Abstract
Dorsoventral (DV) patterning of the spinal cord (SC) is orchestrated by morphogen gradients that specify distinct neural progenitor domains, the dorsal roof plate (RP), ventral floor plate (FP), and delaminating neural crest cells (NCCs). While foundational insights into SC patterning have been gained from animal models, key aspects - such as the role of retinoic acid (RA), dynamics of NCC lineage development, and human-specific features - remain poorly understood due to limitations in existing in vivo and in vitro models. Here, we present a human pluripotent stem cell (hPSC)-derived SC model, termed microfluidic SC-like structures (µSCLSs), generated by applying bioengineered, antiparallel morphogen gradients via a microfluidic platform over micropatterned hPSC colonies. The µSCLS robustly recapitulates complete DV patterning with human specific transcriptional signatures. Using this platform, we uncover a previously unrecognized RA-BMP signaling crosstalk that could explain conflicting reports on the role of RA in SC DV patterning. We further demonstrate lineage-specific ventral migration of NCCs in response to chemoattractant cues, enabling direct visualization and mechanistic dissection of sensory vs. other fate trajectories. This controllable, reproducible, and human-relevant model provides a powerful system for probing human SC development, neural crest biology, and disease modeling with unprecedented resolution.
Similar content being viewed by others
Introduction
During vertebrate neural tube (NT) development, the caudal spinal cord (SC) region is patterned along the dorsal-ventral (DV) axis into 11 distinct neural progenitor domains and two key signaling centers—the dorsal roof plate (RP) and the ventral floor plate (FP) (Fig. 1a)1,2. Each SC progenitor domain gives rise to specific neuronal subtypes that contribute to the formation of the central nervous system (CNS)3. Concurrently, neural crest cells (NCCs) delaminate from the dorsal NT and migrate ventrally (Fig. 1a), forming components of the peripheral nervous system (PNS), including the dorsal root ganglia (DRG) and the sympathetic ganglia4.
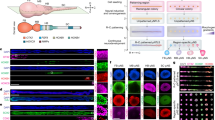
a Schematic of embryonic spinal cord being patterned along the DV axis. b Schematic of microfluidic device with µSCLS embedded in the microfluidic channel. c Protocol for generation of µSCLS. d Representative brightfield images showing development of µSCLS. e Bar plots showing tissue length (left) and projected area (right) of µSCLS from day 2 to 12. nµSCLS = 54. f Representative stitched confocal micrographs showing µSCLS on day 12 immunostained for indicated HOX genes. Similar observations were made in n = 3 independent experiments. g Representative stitched confocal micrograph showing DV patterned µSCLS on day 12 immunostained for indicated markers. Zoom-in views of boxed regions are shown on the right. Arrowheads mark a subset of cells co-expressing PAX3 and PAX6. Intensity maps show relative mean expression levels of indicated markers as a function of relative DV position in µSCLS on day 12. To generate expression heatmaps, nPAX3 = 33, nPAX6 = 33, nOLIG2 = 53 tissues from n = 3 experiments. h Representative confocal micrographs showing NCCs (box I), RP (box II), and FP (box III) in µSCLS on day 12. i Bar plot of patterning efficiencies of µSCLS from three different hPSC lines combined. Different symbols represent each line. n = 3 independent experiments for each cell line. j Representative brightfield image showing neurite-like processes in µSCLS on day 12. Arrowheads mark subset of such processes. Similar observations were made in n = 3 independent experiments. k Representative confocal micrographs showing neurites expressing βIII-TUBULIN. Arrowheads mark subset of neurites. Similar observations were made in n = 3 independent experiments. l Representative confocal micrographs of µSCLS on day 12 immunostained for indicated lineage markers. Top panels show neural crest region, while bottom panels show pMN domain. Green arrowheads mark subset of OLIG2−HB9−ISL1+ sensory neurons. Orange arrowheads mark subset of OLIG2−HB9+ISL1+ motor neuron. White arrowheads mark subset of OLIG2+HB9+ISL1+ early motor neurons. In (e and i), error bars represent mean ± s.d. Scale bars, 500 µm (d, g), 200 µm (j, k), 50 µm (h, l).
