Abstract
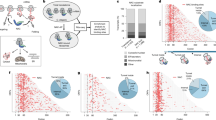
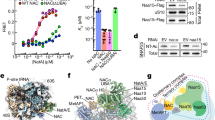
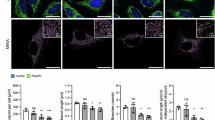
The nascent polypeptide-associated complex (NAC) co-translationally screens all nascent proteins and regulates their access to signal recognition particle (SRP) to ensure the fidelity of protein targeting to the endoplasmic reticulum (ER). However, the mechanism by which NAC prevents the mistargeting of nascent mitochondrial proteins remains unclear. Here, we identify a molecular switch in NAC that allows its central barrel domain to adopt a stabilized conformation on ribosomes exposing a mitochondrial targeting sequence (MTS). Mutations of the MTS on the nascent chain or in the NAC switch region increase NAC barrel dynamics and reduce its binding to the ribosome. This impairs the ability of NAC to prevent mistargeting by SRP and causes ER stress in human cells. Our work reveals how NAC detects nascent mitochondrial proteins early in translation and prevents their promiscuous access to SRP, elucidating the structural basis that underlies this role and providing mechanistic insights into protein targeting fidelity with broader implications for cellular proteostasis.
Similar content being viewed by others
Data availability
The cryo-EM maps and corresponding atomic models generated in this study have been deposited in the PDB and EMDB under accession code: EMD-48552 and PDB-9MR4 [https://doi.org/10.2210/pdb9mr4/pdb] for the OXA1L NAC-RNCMTS structure; EMD-71310 for HSP60 NAC-RNCMTS; EMD-71286 and EMD-71287 for the H59-undocked and -docked barrel states of the OXA1L∆MTS structure, respectively. All data related to this study are available in the main text or the supplementary material. Source data are provided with this paper.
References
Morales-Polanco, F., Lee, J. H., Barbosa, N. M. & Frydman, J. Cotranslational mechanisms of protein biogenesis and complex assembly in eukaryotes. Annu. Rev. Biomed. Data Sci. 5, 67–94 (2022).
Hegde, R. S. & Keenan, R. J. The mechanisms of integral membrane protein biogenesis. Nat. Rev. Mol. Cell Biol. 23, 107–124 (2022).
Gamerdinger, M. & Deuerling, E. Cotranslational sorting and processing of newly synthesized proteins in eukaryotes. Trends Biochem. Sci. 49, 105–118 (2024).
Rabl, L. & Deuerling, E. The nascent polypeptide-associated complex (NAC) as regulatory hub on ribosomes. Biol. Chem. 406, 295–307 (2025).
Truscott, K. N., Brandner, K. & Pfanner, N. Mechanisms of protein import into mitochondria. Curr. Biol. 13, R326–R337 (2003).
Dudek, J., Rehling, P. & van der Laan, M. Mitochondrial protein import: common principles and physiological networks. Biochim. Biophys. Acta BBA Mol. Cell Res. 1833, 274–285 (2013).
Wiedemann, N. & Pfanner, N. Mitochondrial machineries for protein import and assembly. Annu. Rev. Biochem. 86, 685–714 (2017).
Gamerdinger, M., Hanebuth, M. A., Frickey, T. & Deuerling, E. The principle of antagonism ensures protein targeting specificity at the endoplasmic reticulum. Science 348, 201–207 (2015).
Deng, J. M. & Behringer, R. R. An insertional mutation in theBTF3 transcription factor gene leads to an early postimplantation lethality in mice. Transgenic Res. 4, 264–269 (1995).
Markesich, D. C., Gajewski, K. M., Nazimiec, M. E. & Beckingham, K. bicaudal encodes the Drosophila beta NAC homolog, a component of the ribosomal translational machinery. Development 127, 559–572 (2000).
Bloss, T. A., Witze, E. S. & Rothman, J. H. Suppression of CED-3-independent apoptosis by mitochondrial βNAC in Caenorhabditis elegans. Nature 424, 1066–1071 (2003).
Raue, U., Oellerer, S. & Rospert, S. Association of protein biogenesis factors at the yeast ribosomal tunnel exit is affected by the translational status and nascent polypeptide sequence. J. Biol. Chem. 282, 7809–7816 (2007).
Wiedmann, B., Sakai, H., Davis, T. A. & Wiedmann, M. A protein complex required for signal-sequence-specific sorting and translocation. Nature 370, 434–440 (1994).
Liu, Y., Hu, Y., Li, X., Niu, L. & Teng, M. The crystal structure of the human nascent polypeptide-associated complex domain reveals a nucleic acid-binding region on the NACA subunit. Biochemistry 49, 2890–2896 (2010).
Wang, L. et al. Crystal structures of NAC domains of human nascent polypeptide-associated complex (NAC) and its αNAC subunit. Protein Cell 1, 406–416 (2010).
