Abstract
Many insects benefit from ancient nutrient-supplementing endosymbionts. While symbiont losses and replacements occur on evolutionary timescales, their dynamics remain enigmatic due to the lack of experimentally tractable systems. Here, we report on the experimental establishment of the culturable bacterium Sodalis praecaptivus in a grain pest beetle (Oryzaephilus surinamensis) and its effect on the native symbiont Shikimatogenerans silvanidophilus, which produces the tyrosine precursor prephenate. Injection of Sodalis into female beetles led to systemic intracellular infection and efficient transovarial vertical transmission but reduced host survival and reproduction. Interestingly, Sodalis also invaded the host’s bacteriomes, causing irregular morphology and rapid loss of Shikimatogenerans within three beetle generations. Transcriptomics revealed a strong upregulation of host immune effectors upon Sodalis infection, but little reaction from Shikimatogenerans, indicating that the ancient symbiont is incapable of responding adaptively to the introduced competitor. The rapid elimination of the native symbiont in O. surinamensis showcases the fragility of ancient beneficial symbioses and experimentally recapitulates a crucial step towards a functional symbiont replacement.
Similar content being viewed by others
Introduction
Insects commonly engage in intracellular symbioses with microorganisms spanning the continuum from pathogens to mutualists. Associations with beneficial symbionts in particular play an important role in insect adaptation and diversification, because the biosynthetic capabilities of microbial symbionts can provide vital metabolites to the host, such as essential nutrients, digestive or detoxifying enzymes, or protective secondary metabolites, thereby enabling insects to expand into otherwise inaccessible ecological niches1,2. Many interactions between insects and beneficial microorganisms are ancient in origin3,4,5,6 and have experienced tight coevolution. This process commonly results in the erosion of the symbiont genome7 and ultimately leads to mutual metabolic dependence of the symbiotic partners8. Despite the intimacy and functional importance of many intracellular insect symbioses, phylogenetic analyses reveal that ancient beneficial symbionts are occasionally lost9,10 or replaced by other microorganisms11,12,13,14.
Symbiont losses can occur when symbiont-provided benefits no longer outweigh the costs of maintaining them. In insects depending on symbiotic microbes to compensate for nutritional deficiencies2, the loss of a costly symbiont can be advantageous for hosts that acquire the metabolic capacity to perform symbiont-mediated reactions themselves, for example via horizontal gene transfer10. Alternatively, symbiont losses may be catalyzed by changes in the host’s ecological niche—e.g., a switch to a more nutritionally balanced diet—that render symbiont-provided benefits unnecessary15. Finally, a newly acquired symbiont can lead to the demise of the original one by replacing or even expanding the repertoire of benefits provided to the host. Such symbiont replacements are often assumed to be beneficial to the host, as they allow for escaping the rabbit hole of increasing genome erosion and accumulation of slightly deleterious mutations in the symbiont’s genome due to Muller’s ratchet12.
For a replacement of an intracellular symbiont to be successful, four criteria have to be fulfilled: The new symbiont must be able to (i) colonize and persist in the host, and (ii) achieve efficient transmission. Additionally, (iii) the original symbiont must be eliminated, while (iv) the newly acquired symbiont evolves a net benefit and takes over (and potentially extends) the original symbiont’s role. In contrast to unicellular systems, like ciliates, where symbiont replacements have been well studied16 and can be experimentally recapitulated17,18, the sequence of events leading to a replacement of an intracellular symbiont and the speed at which they occur remain obscure in animals, where the soma-germline separation constitutes a considerable barrier for vertical transmission. In insects, symbiont replacements are widespread but rare on ecological timescales, and most intracellular symbioses are intractable, hampering experimental studies that address symbiont losses and replacements despite their importance in understanding the evolutionary dynamics of host-microbe interactions19.
Intracellular associations that may enable experimental symbiont replacements have recently been described in several beetle lineages. Coleoptera, the most species-rich order of insects20, are characterized by the rigid elytra that protect the beetle from desiccation and predation21. To build this protective barrier, however, beetles require large amounts of tyrosine, because tyrosine derivatives like N-acetyldopamine, N-β-alanyldopamine, and melanin are necessary for cuticle sclerotization and melanization22,23. Tyrosine is a semi-essential amino acid that insects cannot synthesize de novo and therefore need to acquire from the diet or from symbiotic microorganisms. Concordantly, to meet their high demands for tyrosine, many beetles associate with intracellular symbionts that supply tyrosine precursors and thereby support cuticle development under nutrient-limited conditions6,9. These tyrosine-supplementing symbioses are promising candidates for establishing experimentally tractable symbiotic systems, since many tyrosine-producing symbionts are not essential for host survival under laboratory conditions24,25,26. This can allow for the functional replacement of a beneficial symbiont with a culturable bacterium, as demonstrated by the experimental replacement of Sitophilus zeamais’ native symbiont Sodalis pierantonius with the culturable congener Sodalis praecaptivus27 that was engineered to overproduce and secrete tyrosine19.
Here, we explored the potential of S. praecaptivus (from here on referred to as Sodalis) to establish successful infections in the saw-toothed grain beetle Oryzaephilus surinamensis harboring the distantly related Bacteroidota symbiont Shikimatogenerans silvanidophilus (from here on referred to as Shikimatogenerans). Shikimatogenerans bacteria are harbored intracellularly in two pairs of bacteriomes and are transmitted vertically from mother to offspring. The symbiotic bacteria provision the tyrosine precursor prephenate to their host, which the beetle converts into tyrosine to support cuticle biosynthesis6,28,29. We demonstrate that, upon injection into adult females, Sodalis establishes systemic intracellular infections in the novel host. Importantly, the new symbiont spontaneously achieves highly efficient vertical transmission to the offspring, a feature that is widely accepted to be crucial for the evolution of beneficial symbioses by aligning host and symbiont fitness interests30,31. However, in the initial encounter between Oryzaephilus and Sodalis, the latter negatively impacts host fitness. We further show that the presence of Sodalis disrupts the native symbiont, affects its morphology, and ultimately leads to its rapid loss from the host population within three beetle generations. Triple RNAseq reveals that the host mounts a strong immune response to Sodalis’ presence in the bacteriome tissue, but Shikimatogenerans is incapable of adjusting its gene expression in response to the intruder. Our results show that infection with a novel facultative symbiont can result in the rapid breakdown of an ancient beneficial symbiosis and lead to the elimination of the native symbiont, thus fulfilling three out of the four criteria necessary for a stable symbiont replacement. Despite the detrimental nature of the novel symbiont, its maintenance and highly efficient vertical transmission may pave the way for a symbiont replacement in the future, thus providing an experimentally tractable system to recapitulate major events in the evolution of beneficial symbioses in insects.
Results
Sodalis establishes systemic infection in O. surinamensis and is vertically transmitted
We tested the potential of fluorescently labeled Sodalis to establish a novel symbiotic interaction with O. surinamensis beetles by microinjecting 10 nl aliquots of a Sodalis suspension (OD600 = 1, i.e., 6.01 × 103 ± 2.79 × 103 CFUs) into the abdomen of reproductively active female adults. After 7 days, we observed that beetles injected with Sodalis exhibited strong fluorescence, indicative of mCherry production by this bacterium (Fig. 1a–d). Uninfected control beetles were devoid of mCherry fluorescence, verifying that the signal observed was not caused by cuticular autofluorescence. As determined via quantitative polymerase chain reaction (qPCR), the population of Sodalis had grown to 1.46 × 107 ± 7.28 × 106 tam gene copy numbers by day 7 and slightly increased even further in the following week (2.38 × 107 ± 9.12 × 106 tam gene copy numbers on day 14, Supplementary Fig. 1).
Brightfield images of injected control (a) and Sodalis-treated females (c) and their corresponding fluorescence microscopy images (b, d) taken 7 days after injection. Note that the distal ends of the elytra were removed for injection, facilitating detection of fluorescence at the posterior end of the abdomen. Brightfield images of first-generation larval offspring of control (e) and Sodalis-injected (g) females and their corresponding fluorescence microscopy images (f, h). Red signal in d, h is indicative of mCherry protein produced by Sodalis. Fluorescence deriving from Sodalis infection was detected in all females injected with Sodalis that survived up to 7 dpi (n = 297) while it was absent from all control beetles surviving up to this timepoint (n = 305). No fluorescence was observed in larvae of the control treatment (n = 1555), but could be detected in a majority of larvae produced by Sodalis-infected females (n = 227). Scale bars = 1 mm.
When screening larval first generation (F1) offspring, we discovered that offspring from females injected with bacteria also showed strong mCherry fluorescence, demonstrating that females harboring Sodalis vertically transmitted the novel symbionts to their offspring (Fig. 1e–h). With fluorescence-in-situ-hybridization (FISH), we confirmed that transmission indeed occurred vertically via the ovaries, since we detected Sodalis cells in young oocytes (Supplementary Fig. 2). Transmission occurred with an efficiency of 93.75% (Supplementary Fig. 3). We did not observe any paternal transmission of Sodalis: Larvae produced by mating pairs of Sodalis-infected virgin males and uninfected virgin females never exhibited mCherry fluorescence (Supplementary Fig. 4). Males harboring Sodalis also did not transmit the bacteria to the female beetles during mating (Supplementary Fig. 4). We were able to maintain Sodalis infected beetles up to the third offspring generation by mating infected males and females, before sacrificing all remaining specimens for microscopy.
