top: Schematic representation of full-length (FL) midnolin, highlighting its domains: ubiquitin-like (Ubl), nuclear localization sequence (NLS), intrinsically disordered region 1 (IDR1), and the C-terminal α-helix (αHelix-C). Bottom: A representative cryo-EM map of the midnolin-26S proteasome in a processing state (MIDN-PSRPT1), showing the midnolin αHelix-C (magenta) interacting with the RPN1(PSMD2) of the 19S regulatory particle. MIDN, short for midnolin throughout the figures. b Local refinement of RPN1 (pink) highlights its interaction with the midnolin αHelix-C (magenta). c Atomic model of midnolin αHelix-C and corresponding Cryo-EM map. d A low-pass filtered map (15 Å) shows midnolin/substrate density (red) at the N-ring entrance of 26S proteasome. e Cryo-EM map of the midnolin/substrate density in the central channel, reaching through the N-ring, the ATPase ring, and into the degradation chamber of the 20S core particle. Density maps in a, b, e were post-processed with EMReady for optimal visualization.
In these reconstructions, we resolved distinct densities corresponding to the inhibitor MG132. This enabled precise docking of the atomic model into the density map, which shows MG132 specifically binds to the catalytic β subunits (β1, β2, β5) of the core particle (CP) (Supplementary Fig. 4). We note that, as an active-site inhibitor, MG132 may allosterically influence the conformational landscape of the 19S regulatory particle17, a consideration relevant to the interpretation of the observed states.
The αHelix-C of midnolin interacts with RPN1 (PSMD2) dynamically
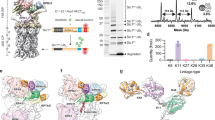
RPN1 is a major docking site on the proteasome for substrates and shuttle factors, particularly those bearing Ubl domains18,19,20. Our cryo-EM analysis revealed that different from previous studies, αHelix-C (not the Ubl domain) of midnolin binds across both the proteasome/cyclosome (PC) repeat toroid domain and the rod-like N-terminal domain of RPN1, a mode distinct from previously characterized Ubl-mediated interactions (Fig. 2a–e). This binding is primarily driven by a hydrophobic core and extensive electrostatic complementarity (Fig. 2a, b), a finding independently corroborated by concurrent structural studies12,13.
a Left: Atomic model of the interaction between RPN1 (pink) and midnolin αHelix-C (magenta). Right: Electrostatic map revealing complementary charge distributions: midnolin αHelix-C exhibits a positive potential (blue), while the opposing RPN1 surface is negative (red). b The interface between midnolin αHelix-C and RPN1 exhibits a tripartite pattern: a central hydrophobic core (circled) flanked by two polar networks. c Superimposition of the atomic models depicting the interaction interface between USP14 (blue) and RPN1 (pink) versus the interaction interface between RPN1 and αHelix-C of midnolin (magenta). The T2 site is indicated by a red dashed circle. d Side-chain interactions between the USP14 UBL domain (blue) and the RPN1 (pink) T2 site, derived from the USP14–RPN1 complex (PDB: 7W3H). The T2 site is indicated by a red dashed circle. e Side-chain interactions between the NUB1 UBL domain (yellow) and the RPN1 (pink) T2 site, derived from the NUB1–RPN1 complex (PDB: 8USB). f, g ITC measurements of binding affinities between MIDN(337-468) or MIDN(337-432) and RPN1. The binding constants (KD values ± standard deviations) and stoichiometries (N) are indicated. KD values and standard deviations were calculated from three independent experiments. h, i MIDN(337-468) and MIDN(337-432) bind to RPN1(PSMD2) with affinities in the same order of magnitude, as measured by surface plasmon resonance; n = 3 independent experiments (one shown). The binding affinity given is the mean of those determined by steady-state analysis of each independent measurement. j–m In vitro peptidase activity of the proteasome. j, l Time course of Suc-LLVY-AMC hydrolysis (relative fluorescence units, RFU) by the 26S proteasome alone (black) or with MBP-MIDN391E/395E/409E (pink) and wild-type MBP-MIDN (red) in (j), and with MBP-MIDN(337-432) (orange), MBP-MIDN(337-468) (blue), and wild-type MBP-MIDN + EGR1(101–200) (red) in (l). k, m Quantification of in vitro peptidase activity from (j, l), shown as RFU change over 90 min. Data are presented as mean ± standard deviation (SD) (n = 3). **** p < 0.0001 (two-tailed one-way ANOVA followed by Tukey’s multiple comparisons test); ns, not significant.
A detailed comparative analysis highlights the distinctiveness of the midnolin αHelix-C–RPN1 interface relative to canonical Ubl domains (Fig. 2b–e). Although it shares a conserved hydrophobic core (anchored by RPN1 Leu426 and Leu465) with T2-site binders like USP14 and NUB114,21, the midnolin αHelix-C establishes a binding mode that spans both the PC repeat toroid and N-terminal domains of RPN1 (Fig. 2c). The interaction interface exhibits a distinctive tripartite pattern. The central portion of the helix engages with RPN1 through an extensive hydrophobic network, where Val392, Leu395, Leu398, and Leu399 of midnolin pack against Val422, Leu426, Cys459, Pro461, Ala464, and Leu465 of RPN1, forming a hydrophobic core (Fig. 2b). This central hydrophobic interface is flanked by two polar interaction networks, especially electrostatic interactions and hydrogen bonds (Fig. 2b). These residues in αHelix-C of midnolin are all highly conserved across species (Supplementary Fig. 5), consistent with our observations that αHelix-C is important for its interaction with RPN1. Furthermore, thermal stability analysis revealed that the binding of midnolin to RPN1 increase its stability (Supplementary Table 2). This sophisticated binding mechanism represents a mode of substrate or shuttle factor recognition by RPN1, expanding our understanding of proteasome-substrate interactions.
To evaluate the binding affinity between them, we purified the C-terminal of midnolin containing αHelix-C and the proteasome subunit RPN1, and measured their binding affinity using isothermal titration calorimetry (ITC) (Fig. 2f, g and Supplementary Table 3). The results demonstrated moderate binding affinities of approximately 130 nM and 146 nM between RPN1 and the two midnolin truncations, respectively. Surface plasmon resonance (SPR) assays further corroborated these values (390 nM and 663 nM, respectively), revealing a dynamic interaction characterized by relatively rapid association and dissociation kinetics (Fig. 2h, i and Supplementary Table 4). Together, these data suggest that midnolin and RPN1 form a moderate binding complex with readily reversible binding, rather than a high-affinity, stable interaction, which may facilitate its self-degradation.
To investigate the role of αHelix‑C in stimulating proteasomal activity, we co‑expressed the MIDN-EGR1(101‑200) complex (the rationale for this construct is detailed later) and its αHelix‑C mutant MIDNK391E/L395E/R409E-EGR1(101‑200) complex. For brevity, EGR1(101–200)-mCherry was referred to as EGR1(101–200). Compared to the wild‑type complex, MIDNK391E/L395E/R409E-EGR1(101‑200) complex significantly reduced the ability to stimulate proteasome activity (Fig. 2j, k). Since αHelix‑C alone is sufficient to bind the proteasome, we further asked whether it is also sufficient to stimulate proteasomal activity. We found that the C‑terminal truncations of midnolin (encompassing αHelix‑C) exhibited no stimulatory effect (Fig. 2l, m). These results indicate that while αHelix‑C is necessary for anchoring midnolin to the proteasome, additional domains are required to drive efficient substrate degradation, a conclusion consistent with previous reports1,13.
Uniform expression of midnolin is dependent on substrate
Structural analysis of the endogenously purified midnolin-proteasome complex revealed substantial complexity, preventing resolution of the Ubl and Catch domains. To overcome this limitation and better understand the mechanism, we sought to establish an in vitro reconstitution system. A key unresolved question is whether midnolin engages its targets after expression or instead constitutively expresses with it. To address this question, we purified the Catch domain of midnolin and its substrate, EGR1(101–200), separately and performed gel filtration assays. When expressed alone, the Catch domain of midnolin exhibits poor stability and solubility, with most of the protein unable to effectively bind its substrate even with a molar excess of substrate (Supplementary Fig. 6a, b). However, under co-expression conditions, the Catch domain displays a more uniform expression profile, suggesting that co-expression facilitates proper folding and enhances the functional performance of midnolin (Supplementary Fig. 6c, d). Compared with the co-expressed MBP-Catch-EGR1(101–200) complex eluted at 14 mL in superdex 200 increase column, the individually expressed MBP-Catch eluted at a volume of 11.5 mL, which suggests MBP-Catch forms oligomer without substrates. Therefore, we co-expressed and purified the full-length midnolin-EGR1(101–200) complex for further biochemical and structural studies (Supplementary Fig. 6e). As expected, the complex shows a single, symmetric peak, whereas midnolin alone shows obvious oligomerization/aggregation, confirming that substrate binding is essential for stabilizing the full-length midnolin (Supplementary Fig. 6e). Additionally, the bioinformatic analysis (Supplementary Fig. 6f) was performed using the public transcriptomic database GEPIA22, which integrates RNA-seq data from projects such as TCGA and GTEx. We directly extracted gene expression values (TPM) for hepatocellular carcinoma and glioblastoma multiforme samples. The results showed that MIDN mRNA levels are significantly and positively correlated with those of its substrate EGR1 in real tumor tissues (p < 1e-5), consistent with the trend observed in our biochemical assays. These findings indicate that midnolin may preferentially co-express with its substrates to achieve optimal activity.