Hsieh, H.-H., Lee, J. H., Chandrasekar, S. & Shan, S. A ribosome-associated chaperone enables substrate triage in a cotranslational protein targeting complex. Nat. Commun. 11, 5840 (2020).
Wegrzyn, R. D. et al. A conserved motif is prerequisite for the interaction of NAC with ribosomal protein L23 and Nascent Chains. J. Biol. Chem. 281, 2847–2857 (2006).
Pech, M., Spreter, T., Beckmann, R. & Beatrix, B. Dual binding mode of the nascent polypeptide-associated complex reveals a novel universal adapter site on the ribosome. J. Biol. Chem. 285, 19679–19687 (2010).
Jomaa, A. et al. Mechanism of signal sequence handover from NAC to SRP on ribosomes during ER-protein targeting. Science 375, 839–844 (2022).
Alamo, M. D. et al. Defining the specificity of cotranslationally acting chaperones by systematic analysis of mRNAs associated with ribosome-nascent chain complexes. PLoS Biol. 9, e1001100 (2011).
Gamerdinger, M. et al. Early scanning of nascent polypeptides inside the ribosomal tunnel by NAC. Mol. Cell 75, 996–1006.e8 (2019).
Lentzsch, A. M. et al. NAC guides a ribosomal multienzyme complex for nascent protein processing. Nature 633, 718–724 (2024).
Gamerdinger, M. et al. NAC controls cotranslational N-terminal methionine excision in eukaryotes. Science 380, 1238–1243 (2023).
Klein, M., Wild, K. & Sinning, I. Multi-protein assemblies orchestrate co-translational enzymatic processing on the human ribosome. Nat. Commun. 15, 7681 (2024).
Gamerdinger, M. et al. Mechanism of cotranslational protein N-myristoylation in human cells. Mol. Cell 85, 2749–2758.e8 (2025).
Zdancewicz, S., Maldosevic, E., Malezyna, K. & Jomaa, A. NAC couples protein synthesis with nascent polypeptide myristoylation on the ribosome. EMBO J. 44, 6320–6342 (2025).
Denk, T. et al. Structural basis of co-translational N-myristoylation in humans. Nat. Commun. 17, 1191 (2026).
Yudin, D. et al. Mechanism of cotranslational modification of histones H2A and H4 by MetAP1 and NatD. Sci. Adv. 11, eaeb1017 (2025).
Zhang, Y. et al. NAC functions as a modulator of SRP during the early steps of protein targeting to the endoplasmic reticulum. Mol. Biol. Cell 23, 3027–3040 (2012).
Nyathi, Y. & Pool, M. R. Analysis of the interplay of protein biogenesis factors at the ribosome exit site reveals new role for NAC. J. Cell Biol. 210, 287–301 (2015).
Powers, T. & Walter, P. The nascent polypeptide-associated complex modulates interactions between the signal recognition particle and the ribosome. Curr. Biol. 6, 331–338 (1996).
Jomaa, A. et al. Molecular mechanism of cargo recognition and handover by the mammalian signal recognition particle. Cell Rep. 36, 109350 (2021).
Kobayashi, K. et al. Structure of a prehandover mammalian ribosomal SRP·SRP receptor targeting complex. Science 360, 323–327 (2018).
Lee, J. H. et al. Receptor compaction and GTPase rearrangement drive SRP-mediated cotranslational protein translocation into the ER. Sci. Adv. 7, eabg0942 (2021).
Costa, E. A., Subramanian, K., Nunnari, J. & Weissman, J. S. Defining the physiological role of SRP in protein-targeting efficiency and specificity. Science 359, 689–692 (2018).
Vögtle, F.-N. et al. Global analysis of the mitochondrial N-proteome identifies a processing peptidase critical for protein stability. Cell 139, 428–439 (2009).
Calvo, S. E. et al. Comparative analysis of mitochondrial N-termini from mouse, human, and yeast. Mol. Cell. Proteom. 16, 512–523 (2017).
von Heijne, G. Mitochondrial targeting sequences may form amphiphilic helices. EMBO J. 5, 1335–1342 (1986).
Roise, D., Horvath, S. J., Tomich, J. M., Richards, J. H. & Schatz, G. A chemically synthesized pre-sequence of an imported mitochondrial protein can form an amphiphilic helix and perturb natural and artificial phospholipid bilayers. EMBO J. 5, 1327–1334 (1986).
Abe, Y. et al. Structural basis of presequence recognition by the mitochondrial protein import receptor Tom20. Cell 100, 551–560 (2000).
Shanmuganathan, V. et al. Structural and mutational analysis of the ribosome-arresting human XBP1u. eLife 8, e46267 (2019).
Feng, Q. & Shao, S. In vitro reconstitution of translational arrest pathways. Methods 137, 20–36 (2018).