Sodalis infection negatively affects host fitness
Following the successful establishment of Sodalis in O. surinamensis beetles, we investigated the impact on host fitness. While 80% of control beetles survived the observation period of 80 days, Sodalis infection significantly reduced beetle survival (Cox mixed-effects model, df = 1, Χ2 = 214.31, P < 0.001, Fig. 2a). Almost no beetles harboring the novel symbiont survived the observation period, and over 50% of individuals had died by day 50. We also found beetles carrying Sodalis to produce significantly fewer larvae than beetles of the control treatment (zero-inflated negative binomial model, z = −9.46, P < 0.001, Fig. 2b). For the F1 adults, we investigated cuticle melanization and thickness as indicators for tyrosine availability and proxies for host fitness with respect to protection from predation and desiccation. Beetles harboring Sodalis were significantly lighter than their control counterparts in both abdomen (Wilcoxon rank sum test, W = 741, P < 0.001) and thorax (two sample t-test, df = 57, t = 5.76, P < 0.001), suggesting a reduced amount of tyrosine available for melanization (Fig. 2c). In contrast to melanization, cuticle thickness was not affected by Sodalis infection, independent of whether we measured in individual sections (Wilcoxon rank sum test, W = 90, P = 0.13, Fig. 2d) or whole beetles (two-sample t-test, df = 20. t = −0.85, P = 0.41, Supplementary Fig. 5).
a Survival probability (mean and 95% confidence interval) of control (light blue) and Sodalis injected beetles (purple). The presence of Sodalis negatively affected beetle survival probability over the observation period (Cox mixed-effects model, ANOVA, P < 2.2 × 10−16). b The number of larvae produced by each female injected with Sodalis (purple) was significantly lower than in control females (light blue, zero-inflated negative binomial model, Wald z-test, P < 2.2 × 10−16). c 7-day old adults of first-generation offspring had significantly less melanized cuticles of both thorax and abdomen when they harbored Sodalis (thorax: two-sided two sample t-test, P = 3.49 × 10−7, abdomen: two-sided Wilcoxon rank sum test, P = 1.23 × 10−10). d Cuticle thickness in thorax sections did not differ in 7-day old adults of first-generation offspring between both treatments (two-sided Wilcoxon rank sum-test, P = 0.13). Statistical differences indicated by asterisks (***P < 0.001). n.s.: not significant. Source Data are provided as a Source Data file.
Sodalis-infected beetles lose their native symbiont within three generations
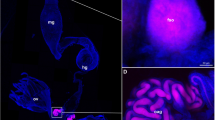
We performed FISH to investigate the localization of Sodalis in O. surinamensis across three different host generations. While Sodalis was consistently absent from control beetles (Fig. 3a, b, e, f, i, and Supplementary Figs. 6–8), it occupied various tissues in injected beetles and their offspring. In the parental beetle generation, the novel symbiont was found in the hemolymph and fatbody across the whole host body, the ventral nerve cord, and was also highly abundant in the ovaries. We also found Sodalis in the bacteriomes, even though it was less abundant than in the other tissues (Fig. 3c, d). In the first offspring generation, Sodalis colonized the same tissues, and we additionally observed the sporadic presence of bacteria in the male testes, although the bacteria—when present—seemed to be less abundant compared to female ovaries (Supplementary Fig. 2). In contrast to the parental generation, Sodalis was more abundant in the bacteriomes, even though its density varied even within the organs—while some bacteriocytes harbored high loads of Sodalis, other bacteriocytes of the same bacteriome were apparently uninfected (Fig. 3g, h, and Supplementary Fig. 7). We could not detect bacteriomes and the native symbiont Shikimatogenerans in two of the twelve Sodalis-infected F1 beetles we sampled for microscopy (Supplementary Fig. 7). In the third offspring generation, all six Sodalis-infected specimens (five adults and one pupa) were devoid of the native symbiont (Fig. 3j, and Supplementary Fig. 8). The seeming absence of bacteriomes in the respective F1 and F3 individuals either indicates that the beetles fail to develop these organs following the loss of Shikimatogenerans or that we failed to detect empty and thus atrophied bacteriomes. In contrast to Sodalis-infected F3 individuals, control beetles of the third and subsequent offspring generations retained fully developed bacteriomes (Fig. 3i, and Supplementary Fig. 8). The observed loss of Shikimatogenerans in Sodalis-treated beetles compared to control beetles could not be explained by chance (Fisher’s exact test, P < 0.001).
FISH micrographs of control (a, b) and Sodalis injected beetles (c, d) of the parental generation, b and d are closeups of the areas marked with a frame in a and c. In 7-day old beetles of the first offspring generation, Shikimatogenerans displayed globular or rod-shaped morphology in Sodalis-free control beetles (e, f), while symbiont cells were aberrantly shaped upon colocalization with Sodalis (g, h). f and h are closeups of the areas marked with a frame in e and g. Arrows in g highlight bacteriocytes that are almost uninfected with Sodalis. Arrows in h highlight particularly altered Shikimatogenerans morphology, while the dashed area highlights regularly shaped symbiont cells in a practically uninfected bacteriocyte of the same bacteriome. In adults of the third offspring generation, Shikimatogenerans was present in the control treatment (i), while it was lost in adults harboring Sodalis (j). DNA was stained with DAPI (cyan), while the probes OsurSym16S-Cy5 and Sod-FISH-Cy3 were used to label Shikimatogenerans (yellow) and Sodalis (magenta), respectively. Observations were replicated independently with similar results (Parental generation: n = 5 for either treatment, F1: n = 7 for control and n = 10 for Sodalis-infected beetles, F3(+): n = 10 for control and n = 6 for Sodalis-infected beetles). Scale bars represent 50 µm, gl= gut lumen.
Sodalis affects Shikimatogenerans and bacteriome phenotype
Given the loss of Shikimatogenerans and the apparent lack of bacteriomes in F3 adults following Sodalis infection, we further investigated the impact of Sodalis on bacteriome size and Shikimatogenerans phenotype. Using µCT scans, we examined the influence of Sodalis on the bacteriomes of F1 beetles. We found that the mean bacteriome volume did not differ between beetles harboring Sodalis and uninfected beetles (Wilcoxon rank sum test, W = 55, P = 1; Fig. 4a–c). However, we observed an increase in the variance of bacteriome volumes in beetles harboring Sodalis, with one bacteriome per pair (dorsal/ventral) being bigger than the other (two sample t-test, df = 19, t = −5.32, P < 0.001, Fig. 4d). Despite the increased variance in bacteriome volume, the titer of Shikimatogenerans in F1 adults was unaffected by the presence of Sodalis and did not differ from the titer we observed in control beetles (Wilcoxon rank sum test, W = 234, p = 0.19; Fig. 4e). We also did not observe differences in titers in the parental generation (0 dpi: Wilcoxon rank sum test, W = 119.5, P = 0.79; 7 dpi: t-test, df = 27, t = 0.57, P = 0.57; 14 dpi: Wilcoxon rank sum test, W = 142, P = 0.11, Supplementary Fig. 1). However, in bacteriocytes harboring abundant Sodalis, we observed altered morphologies of the beetles’ native symbiont Shikimatogenerans: Colocalization of the two microbial partners led to enlarged and irregularly shaped Shikimatogenerans cells as compared to globular or rod-shaped Shikimatogenerans in bacteriocytes of uninfected control beetles (Fig. 3g, h). Quantifying the effect of Sodalis on Shikimatogenerans cell length, we detected a significant increase in cell length of at least 30% in the presence of Sodalis (two-sample t-test, df = 16, p = −5.71, P < 0.001; Fig. 4f).
3D reconstruction of 7-day old adult F1 beetle offspring belonging to the control (a) and Sodalis-infected (b) treatment. Dorsal and ventral bacteriomes were reconstructed based on µCT scans and are labelled in blue/green and yellow/orange, respectively. Scale bar represents 1 mm. c The presence of Sodalis did not lead to a change in mean bacteriome volume (two-sided Wilcoxon rank sum test, P = 1.0). d Mean variance of bacteriome volume within a beetle was higher when beetles harbored Sodalis (two-sample t-test, P = 3.95 × 10−5). e 16S copy number of Shikimatogenerans did not differ between F1 beetles of the control and Sodalis infected treatment (two-sided Wilcoxon rank sum test, P = 0.19). f Upon colocalization of bacteriocytes with Sodalis, the mean length of Shikimatogenerans cells significantly increased (two-sample t-test, P = 3.23 × 10−5). Asterisks indicate statistical significance. (***P < 0.001). Source Data are provided as a Source Data file.
The host beetle, but not Shikimatogenerans, transcriptionally responds to Sodalis infection
To understand how the Sodalis infection affected host and symbiont transcriptional responses in the bacteriomes of O. surinamensis, we analyzed gene expression of O. surinamensis and Shikimatogenerans in the bacteriomes of adult Sodalis-injected versus control beetles, as well as their offspring 7 days after reaching adulthood. O. surinamensis showed a significant upregulation of many immune system genes in the presence of Sodalis (Fig. 5a, b, and Supplementary Fig. 9a). In both generations, the genes with the highest fold change difference and significance were immune genes such as antimicrobial peptides (AMPs), peptidoglycan recognition proteins LB (PGRP-LB) and S1 (PGRP-S1), serine protease inhibitors (serpins) and proteases. We also detected differentially expressed genes involved in amino acid metabolism: Dopamine beta-hydroxylase was upregulated in Sodalis-infected bacteriomes of the parental generation and followed the same albeit non-significant trend in the first offspring generation (Supplementary Fig. 9). Furthermore, a Kynurenine/alpha-aminoadipate aminotransferase (AADAT) and a phenylananine-4-hydroxylase (phhA) were upregulated in the F1 generation of Sodalis-infected beetles. No genes in Shikimatogenerans demonstrated differential expression in accordance with a definition of statistical significance involving at least a log2-fold change of normalized transcripts over |1| in Shikimatogenerans and a corrected p-value threshold of 0.05 (Fig. 5c, d). Yet, we note that the gene showing the highest level of differential expression encodes a proline dehydrogenase (PRODH), which was significantly differentially expressed with a log2-fold change of 0.75 (i.e., a 1.68-fold upregulation) in the F1 generation. Besides PRODH, we observed a total of 13 other genes to be significantly differentially expressed, albeit below a log2-fold change of 1 in the parental (upregulated: not annotated; downregulated: groES, MTFMT, rsml) and the F1 generation (upregulated: mvaK1, rpoB, rpoC, sufB, sufC; downregulated: rpmg, prtC, recJ). We also investigated the 100 most highly expressed protein-coding genes in Sodalis in both generations. In the parental generation, 20 of the 100 most highly expressed genes were predicted to be involved in the flagellar apparatus or its regulation (Supplementary Fig. 10a). By contrast, the respective motility genes were significantly downregulated in the first offspring generation (Supplementary Fig. 10b). We also detected genes of the type three secretion system (T3SS, e.g., sseD, sseB, yscU and yscF) and genes involved in quorum sensing (e.g., expI, expR, hfq and secG) to be highly expressed in both beetle generations (Supplementary Fig. 10a). Additionally, several genes encoding metabolism regulators, namely ptsH, rpoH, crp and crr were identified among the top 100 genes in both generations.