The Ubl and catch domains direct substrate to the pore entrance
To elucidate the molecular mechanism of midnolin-mediated substrate delivery to the proteasome entry port, we purified human proteasomes via an mCherry-hexahistidine-twin-strep-tagged RPN11 (Supplementary Fig. 6g) and incubated them with purified midnolin-EGR1(101–200) complex before plunge-freezing. We determined the cryo-EM structure of the 26S proteasome in complex with midnolin and EGR1, revealing midnolin bound to the proteasome with both the Ubl and Catch domains resolved (Supplementary Fig. 7). In a subset of particles, we observed additional density on the RPN11 subunit. Focused refinement of 19S proteasome enables clear visualization of the Ubl domain bound to RPN11 (Fig. 3a). The substrate, translocated into the ATPase AAA+ ring near RPT5 (Fig. 3b), suggests a state at the initial stage of substrate degradation. Adjacent to the Ubl domain, above the proteasome entry port, weaker but discernible density indicated the presence of the Catch domain. To improve this region, we collected additional data and performed 3D classification, isolating a subset of particles with improved density (Fig. 3c, d and Supplementary Fig. 7). Rigid-body docking of the predicted Catch–EGR1 model into the refined map yielded a strong fit, allowing unambiguous placement of the Catch domain (Fig. 3e and Supplementary Fig. 8). Additionally, continuous density for the interdomain linker between the Ubl and Catch domains confirmed their spatial arrangement, further validating our structural model (Fig. 3c, d). This interpretation is further corroborated by our in silico structural model of the midnolin-proteasome-substrate complex, predicted using AlphaFold3 (Supplementary Fig. 9). The high conformational dynamics of the Catch domain are consistent with its role in delivering substrates to the proteasome entry port. Although the Catch domain was resolved at relatively lower resolution, our integrated analysis combining AlphaFold3 predictions with the cryo-EM density map demonstrate that the substrate’s C-terminus might be positioned directly above the proteasome entry port, an optimal configuration for efficient substrate degradation (Fig. 3e).
a Cryo-EM map of MIDN-PSUbl (Ubl bound state), showing the midnolin Ubl domain (magenta) interacting with RPN11 (PSMD14) of the 19S regulatory particle. b Cryo-EM map of the midnolin/substrate density in the central channel, reaching the ATPase ring. c Cryo-EM map of MIDN-PSUbl_Catch (Ubl and Catch domains bound state). The midnolin Ubl and Catch domains (magenta) interacts with RPN11 (PSMD14) of the 19S regulatory particle and the Catch domain is positioned above the proteasome entry port. d A cut-through of the cryo-EM map of the midnolin/substrate density in the central channel, reaching the ATPase ring. e Atomic model showing the interaction between the 19S proteasome (surface) and the Ubl and Catch domains of midnolin. f The atomic model illustrates the binding interface between Ubl and the deubiquitinating enzyme RPN11. For comparison, the experimentally determined complex of canonical ubiquitin with RPN11 (PDB: 6MSG) is superimposed. g The molecular basis of Ubl (magenta) interaction with the RPN11 (light pink). h Time course of Suc-LLVY-AMC hydrolysis for the 26S proteasome alone (black), or with MBP-MIDNP100E + EGR1(101–200) (dark green), with MBP-MIDNP100E/V102E/104E + EGR1(101–200) (light green), with wild-type MBP-MIDN + EGR1(101–200) (red). The y-axis indicates relative fluorescence units (RFU). i Quantification of in-vitro peptidase activity results in (h), shown as RFU change over 90 min. Data are presented as mean ± SD (n = 3). **** p < 0.0001, *** p < 0.001, obtained by two-tailed one-way ANOVA followed by Tukey’s multiple comparisons test.
Structural analysis demonstrated that the Ubl domain interacts with RPN11 in a manner reminiscent of ubiquitin binding (Fig. 3f). The interaction is stabilized by both hydrophobic and polar contacts (Fig. 3g). Specifically, a hydrophobic core is formed by Ala72, Leu74, Val99, Pro100, Val102 and Ala104 of Ubl together with Met54, Leu56, Met75, Pro89, Trp111 and Phe133 in RPN11. The interaction network may be further reinforced by specific salt bridges, notably between Asp77 of Ubl and Lys94 in RPN11. To validate the functional role of these key residues, we mutated the core hydrophobic residues within the Ubl domain and assessed their impact on proteasome activity stimulation. The results demonstrated that both the MIDNP100E/V102E/A104E and MIDNP100E mutants exhibited a significant reduction in their ability to stimulate proteasome activity compared to the wild-type complex (Fig. 3h, i). This finding directly confirms that these hydrophobic residues are essential for the Ubl domain’s function in stimulating proteasomal activities.
The Catch domain is required for efficient substrate engagement and degradation
The αHelix-C of midnolin anchors to the proteasome by binding RPN1, while its Ubl domain engages RPN11 to position the Catch domain near the proteasome entry port for substrate degradation. To develop targeted degraders based on this mechanism, we wondered whether the core elements (αHelix-C and the Ubl domain) are sufficient to stimulate proteasome degradation activity. To test this, we constructed and purified a midnolin variant MIDN∆(112-336) which is deficient in the Catch domain (Fig. 4a and Supplementary Fig. 10). This midnolin variant maintained its interaction with endogenous proteasome, and we determined its high-resolution structure (Supplementary Figs. 10, 11). The Catch-deficient variant MIDN∆(112-336) demonstrated that approximately one-third of the 26S proteasome particles (29,705 particles in 84,164 particles for 26S) were in substrate-free resting states and two third in processing states (Fig. 4b). Compared with full-length MIDN purified complexes, the MIDNΔ(112–336)-proteasome sample lacking the Catch domain exhibits a significantly higher proportion of particles in the substrate-free resting state (RS). This comparison indicates that the Catch domain, together with substrate binding, plays a role in stabilizing the MIDN-proteasome complex and maintaining it in an active, substrate-processing state. We found that the substrate in the processing state structurally resembles midnolin itself, with its N-terminal disordered region (intrinsically disordered region 1, IDR1) extending into the motor domain and its Ubl domain docked at the proteasome entry port (Fig. 4c–f). This structural observation is consistent with previous biochemical evidence demonstrating that midnolin undergoes proteasomal degradation after substrate delivery3.
a Schematic representation of truncations introduced into the midnolin. bThe population of the processing state (PS) and the resting state (RS) proteasome pulled down by MIDN FL and MIDNΔ(112-336). c A low-pass filtered map (15 Å) shows the Ubl domain of midnolin density (magenta) at the N-ring entrance of 26S proteasome in the MIDNΔ(112-336)-26S proteasome complex. The model for midnolin is shown as cartoon. d Cryo-EM map of the midnolin IDR1 density in the central channel, translocating into the ATPase ring. The map was post-processed with EMReady for optimal visualization. e Atomic model of midnolin IDR1 and corresponding Cryo-EM map. f Cryo-EM density maps for pore-1 loop residues of RPTs interacting with IDR1. g Time course of Suc-LLVY-AMC hydrolysis for the 26S proteasome alone (black), or with Catch-deleted mutant [MBP-MIDN∆(112-336)] (purple), with Catch-replaced midnolin [MBP-MIDN(1-111)-EGR1(101–200)-MIDN(337-468)], with wild-type MBP-MIDN + EGR1(101–200) (red). h Quantification of in-vitro peptidase activity results in (g), shown as RFU change over 90 min. Data are presented as mean ± SD (n = 3). **** p < 0.0001, obtained by two-tailed one-way ANOVA followed by Tukey’s multiple comparisons test; ns, not significant.
