Abstract
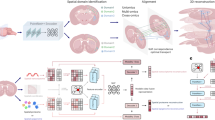
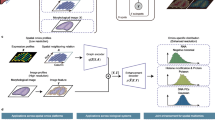
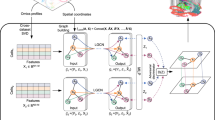
Advances in spatially resolved technologies enable the characterization of tissues at molecular resolution by preserving spatial information. However, integrating and aligning spatial-omics data across different platforms and modalities remains challenging. Flexible tools for slice alignment, stitching and slice-to-volume 3D reconstruction are still lacking because available spatial-omics datasets are affected by partial overlapping, local non-rigid deformations, and large-scalability. Here we propose GEASO (Graph-based Elastic Alignment for Spatial-Omics data), a network-based algorithm for slice alignment, stitching and slice-to-volume 3D reconstruction. GEASO learns consistent spot features with graph neural network, and performs elastic registration to address rigid transformation and local deformation of slices by exploiting topological structure of spot connectivity graphs. GEASO also adopts acceleration strategies to enable its application to large-scale datasets. Experiment results demonstrate that GEASO outperforms state-of-the-art baselines in alignment, stitching and 3D reconstruction of slices across various platforms, modalities and tissues, providing a versatile tool for analyzing spatial-omics data.
Similar content being viewed by others
Data availability
All datasets analyzed in this paper are published datasets and available for public download. The human dorsolateral prefrontal cortex data10 used in this study are available in the spatialLIBD database (http://spatial.libd.org/spatialLIBD). The mouse brain (8 months) data41 and mouse hippocampus data42 used in this study are available in the Single Cell Portal database under accession codes SCP1375 and SCP1830. The serial mouse brain data43 used in this study are available in the Brain Image Library database (https://doi.brainimagelibrary.org/doi/10.35077/act-bag). The human metastatic lymph node data45 and mouse brain (Spatial CUT&Tag-RNA-seq) data57 used in this study are available in the GEO database under accession codes GSE251926 and GSE165217. The human breast cancer data4 used in this study are available in the 10 × Genomics datasets database (https://www.10xgenomics.com/resources/datasets). The whole mouse embryo data51 used in this study are available in the Spateo database (http://spateodata.aristoteleo.com). The spatial multi-omics mouse brain data55 used in this study are available in the Single Cell Portal database under accession code SCP1835. The processed data generated in this study have been deposited in the Zenodo database (https://doi.org/10.5281/zenodo.18811760). Source data are provided with this paper.
Code availability
The code for GEASO algorithm is implemented in Python and detailed tutorials are freely available at https://github.com/xkmaxidian/GEASO67. The source code of GEASO is released under the MIT License.
References
Marx, V. Method of the year: spatially resolved transcriptomics. Nat. Methods 18, 9–14 (2021).
Ståhl, P. L. et al. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 353, 78–82 (2016).
Vickovic, S. et al. High-definition spatial transcriptomics for in situ tissue profiling. Nat. Methods 16, 987–990 (2019).
Marco Salas, S. et al. Optimizing xenium in situ data utility by quality assessment and best-practice analysis workflows. Nat. Methods 22, 813–823 (2025).
Stickels, R. R. et al. Highly sensitive spatial transcriptomics at near-cellular resolution with slide-seqv2. Nat. Biotechnol. 39, 313–319 (2021).
Wei, X. et al. Single-cell stereo-seq reveals induced progenitor cells involved in axolotl brain regeneration. Science 377, eabp9444 (2022).
Andersson, A. et al. Spatial deconvolution of HER2-positive breast cancer delineates tumor-associated cell type interactions. Nat. Commun. 12, 6012 (2021).
Berglund, E. et al. Spatial maps of prostate cancer transcriptomes reveal an unexplored landscape of heterogeneity. Nat. Commun. 9, 2419 (2018).
Moncada, R. et al. Integrating microarray-based spatial transcriptomics and single-cell RNA-seq reveals tissue architecture in pancreatic ductal adenocarcinomas. Nat. Biotechnol. 38, 333–342 (2020).
Maynard, K. R. et al. Transcriptome-scale spatial gene expression in the human dorsolateral prefrontal cortex. Nat. Neurosci. 24, 425–436 (2021).
Farah, E. N. et al. Spatially organized cellular communities form the developing human heart. Nature 627, 854–864 (2024).
Maniatis, S. et al. Spatiotemporal dynamics of molecular pathology in amyotrophic lateral sclerosis. Science 364, 89–93 (2019).
Cable, D. M. et al. Cell type-specific inference of differential expression in spatial transcriptomics. Nat. Methods 19, 1076–1087 (2022).
Wu, L. et al. An invasive zone in human liver cancer identified by stereo-seq promotes hepatocyte–tumor cell crosstalk, local immunosuppression and tumor progression. Cell Res. 33, 585–603 (2023).