Transcriptional responses of O. surinamensis and its ancient Shikimatogenerans symbiont to infection by Sodalis in the bacteriomes. Volcano plots of differential gene expression analysis results for O. surinamensis (a, b) and Shikimatogenerans (c, d) in the bacteriomes of the parental (a, c) and the first offspring generation (b, d), respectively. Lines represent the thresholds for gene expression regarding log2 fold change and -log10 of false discovery rate (a, b) or adjusted p-value (c, d, Wald test with Benjamini–Hochberg correction for multiple testing). Source Data are provided as a Source Data file.
Discussion
Intracellular symbioses with bacteria fundamentally impact insect ecology and evolution and can remain stable for hundreds of millions of years. Occasionally, however, even ancient and metabolically tightly integrated symbionts can be lost when their benefits deteriorate or become obsolete10 or when other microorganisms colonizing the host outcompete the native symbionts. In the latter case, the acquired microorganisms can functionally replace the original symbiont and/or provide additional benefits, occasionally expanding the host’s ecological niche space. Based on phylogenetic reconstructions, losses and replacements of ancient intracellular symbionts have been described across a variety of insect taxa, including spittlebugs, cicadas, aphids, and beetles9,10,11,12,32,33,34. The prerequisites for a symbiont replacement comprise (i) the establishment and persistence in a host, (ii) reliable vertical transmission, (iii) loss of the original symbiont, and (iv) a net benefit provided by the new symbiont. Yet the early steps and dynamics of these replacement events remain poorly understood, due to the lack of tractable study systems. Here, we describe the establishment of such a tractable symbiosis and report on the rapid loss of the host’s ancient beneficial symbiont upon establishment of a novel vertically transmitted bacterium. Our findings thus recapitulate the collapse of an ancient beneficial symbiosis and an important step towards a symbiont replacement, meeting three out of four necessary prerequisites, namely (i)–(iii), but not (iv).
To replace an ancient symbiont, an invading microbe like Sodalis must be capable of establishing in its new host by withstanding the insect’s immune system. Most intracellular beneficial symbionts likely evolved from pathogenic bacteria2 that employ two general strategies for coping with host immunity: (i) evasion of host immunity, i.e., preventing the host from mounting an immune response35,36,37, or (ii) resistance to host immune effectors36. Following infection with Sodalis, AMPs, PGRP-LB and other PGRPs, serpins, as well as other host immune effectors (e.g., dual oxidases (DUOX) producing reactive oxygen species, see Supplementary Fig. 9) were upregulated in the host bacteriome. Thus, the novel symbiont elicits an immune response but can persist despite the upregulation of host effectors, indicating resistance to host immunity. Indeed, the resistance of S. praecaptivus and its congener S. glossinidius to different AMPs has been demonstrated in vitro as well as in vivo and is mediated by the PhoP-PhoQ two-component system that regulates the expression of genes involved in modifying bacterial lipopolysaccharide (LPS) and thereby prevents damage by AMPs in vivo38,39,40. Apart from the upregulation of AMPs in Sodalis-infected bacteriomes, we also detected the upregulation of PGRP-LB and serpins in both investigated beetle generations. PGRP-LB is a negative regulator of the Imd immune pathway and plays a crucial role in the maintenance of the symbiosis between Sitophilus zeamais and S. pierantonius41,42. In the weevil bacteriomes, PGRP-LB is highly expressed and cleaves the peptidoglycan of S. pierantonius into non-immunogenic fragments, preventing an immune response that would lead to the elimination of the symbiont42. In contrast to S. pierantonius, Shikimatogenerans lacks a cell wall29—accordingly, PGRP-LB is not constitutively expressed in control O. surinamensis bacteriomes (Supplementary Fig. 9b) but might rather regulate host immunity after pathogen infection. The upregulation of PGRP-LB and serpins in the presence of Sodalis might modulate the immune response to prevent a negative impact of immunity on the host or damage to the native symbiont Shikimatogenerans by the beetle’s immune effectors. Despite the elevated expression of negative regulators of the host immune response, we observed a significant upregulation of effectors involved in the Imd pathway, likely incurring a cost on the insect host and potentially causing or contributing to the observed detrimental impacts on the native Shikimatogenerans symbiont.
After successful establishment, newly acquired symbionts need to secure their continuous association with the host by achieving efficient transmission43. Although not native to O. surinamensis, Sodalis readily invaded the host’s germline and infected the developing eggs, thereby establishing a highly efficient transovarial vertical transmission route. Sodalis’ motility genes were among the most highly expressed genes in bacteriomes of newly infected beetles, but not in the following generation, suggesting a role of motility in the initial infection, as seen in other systems44,45,46. Additionally, successful colonization and transmission likely depend on Sodalis’ T3SS that allows for the invasion of host cells in bacterial pathogens and some beneficial symbionts47,48,49. Indeed, genes encoding for T3SS components (e.g., sseD, sseB, yscU and yscF) were among the most highly expressed genes in Sodalis in both P and F1 beetle generations (Supplementary Fig. 10a). Our study provides further support for the notion that efficient vertical transmission can occur spontaneously in new host-symbiont combinations. However, it also demonstrates that there is a strong host component contributing to transmission, since the same Sodalis strain is transmitted at lower efficiency in another beetle host, S. zeamais, that natively harbors a Sodalis symbiont19, and fails to achieve any vertical transmission following injection into Drosophila melanogaster50.
In insects that already harbor intracellular symbionts, newly acquired vertically transmitted microbes can coinhabit their new niche with the host’s native symbionts, providing opportunities for symbiont-symbiont interactions that can determine the outcome of the coinfection. Although symbiont replacements are widespread across insects, little is known about the initial interactions between new and native symbiont. Limited host resources could drive microbial competition for nutrients, which might disrupt the tightly regulated metabolite exchange between native symbiont and host and could ultimately lead to the exclusion of one microbial partner, as observed in this study. Nutritional competition among intracellular bacteria residing in the same host is an understudied phenomenon, especially in the context of symbiont replacements. Yet, metabolic models predict that obligate Portiera aleyrodidarum and facultative Hamiltonella defensa symbionts compete for nutrients supplied by their whitefly (Bemisia tabaci) host, thereby limiting Portiera growth and resulting in a deficit of nutrients supplied to the insect51. Similarly, competition for nutrients is suggested to explain the negative effect of Spiroplasma or Serratia symbiotica on the abundance of Wolbachia pipientis in D. melanogaster52 or Buchnera aphidicola in Acyrthosiphon pisum53, respectively.
Since competition for resources among coinfecting microbes seems to occur across different insect systems, it may also explain the rapid loss of Shikimatogenerans upon introduction of Sodalis: The native symbiont Shikimatogenerans aids its beetle host by supplying it with prephenate, a precursor of tyrosine needed for robust cuticle synthesis and melanization29. Due to a highly eroded genome, the symbiont is not able to produce any other nutrients, such as vitamins, amino acids, fatty acids or nucleotides29. Thus, the host needs to supply these metabolites to the symbiont to enable the biosynthesis of prephenate. The bacteriomes are therefore expected to contain high concentrations of host-provided nutrients that could be prone to exploitation by bacteria like Sodalis that invade the symbiont-bearing organs. Exemplary for the exploitation of host-supplied nutrients is the depletion of tyrosine by Sodalis, as evidenced by a decrease in beetle melanization and the upregulation of host phhA in the bacteriomes, mediating the conversion of phenylalanine into tyrosine. Similarly, AADAT expression was upregulated in bacteriomes following Sodalis infection. Analogous to a glutamate oxaloacetate transaminase in Rhynchophorus ferrugineus weevils54, promiscuous AADAT55 can be involved in converting prephenate and its derivative 4-hydroxyphenylpyruvate into tyrosine. These findings are indicative of a tyrosine starvation response in the beetle56 and suggest the disruption of host-symbiont metabolite exchange via nutritional competition between Sodalis and Shikimatogenerans.
Apart from the host, we also detected signs of nutrient depletion in the ancient symbiont. Interestingly, in Shikimatogenerans, the most highly upregulated transcript encodes a PRODH (log2-fold change = 0.75), that is involved in proline catabolism leading to the production of glutamate which can serve as a donor of amino groups or be used for protein biosynthesis. Upregulation of PRODH is therefore consistent with amino acid starvation and corroborates the hypothesis that Sodalis infection disturbs the metabolic exchange between host and native symbiont. Beyond PRODH, however, our RNAseq data indicate that Shikimatogenerans appears to be limited in its ability to modulate its gene expression. Like many other ancient and genome-eroded insect endosymbionts, the Shikimatogenerans gene inventory lacks protein-coding genes that are predicted to serve as transcriptional regulators (but see Carvalho et al.57). Yet, we note that transcription could be modulated through other means such as non-coding small RNAs and interactions between metabolites and transcription machinery.
Facilitated by the symbiont’s reduced capacity to modulate its gene expression and respond to stressors, as well as the absence of a bacterial cell wall to protect cell integrity58, changes in the nutritional and/or physiochemical environment in the bacteriome caused by Sodalis (and potentially exacerbated by the host immune response) could result in the observed morphological aberrations and the ultimate elimination of the native symbiont. Still, the outcome of symbiont-symbiont interactions seems highly dependent on the combination of partners. While in O. surinamensis, Sodalis infection causes the rapid loss of Shikimatogenerans, the same strain failed to stably coexist with the more recently acquired congener S. pierantonius in S. zeamais due to reduced transmission efficiency in the presence of the native weevil symbiont19.