To clarify the core function of the Catch domain, we engineered a "Catch-replaced midnolin" construct MBP-MIDN(1-111)-EGR1(101–200)-MIDN(337-468), where the native Catch domain is replaced by the known substrate fragment EGR1(101–200), covalently tethered via a flexible linker. This chimera retains constitutive substrate binding but lacks the native Catch interface. We then compared the stimulated proteasome activity of three proteins: wild-type full-length midnolin (MBP-MIDN + EGR1(101–200)), Catch-deleted mutant (MIDNΔ(112-336), Catch-replaced midnolin. If the role of the Catch domain is to bind substrate, the Catch-replaced midnolin should stimulate proteasome activity as efficiently as wild-type because the substrate is already effectively "provided". Conversely, if the Catch domain possesses an additional "positioning" or "optimized delivery" function, then the Catch-replaced midnolin should show markedly lower stimulated efficiency, resembling or only slightly exceeding that of Catch-deleted mutant. The results show that the Catch-replaced midnolin, despite having the substrate pre-bound, showed a significantly lower degradation efficiency than the full-length, much closer to the Catch-deleted mutant MIDNΔ(112-336) (Fig. 4g, h). These results demonstrate that the Catch domain provides a specific function beyond simple substrate binding, consistent with its proposed role in actively positioning the substrate for optimal delivery to the proteasome.
Together, these results indicate that the Catch domain is crucial for engaging substrates and facilitating their efficient degradation, although midnolin itself can still be engaged by the 26S proteasome even in the absence of the Catch domain in a less efficient manner.
The human midnolin-26S proteasomes are captured in four continuous states
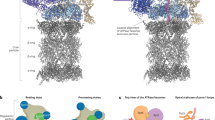
Our current understanding of proteasomal translocation comes from cryo-EM studies of 26S proteasomes with substrates in processing states. These studies revealed that the AAA+ motor (RPT1–6) adopts right-handed spiral-staircase conformations, with typically 4–5 RPT subunits engaging the substrate via their pore-1 loops and 1–2 subunits (seam subunits) positioned off the substrate22. These observations led to the ‘hand-over-hand’ translocation model, wherein the sequential ATP hydrolysis cycle coordinates the movement of the seam subunit to drive stepwise substrate translocation. Our cryo-EM structures capture the midnolin-proteasome complex within this dynamic cycle, providing snapshots of specific substrate-engaged translocation intermediates14,15,16,21. Within this dynamic framework, our study reveals four sequential translocation states of the endogenous human midnolin-26S proteasome complex. We classify these states as MIDN-PSRPT6, -PSRPT2, -PSRPT1, and -PSRPT5, following the established nomenclature based on the highest substrate-engaged RPT subunit21 (Fig. 5a). Conformational analysis shows the progressive advancement of the spiral staircase. For instance, in MIDN-PSRPT6, RPT6 occupies the top substrate-bound position, while seam subunits RPT1 and RPT2 are disengaged (Fig. 5b, c). This pattern shifts sequentially through the subsequent states.
a Cryo-EM density maps of the AAA+ motor in all processing-state (PS) conformations, pseudo-colored by RPT subunit identity. Substrate-disengaged seam subunits exhibit reduced local resolution, likely attributable to heightened conformational flexibility and heterogeneous vertical positioning along the spiral staircase architecture (spanning the base to apex). Notably, PSRPT6 and PSRPT2 display pronounced inter-subunit density discontinuities at the seam, a feature consistent with the presence of two substrate-disengaged seam subunits undergoing continuous dynamic motions. b Conformational landscape of the proteasomal AAA+ motor with four distinct spiral-staircase registers during asymmetric ATP hydrolysis and active substrate processing. Atomic models show the AAA+ ATPase domains in different colors with the engaged substrate in the central channel, bound ATP depicted in yellow, ADP in cyan, magnesium ions in green. The states are organized into a logical sequence according to event-driven transitions and their associated processes. The schematics depict the spiral-staircase arrangements and RPT contacts with the substrate. c Spiral staircase organization of pore-1 loop aromatic residues (tyrosine for RPT1-4, phenylalanine for RPT5 and RPT6) across processing states. Measured distances between the substrate backbone and the Cα atom of the pore-1 loop aromatic residue are annotated for disengaged seam subunits. These seam subunits occupy variable vertical positions, with some localized near the staircase base and others positioned toward the apex. The spatial assignment of pore-loop residues in these subunits remains approximate due to their elevated mobility and resultant lower resolution. d, e Cryo-EM densities and atomic models around ADP binding site in RPT3. The density of the magnesium ion at the ADP binding site in PSRPT1 (d) is visible when compared to PSRPT5 (e). f Structural superimposition shows the movement of the pore-1 loop between PSRPT1 and PSRPT5. Specifically, Lys222, Phe223 in RPT6 and their coordinated substrate residue shift downwards by 2.4 Å and 2.3 Å, respectively.
A notable transition occurs between MIDN-PSRPT1 and MIDN-PSRPT5. In MIDN-PSRPT1, RPT3 retains a bound Mg²⁺ ion, while in MIDN-PSRPT5, this Mg²⁺ is released (Fig. 5d, e). Concomitantly, the entire AAA+ ring undergoes a ~ 2.4 Å downward rigid-body shift (Fig. 5f). The spatiotemporal correlation between Mg²⁺ release and ring movement suggests that Mg²⁺ dissociation from RPT3 may act as a coordinated trigger, working in concert with ATP hydrolysis to facilitate the stepwise substrate translocation observed in our structural continuum (Supplementary Movie 1). Collectively, these four sequential states provide a structural visualization of the hand-over-hand cycle14,15,16 and indicate a potential regulatory role for Mg²⁺ release in coordinating the sequential rearrangements of the ATPase subunits.
Working mode for midnolin-mediated ubiquitin-independent substrate degradation
Based on our results, we propose a working model for midnolin (Fig. 6a). Under specific stimulation conditions, midnolin may rapidly express in conjunction with substrates. After the substrate exerts its function, midnolin utilizes its αHelix-C to target the substrate to the 26S proteasome, where it binds to the RPN1 subunit. Subsequently, its Ubl domain interacts with RPN11, positioning the Catch domain at the entrance of the 26S proteasome N-ring. This proximity allows for preferential degradation of the substrate, followed by the degradation of midnolin itself possibly. This mechanism not only facilitates efficient substrate degradation but also optimizes the utilization of the limited proteasome, thereby maximizing resource efficiency. This is important for the proteasome to degrade various substrates.
a MIDN may bind to substrates in the cytoplasm, and direct the complexes into the nucleus. The αHelix-C of MIDN binds to RPN1, promoting the interaction between MIDN and the 19S regulatory particle of the proteasome within the nucleus. Subsequently, the Ubl of MIDN binds to the proteasomal component RPN11, positioning the Catch domain above the ATPase ring to initiate substrate degradation. The substrate is translocated into the 20S core particle for degradation. After substrate translocation is complete, the disordered regions of MIDN guide itself into the 20S core particle for degradation. The proteasome then returns to a substrate-free state and continues to perform protein degradation functions within the nucleus. b Conformational dynamics of the AAA+ motor domain during substrate processing. Four sequential states of the human midnolin–26S proteasome reveal the mechanism by which ATP hydrolysis and Mg²⁺ release collaboratively drive substrate translocation. In the model, the AAA+ ATPase subunits are colored differently, with the bound substrate positioned within the central channel, illustrating a helical staircase arrangement and contacts between the RPT subunits and the substrate. Bound ATP is shown in orange, and ADP in cyan. The states are arranged in a cycle according to a logical sequence of events and transitions. Key events during proteasomal substrate degradation, including magnesium ion release, ATP hydrolysis (H), and ADP-to-ATP exchange (E), are indicated.
Discussion
Protein homeostasis, essential for cellular physiology, relies on the proteasome to regulate protein turnover23,24,25. While ubiquitin-dependent degradation has been well characterized, emerging evidence highlights ubiquitin-independent pathways that expand the functional repertoire of proteasomal regulation26,27. In this study, we uncover a non-canonical degradation mechanism mediated by midnolin, which directly targets IEG proteins to the proteasome. Through structural and biochemical analyses, we elucidate how midnolin facilitates the degradation of IEG proteins without ubiquitination and propose a working model for this process.
In this model, substrate binding to the Catch domain appears to stabilize midnolin, particularly the Catch domain, into a soluble and functional conformation. This substrate-occupied complex constitutes a stable "ready-to-degrade" module, primed for proteasomal engagement. This proposed mechanism, in which specificity is determined by the initial engagement of the substrate with midnolin, functionally parallels ubiquitination. This mechanism acts as a "midnolin tag" that marks the substrate for subsequent proteasomal delivery and degradation. The Catch domain of midnolin specifically recognizes a β-strand degron in IEG proteins by forming an FG zipper3,13. Our structural data show that the Catch domain is positioned directly above the proteasome entry port, suggesting that it may ensure efficient delivery and degradation of only its captured substrates through a spatial selection mechanism, thereby preventing off-target degradation of non-specific proteins. Furthermore, the possible co-expression mechanism of midnolin with its substrates may contribute to an additional regulation, ensuring that this pathway is activated only when needed. On the other hand, in the absence of substrates, the Catch domain of midnolin exhibits a deficiency in a central β-sheet structure, which predisposes it to instability and subsequent aggregation. Although midnolin can express independently, its capacity to degrade substrates is probably rather limited on its own. It has been reported that midnolin is required for the DNA-binding activity of EGR128, which is consistent with our observation that midnolin co-expresses with substrates. Whether the DNA-binding activity of other substrates is also dependent on midnolin deserves further investigation.