Hu, J. et al. Spagcn: Integrating gene expression, spatial location and histology to identify spatial domains and spatially variable genes by graph convolutional network. Nat. Methods 18, 1342–1351 (2021).
Zhao, E. et al. Spatial transcriptomics at subspot resolution with bayesspace. Nat. Biotechnol. 39, 1375–1384 (2021).
Cable, D. M. et al. Robust decomposition of cell type mixtures in spatial transcriptomics. Nat. Biotechnol. 40, 517–526 (2022).
Li, J., Chen, S., Pan, X., Yuan, Y. & Shen, H.-B. Cell clustering for spatial transcriptomics data with graph neural networks. Nat. Comput Sci. 2, 399–408 (2022).
Dong, K. & Zhang, S. Deciphering spatial domains from spatially resolved transcriptomics with an adaptive graph attention auto-encoder. Nat. Commun. 13, 1–12 (2022).
Wang, Y., Liu, Z. & Ma, X. Mucst: restoring and integrating heterogeneous morphology images and spatial transcriptomics data with contrastive learning. Genome Med 17, 21 (2025).
Ueda, H. R. et al. Tissue clearing and its applications in neuroscience. Nat. Rev. Neurosci. 21, 61–79 (2020).
Richardson, D. S. & Lichtman, J. W. Clarifying tissue clearing. Cell 162, 246–257 (2015).
Xie, P. et al. Digital reconstruction of full embryos during early mouse organogenesis. Cell 188, 4754–4772 (2025).
Zeira, R., Land, M., Strzalkowski, A. & Raphael, B. J. Alignment and integration of spatial transcriptomics data. Nat. Methods 19, 567–575 (2022).
Liu, X., Zeira, R. & Raphael, B. J. Partial alignment of multislice spatially resolved transcriptomics data. Genome Res 33, 1124–1132 (2023).
Klein, D. et al. Mapping cells through time and space with Moscot. Nature 638, 1065–1075 (2025).
Xia, C.-R., Cao, Z.-J., Tu, X.-M. & Gao, G. Spatial-linked alignment tool (slat) for aligning heterogenous slices. Nat. Commun. 14, 7236 (2023).
Gao, J., Huang, X. & Li, J. Unsupervised graph alignment with Wasserstein distance discriminator. In Proceedings of the 27th ACM SIGKDD Conference on Knowledge Discovery & Data Mining, 426–435 (2021).
Li, H. et al. Santo: a coarse-to-fine alignment and stitching method for spatial omics. Nat. Commun. 15, 6048 (2024).
Xu, H. et al. Spacel: deep learning-based characterization of spatial transcriptome architectures. Nat. Commun. 14, 7603 (2023).
Clifton, K. et al. Stalign: Alignment of spatial transcriptomics data using diffeomorphic metric mapping. Nat. Commun. 14, 8123 (2023).
Tang, Z. et al. Search and match across spatial omics samples at single-cell resolution. Nat. Methods 21, 1818–1829 (2024).
Qiu, X. et al. Spatiotemporal modeling of molecular holograms. Cell 187, 7351–7373 (2024).
Tajdari, F., Huysmans, T., Yang, Y. & Song, Y. Feature preserving non-rigid iterative weighted closest point and semi-curvature registration. IEEE Trans. Image Process. 31, 1841–1856 (2022).
Hirose, O. Acceleration of non-rigid point set registration with downsampling and Gaussian process regression. IEEE Trans. Pattern Anal. Mach. Intell. 43, 2858–2865 (2020).
Wang, L. et al. Equivariant local reference frames with optimization for robust non-rigid point cloud correspondence. IEEE Transactions on Image Processing (2025).
Liu, M. et al. Lcnet: a robust and accurate non-rigid 3d point set registration approach for image-guided liver surgery. IEEE Transactions on Medical Robotics and Bionics (2025).
Yuille, A. L. & Grzywacz, N. M. A mathematical analysis of the motion coherence theory. Int. J. Computer Vis. 3, 155–175 (1989).
Dempster, A. P., Laird, N. M. & Rubin, D. B. Maximum likelihood from incomplete data via the em algorithm. J. R. Stat. Soc.: Ser. B (Methodol.) 39, 1–22 (1977).
Tzikas, D. G., Likas, A. C. & Galatsanos, N. P. The variational approximation for Bayesian inference. IEEE Signal Process. Mag. 25, 131–146 (2008).
Zeng, H. et al. Integrative in situ mapping of single-cell transcriptional states and tissue histopathology in a mouse model of alzheimer’s disease. Nat. Neurosci. 26, 430–446 (2023).
Shi, H. et al. Spatial atlas of the mouse central nervous system at molecular resolution. Nature 622, 552–561 (2023).
Zhang, M. et al. Molecularly defined and spatially resolved cell atlas of the whole mouse brain. Nature 624, 343–354 (2023).
Wang, Q. et al. The Allen Mouse Brain Common Coordinate Framework: a 3d reference atlas. Cell 181, 936–953 (2020).
Schott, M. et al. Open-st: high-resolution spatial transcriptomics in 3d. Cell 187, 3953–3972 (2024).