While the introduction of Sodalis into Oryzaephilus beetles meets three out of four necessary prerequisites for a symbiont replacement, Sodalis did not provide a net benefit to its novel host and, thus, remained pathogenic in our experimental setup. Generally, it is assumed that ancient symbionts are replaced because the benefits they provide deteriorate over time due to genome erosion, resulting in selective pressures on the host to favor symbionts that provide the benefits more efficiently59. Thus, when a novel symbiont replaces its host’s ancient symbiont, it is commonly expected that it will evolve necessary beneficial properties before the native obligate symbiont is eliminated. Still, under certain conditions, hosts might be able to tolerate the loss of an ancient symbiont before establishing a mutually beneficial relationship with another microbe. This particularly applies to cases where the novel microbe is the main driver of the native symbiont’s elimination and where the ancient symbiont is not essential for host survival and provides context-dependent benefits to the host. While novel microbes as drivers of symbiont losses have not previously been described in the literature, context-dependent symbiont benefits are common in insect-microbe symbioses6,25,60,61,62,63. This also applies to the Oryzaephilus system, where beetle hosts can survive without Shikimatogenerans, at least under conditions of low desiccation stress and antagonist pressure. We hypothesize that Sodalis could be capable of evolving a benefit to the host and functionally replacing the native symbiont, based on several factors: First, Sodalis should experience high selection pressure to attenuate its virulence and reduce the negative impact on host fitness due to its strictly vertical transmission route. The invasion of reproductive tissues and the ability to achieve vertical transmission are important steps towards evolving a mutualistic relationship with a host, since vertical transmission aligns host and symbiont fitness and thereby selects for symbionts with host-beneficial phenotypes30,31. As such, vertical transmission is generally thought to be a key driver in the evolution of beneficial intracellular symbioses2. Second, given its metabolic potential, it is conceivable that S. praecaptivus could evolve a beneficial phenotype, e.g., via leakage of metabolites like essential amino acids or vitamins that can lead to the establishment of beneficial cross-feeding in bacteria-bacteria7,64 and bacteria-eukaryote interactions65. Concordantly, previous studies revealed that mutualisms can evolve rapidly, as observed in Escherichia coli where single mutations in either cyaA or crp, two genes involved in carbon catabolite repression (CRR), were sufficient to render it a mutualist of stink bugs66. Interestingly, crp as well as crr and ptsH, which are also involved in CRR, are among the 100 most highly expressed genes of Sodalis in the beetle host and could be potential targets for experimentally manipulating Sodalis to become more benign. Third, the potential of Sodalis to form lasting beneficial symbioses is supported by the high number of Sodalis-allied symbionts in various insect taxa67,68,69,70,71. While functionally replacing a highly efficient and streamlined ancient symbiont is not trivial, many Sodalis symbionts have apparently succeeded in substituting other intracellular insect symbionts11,34. The success of this scenario, however, will highly depend on the speed at which virulence attenuation can be achieved and beneficial traits evolve.
In conclusion, the elimination of Shikimatogenerans following the introduction of Sodalis in O. surinamensis showcases the fragility of ancient intracellular symbioses and reveals that symbiont losses can occur rapidly upon acquisition of a novel symbiont. We experimentally demonstrate that newly acquired microbes can drive losses of ancient intracellular symbionts, potentially leading to the replacement of non-obligate or context-dependent host-beneficial microbes. Given that Sodalis is genetically tractable, the Oryzaephilus-Sodalis symbiosis allows for the investigation of symbiont elimination and replacement as well as the transition from pathogenicity to mutualism on a molecular level. Such experimentally and genetically tractable systems are urgently needed to deepen our understanding of the establishment and maintenance of animal-microbe symbioses and to study the evolution of host-beneficial interactions in real time.
Methods
Insect cultures and bacterial strains
O. surinamensis beetles, harboring the intracellular symbiont S. silvanidophilus, were originally provided by the Julius-Kühn-Institute/Federal Research Centre for Cultivated Plants (Berlin, Germany) in 2014. Since then, these beetles have been reared on organic oat flakes (Huber Mühle, Germany) under controlled conditions of 27–28 °C and 60% relative humidity (rH) following a day/night cycle of 16/8 h, respectively.
As a culturable symbiont, we used a strain of S. praecaptivus (strain MC1, identifier: CD2555, Su et al.19) that expresses fluorescent mCherry protein, enabling us to observe infection of beetles in vivo, since infected beetles exhibit strong fluorescence.
Establishing Sodalis in O. surinamensis
Sodalis was introduced into O. surinamensis by microinjecting a bacterial suspension into the hemolymph of female adults, following established protocols47. Briefly, we grew Sodalis in liquid LB medium overnight, at 28 °C and 220 rpm. After incubation, we concentrated bacteria to OD600 = 1.0 in physiological saline (0.85% NaCl (w/v)) under sterile conditions. Using a Flaming/Brown micropipette puller (Model P-1000, Sutter Instruments, USA), we pulled needles from 3.5” glass capillaries (#3-000-203-G/X, Drummond, USA) according to the following settings: heat=512, pull=45, velocity=75, delay=90, pressure=500. Oil-filled needles were mounted to a Nanoject III microinjector (Drummond, USA) and loaded with bacterial suspension. 10 nl of suspension with OD600 = 1 (translating to 6.01 × 103 ± 2.79 × 103 CFUs) was injected into the abdomen of fully melanized O. surinamensis females at a speed of 10 nl/s. Beetles of the control treatment were injected with physiological saline. To facilitate injections, we cut off the posterior third of the beetles’ elytra the day before the injections and immobilized the insects on a cold metal plate during the procedure. Injected females were of undefined age. However, since they were fully melanized, we considered them to be at least 2 weeks old and therefore at a reproductive age72.
After the injections, we transferred each beetle into a fluon-lined well of a 48-well plate (Greiner Bio-One, Germany) filled with oat flakes and incubated the plates at 27–28 °C and 60% rH. Injections were performed in seven batches with 50 beetles per treatment. Two specimens per batch were immediately stored in ethanol for qPCR (see “qPCR” section).
To assess the establishment of Sodalis in O. surinamensis, we inspected live female beetles for mCherry fluorescence using a Leica M165FC fluorescence stereomicroscope (Wetzlar, Germany) 7 days after injections were performed.
Monitoring survival and fitness correlates after Sodalis infection
To investigate the influence of Sodalis on host survival, we monitored the survival of Sodalis-injected and control beetles for a period of 80 days. Beetle survival was monitored every 24 h for the first 3 weeks and twice a week thereafter. 0, 7, and 14 days post injection (dpi), a subset of infected females was processed for additional analyses, see the following sections for more details on the sampling process.
During the observation period, we counted the number of first offspring generation (F1) larvae produced by each injected female as a proxy for fitness. We checked for the presence of larvae on a daily basis for the first 3 weeks, and once a week thereafter. When we found larvae in a well, each larva was carefully moved to a fluon-lined well of a new 48-well plate using a toothpick. Survival and development of isolated larvae were monitored every 2 days, and we switched to daily observations once the larvae neared pupation to be able to assess the exact date of adult emergence. After 28 days, we stopped moving larvae of the control treatment to new wells, since we had already collected enough for the coming sampling. Instead of moving the larvae, we now transferred the female beetles to new plates, leaving the larvae in the wells. We counted larvae on the day of transfer and 10 days after, to account for individuals that might have hatched from eggs after the transfer of the adult female. For both treatments, we let isolated F1 larvae develop into adults, recording the day adults emerged from the pupa. Seven days after eclosion, we fixated F1 adults for further analyses. For more details, see the following sections. For all analyses concerning the F1 offspring of Sodalis-treated beetles, we only included individuals that were confirmed to harbor Sodalis based on mCherry fluorescence observed under a Leica M165FC fluorescence stereomicroscope.
We transferred excess F1 beetles of both treatments that were not needed for further analyses to small plastic boxes, supplied with oats. Offspring of these beetles were transferred to a new box, where they developed into F2 adults and reproduced. F3 offspring larvae were transferred into a 48-well plate, where they developed into the pupal stage and subsequently adulthood. The few surviving Sodalis-infected F3 beetles (five adults and one pupa) were fixated for FISH to investigate the localization of Sodalis and its impact on Shikimatogenerans. Uninfected controls showed much higher survival, so only a subset of the beetles was maintained. To confirm the long-term maintenance of Shikimatogenerans in control beetles, individual beetles from approximately generations 8-10 were fixated for FISH.
Sodalis transmission efficiency experiments
Late instar larvae and pupae were separated from the main O. surinamensis lab colony and reared individually in fluon-lined wells of a 48-well plate supplied with oat flakes until adults emerged. A subset of virgin female and male adults was injected with 10 nl of Sodalis suspended in 0.85% NaCl solution (OD600 = 1.0) and kept in individual wells for 1 week to let beetles recover from the injection and wait for Sodalis to establish. After a week, we checked for the presence of mCherry fluorescence, indicative of Sodalis infection, in the injected beetles. Using infected beetles, two mating groups were set up to assess maternal and paternal Sodalis transmission, respectively. We placed one male and one female beetle in each well of a 24-well plate (Greiner Bio-One, Germany)—Sodalis infected females were placed with uninfected males, and uninfected females were placed in wells with Sodalis infected males. The mating pairs were kept in the wells for 21 days, after which we checked both adult beetles for Sodalis presence (i.e., mCherry fluorescence) to see if bacteria were transmitted horizontally between individuals. At the same time, we separated the produced larvae from the parents and screened for Sodalis by fluorescence stereomicroscopy. Using the number of infected and non-infected larval offspring, we calculated the mean transmission efficiency of Sodalis for the two groups.
Examining symbiont titers via qPCR
To study the bacterial load and its dynamics in O. surinamensis, we sampled beetles of the parental generation and F1 adults. Immediately after injection, 7 and 14 dpi, adults of the parental generation were fixed in 70% ethanol and stored at −20 °C until DNA extraction. For the F1 offspring, we sampled adults 7 days after eclosion. Prior to DNA extraction, samples were placed in individual Eppendorf tubes, and the remaining ethanol was evaporated in a vacuum concentrator (Concentrator 5301, Eppendorf, Germany). We extracted DNA of individual beetles using the Epicentre MasterPure complete DNA and RNA purification kit (Lucigen, USA) according to the manufacturer’s instructions. DNA was eluted in 30 µl of low TE buffer.