The moderate binding affinity of midnolin’s αHelix-C to RPN1 is not a defect but rather part of midnolin’s self-degradation mechanism. After delivering the substrate, midnolin needs to dissociate from the proteasome and enter the degradation pathway. The moderate binding of αHelix-C to RPN1 likely facilitates this process, allowing midnolin to rapidly detach from RPN1 and proceed to subsequent degradation steps. Additionally, IDR1 may guide midnolin itself into the proteasome for degradation after substrate delivery. This self-sacrificial mechanism not only optimizes the utilization efficiency of the proteasome to degrade other substrates but also prevents excessive accumulation of midnolin within the cell, reflecting the cell’s fine-tuned regulation of resource utilization.
Our findings indicate that ATP hydrolysis plays a role in the displacement of the proteasome’s AAA+ module, with Mg²⁺ dissociation also actively facilitating substrate translocation (Fig. 6b). The conformational asymmetries identified in our study align with those previously observed in human proteasomes bound to USP14 during ATPγS-inhibited Sic1-substrate degradation14, or in yeast proteasomes inhibited by deubiquitinase16, where PSRPT1 and PSRPT5-equivalent states ED2 and 5 T predominantly characterized the particle distributions. Using non-hydrolysable ATP analogue ATPγS or inhibitors targeting RPN11 to halt or trap substrate processing intermediates could potentially skew high-resolution structural determinations towards specific subsets of motor states. However, we captured endogenous substrate-proteasome complexes with hydrolysable ATP. Consistent with the well-established asymmetric mechanism of the proteasomal AAA+ motor14,21, our structures capture consecutive spiral-staircase conformations (PSRPT6, PSRPT2, PSRPT1, PSRPT5) during midnolin-mediated substrate translocation. This confirms that the hand-over-hand translocation mechanism is operational in this ubiquitin-independent pathway. The specific distribution and progression of these states in our dataset provide a detailed view of the mechanochemical cycle in the context of midnolin engagement, although the general principle of asymmetry is pervasively established across AAA+ ATPases. However, it has to be noted that the proteasomes were halted using MG132, which could also skew the distribution of different states. It has been shown by Haselbach et al.17 that inhibitors acting on the 20S core particle can achieve a long-range allosteric effect that modulates the conformational landscape of the 19S particle. Future studies using alternative stabilization methods or time-resolved approaches will be valuable to elucidate the native conformational dynamics of the midnolin-proteasome complex in the absence of core particle inhibition.
The significance of directly targeting the proteasome for degradation is not only reflected in its efficient regulation of the intracellular environment but also provides a strategy for treating drug-resistant diseases29,30,31,32. The development of many diseases is associated with imbalances in protein homeostasis, such as neurodegenerative diseases, cancer, and autoimmune diseases33,34,35,36. By modulating the degradation pathway that directly targets the proteasome, ideas for the treatment of these diseases may be provided. For instance, by designing small molecules or peptides capable of selectively binding to midnolin and proteins of interest or by engineering midnolin to interact with proteins of interest, it becomes possible to directly target proteins to the proteasome. The requirement of the Catch domain for maximal degradation efficiency, in addition to αHelix-C and Ubl domain functions, offers valuable insights for designing next-generation protein degradation platforms based on midnolin architecture. This approach can lead to the rapid degradation of pathogenic proteins, potentially alleviating disease symptoms. This has implications for therapeutic development targeting undruggable nuclear proteins such as MYC and p5337,38,39,40,41.
In summary, this study not only provides a structural basis for understanding the function of midnolin within the cell but also opens up avenues for exploring the role of the proteasome in non-canonical degradation pathways. Future efforts to engineer proteasome-targeting midnolin chimeras may unlock strategies against diseases rooted in proteostasis dysfunction.
Methods
Plasmids, genes, and site-directed mutagenesis
All cDNAs (unless noted otherwise) used in this study were of Homo sapiens origin, and generated from mRNAs extracted from HEK293T cells. Gene sequences used in this study are available from UniProt. The full-length cDNA of MIDN was cloned into the pFastBac Dual CMV vector, with the C-terminal mCherry, 6*His, and twin-strep tags separated by a tobacco etch virus protease cleavage site. Mutants of MIDN were generated by site-directed mutagenesis. The primers used in this study have been listed in Supplementary Table 7.
Expression and purification of human MIDN-proteasome complex
MIDN and its truncated mutants were expressed in HEK293F cells using the BacMam system. Recombinant baculoviruses were generated in Sf9 insect cells through the Bac-to-Bac baculovirus expression system. For protein production, HEK293F cells were cultured at a density of 2.5 × 10⁶ cells/mL in 1% serum medium at 37 °C under 5% CO2 atmosphere with orbital agitation. Following transfection with recombinant baculovirus, 10 mM sodium butyrate (303410-500 G, Sigma-Aldrich) was added after 12 h to enhance protein expression. To promote complex yield, we supplemented 10 µM MG132 (T2154, TargetMol) and 20 ng/mL PMA (HY-18739, MedChem Express) 6 h prior to harvest. Cells were harvested by centrifugation at 3500 x g for 15 min. Pellets were resuspended in ice-cold lysis buffer (50 mM HEPES (pH 7.4), 10% (v/v) glycerol, 5 mM MgCl2, 5 mM ATP, 1 mM Tris(2-carboxyethyl)phosphine (TCEP), 0.5% (v/v) NP-40) supplemented with 1 mM PMSF. Cell disruption was performed via sonication (150 W output) under ice-cold conditions using 4 s pulses alternating with 6 s intervals for 6 min. Lysates were clarified by centrifugation at 46,500 × g for 40 min under 4 °C. For affinity purification, clarified lysates were loaded onto a pre-equilibrated immobilized metal affinity chromatography column (Ni²⁺-IDA resin, SA052100; Smart-Lifescience) at 4 °C using gravity flow. The column was washed with 10 column volumes of wash buffer (lysis buffer + 20/40 mM imidazole), and His-tagged proteins were eluted with elution buffer (lysis buffer + 300 mM imidazole). The eluate was subsequently loaded onto a Strep-Tactin XT column (2-5010-010; IBA Lifesciences) pre-equilibrated with lysis buffer. After washing with 30 mL of lysis buffer under gravity flow, bound proteins were competitively eluted using lysis buffer containing 50 mM biotin (A84479, innochem) under gravity flow. The eluate was concentrated to 500 μL using a 100 kDa cutoff concentrator. Finally, a Superose 6 Increase 10/300 GL column was equilibrated using the size-exclusion buffer (30 mM HEPES, 10% (v/v) glycerol, 1 mM MgCl2, 0.6 mM ATP, 0.5 mM TCEP, 60 mM NaCl, pH 7.4) and a 0.5 mL injection loop was washed thoroughly with the size-exclusion buffer. The protein sample was centrifuged at 11,200 x g for 5 min at 4 °C to remove any debris. We manually injected 0.5 mL of the concentrated eluted protein into the loop and then fractionated it on a Superose 6 Increase column using size-exclusion buffer. Fractions containing intact 26S proteasomes were collected, pooled, and concentrated using Amicon Ultra-15, 10 kDa MWCO (6200 × g, 10 min intervals) to a final volume of around 50 μL at 0.3 mg/mL. The samples were flash-frozen in liquid nitrogen, and stored at −80 °C. Protein concentration was determined using a Bradford assay (BCA) kit (20201ES76, Yeasen). Graphene-supported grids were used for data collection for the sample.