Lee, J. H. et al. Sequential activation of human signal recognition particle by the ribosome and signal sequence drives efficient protein targeting. Proc. Natl. Acad. Sci. USA. 115, E5487–E5496 (2018).
Yoshida, H., Matsui, T., Yamamoto, A., Okada, T. & Mori, K. XBP1 mRNA is induced by ATF6 and spliced by IRE1 in response to ER stress to produce a highly active transcription factor. Cell 107, 881–891 (2001).
Qi, L., Yang, L. & Chen, H. Chapter Eight - Detecting and Quantitating Physiological Endoplasmic Reticulum Stress. in Methods in Enzymology (ed. Conn, P. M.) vol. 490 137–146 (Academic Press, 2011).
Klein, M. A., Wild, K., Kišonaitė, M. & Sinning, I. Methionine aminopeptidase 2 and its autoproteolysis product have different binding sites on the ribosome. Nat. Commun. 15, 716 (2024).
Wild, K. et al. MetAP-like Ebp1 occupies the human ribosomal tunnel exit and recruits flexible rRNA expansion segments. Nat. Commun. 11, 776 (2020).
Aitken, C. E., Marshall, R. A. & Puglisi, J. D. An oxygen scavenging system for improvement of dye stability in single-molecule fluorescence experiments. Biophys. J. 94, 1826–1835 (2008).
Shen, K., Arslan, S., Akopian, D., Ha, T. & Shan, S. Activated GTPase movement on an RNA scaffold drives co-translational protein targeting. Nature 492, 271–275 (2012).
Preus, S., Noer, S. L., Hildebrandt, L. L., Gudnason, D. & Birkedal, V. iSMS: single-molecule FRET microscopy software. Nat. Methods 12, 593–594 (2015).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Punjani, A. & Fleet, D. J. 3D variability analysis: Resolving continuous flexibility and discrete heterogeneity from single particle cryo-EM. J. Struct. Biol. 213, 107702 (2021).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. Sect. Struct. Biol. 75, 861–877 (2019).
Weis, D. et al. Biallelic Cys141Tyr variant of SEL1L is associated with neurodevelopmental disorders, agammaglobulinemia, and premature death. J. Clin. Investig. 134, (2024).
Wang, H. H. et al. Hypomorphic variants of SEL1L-HRD1 ER-associated degradation are associated with neurodevelopmental disorders. J. Clin. Investig. 134, (2024).
Acknowledgements
We thank the Jomaa, Shan, and Qi lab members for the helpful discussions. Cryo-EM data collection was conducted at the molecular electron microscopy core facility (RRID:SCR_019031) at the University of Virginia (UVA) School of Medicine, which was built with NIH grant G20-RR31199. We thank Michael Purdy for assisting with the cryo-EM data collection and additional computational support. We thank Sarah Marks, Kinga Malezyna, and Travis Bishop for assisting with ribosome isolations, NAC purifications, and other biochemical assays. We also thank Shuangcheng Alivia Wu for initial help with preparation of NACβ-KO cells. This work was supported by NIH grant 1R35GM160490, the Owens Family Foundation, by the Searle Scholars Program, Grant #: SSP-2023-106, and aided by Grant # 134088-IRG-19-143-33-IRG from the American Cancer Society and the University of Virginia Comprehensive Cancer Center to A.J., by National Institutes of Health grant R35 GM136321 and National Science Foundation grant 2219287 to S.S., by National Institutes of Health grant R01DK120047, R01DK120330, R35GM130292 to L.Q. We acknowledge the cellular and molecular biology training program at UVA for support provided to E.M. through NIH T32GM139787-3 and the medical scientist training program for support provided to Z.J.L. In addition, the National Ataxia Foundation is acknowledged for support provided to L.L.L and Z.J.L through Post- and Pre-doctoral Fellowships NAF 918037 and 1036307, respectively.
Author information
Authors and Affiliations
Contributions
This study was conceived by A.J., S.S., E.M. and L.Q. Ribosomes were purified by E.M. and R.J.G. for structural and single molecule experiments, respectively. Y.P. provided materials used in various biochemical experiments throughout the study (site directed point mutagenesis, plasmid preps, cultured cells for ribosome isolations, and assisted with the NAC purifications). E.M. collected, processed cryo-EM data, and built the atomic model, and assembled the structural snapshots. R.J.G. conducted the single molecule total internal reflection microscopy studies. L.L.L., L.E.Z. and Z.J.L. generated the NACβ-KO cells and conducted the cell-based experiments. A.J., S.S. and L.Q. supervised the work. E.M., A.J., S.S. and R.J.G. wrote the manuscript. All authors contributed to data analysis and the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Andrei Korostelev and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Maldosevic, E., Gora, R.J., Lin, L.L. et al. A molecular switch in NAC prevents mitochondrial protein mistargeting by SRP. Nat Commun (2026). https://doi.org/10.1038/s41467-026-71061-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-71061-3