Janesick, A. et al. High resolution mapping of the tumor microenvironment using integrated single-cell, spatial and in situ analysis. Nat. Commun. 14, 8353 (2023).
Yoshiura, K. -i et al. A SNP in the ABCC11 gene is the determinant of human earwax type. Nat. Genet. 38, 324–330 (2006).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn Res 12, 2825–2830 (2011).
Wang, L. et al. Extracellular matrix protein 1 (ECM1) is over-expressed in malignant epithelial tumors. Cancer Lett. 200, 57–67 (2003).
Rebbeck, C. A. et al. Gene expression signatures of individual ductal carcinoma in situ lesions identify processes and biomarkers associated with progression towards invasive ductal carcinoma. Nat. Commun. 13, 3399 (2022).
Cheng, M. et al. Three-dimension transcriptomics maps of whole mouse embryo during organogenesis. bioRxiv https://doi.org/10.1101/2024.08.17.608366 2024–08 (2024).
Sunkin, S. M. et al. Allen brain atlas: an integrated spatio-temporal portal for exploring the central nervous system. Nucleic Acids Res 41, D996–D1008 (2012).
Wang, Y., Liu, Z. & Ma, X. Mnmst: topology of cell networks leverages identification of spatial domains from spatial transcriptomics data. Genome Biol. 25, 133 (2024).
Wang, X. et al. Three-dimensional intact-tissue sequencing of single-cell transcriptional states. Science 361, eaat5691 (2018).
Zeng, H. et al. Spatially resolved single-cell translatomics at molecular resolution. Science 380, eadd3067 (2023).
Wake, H., Lee, P. R. & Fields, R. D. Control of local protein synthesis and initial events in myelination by action potentials. Science 333, 1647–1651 (2011).
Zhang, D. et al. Spatial epigenome–transcriptome co-profiling of mammalian tissues. Nature 616, 113–122 (2023).
Chen, M., Wei, Z., Huang, Z., Ding, B. & Li, Y. Simple and deep graph convolutional networks. In Proc. of the 37th International Conference on Machine Learning. Vol. 119, 1725–1735 (PMLR, 2020).
Zhang, H., Wu, Q., Yan, J., Wipf, D. & Yu, P. S. From canonical correlation analysis to self-supervised graph neural networks. Adv. Neural Inf. Process. Syst. 34, 76–89 (2021).
Li, L., Li, Z., Zakharchenko, V., Chen, J. & Li, H. Advanced 3d motion prediction for video-based dynamic point cloud compression. IEEE Trans. Image Process. 29, 289–302 (2019).
Williams, C. & Seeger, M. Using the nyström method to speed up kernel machines. Adv. Neural Inf. Process. Syst. 13, 661–667 (2000).
Xiao, Z. et al. 3d reconstruction of a gastrulating human embryo. Cell 187, 2855–2874 (2024).
Hu, Y. et al. Benchmarking clustering, alignment, and integration methods for spatial transcriptomics. Genome Biol. 25, 212 (2024).
Wolf, F. A., Angerer, P. & Theis, F. J. Scanpy: large-scale single-cell gene expression data analysis. Genome Biol. 19, 1–5 (2018).
Wu, T. et al. clusterprofiler 4.0: a universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Palla, G. et al. Squidpy: a scalable framework for spatial omics analysis. Nat. Methods 19, 171–178 (2022).
Yu, W., Xiaoke, M. & Zaiyi, L. Network model for alignment, stitching and slice-to-volume 3d reconstruction of large-scale spatially resolved slices. https://doi.org/10.5281/zenodo.18815221 (2026).
Acknowledgements
We thank the members of the Ma Lab for helpful discussions, and appreciate the researchers who provided us with the source code for comparison. This work was supported by the Noncommunicable Chronic Diseases-National Science and Technology Major Project (Grant Nos. 2024ZD0531100 and 2024ZD0531103 to X.M), the Joint Funds of the National Natural Science Foundation of China (Grant No. U22A20345 to X.M), the Natural Science Basic Research Program of Shaanxi (Grant No. 2025JC-QYCX-057 to X.M), the Xidian University Specially Funded Project for Interdisciplinary Exploration (Grant No. TZJHF202507 to X.M), and the R&D-Oriented Science and Technology Program Projects of Guyuan City (Grant No. 2025GKJYF0002 to Y.W.).
Author information
Authors and Affiliations
Contributions
Z.L. and X.M. conceived and designed the study. X.M., Z. L. and Y.W. performed the research. Y.W. collected and constructed the benchmark datasets and models. Y.W., Z. L. and X.M. completed the downstream analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Peer review
Peer review information
Nature Communications thanks Yanxiang Deng, who co-reviewed with Liran Mao, and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, Y., Liu, Z. & Ma, X. Network model for alignment, stitching and slice-to-volume 3D reconstruction of large-scale spatially resolved slices. Nat Commun (2026). https://doi.org/10.1038/s41467-026-71042-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-71042-6