Expression, purification, and in vitro incubation of human 26S proteasome
To purify the human 26S proteasome, we employed the BacMam system for large-scale expression of the RPN11 protein in mammalian HEK293F cells For purification, HEK293F cells infected with the virus were cultured for 48 h, then harvested via centrifugation at 3500 x g. The cell pellet was resuspended in lysis buffer (40 mM HEPES, 100 mM NaCl, 1% (v/v) NP-40, 1 mM MgCl2, 5 mM ATP, 1 mM TCEP, 1 × Cocktail, 1 mM PMSF, pH 7.4) and subjected to ice-cold sonication (150 W, 4 s pulse/6 s interval for 6 min). The lysate was transferred to a pre-cooled centrifuge tube and centrifuged at 46,500 × g for 20 min at 4 °C to collect the supernatant. Intact proteasomes were obtained by sequential Strep and His affinity purification using the C-terminal twin-Strep and 6×His tags on RPN11. The lysates were applied to a pre-equilibrated Strep column, washed, and then the bound protein was eluted and collected. Then, the eluate was loaded onto a pre-equilibrated IDA column at 4 °C, where the His-tagged target protein bound to the column. Contaminants were removed using a buffer (40 mM HEPES, 100 mM NaCl, 1% (v/v) NP-40, 1 mM MgCl2, 5 mM ATP, 1 mM TCEP, 1 × Cocktail, pH 7.4) containing 20 mM and 40 mM imidazole, followed by protein elution with 300 mM imidazole. The eluent was collected and concentrated to 500 μL using a 100 kDa cutoff concentrator. The protein sample was centrifuged at 11,200 × g for 5 min at 4 °C, then fractionated using a Superose 6 column with size-exclusion buffer (50 mM HEPES, 50 mM NaCl, 1 mM MgCl2, 2.5 mM ATP, 1 mM TCEP, pH 7.4). Fractions containing intact 26S proteasomes were collected. Protein concentration was determined using a Bradford assay kit. The complex was concentrated to approximately 4 mg/mL in size-exclusion buffer, flash-frozen in liquid nitrogen, and stored at −80 °C.
The co-expressed and purified EGR1(101–200)-mCherry complexed with MBP-midnolin were mixed with the purified 26S proteasome at a 50:1 molar ratio, incubated on ice for 30 s, and immediately prepared for cryo-EM analysis. 3 µL of the sample was applied onto a freshly glow-discharged (at 20 mA for 70 s, negative charge) Quantifoil R1.2/1.3 300-mesh Au grid. Grids were blotted for 3 s under 100% humidity at 4 °C using a Vitrobot (Thermo Fisher Scientific), then plunge-frozen in liquid ethane cooled by liquid nitrogen. Frozen grids were stored in liquid nitrogen until data collection.
Negative stain electron microscopy
For negative stain electron microscopy, a regular carbon-coated grid was grasped and subjected to glow discharge using plasma (PELCO easiGlow). The protein sample (3 µL, 0.05 mg/mL) was then pipetted onto the grid and allowed to sit for 2 min. Following this, 3 µL of uranyl acetate stain was applied to the grid and left to stand for another 2 min. Data collection was carried out using a transmission electron microscope (JEM1400PLUS).
Cryo-EM sample preparation and data collection
The purified samples were centrifuged at 11,200 × g for 5 min at 4 °C before cryo-EM sample preparation. Aliquots of 3.0 µL samples were applied to glow-discharged grids (Quantifoil, 300 mesh, R1.2/1.3, Au). The plasma treatment conditions were set at 20 mA for 70 s, followed by a 10-s holding period. Graphene-supported mesh was purchased from Shuimu Biosciences Ltd. Using a Vitrobot (Thermo Fisher Scientific), glow discharged grids were placed at 100% humidity and samples were applied and immediately blotted for 3 s before plunge freezing in liquid ethane. Grids were clipped and transferred to transmission electron microscope. All cryo-EM data acquisition was performed on a 300 kV Titan Krios transmission electron microscope (Thermo Fisher Scientific) equipped with a Gatan K3 direct electron detector and a GIF Quantum energy filter, operating in EPU mode. All movie stacks were fractioned into 32 frames with a total electron dose 40 e-/Å2. Images were taken at a nominal magnification of 105,000 with a pixel size of 0.85 Å in super resolution mode with a defocus ranging from −0.8 and −1.8 µm.
Data processing of 26S Proteasome-MIDN-FL dataset
This integrated cryo-EM datasets collected from four independently frozen grids of the same biological sample. Following patch-based motion correction and contrast transfer function (CTF) estimation, micrographs exhibiting severe ice contamination or suboptimal CTF fitting were systematically excluded. An initial template for particle picking was generated using a representative subset of ~20 micrographs. Template matching from 37,413 qualified micrographs yielded 2,067,075 raw particles in the initial round. After iterative 2D classification, 461,310 high-quality particles were retained for ab initio reconstruction. From this pool, 161,296 particles with well-defined features underwent further 3D classification after global refinement (I = 20 classes). Particles from Classes 0, 5, 6, and 19, which exhibited features of substrate-engaged states, were selected. To improve the resolution of the AAA+ motor, these particles underwent local refinement using a soft mask focused on the AAA+ ATPase domain, yielding two locally refined proteasome complex structures at 3.45 Å (PSRPT1, PDB: 9MBP, EMD-63766, 40,572 particles) and 3.66 Å (PSRPT5, PDB: 9MBQ, 23,186 particles), respectively.
To resolve the interaction between midnolin’s αHelix-C and RPN1 at high resolution, we selected particles from Classes 0 and 1, which represented substrate-engaged complexes. Notably, 63,758 particles produced a high-resolution RPN1 subunit structure (PDB: 9MBO, EMD-63775, 3.07 Å) after local refinement. For Classes 1–4 and 7–18 (combined 97,538 particles), further 3D classification (20 classes) was performed. Subsequent selection of 9,022 particles from Class 11 through mask focused on 19S generated a 4.48 Å map (PSRPT6, PDB: 9U4M, EMD-63850). Additionally, 25,990 particles selected from Classes 0, 1, and 10 were processed identically to obtain an improved 19S structure (PSRPT2, PDB: 9U3L, EMD-63817) at 3.72 Å resolution. The other classes (Classes:2–9 and 12–19) represented unassigned processing states. The final maps were further processed using EMReady42. Model refinement and statistical analyses were performed based on the raw (EMReady-unprocessed) density maps.
Data processing of 26S proteasome-MIDN∆ (112-336) dataset
The workflow was similar to the processing pipeline established for the 26S Proteasome-MIDN-FL dataset. From 19,307 acquired movies, 18,890 micrographs meeting quality thresholds were selected for particle picking using Topaz Pick (v0.2.5). This procedure generated 1,646,618 initial particle extracts, which underwent iterative 2D classification to yield 261,241structurally homogeneous particles suitable for ab initio reconstruction. Particles from Classes 0 and 2 were subjected to 3D classification comprised 10 distinct classes, which were systematically categorized into two functional states: Classes 1–6 and 8 (total 7 classes) representing the substrate processing state containing 54,459 particles, while Classes 0 and 9 were identified as the resting state comprising 29,705 particles. Particles in class 8 were subjected to sequential non-uniform (NU) 3D refinement, ultimately yielding a consensus proteasome structure with a global resolution of 3.42 Å (MIDN-PS ∆(112-336), PDB: 22MM, EMD-68472). Particles in class 9 were subjected to NU 3D refinement, ultimately yielding a consensus proteasome structure with a global resolution of 3.17 Å (Resting state). To enhance the local resolution of the 19S regulatory particle, we performed signal subtraction of the 20S core particle density followed by local refinement using a soft mask focused on 19S. This local refinement using 22,379 optimally aligned particles achieved a high-resolution 3.35 Å reconstruction of the 19S regulatory complex. The final maps were further processed using EMReady42. Model refinement and statistical analyses were performed based on the raw (EMReady-unprocessed) density maps.
Data processing of MIDN-EGR1-human 26S proteasome complex
Data processing of MIDN-PSUbl
This integrated three datasets of cryo-electron microscopy images acquired from the same frozen specimen. Following patch-based motion correction and CTF estimation, micrographs exhibiting severe ice contamination or suboptimal CTF fitting were systematically excluded. Particle extraction was performed using Topaz Pick (v0.2.5), which generated three sets of initial particle extracts. These extracts underwent iterative 2D classification, resulting in three distinct particle sets. Based on the resulting 2D classifications, the particles were re-extracted.
Due to structural heterogeneity, the re-extracted particles were subjected to heterogeneous refinement, yielding three different 3D structures. A total of 305,216 particles from Class 0 and Class 1 of these three groups were selected for NU 3D refinement. Prior to symmetry expansion, the selected particles were subjected to Non-uniform refinement with C2 symmetry imposed. The resulting alignment parameters were then used to perform symmetry expansion, doubling the particle count. The expanded particles were subsequently re-extracted, re-centered, and used for downstream focused classification and refinement steps. Since the 19S regulatory particle in the resulting 26S proteasome structure was not centered, symmetry expansion was applied to generate 610,432 particles. Masking was performed to generate a map of the 19S regulatory particles, and the particles were realigned to center the 19S RP within the box. Subsequently, 610,019 particles were re-extracted and subjected to local refinement and masking to generate a map of the 19S regulatory particles. From this pool, particles with well-defined features underwent 3D classification after global refinement using a 3 Å filter resolution (10 classes). Class 7 (68,581 particles) was selected for local refinement, and masking was applied to generate a map of the Ubl domain. This was followed by an additional round of 3D classification, from which Class 0 (24,852 particles) was selected for local refinement, ultimately, a consensus proteasome structure containing a Ubl domain was determined at a global resolution of 3.54 Å (MIDN-PSUbl, PDB:9W39, EMD-65595).