For qPCR, we combined 19 µl of mastermix (10 µl Blue S’Green qPCR mix (Biozym, Germany), 7.4 µl nuclease-free water, 0.8 µl of each primer) with 1 µl of template DNA in the wells of a 96-well plate (Bio-Rad, USA). We amplified all samples on a CFX Connect cycler (Bio-Rad, USA) according to the following conditions: initial denaturation for 15 min at 95 °C, 60 cycles 95 °C for 15 s, 60 °C for 30 s and 72 °C for 30 s followed by a final denaturation step at 95 °C for 15 s. Then a melting curve was recorded by incrementing the temperature by 0.5 °C every 5 s from 55 °C up to 95 °C. To allow for absolute quantification of input material, we included a set of standards ranging from 1 to 10−7 ng/µl of amplicon on each plate. Using the standards, the qPCR program (CFX Manager v3.1, Bio-Rad, USA) assessed the starting quantities of the respective fragment in our samples, from which we calculated the corresponding copy number. To amplify the 16S gene of Shikimatogenerans we used the OsurSym_fwd2 (5′-GGCAACTCTGAACTAGCTACGC-3′) and mod. CFB563_rev (5′-GCACCCTTTAAACCCAAT-3′) primers6. For Sodalis, we amplified part of the tam gene using the primers SP_tam3_fwd (5′-CGTCTTAGCGGTACAAATGCC-3′) and SP_tam3_reverse (5′-GCAGATCGTAAT-AGTCGGCG-3′) modified from Medina Munoz et al.73.
Melanization
For the F1 offspring, we analyzed cuticle coloration as a measurement of melanization that may reflect tyrosine availability to the host6. We collected F1 adults 7 days after eclosion and immobilized them on a cold metal plate. Using a Leica M165 FC stereomicroscope (Leica, Germany), we obtained pictures of the beetles with standardized settings and illumination conditions (2× magnification, 293,5 ms exposure time, 50 gain, 6.2 MP resolution) that we used to analyze the color of beetle thoraces and abdomens. Pictures were imported into the Natsumushi software v1.1074, and average red values were extracted from circular areas for both thorax (radius = 326.7 pixels) and abdomen (radius = 388.9 pixels). Average red values were transformed to inverse red values and compared between treatments. Specimens photographed for the assessment of melanization were then fixated for cuticle thickness measurements.
Cuticle thickness
Besides melanization, we also examined the cuticle thickness of F1 adults as a proxy of tyrosine availability. Cuticle thickness was assessed using micro-computed tomography (µCT) scans of adult F1 beetles. To this end, we fixated 7 d old adults in 4% paraformaldehyde (PFA) in 80% ethanol and removed their heads to facilitate entry of the fixative into the body cavity. Samples were stored in fixative at 4 °C until we prepared them for µCT according to Janke et al.75. Briefly, we transferred beetles to absolute methanol to wash out the fixative and incubated the beetles for 24 h. Afterwards, beetles were moved to 1% iodine in absolute methanol and incubated at room temperature for another 24 h to enhance the contrast of internal tissues. Beetle specimens were then washed in absolute ethanol (denatured) for 1 h on a shaker (Unimax1010, Heidolph, Germany) to remove excess iodine. The washing step was repeated twice, followed by 3 washing steps in 100% absolute ethanol on a shaker for 1 h each. Washed beetles were dried in a critical point drier (Leica EM CPD300, Leica, Germany) with medium CO2 supply with a delay of 20 min, 20 exchange cycles, heating at medium speed and slow gas exhaust. Dried specimens were mounted in translucent 10 µl pipette tips with pipette tip pieces as spacers in between samples.
Pipette tips with samples were placed on the sample holder and scanned in an X-ray microtomograph (Skyscan 1272, Bruker, USA). We created 360° scans with 0.2° rotation steps, the voltage and electric current were adjusted to 35–40 kV and 135–200 mA, respectively, to achieve a signal attenuation of 35%. Final pixel size of the scans was 2 µm with a resolution of 2452 × 1640.
Based on the scans, we performed 3D reconstruction of the scanned specimens using the NRecon software (v2.1.0.1, Bruker, USA) with the following settings: ring artefacts reduction = 10 and beam hardening correction = 20%. 3D reconstructions were imported into Dragonfly76 (v2022.2, Comet Technologies Canada Inc., Montreal, Canada; software available at https://www.theobjects.com/dragonfly) where we measured beetle cuticle thickness in two ways: First, we measured mean cuticle thickness across the whole insect’s body (excluding the head, which had been removed to facilitate fixation), by extracting the cuticle from the 3D reconstruction and applying a thickness mesh. The mesh was then used to extract mean cuticle thickness. Additionally, we measured the mean cuticle thickness in a single section of the thorax, between the 3rd and 4th lateral spike. Thickness was measured in a dorsal, ventral and sagittal position of the section and the individual measurements were averaged.
Symbiont localization via fluorescence-in-situ-hybridization (FISH)
To examine localization of Sodalis in the beetles, we fixated adult beetles of the parental generation 7 dpi and of the F1 generation 7 days after eclosion in 4% PFA in 80% butanol for FISH. Sample preparation was conducted according to Weiss77: The extremities of fixated beetles were removed to facilitate fixation of the remaining tissues, and samples were pre-embedded in 1% agar blocks. Beetle specimens were then washed in 80% tertiary butanol and subsequently dehydrated in a series of butanols with ascending concentrations (90%, 96% and 100%). After a final dehydration step in acetone, samples were infiltrated with, and finally embedded in, HistoCURE 8100 (Morphisto, Germany) according to the manufacturer’s instructions. Embedded beetles were cut into 8 µm thin sagittal sections with a Leica Histocore AUTOCUT microtome equipped with glass knives. Sections were placed on silanized glass slides that were covered with 100 µl of hybridization mix consisting of hybridization buffer (0.9 M NaCl, 0.02 M Tris/HCl, pH 8.0, 0.01% SDS), 0.5 µM fluorescently labeled oligonucleotide probes and 0.5 mg/ml 4′,6-diamidino-2-phenylindole (DAPI). As probes, we used one probe specific for Shikimatogenerans (OsurSym_16S_Cy5, 5′-AGCAGCCCATTGGACTAACC-3′), one specific for Sodalis (Sod_FISH-Cy3, 5′-TCCGCTGACTCTCGCGAGAT-3′; Shalom et al.78), and a general probe targeting eubacteria (EUB338-RhodamineGreen, 5′-GCTGCCTCCCGTAGGAGT-3′; Amann et al.79). We applied cover glasses and incubated the slides in a humid chamber at 50 °C overnight. The next day, we removed the cover glasses, transferred the slides to wash buffer (0.1 M NaCl, 0.02 Tris/HCl, 5 mM EDTA, 0.01% SDS) and incubated them at 50 °C for 2 h. Afterwards, we washed the slides in distilled water for 20 min and mounted them with Vectashield (VectorLabs, USA). Sections were imaged with a Leica THUNDER imager DMi8 equipped with a DFC9000GT camera and CYR71010 and DFT51010 filter cubes. Pictures were processed with small volume computational clearing in the Leica Application Suite X software (Version 3.8.2.27713, Leica, Germany). Besides beetles of the parental and F1 generations, we also performed FISH on individuals of F3 and the following generations.
Measuring the impact of Sodalis on Shikimatogenerans cell length and bacteriome volume
In each beetle sample used for FISH, we assessed the effect of Sodalis on the cell size of Shikimatogenerans. To provide a conservative estimate of this effect, we pre-selected Shikimatogenerans cells that resembled the shape of the symbiont in the control treatment the most. Among these selected cells, we measured the length of 10 random Shikimatogenerans cells in one picture of a random bacteriome. The size of 10 cells was averaged per sample and then used to compare cell sizes between the two treatments. Besides Shikimatogenerans cell length, we also investigated the impact of Sodalis on bacteriome volume. Using Dragonfly (v2022.2), we marked the bacteriomes in each section as a region of interest (ROI) and interpolated between the sections to obtain the entire bacteriome volume. We manually corrected ROIs to only cover the bacteriomes and none of the surrounding tissues. For each of the bacteriomes, we extracted the number of voxels each ROI covered and calculated the total volume of each bacteriome accordingly (1 voxel = pixelsize3).
Gene expression in the bacteriome: RNASeq analysis of O. surinamensis, Shikimatogenerans and Sodalis
Sample collection, RNA extracting, and sequencing
To investigate gene expression of Shikimatogenerans, Sodalis and O. surinamensis, we dissected bacteriomes of adults of the parental generation 7 days after injection, and of F1 adults 7 days after eclosion. Both control and Sodalis infected beetles were dissected in sterile 1.5× phosphate-buffered saline, and bacteriomes were transferred into a drop of Trizol in the lid of a reaction tube. Per treatment group, we dissected 7 replicates of pooled bacteriomes, making sure that for each pool, at least 15 bacteriomes were collected (out of 20 possible for the five beetles, since there are four bacteriomes in each individual beetle). For the parental generation, we pooled the bacteriomes of 5 beetles in 100 µl of Trizol and added 150 µl Trizol before RNA extraction, while for the F1, we collected bacteriomes of individual beetles in 50 µl of Trizol and pooled them just before extracting RNA. Bacteriomes in Trizol were manually homogenized using tweezers and collected at the bottom of the tube by spinning down the Trizol. Bacteriomes were stored at −80 °C until RNA extractions. Tweezers were cleaned with 70% ethanol in between dissections, and the buffer was changed between samples. RNA was extracted using the Directzol RNA Microprep kit (Zymo Research, USA) according to the manufacturer’s instructions. We eluted RNA in 15 µl of nuclease-free water and stored it at −80 °C until sequencing. Following the extraction, RNA quality and quantity were assessed by capillary electrophoresis using an Agilent Bioanalyzer PicoChip (Agilent, USA). A total RNA library was prepared with a Zymo-Seq RiboFree Total RNA Library kit (Zymo Research, USA) as outlined in the manufacturer’s instructions. Total RNA inputs ranged from 7 to 15 ng and library preparation was adjusted according to the protocol recommendations for low input amounts. Library fragments passing the universal depletion step were then amplified by PCR to convert single-stranded cDNAs into double-stranded cDNAs. We sequenced 2 × 150 bp paired reads on an Illumina NextSeq 2000 sequencer with a depth of approximately 50,000,000 reads per library. RNA quality control, rRNA depletion, library preparation and sequencing were performed at the Max Planck Genome Center in Cologne.