Data processing of MIDN-PSUbl_catch
An additional cryo-EM dataset was collected from the same specimen. Following patch-based motion correction and CTF estimation, micrographs with significant ice contamination or suboptimal CTF fits were systematically discarded. Initial particle picking was performed using Topaz (v0.2.5), yielding a set of raw particle extracts. Subsequent iterative rounds of 2D classification and particle re-extraction resulted in a final set of 68,908 particles. Owing to structural heterogeneity, these particles were subjected to heterogeneous refinement, which produced three distinct 3D classes. Particles from Class 0 and Class 1 across these groups (totaling 52,879 particles) were pooled and processed through NU 3D refinement. The resulting consensus 26S proteasome map exhibited a decentered 19S. To address this, symmetry expansion was applied, generating 105,647 particles. A mask focused on the 19S region was then used to realign the particles, centering the 19S within the box. These processed particles (105,647) were subsequently combined with the 610,019 particles derived from earlier processing stages (as described in the preceding section). The combined pool of 715,666 particles underwent high-resolution 3D classification into 10 classes. Classes 0 and 1 from this classification were selected for local refinement, followed by an additional round of 3D classification. From this subsequent classification, Classes 0 and 2 were chosen for further 3D classification. To enrich for a specific state, particles from all classes of this refinement were merged with a reference set from UBL ("seed1"). After duplicate removal, another round of 3D classification was performed. Particles from Class 0 were selected, and any duplicates overlapping with the seed1 set were excluded, yielding 16,606 particles. These were then merged back with the original seed1 particles, creating a combined set of 41,458 particles for another round of 3D classification (10 classes). Class 1 from this classification was selected for local refinement. The entire set of 41,458 particles then underwent a final round of 3D classification facilitated by a second reference ("seed2"). Particles from Class 0 (7251 particles) of this last classification were subjected to local refinement, ultimately yielding a consensus proteasome structure containing a Catch domain at an overall resolution of 4.97 Å (MIDN-PSUbl_catch, PDB:9WBG, EMD-65839). The final maps were further processed using EMReady42. Model refinement and statistical analyses were performed based on the raw (EMReady-unprocessed) density maps.
Model building and refinement
The data processing procedures were carried out using the cryoSPARC software v4.3.143. The density maps were further enhanced by B-factor sharpening in cryoSPARC and additional sharpening with EMReady42. A previously published proteasome model (see Supplementary Table 1) was initially placed into the cryo-EM density map using UCSF Chimera. The AlphaFold-predicted midnolin structure44 was then rigid-body fitted into the remaining density in Chimera. The combined model was subsequently manually adjusted and rebuilt in COOT (version 0.9.4.1). The model underwent iterative optimization through real-space refinement, employing both the Phenix software suite (version 1.20)45 and Coot46 for structural refinement and building. Model refinement and statistical analyses were performed based on the raw (EMReady-unprocessed) density maps. All figures were crafted using PyMOL 2.4.1 (Schrödinger) and ChimeraX47.
AlphaFold3 prediction
AlphaFold3 predictions of midnolin-RPN1-RPN11-ATPase, or complexed with EGR1 (101–200), IRF4(131-248) and FosB (231-338) were conducted by the online AlphaFold3 server website with random seed. The UniProt sequence ID are provided: RPT1 (P35998), RPT2 (P62191), RPT6 (P62195), RPT3 (P43686), RPT4 (P62333), RPT5 (P17980), RPN11 (O00487), RPN1 (Q13200), MIDN (Q504T8), EGR1 (P18146), IRF4 (Q15306), FosB (P53539). The N-terminal 10 residues of RPN11 (MDRLLRLGGG) were omitted in all the predictions. In addition to the proteins, four ATP molecules, two ADP molecules, and one zinc ion were also added. The resulting structural models have been deposited in ModelArchive (https://www.modelarchive.org/) under accession codes ma-bn2mb, ma-xuxb0, ma-vfr8j, and ma-ue0q7, respectively.
Protein expression and purification in E. coli
The bacterial-expressed proteins were produced in BL21(DE3) E. coli cells (TransGen Biotech) cultured in LB medium (comprising 5 g/L yeast extract, 10 g/L tryptone, and 10 g/L NaCl). Protein expression was induced by adding 0.1 mM IPTG (INALCO) when the OD600 reached 0.8, followed by overnight incubation at 16 °C. The bacteria were resuspended in 40 mL of TBS buffer (20 mM Tris-HCl, 137 mM NaCl, pH 7.4) after centrifugation at 4000 × g for 10 min and then subjected to rapid freezing in liquid nitrogen. After thawing by rotation, PMSF (working concentration: 1 mM) was added to the sample, which was then subjected to ultrasonic disruption on ice (150 W, pulse 4 s/interval 6 s, 10 min). The lysate was transferred to a pre-cooled centrifuge tube and centrifuged at 15,616 × g for 20 min at 4 °C to collect the supernatant. Clarified lysates were loaded onto a pre-equilibrated immobilized metal affinity chromatography column (Ni²⁺-IDA resin) equilibrated with buffer A (20 mM Tris–HCl, 150 mM NaCl, 0.5 mM TCEP). The protein-containing supernatant was loaded onto the column. Following binding of the target molecules to the column, impurities were sequentially removed by washing with buffer A containing 20 mM, 40 mM, and 60 mM, His-tagged proteins were eluted with elution buffer (buffer A + 300 mM imidazole). The eluate was concentrated to 500 μL using a 30 or 50 kDa cutoff concentrator. To remove contaminating nucleic acids, the eluates of MBP-MIDN, MIDN(337–468)-mCherry, MIDN(337–432)-mCherry, and MBP-MIDN-EGR1(101–200)-mCherry were treated with Benzonase nuclease (Cat. No. SSNP01-50KU, Sino Biological) in the presence of 2 mM MgCl₂ for 2 h at 4 °C before SEC. Finally, a Superdex 200 Increase 10/300 GL column was equilibrated using the buffer B (20 mM HEPES, 150 mM NaCl, 0.5 mM TCEP, pH 7.4) and a 0.5 mL injection loop was washed thoroughly with the buffer B. The protein sample was centrifuged at 11,200 × g for 5 min at 4 °C to remove any debris. The loop was then manually injected with 0.5 mL of concentrated eluted protein. Then fractionated using a Superdex 200 Increase 10/300 GL column with buffer B. Fractions were collected, pooled, and concentrated using a 30 or 50 kDa cutoff concentrator (3500 x g, 20 min intervals) based on chromatographic peaks and Coomassie Brilliant Blue staining results of Tris-glycine gels. The samples were flash-frozen with liquid nitrogen in aliquots of 30 μL, and stored at −80 °C. Protein concentration was determined using Nanodrop.
Co-expression and purification of EGR1(101–200) and Catch
Catch domain with His-MBP tag was cloned into pET-28a(+) vector. EGR1 (101–200)-mCherry without any other tag was cloned into ORF2 of pETDuet-1 vector. A mixture of 200 ng Catch and 200 ng EGR1 (101–200) plasmids was co-transformed into 10 μL of E. coli BL21 (DE3). E. coli BL21 (DE3) were incubated on ice for 20 min, heat-shocked at 42 °C for 45 s in a water bath, and immediately returned to ice for 2 min. Subsequently, the transformed cells were resuspended in LB medium supplemented with 50 μg/mL ampicillin and 25 μg/mL kanamycin, followed by incubation at 37 °C with stirring until the OD600 reached 0.8. The culture was cooled to 16 °C, and protein expression was induced by adding 0.1 mM IPTG overnight. Cells were harvested by centrifugation at 4000 × g for 15 min at 4 °C. The pellets were flash-frozen in liquid nitrogen, and stored at −80 °C. The purification steps are the same as those for protein expression in E. coli mentioned above.
Size-exclusion chromatography
To validate the assembly characteristics of the substrate proteins EGR1 and midnolin, we employed two approaches: (1) in vitro reconstitution by incubating Catch with EGR1(101–200), and (2) co-expression, combined with size-exclusion chromatography (SEC) to assess the complex formation. For in vitro reconstitution: Catch and EGR1(101–200) were individually expressed and purified using the E. coli-based method described above. A mixture containing 1 mg of EGR1(101–200) and 600 μg of Catch was incubated on ice for 30 min. The sample was then injected into a pre-equilibrated Superdex 200 Increase 10/300 GL column with Buffer B. Elution fractions (0.5 mL per tube) were collected during SEC separation. Elution fractions were analyzed by 10% SDS-PAGE followed by Coomassie Brilliant Blue staining. For co-expression analysis: the recombinant Catch-EGR1(101–200) complex was co-expressed in E. coli BL21(DE3) using the protocol above. The complex was purified via the same method under Buffer B conditions. Eluted fractions were similarly collected and subjected to 10% SDS-PAGE analysis with Coomassie Brilliant Blue staining.
Isothermal titration calorimetry (ITC)
All measurements were done in ITC buffer containing 20 mM HEPES, pH 8.0, 150 mM NaCl. Each titration consisted of 20 successive injections (the first at 0.4 μL and the remaining 19 at 2 μL). The titrant, 100 μM MIDN(337-468) or MIDN(337-432), and the 10 μM RPN1 analyte were dissolved in ITC buffer. Before measuring the protein–protein interactions, we performed blank titrations by injecting MIDN(337-468) or MIDN(337-432) into buffer to exclude non-specific binding signals. Reference Power was set to 5 μcal/s; Initial Delay is 60 s; The injection interval was 120 s to ensure that the signal returns to baseline; All titrations were conducted at 25 °C with stirring to ensure thorough mixing. The thermal signal curve was subsequently fitted using MicroCal PEAQ-ITC Analysis Software. The resulting blank heat effects were subtracted from the raw thermograms before fitting the binding isotherms. Binding isotherms were fitted to a single-site binding model to derive thermodynamic parameters: dissociation constant (KD), enthalpy change (ΔH), entropy change (ΔS), and binding stoichiometry (N). Additionally, all ITC experiments were performed in triplicate, and the data are presented as mean ± SD.
Surface plasmon resonance (SPR)
All surface plasmon resonance (SPR) experiments were performed at 25 °C on an INTERBIO MI-S200F instrument using CM5 sensor chips and running buffer (10 mM HEPES, 150 mM NaCl, 0.05% Tween-20, pH 7.4) that had been degassed and 0.22 µm-filtered. After a 5-min deionised-water rinse, the carboxylated surface was activated with a freshly mixed 1:1 solution of 0.4 M EDC and 0.1 M NHS for 7 min at 10 µL min⁻¹. Ligand proteins diluted in 10 mM sodium acetate (pH 4.5) were immobilized by continuous flow until the target response was reached; residual NHS-esters were blocked with 1 M ethanolamine-HCl (pH 8.5) for 7 min. A reference flow cell treated identically but receiving no ligand served as the blank for subsequent double-referencing.
Three binary interactions were analysed in succession. (i) RPN1 (ligand, 498 µM stock) was coupled to 759 RU; analyte MIDN(337-468) (170 µM stock) was injected for 120 s at 30 µL min⁻¹ followed by 300 s dissociation, using concentrations of 1.25, 0.625, 0.3125, 0.156, 0.078, and 0.039 µM. (ii) The same surface was regenerated with 20 mM NaOH (30 s) and reused for MIDN(337-432) (464 µM stock) with an identical concentration series. (iii) A second CM5 chip was prepared with 26S proteasome (161 nM stock) immobilized to 1741 RU; MIDN Δ(112-336) (90 µM stock) was titrated at 3, 1.5, 0.75, 0.375, 0.187, and 0.093 µM with regeneration by 10 mM glycine-HCl pH 1.5 (30 s). Each concentration was injected once in randomized order and the complete sequence was repeated in triplicate on independent chips. Sensorgrams were processed with the instrument software, solvent-corrected, and globally fitted to a 1:1 Langmuir model to extract ka, kd and KD values; reported constants are mean ± SD of the three independent experiments.
Proteasome stimulating activity assay
Proteasome stimulating activity of MBP-MIDN + EGR1(101–200), MBP-MIDNP100E + EGR1(101–200), MBP-MIDNP100E/V102E/A104E + EGR1(101–200), MBP-MIDNK391E/L395E/R409E + EGR1(101–200), MBP-MIDN∆(112-336), MBP-MIDN(1-111)-EGR1(101–200)-MIDN(337-468), MBP-MIDN(337-432) and MBP-MIDN(337-468) toward 26S human proteasomes were measured using fluorogenic substrate Suc-LLVY-AMC (South Bay Bio)12. Briefly,1 nM human 26S proteasomes were incubated with 100 nM of substrate in buffer (50 mM HEPES pH 7.5, 100 mM KCl, 0.5 mM MgCl2, 1 mM TCEP, 0.2 mM ATP, and 25 ng/μL BSA) for 5 min at room temperature. 10 μM Suc-LLVY-AMC was added to the reaction mixture. For time course analyses, proteasome stimulating activity was measured immediately after substrate addition and for up to 1.5 h at 37 °C using a Thermo Fisher Scientific multimode plate reader (excitation, 345 nm; emission, 445 nm).
Multiple sequence alignment
Multiple sequence alignment was performed using ESPript (Version 3.2)48.
Thermal stability analysis of proteins using nanoDSF
To assess the thermal stability of various proteins, we employed nanoDSF (nano Differential Scanning Fluorimetry) using the Prometheus NT.48 instrument (NanoTemper Technologies GmbH, Munich, Germany). This method enables the evaluation of protein thermal stability in a label-free manner by monitoring changes in the intrinsic fluorescence of tryptophan residues. We purified and obtained different protein samples. Then, 10 µL of each protein sample was loaded into high-purity glass capillaries, and the capillaries were placed into the Prometheus NT.48 instrument. The temperature gradient was set to increase from 20 °C to 95 °C at a rate of 1.0 °C /min. The intrinsic fluorescence of tryptophan residues was monitored at excitation wavelengths of 280 nm and 350 nm, with emission detected at 330 nm and 350 nm, respectively. The ratio of fluorescence intensities at 350 nm/330 nm was used to track protein unfolding. The melting temperature (Tm), defined as the midpoint of the protein unfolding transition, was determined from the fluorescence ratio curve. Each measurement was performed in triplicate to ensure reproducibility and accuracy.
Statistical analysis
All statistical analyses and data visualizations were performed using GraphPad Prism (version 10.1.2). For the in vitro peptidase activity assays, data are presented as mean ± SD from three independent experiments (n = 3). Statistical significance was determined by two-tailed one-way ANOVA followed by Tukey’s multiple comparisons test. The exact p values and significance thresholds (***p < 0.001, ****p < 0.0001) are detailed in the respective figure legends. ns, not significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The atomic model coordinates and cryo-EM maps have been deposited in the Protein Data Bank (PDB) and the Electron Microscopy Data Bank (EMDB), respectively, under the accession codes: 9MBO and EMD-63775 (RPN1-MIDN), 9MBP and EMD-63776 (MIDN-PSRPT1), 9MBQ and EMD-63777 (MIDN-PSRPT5), 9U3L and EMD-63817 (MIDN-PSRPT2), 9U4M and EMD-63850 (MIDN-PSRPT6), 9U7R and EMD-63943 (RS), 9W39 and EMD-65595 (MIDN-PSUbl), 9WBG and EMD-65839 (MIDN-PSUbl_Catch), 22MM and EMD-68472 (MIDN-PSΔ(112-336)). Additionally, for the previously published structures used in this study, the atomic coordinate of the USP14–RPN1 complex is under the accession code 7W3H, the NUB1–RPN1 complex is under the accession code 8USB, the canonical ubiquitin with RPN11 is under the accession code 6MSG, the initial model used for building of MIDN-PSRPT1 is under the accession code 6MSK, the initial model used for building of MIDN-PSRPT2 is under the accession code 6MSJ, the initial model used for building of PSRPT6 is under the accession code 9E8K, the TXNL1-bound proteasome state PSRPT2 is under the accession code 9E8O, the TXNL1-bound proteasome state PSRPT1 is under the accession code 9E8J, and the TXNL1-bound proteasome state PSRPT5 is under the accession code 9E8G. Source data are provided with this paper.
References
Arkinson, C., Dong, K. C., Gee, C. L. & Martin, A. Mechanisms and regulation of substrate degradation by the 26S proteasome. Nat. Rev. Mol. Cell Biol. 26, 104–122 (2025).
Bard, J. A. M. et al. Structure and function of the 26S proteasome. Annu. Rev. Biochem. 87, 697–724 (2018).
Gu, X. et al. The midnolin-proteasome pathway catches proteins for ubiquitination-independent degradation. Science 381, eadh5021 (2023).
Du, J. et al. Stuxnet facilitates the degradation of polycomb protein during development. Dev. Cell 37, 507–519 (2016).
Jin, W. L. Midnolin-proteasome pathway for protein degradation. MedComm 4, e450 (2023).
Gallo, F. T., Katche, C., Morici, J. F., Medina, J. H. & Weisstaub, N. V. Immediate early genes, memory and psychiatric disorders: focus on c-Fos, Egr1 and Arc. Front. Behav. Neurosci. 12, 79 (2018).
Shaffer, A. L. et al. IRF4 addiction in multiple myeloma. Nature 454, 226–231 (2008).