Read processing and mapping
For the analysis of Sodalis and Shikimatogenerans gene expression, all reads were trimmed using Trim Galore v0.6.1080 for a Phred quality score threshold of 30 and minimum read length of 50 bp. Potential remaining rRNA reads were removed using Sortmerna v4.3.681 against the default database smr_v4.3_default_db. Of the on average 93.7 million total reads, an average of 82.8 million reads were retained after trimming and rRNA removal and used in the subsequent analysis. For the host gene expression analysis, RNASeq reads were trimmed, mapped and counted against the O. surinamensis genome using the NF-CORE/RNASeq pipeline v3.1882 with the trimgalore, remove_ribo_rna and star_salmon options.
The published genomes of S. silvanidophilus (GCF_018200315.1) and S. praecaptivus (GCA_000517425.1) were obtained, and genes were predicted and annotated using Bakta v1.9.383 with default parameters. The translated CDSs were then used to obtain KEGG functional annotations84 using KEGGREST v1.40.185 in R v4.3.386. Reads were mapped using Bowtie2 v2.4.187 from end to end (sensitive mode), allowing for no discordant or mixed reads. The BAM files were sorted and indexed using Samtools version 1.1388. On average, 43.9% of reads mapped to the genome of Shikimatogenerans, and 0.37% of reads mapped to the genome of Sodalis (averaged across infected samples, control samples had 0.00% mapping). The aligned reads were counted with FADU v1.8.389 using the GFF files from the above-mentioned genomes for gene features.
The O. surinamensis genome was assembled using previously published single-molecule long reads (SRR12881567–SRR12881568) obtained by sequencing genomic DNA on an Oxford Nanopore (ONT) MinION Mk1B sequencer. The ONT raw data were base-called using GUPPY v4.0.1190 with a high-accuracy option (dna_r9.4.1_450bps_hac.cfg model), generating 11.7 Gb of sequence reads with an N50 of 24.5 kb. Genome assembly was performed using Flye v2.9.291 in meta-mode, followed by four iterations of polishing using Racon v1.4.1392 and one round of error correction using Medaka v1.0.3 (https://nanoporetech.github.io/medaka). For final error correction, we used ntEdit v1.3.293 with paired-end (2 × 150 bp) Illumina data generated from the same DNA used for ONT sequencing. PurgeDups was used to remove duplications (heterozygous regions) and generate a haploid genome for downstream analysis. The relative contig coverage, GC content and contig taxonomic classification were scanned after genome assembly using BlobTools v1.0 and Taxon Analysis94 to enable the identification and removal of potential microbial symbionts or contaminants. The completeness of the genome was assessed using BUSCO v5.2.195, utilizing the insect_odb10 database. The assembled haploid genome contained 1770 contigs with a total genome size of 142 Mb and an N50 contig size of 1,5 Mb. A genome completeness assessment using BUSCO resulted in 99.3% BUSCO complete and single-copy orthologs and low duplication levels (0.4% duplicated BUSCO). Genome annotation of O. surinamensis was performed using a combination of RNASeq data obtained from beetle bacteriomes (RNA extraction with Zymo Direct-zol RNA Miniprep kit, rRNA depletion and poly(A)-enrichment, 2 × 150 bp paired-end sequencing with a depth of 25,000,000 reads on a NextSeq 2000 device) and the reference protein database Arthropoda ODB10. Briefly, reads were trimmed using TrimGalore v0.6.1080 and read length and quality thresholds set as 50 bp and 30, respectively. Reads were then mapped against the O. surinamensis genome using HiSAT2 (v2.2.1) with default parameters except for the intron length which was set to 10 kb and the “dta” option. Mapped samples were then merged, and the resulting file was processed with samtools v1.1788 using the subcommands “collate”, “fixmate”, “sort” and “markdup” with default parameters. The final BAM file, along with the protein reference database, was then provided to BRAKER v3.0.796,97,98,99,100,101,102,103,104 to perform gene model prediction. BRAKER predictions were post-processed to fix overlapping genes and retain only the longest isoform using AGAT v1.5.1105. Prior to annotation, the genome of O. surinamensis was softmasked using the RepeatMasker suite. A first round of masking using RepeatMasker v4.1.5106 was performed based on the insect lineage and the default database of RepeatMasker. In parallel, de novo prediction of repeats was performed using RepeatModeler v2.0.4107 with default parameters. A second round of softmasking was then performed using the custom repeats library produced by RepeatModeler.
Finally, nucleotide coding sequences and proteins were extracted with GFFREAD v0.12.8108. Functional information was obtained EggNOG (eggNOG-Mapper 2.1.0 with EggNOG 5.0.2, Huerta-Cepas et al.109) within OmicsBox v3.4.1 and InterProScan110.
Differential gene expression analysis
Gene expression was compared between control and Sodalis-infected beetles for each generation, except for Sodalis, in which gene expression was compared across generations for the Sodalis-infected treatments. For the host gene expression analysis, counting matrices were then subjected to differential expression analysis using EdgeR v4.2.2111 in R v4.4.0 using a quasi-likelihood test and an FDR threshold of 0.05 and a log2FoldChange of ≥|1|. Prior to differential expression estimation, genes with low variation were removed through the filterByExpr function of the EdgeR package. Out of the total 13,283 genes, 7811 and 7689 were kept in the analysis of the parental and first generations, respectively. A total of 150 and 46 genes were identified as differentially expressed in the parental and first generations, respectively.
For the bacterial gene expression analysis, differential gene expression was assessed using DESeq2 v1.40.2112 in R version 4.3.3113. Genes with fewer than a total of 20 reads across samples were excluded from the analysis. For Shikimatogenerans, 276 protein-coding genes were analyzed, whereas for Sodalis 4362 out of 4657 total genes were kept. For all differential gene expression analyses, the function lfcshrink was run using the adaptive Student’s t prior shrinkage estimator from the package apeglm v1.22.1114. Genes were considered differentially expressed if the adjusted p-value was ≤0.05 and the log2FoldChange was ≥|1|. No genes were identified as differentially expressed for Shikimatogenerans. Conversely, Sodalis differentially expressed 168 genes across generations. Additionally, Sodalis genes were sorted according to the average TPM values obtained from the FADU outputs for each generation, and the top 100 genes were selected out of the 2946 protein-coding genes annotated with BlastKoala v3.0.
Statistics
We analyzed the survival of injected beetles with cox mixed-effect models v2.2.20 (coxme package115). Our models included treatment as an explanatory variable while the injection batch was included as a random factor. An effect of treatment was tested using a type II analysis of variance (ANOVA, car package v3.1.2116). The number of larvae produced by injected females was analyzed with a zero-inflated negative binomial model (pscl package v1.5.9117) using treatment as an explanatory variable. Bacteriome absence and presence in F3 individuals was compared between the two treatments using a Fisher’s exact test (stats package v4.2.386). Remaining fitness parameters were compared between the two treatments using two-sided parametric two sample t-tests or non-parametric Wilcoxon rank sum tests (stats package v4.2.386). Assumptions of normality and homogeneity of variance were verified using the Shapiro–Wilk test and Levene’s test, respectively (stats package v4.2.386). Figures for survival and fitness parameters were created with ggplot2 v3.5.1118.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The annotated beetle and symbiont genomes and RNAseq data are available in the Edmond open data repository of the Max Planck Society under https://doi.org/10.17617/3.MUV1MF. RNAseq data generated in this study has also been deposited in the NCBI Sequence Read Archive under accession number PRJNA1423255 [http://www.ncbi.nlm.nih.gov/bioproject/1423255]. The bacterial genomes used in this study were obtained from NCBI using accession numbers GCF_018200315.1 [https://www.ncbi.nlm.nih.gov/datasets/genome/GCF_018200315.1/] (Shikimatogenerans silvanidophilus) and GCA_000517425.1 [https://www.ncbi.nlm.nih.gov/datasets/genome/GCF_000517425.1/] (Sodalis praecaptivus). Single-molecule long reads for assembling the Oryzaephilus surinamensis genome were obtained from the NCBI Sequence Read Archive under accession numbers [SRR12881567]-[SRR12881568]. RNAseq data used for annotation of the Oryzaephilus surinamensis genome is available in the NCBI Sequence Read Archive under accession number PRJNA1423469 [http://www.ncbi.nlm.nih.gov/bioproject/1423469]. Source data are provided with this paper.
References
Moran, N. A. Symbiosis as an adaptive process and source of phenotypic complexity. Proc. Natl. Acad. Sci. USA. 104, 8627–8633 (2007).
Kaltenpoth, M., Flórez, L. V., Vigneron, A., Dirksen, P. & Engl, T. Origin and function of beneficial bacterial symbioses in insects. Nat. Rev. Microbiol. 23, 551–567 (2025).
Munson, M. A. et al. Evidence for the establishment of aphid eubacterium endosymbiosis in an ancestor of four aphid families. J. Bacteriol. 173, 6321–6324 (1991).
Clark, M. A., Moran, N. A. & Baumann, P. Sequence evolution in bacterial endosymbionts having extreme base compositions. Mol. Biol. Evol. 16, 1586–1598 (1999).
Moran, N. A., Tran, P. & Gerardo, N. M. Symbiosis and insect diversification: an ancient symbiont of sap-feeding insects from the bacterial phylum Bacteroidetes. Appl Environ. Microbiol. 71, 8802–8810 (2005).
Engl, T. et al. Ancient symbiosis confers desiccation resistance to stored grain pest beetles. Mol. Ecol. 27, 2095–2108 (2018).
Moran, N. A., McLaughlin, H. J. & Sorek, R. Dynamics and time scale of ongoing genomic erosion in symbiotic bacteria. Science. 323, 379–382 (2009).
Luan, J. B. et al. Metabolic coevolution in the bacterial symbiosis of whiteflies and related plant sap-feeding insects. Genome Biol. Evol. 7, 2635–2647 (2015).
Wierz, J. C., Gimmel, M. L., Huthmacher, S., Engl, T. & Kaltenpoth, M. Evolutionary history of tyrosine-supplementing endosymbionts in pollen-feeding beetles. ISME J. 18, wrae080 (2024).
Kirsch, R. et al. Symbiosis and horizontal gene transfer promote herbivory in the megadiverse leaf beetles. Curr. Biol. 35, 640–654 (2025).
Koga, R., Bennett, G. M., Cryan, J. R. & Moran, N. A. Evolutionary replacement of obligate symbionts in an ancient and diverse insect lineage. Environ. Microbiol. 15, 2073–2081 (2013).