Klein, U. et al. Transcription factor IRF4 controls plasma cell differentiation and class-switch recombination. Nat. Immunol. 7, 773–782 (2006).
Mittrucker, H. W. et al. Requirement for the transcription factor LSIRF/IRF4 for mature B and T lymphocyte function. Science 275, 540–543 (1997).
Healy, S., Khan, P. & Davie, J. R. Immediate early response genes and cell transformation. Pharm. Ther. 137, 64–77 (2013).
Tsukahara, M., Suemori, H., Noguchi, S., Ji, Z. S. & Tsunoo, H. Novel nucleolar protein, midnolin, is expressed in the mesencephalon during mouse development. Gene 254, 45–55 (2000).
Peddada, N. et al. Structural insights into the ubiquitin-independent midnolin-proteasome pathway. Proc. Natl. Acad. Sci. USA 122, e2505345122 (2025).
Nardone, C. et al. Structural basis for the midnolin-proteasome pathway and its role in suppressing myeloma. Mol. Cell 85, 2597–2609 e2511 (2025).
Zhang, S. et al. USP14-regulated allostery of the human proteasome by time-resolved cryo-EM. Nature 605, 567–574 (2022).
Dong, Y. et al. Cryo-EM structures and dynamics of substrate-engaged human 26S proteasome. Nature 565, 49–55 (2019).
de la Pena, A. H., Goodall, E. A., Gates, S. N., Lander, G. C. & Martin, A. Substrate-engaged 26S proteasome structures reveal mechanisms for ATP-hydrolysis-driven translocation. Science 362, eaav0725 (2018).
Haselbach, D. et al. Long-range allosteric regulation of the human 26S proteasome by 20S proteasome-targeting cancer drugs. Nat. Commun. 8, 15578 (2017).
Chen, X. et al. Structures of Rpn1 T1:Rad23 and hRpn13:hPLIC2 Reveal Distinct Binding Mechanisms between Substrate Receptors and Shuttle Factors of the Proteasome. Structure 24, 1257–1270 (2016).
Shi, Y. et al. Rpn1 provides adjacent receptor sites for substrate binding and deubiquitination by the proteasome. Science 351, https://doi.org/10.1126/science.aad9421 (2016).
Martinez-Fonts, K. et al. The proteasome 19S cap and its ubiquitin receptors provide a versatile recognition platform for substrates. Nat. Commun. 11, 477 (2020).
Arkinson, C., Gee, C. L., Zhang, Z., Dong, K. C. & Martin, A. Structural landscape of the degrading 26S proteasome reveals conformation-specific binding of TXNL1. Nat. Struct. Mol. Biol. 32, 2403–2415 (2025).
Tang, Z. et al. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 45, W98–w102 (2017).
Bhattacharyya, S., Yu, H., Mim, C. & Matouschek, A. Regulated protein turnover: snapshots of the proteasome in action. Nat. Rev. Mol. Cell Biol. 15, 122–133 (2014).
Ford, P. W., Narasimhan, M. & Bennett, E. J. Ubiquitin-dependent translation control mechanisms: degradation and beyond. Cell Rep. 43, 115050 (2024).
Carroll, E. C., Greene, E. R., Martin, A. & Marqusee, S. Site-specific ubiquitination affects protein energetics and proteasomal degradation. Nat. Chem. Biol. 16, 866–875 (2020).
Negi, H., Ravichandran, A., Dasgupta, P., Reddy, S. & Das, R. Plasticity of the proteasome-targeting signal Fat10 enhances substrate degradation. eLife 13, https://doi.org/10.7554/eLife.91122 (2024).
Chen, C. et al. Ubiquitin like protein FAT10 repressed cardiac fibrosis after myocardial ischemic via mediating degradation of Smad3 dependent on FAT10-proteasome system. Int. J. Biol. Sci. 19, 881–896 (2023).
Chiba, A. et al. Midnolin, a genetic risk factor for Parkinson’s disease, promotes neurite outgrowth accompanied by early growth response 1 activation in PC12 cells. Mol. Cell Biol. 44, 516–527 (2024).
Bashore, C. et al. Targeted degradation via direct 26S proteasome recruitment. Nat. Chem. Biol. 19, 55–63 (2023).
Kim, J. et al. Targeted protein degradation directly engaging lysosomes or proteasomes. Chem. Soc. Rev. 53, 3253–3272 (2024).
Rani, N., Aichem, A., Schmidtke, G., Kreft, S. G. & Groettrup, M. FAT10 and NUB1L bind to the VWA domain of Rpn10 and Rpn1 to enable proteasome-mediated proteolysis. Nat. Commun. 3, 749 (2012).
Tsai, J. M., Nowak, R. P., Ebert, B. L. & Fischer, E. S. Targeted protein degradation: from mechanisms to clinic. Nat. Rev. Mol. Cell Biol. 25, 740–757 (2024).
Kumar, D. & Hassan, M. I. Targeted protein degraders march towards the clinic for neurodegenerative diseases. Ageing Res. Rev. 78, 101616 (2022).
Mannion, J. et al. A RIPK1-specific PROTAC degrader achieves potent antitumor activity by enhancing immunogenic cell death. Immunity 57, 1514–1532.e1515 (2024).
Popow, J. et al. Targeting cancer with small-molecule pan-KRAS degraders. Science 385, 1338–1347 (2024).
Ackerman, L. et al. IRAK4 degrader in hidradenitis suppurativa and atopic dermatitis: a phase 1 trial. Nat. Med. 29, 3127–3136 (2023).
Henley, M. J. & Koehler, A. N. Advances in targeting ‘undruggable’ transcription factors with small molecules. Nat. Rev. Drug Discov. 20, 669–688 (2021).
Hassin, O. & Oren, M. Drugging p53 in cancer: one protein, many targets. Nat. Rev. Drug Discov. 22, 127–144 (2023).
Whitfield, J. R. & Soucek, L. MYC in cancer: from undruggable target to clinical trials. Nat. Rev. Drug Discov. https://doi.org/10.1038/s41573-025-01143-2 (2025).
Xie, X. et al. Recent advances in targeting the “undruggable” proteins: from drug discovery to clinical trials. Signal Transduct. Target Ther. 8, 335 (2023).
Speltz, T. E. et al. Targeting MYC with modular synthetic transcriptional repressors derived from bHLH DNA-binding domains. Nat. Biotechnol. 41, 541–551 (2023).
He, J., Li, T. & Huang, S. Y. Improvement of cryo-EM maps by simultaneous local and non-local deep learning. Nat. Commun. 14, 3217 (2023).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Tunyasuvunakool, K. et al. Highly accurate protein structure prediction for the human proteome. Nature 596, 590–596 (2021).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. Sect. D. Struct. Biol. 74, 531–544 (2018).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. Biol. Crystallogr. 60, 2126–2132 (2004).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Gouet, P., Robert, X. & Courcelle, E. ESPript/ENDscript: extracting and rendering sequence and 3D information from atomic structures of proteins. Nucleic Acids Res. 31, 3320–3323 (2003).
Acknowledgements
We thank the cryo-EM platform of Peking University Health Science Center with the assistance of Dr. Dandan Chen and Dr. Lihong Chen for data collection. We thank the cryo-EM facility from Shuimu BioSciences for data collection. We thank Beijing Inter-bioTech CO., LTD for the experimental assistance with SPR. The authors thank Dr. Jing Wang from the State Key Laboratory of Natural and Biomimetic Drugs (Peking University, Beijing, China) for assistance with the ITC assay; Prof. Hui Liang from Peking University for help with functional assays; Dr. Lei Zhang from Peking University Medical and Health Analysis Center for negative staining electron microscopy data collection. This work was supported by grants including the National Natural Science Foundation of China (grant 32571452, 32171224 to L.L., grant 22595464, 22378004 to Z.L.), the Beijing Natural Science Foundation (grant L242055 to Z.L.), the Fundamental Research Funds for the Central Universities (BMU2022RCZX008 to L.L.) and Lam Chung Nin Foundation for Systems Biomedicine.
Author information
Authors and Affiliations
Contributions
Conceptualization: L.L.; Methodology: L.L., Z.D., and P.Z.; Investigation: C.Z., L.Q., and L.Z.; Visualization: C.Z., L.Q., Z.D., P.Z., and A.Y.; Funding acquisition: L.L and Z.L.; Project administration: L.L.; Supervision: L.L. and Z.L.; Writing-original draft: L.L., C.Z., and L.Q.; Writing-review & editing: L.L.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Marta Carroni who co-reviewed with Piotr Draczkowski; Rui Fang, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhu, C., Qin, L., Dai, Z. et al. Structural dynamics of the midnolin-proteasome during ubiquitin-independent substrate turnover. Nat Commun 17, 2800 (2026). https://doi.org/10.1038/s41467-026-71002-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-71002-0