Bennett, G. M. & Moran, N. A. Heritable symbiosis: The advantages and perils of an evolutionary rabbit hole. Proc. Natl. Acad. Sci. USA. 112, 10169–10176 (2015).
Husnik, F. & McCutcheon, J. P. Repeated replacement of an intrabacterial symbiont in the tripartite nested mealybug symbiosis. Proc. Natl. Acad. Sci. USA. 113, E5416–5424 (2016).
Sudakaran, S., Kost, C. & Kaltenpoth, M. Symbiont acquisition and replacement as a source of ecological innovation. Trends Microbiol. 25, 375–390 (2017).
Cornwallis, C. K. et al. Symbioses shape feeding niches and diversification across insects. Nat. Ecol. Evol. 7, 1022–1044 (2023).
Boscaro, V. et al. Symbiont replacement between bacteria of different classes reveals additional layers of complexity in the evolution of symbiosis in the ciliate Euplotes. Protist. 169, 43–52 (2018).
Vannini, C., Sigona, C., Hahn, M., Petroni, G. & Fujishima, M. High degree of specificity in the association between symbiotic betaproteobacteria and the host Euplotes (Ciliophora, Euplotia). Eur. J. Protistol. 59, 124–132 (2017).
Fujishima, M. & Heckmann, K. Intra- and interspecies transfer of endosymbionts in Euplotes. J. Exp. Zool. 230, 339–345 (1984).
Su, Y. et al. Rational engineering of a synthetic insect-bacterial mutualism. Curr. Biol. 32, 3925–3938 (2022).
Grimaldi, D. & Engel, M. S. Evolution of the Insects (Cambridge University Press, 2005).
Linz, D. M., Hu, A. W., Sitvarin, M. I. & Tomoyasu, Y. Functional value of elytra under various stresses in the red flour beetle, Tribolium castaneum. Sci. Rep. 6, 34813 (2016).
Andersen, S. O. in Comprehensive Molecular Insect Science (ed L. I. Gilbert) 145–170 (Elsevier, 2005).
Andersen, S. O. Insect cuticular sclerotization: a review. Insect Biochem. Mol. Biol. 40, 166–178 (2010).
Duplais, C. et al. Gut bacteria are essential for normal cuticle development in herbivorous turtle ants. Nat. Commun. 12, 676 (2021).
Kanyile, S. N., Engl, T., Heddi, A. & Kaltenpoth, M. Endosymbiosis allows Sitophilus oryzae to persist in dry conditions. Front. Microbiol. 14, 1199370 (2023).
Kanyile, S. N., Engl, T. & Kaltenpoth, M. Nutritional symbionts enhance structural defence against predation and fungal infection in a grain pest beetle. J. Exp. Biol. 225, jeb243593 (2022).
Chari, A. et al. Phenotypic characterization of Sodalis praecaptivus sp. nov., a close non-insect-associated member of the Sodalis-allied lineage of insect endosymbionts. Int. J. Syst. Evol. Microbiol. 65, 1400–1405 (2015).
Koch, A. Die Symbiose von Oryzaephilus surinamensis L. (Cucujidae, Coleoptera). Z. für. Morphologie und Ökologie der Tiere. 23, 389–424 (1931).
Kiefer, J. S. T. et al. Inhibition of a nutritional endosymbiont by glyphosate abolishes mutualistic benefit on cuticle synthesis in Oryzaephilus surinamensis. Commun. Biol. 4, 554 (2021).
Ewald, P. W. Transmission modes and evolution of the parasitism-mutualism continuum. Ann. N. Y. Acad. Sci. 503, 295–306 (1987).
Frank, S. A. Models of symbiosis. Am. Nat. 150, S80–99 (1997).
Matsuura, Y. et al. Recurrent symbiont recruitment from fungal parasites in cicadas. Proc. Natl. Acad. Sci. USA. 115, E5970–E5979 (2018).
Chong, R. A. & Moran, N. A. Evolutionary loss and replacement of Buchnera, the obligate endosymbiont of aphids. ISME J. 12, 898–908 (2018).
Lefevre, C. et al. Endosymbiont phylogenesis in the Dryophthoridae weevils: evidence for bacterial replacement. Mol. Biol. Evol. 21, 965–973 (2004).
Hurst, G. D., Anbutsu, H., Kutsukake, M. & Fukatsu, T. Hidden from the host: Spiroplasma bacteria infecting Drosophila do not cause an immune response, but are suppressed by ectopic immune activation. Insect Mol. Biol. 12, 93–97 (2003).
Anbutsu, H. & Fukatsu, T. Evasion, suppression and tolerance of Drosophila innate immunity by a male-killing Spiroplasma endosymbiont. Insect Mol. Biol. 19, 481–488 (2010).
Zhang, Q., Wei, X., Fang, W., Huang, X. & Zhang, X. The secretory protein COA1 enables Metarhizium robertsii to evade insect immune recognition during cuticle penetration. Commun. Biol. 7, 1220 (2024).
Hu, Y. & Aksoy, S. An antimicrobial peptide with trypanocidal activity characterized from Glossina morsitans morsitans. Insect Biochem. Mol. Biol. 35, 105–115 (2005).
Pontes, M. H., Smith, K. L., De Vooght, L., Van Den Abbeele, J. & Dale, C. Attenuation of the sensing capabilities of PhoQ in transition to obligate insect-bacterial association. PLoS Genet. 7, e1002349 (2011).
Clayton, A. L., Enomoto, S., Su, Y. & Dale, C. The regulation of antimicrobial peptide resistance in the transition to insect symbiosis. Mol. Microbiol. 103, 958–972 (2017).
Anselme, C., Vallier, A., Balmand, S., Fauvarque, M. O. & Heddi, A. Host PGRP gene expression and bacterial release in endosymbiosis of the weevil Sitophilus zeamais. Appl. Environ. Microbiol. 72, 6766–6772 (2006).
Maire, J. et al. Weevil pgrp-lb prevents endosymbiont TCT dissemination and chronic host systemic immune activation. Proc. Natl. Acad. Sci. USA. 116, 5623–5632 (2019).
Bright, M. & Bulgheresi, S. A complex journey: transmission of microbial symbionts. Nat. Rev. Microbiol. 8, 218–230 (2010).
Rio, R. V. et al. Insight into the transmission biology and species-specific functional capabilities of tsetse (Diptera: Glossinidae) obligate symbiont Wigglesworthia. mBio. 3, e00240–11 (2012).
Santos-Garcia, D. et al. The genome of Cardinium cBtQ1 provides insights into genome reduction, symbiont motility, and its settlement in Bemisia tabaci. Genome Biol. Evol. 6, 1013–1030 (2014).
Lee, J. B. et al. Bacterial cell motility of Burkholderia gut symbiont is required to colonize the insect gut. FEBS Lett. 589, 2784–2790 (2015).
Dale, C., Young, S. A., Haydon, D. T. & Welburn, S. C. The insect endosymbiont Sodalis glossinidius utilizes a ype III secretion system for cell invasion. PNAS. 98, 1883–1888 (2000).
Deng, W. et al. Assembly, structure, function and regulation of type III secretion systems. Nat. Rev. Microbiol. 15, 323–337 (2017).
Teulet, A., Camuel, A., Perret, X. & Giraud, E. The versatile roles of type III secretion systems in rhizobium-legume symbioses. Annu. Rev. Microbiol. 76, 45–65 (2022).
Su, Y., Lin, H. C. & Dale, C. Protocol to establish a genetically tractable synthetic symbiosis between Sodalis praecaptivus and grain weevils by insect egg microinjection. STAR Protoc. 4, 102156 (2023).
Ankrah, N. Y. D., Luan, J. & Douglas, A. E. Cooperative metabolism in a three-partner insect-bacterial symbiosis revealed by metabolic modeling. J. Bacteriol. 199, e00872–16 (2017).
Goto, S., Anbutsu, H. & Fukatsu, T. Asymmetrical interactions between Wolbachia and Spiroplasma endosymbionts coexisting in the same insect host. Appl. Environ. Microbiol. 72, 4805–4810 (2006).
Koga, R., Tsuchida, T. & Fukatsu, T. Changing partners in an obligate symbiosis: a facultative endosymbiont can compensate for loss of the essential endosymbiont Buchnera in an aphid. Proc. Biol. Sci. 270, 2543–2550 (2003).
Huang, Y., Feng, Z. F., Li, F. & Hou, Y. M. Host-encoded aminotransferase import into the endosymbiotic bacteria Nardonellaof Red Palm Weevil. Insects. 15, 35 (2024).
Halary, S., Duperron, S. & Boudier, T. Direct image-based correlative microscopy technique for coupling identification and structural investigation of bacterial symbionts associated with metazoans. Appl. Environ. Microbiol. 77, 4172–4179 (2011).
Vigneron, A. et al. Insects recycle endosymbionts when the benefit is over. Curr. Biol. 24, 2267–2273 (2014).
Carvalho, A. S. P. et al. Symbionts with eroded genomes adjust gene expression according to host life-stage and environment. EMBO Rep. 26, 4656–4674 (2025).
Cabeen, M. T. & Jacobs-Wagner, C. Bacterial cell shape. Nat. Rev. Microbiol. 3, 601–610 (2005).
McCutcheon, J. P., Boyd, B. M. & Dale, C. The life of an insect endosymbiont from the cradle to the grave. Curr. Biol. 29, R485–R495 (2019).
Florez, L. V., Biedermann, P. H., Engl, T. & Kaltenpoth, M. Defensive symbioses of animals with prokaryotic and eukaryotic microorganisms. Nat. Prod. Rep. 32, 904–936 (2015).
Engl, T. et al. Evolutionary stability of antibiotic protection in a defensive symbiosis. Proc. Natl. Acad. Sci. USA. 115, E2020–E2029 (2018).
Oliver, K. M., Degnan, P. H., Hunter, M. S. & Moran, N. A. Bacteriophages encode factors required for protection in a symbiotic mutualism. Science. 325, 992–994 (2009).
Scarborough, C. L., Ferrari, J. & Godfray, H. C. J. Aphid protected from pathogen by endosymbiont. Science. 310, 1781 (2005).
Pande, S. et al. Fitness and stability of obligate cross-feeding interactions that emerge upon gene loss in bacteria. ISME J. 8, 953–962 (2014).
Scarinci, G. et al. Enhanced metabolic entanglement emerges during the evolution of an interkingdom microbial community. Nat. Commun. 15, 7238 (2024).
Koga, R. et al. Single mutation makes Escherichia coli an insect mutualist. Nat. Microbiol. 7, 1141–1150 (2022).
Rogowska-van der Molen, M. A. et al. From eggs to guts: symbiotic association of Sodalis nezarae sp. nov. with the southern green shield bug Nezara viridula. FEMS Microbiol. Ecol. 101, fiaf017 (2025).
Santos-Garcia, D., Silva, F. J., Morin, S., Dettner, K. & Kuechler, S. M. The All-Rounder Sodalis: a new bacteriome-associated endosymbiont of the lygaeoid bug Henestaris halophilus (Heteroptera: Henestarinae) and a critical examination of its evolution. Genome Biol. Evol. 9, 2893–2910 (2017).
Boyd, B. M. et al. Two bacterial genera, Sodalis and Rickettsia, associated with the seal louse Proechinophthirus fluctus (Phthiraptera: Anoplura). Appl. Environ. Microbiol. 82, 3185–3197 (2016).
Fukatsu, T. et al. Bacterial endosymbiont of the slender pigeon louse, Columbicola columbae, allied to endosymbionts of grain weevils and tsetse flies. Appl Environ. Microbiol. 73, 6660–6668 (2007).
Dale, C. & Maudlin, I. Sodalis gen. nov. and Sodalis glossinidius sp. nov., a microaerophilic secondary endosymbiont of the tsetse fly Glossina morsitans morsitans. Int. J. Syst. Evolut. Microbiol. 49, 267–275 (1999).
Engl, T., Schmidt, T. H. P., Kanyile, S. N. & Klebsch, D. Metabolic cost of a nutritional symbiont manifests in delayed reproduction in a grain pest beetle. Insects. 11, 717 (2020).
Medina Munoz, M., Spencer, N., Enomoto, S., Dale, C. & Rio, R. V. M. Quorum sensing sets the stage for the establishment and vertical transmission of Sodalis praecaptivus in tsetse flies. PLoS Genet. 16, e1008992 (2020).
Tanahashi, M. & Fukatsu, T. Natsumushi: image measuring software for entomological studies. Entomol. Sci. 21, 347–360 (2018).
Janke, R. S., Moog, S., Weiss, B., Kaltenpoth, M. & Florez, L. V. Morphological adaptation for ectosymbiont maintenance and transmission during metamorphosis in Lagria beetles. Front. Physiol. 13, 979200 (2022).
Comet Technologies Canada Inc., Montreal Canada, Dragonfly V2022.2.
Weiss, B. Techniques of Insect Histology. A Guideline for the Preparation of Insects for Light Microscopic Analysis (Shaker Verlag, 2023).
Shalom, S. R., Weiss, B., Lalzar, M., Kaltenpoth, M. & Chiel, E. Abundance and localization of symbiotic bacterial communities in the fly parasitoid Spalangia cameroni. Appl. Environ. Microbiol. 88, e02549–02521 (2022).
Amann, R. I. et al. Combination of 16S rRNA-targeted oligonucleotide probes with flow cytometry for analyzing mixed microbial populations. Appl. Environ. Microbiol. 56, 1919–1925 (1990).
FelixKrueger/TrimGalore: v0.6.10 - add default decompression path (0.6.10) (Zenodo, 2023).
Kopylova, E., Noe, L. & Touzet, H. SortMeRNA: fast and accurate filtering of ribosomal RNAs in metatranscriptomic data. Bioinformatics. 28, 3211–3217 (2012).
Ewels, P. A. et al. The nf-core framework for community-curated bioinformatics pipelines. Nat. Biotechnol. 38, 276–278 (2020).
Schwengers, O. et al. Bakta: rapid and standardized annotation of bacterial genomes via alignment-free sequence identification. Microbial Genomics. 7, 000685 (2021).
Kanehisa, M. G. S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27–30 (2000).
Kanehisa, M., Sato, Y. & Morishima, K. BlastKOALA and GhostKOALA: KEGG tools for functional characterization of genome and metagenome sequences. J. Mol. Biol. 428, 726–731 (2016).
R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, Austria, 2024).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods. 9, 357–359 (2012).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience. 10, giab008 (2021).
Chung, M. A. et al. FADU: a quantification tool for prokaryotic transcriptomic analyses. mSystems. 6, e00917–00920 (2021).
Wick, R. R., Judd, L. M. & Holt, K. E. Performance of neural network basecalling tools for Oxford Nanopore sequencing. Genome Biol. 20, 129 (2019).
Kolmogorov, M. et al. metaFlye: scalable long-read metagenome assembly using repeat graphs. Nat. Methods. 17, 1103–1110 (2020).
Vaser, R., Sovic, I., Nagarajan, N. & Sikic, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res. 27, 737–746 (2017).
Warren, R. L. et al. ntEdit: scalable genome sequence polishing. Bioinformatics. 35, 4430–4432 (2019).
Laetsch, D. R. & Blaxter, M. L. BlobTools: interrogation of genome assemblies. F1000Research. 6, 1287 (2017).
Manni, M., Berkeley, M. R., Seppey, M., Simao, F. A. & Zdobnov, E. M. BUSCO update: novel and streamlined workflows along with broader and deeper phylogenetic coverage for scoring of eukaryotic, prokaryotic, and viral genomes. Mol. Biol. Evol. 38, 4647–4654 (2021).
Stanke, M., Diekhans, M., Baertsch, R. & Haussler, D. Using native and syntenically mapped cDNA alignments to improve de novo gene finding. Bioinformatics. 24, 637–644 (2008).
Stanke, M., Schoffmann, O., Morgenstern, B. & Waack, S. Gene prediction in eukaryotes with a generalized hidden Markov model that uses hints from external sources. BMC Bioinform. 7, 62 (2006).
Gabriel, L., Hoff, K. J., Bruna, T., Borodovsky, M. & Stanke, M. TSEBRA: transcript selector for BRAKER. BMC Bioinform. 22, 566 (2021).
Bruna, T., Lomsadze, A. & Borodovsky, M. GeneMark-EP+: eukaryotic gene prediction with self-training in the space of genes and proteins. NAR Genom. Bioinform. 2, lqaa026 (2020).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods. 12, 59–60 (2015).
Lomsadze, A., Ter-Hovhannisyan, V., Chernoff, Y. O. & Borodovsky, M. Gene identification in novel eukaryotic genomes by self-training algorithm. Nucleic Acids Res. 33, 6494–6506 (2005).
Iwata, H. & Gotoh, O. Benchmarking spliced alignment programs including Spaln2, an extended version of Spaln that incorporates additional species-specific features. Nucleic Acids Res. 40, e161–e161 (2012).
Gotoh, O., Morita, M. & Nelson, D. R. Assessment and refinement of eukaryotic gene structure prediction with gene-structure-aware multiple protein sequence alignment. BMC Bioinform. 15, 189 (2014).
Gabriel, L. et al. BRAKER3: fully automated genome annotation using RNA-seq and protein evidence with GeneMark-ETP, AUGUSTUS, and TSEBRA. Genome Res. 34, 769–777 (2024).
NBISweden/AGAT: AGAT v. v1.5.1 (Zenodo, 2025).
Tarailo-Graovac, M. & Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Curr. Protoc. Bioinform. 4, 4 10 11–14 10 14 (2009).
Flynn, J. M. et al. RepeatModeler2 for automated genomic discovery of transposable element families. Proc. Natl. Acad. Sci. USA. 117, 9451–9457 (2020).
Pertea, G. & Pertea, M. GFF Utilities: GffRead and GffCompare. F1000Res. 9, 304 (2020).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 47, D309–D314 (2019).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics. 30, 1236–1240 (2014).
Chen, Y., Chen, L., Lun, A. T. L., Baldoni, P. L. & Smyth, G. K. edgeR v4: powerful differential analysis of sequencing data with expanded functionality and improved support for small counts and larger datasets. Nucleic Acids Res. 53, gkaf018 (2025).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Ripley, B. D. Spatial statistics in R. R. N. 1, 14–15 (2001).
Zhu, A., Ibrahim, J. G. & Love, M. I. Heavy-tailed prior distributions for sequence count data: removing the noise and preserving large differences. Bioinformatics. 35, 2084–2092 (2019).
Therneau, T. M. coxme: mixed effects Cox models https://CRAN.R-project.org/package=coxme (The Comprehensive R Archive Network, 2024).
Fox, J. & Weisberg, S. An R Companion to Applied Regression (Sage, Thousand Oaks CA, 2019).
Zeileis, A., Kleiber, C. & Jackman, S. Regression models for count data in R. J. Stat. Softw. are. 27, 1–25 (2008).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer-Verlag New York, 2016).
Acknowledgements
We thank Freya Weituschat for assistance with beetle rearing and Bruno Hüttel and the Max Planck-Genome-Centre Cologne (http://mpgc.mpipz.mpg.de/home/) for performing library preparation and sequencing. We gratefully acknowledge financial support from the Max Planck Society, the European Research Council through an ERC Consolidator Grant (ERC CoG 819585 “SYMBeetle” to M.K.), and the U.S. National Science Foundation (DEB1926738 to C.D.).
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
R.K. and M.K. conceived the study and designed the experiments. C.D and T.E. provided conceptual support. C.D. provided Sodalis praecaptivus strain MC1. R.K. performed the fitness experiments, microscopy and µCT analysis as well as respective data acquisition and analysis. For the triple RNASeq analysis, R.K. dissected bacteriomes and extracted RNA. H.V. and J.K. assembled and annotated the O. surinamensis genome. A.S.P.C. and J.K. performed the gene expression analyses. M.K. and T.E. provided supervision. R.K. and M.K. drafted the original manuscript. All authors reviewed and edited the final manuscript version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Krüsemer, R., Carvalho, A.S.P., Keller, J. et al. Experimental Sodalis infection eliminates ancient insect symbiont. Nat Commun 17, 3153 (2026). https://doi.org/10.1038/s41467-026-71143-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-71143-2