Classic animal studies have demonstrated that DV patterning of the SC is largely orchestrated by opposing gradients of morphogens secreted from adjacent tissues: dorsally derived BMPs and WNTs, and ventrally derived SHH (Fig. 1a)5,6,7,8,9,10. These morphogens induce the expression of specific transcription factors (TFs) within SC progenitor domains in a dose-dependent manner, activating lineage-specific differentiation programs (Fig. 1a). Some TFs, such as OLIG2 and IRX3, also engage in mutually repressive interactions that help stabilize domain identities, sharpen boundaries, and ensure the segregation of distinct progenitor pools.
Retinoic acid (RA), secreted from the somites flanking the SC, also plays a role in DV patterning, although its precise function remains unclear. Explant studies have shown that RA can upregulate both dorsal and ventral TFs in the SC11,12,13,14; notably, both OLIG2 and IRX3 are induced by RA treatment12,14. Disruption of RA signaling in vivo results in SC ventralization, suggesting a potential dorsalizing effect of RA—although this interpretation is confounded by the concurrent impairment of RP development11,12,14.
Shortly after NCCs delamination, these cells undergo lineage bifurcation into sensory or sympathetic fates15,16. Sensory neuron precursors halt their migration near the NT to form the DRG, while sympathetic neuron precursors continue migrating ventrally to the dorsal aorta17,18. Although several chemoattractants guiding sympathetic precursor migration have been identified19,20, the mechanisms governing sensory precursor arrest remain poorly understood. In vivo studies are limited by significant technical challenges, including imaging constraints, physiological complexity, and difficulties in controlling gene function or signaling dynamics. Therefore, an in vitro model that faithfully recapitulates NCC development—and is amenable to live imaging and genetic or chemical perturbation—would provide a powerful experimental tool for dissecting the cellular mechanisms underlying NCC migration and lineage development.
Recent studies have also identified human-specific features of SC development, such as unique gene expression profiles and the early emergence of oligodendrocyte progenitors21,22. These findings underscore the need for human-relevant SC models that are controllable, reproducible, ethically accessible, and compatible with hypothesis-driven research and therapeutic development23,24,25.
Human pluripotent stem cell (hPSC)-based developmental models have emerged as powerful, human-relevant platforms for studying development, physiology, and disease. However, most current models rely on uniform culture conditions without externally imposed signaling gradients, making regional patterning and tissue organization heavily dependent on self-organization. As a result, hPSC-derived SC-like tissues often exhibit inconsistent, incomplete patterning and low efficiency26,27,28,29. More robust models have recently been developed by introducing bioengineered gradients—such as those generated by microfluidics or synthetic organizers—to hPSC-derived neural tissues30,31,32,33. Notably, a recent model using synthetic organizers successfully recapitulated full DV patterning of the SC33. However, this approach has some limitations: the use of synthetic organizers restricts experimental control, the free-floating culture format hinders high-resolution imaging, and ventral migration of NCCs was not observed33.
In this study, we present a fully patterned and controllable human SC model, generated by applying microfluidic gradients of exogeneous morphogens to hPSC-derived SC-like tissues. We refer to these as microfluidic SC-like structures (µSCLSs). The µSCLS exhibits complete DV patterning—including all 11 progenitor domains as well as the dorsal RP and ventral FP—and recapitulates human-specific gene expression profiles. Using this controllable platform, we investigate the previously ambiguous role of RA in SC DV patterning and uncover a crosstalk between RA and BMP signaling. Furthermore, we demonstrate that NCCs within the µSCLS undergo lineage-specific ventral migration driven by chemoattraction, faithfully modeling in vivo behavior.
Results
DV patterned µSCLS
To generate DV patterned µSCLSs, we designed a microfluidic device composed of a central channel connected to two medium reservoirs, into which dorsalizing and ventralizing factors could be introduced, respectively (Fig. 1b and Supplementary Fig. 1a; see “Methods”). The reservoir containing dorsalizing factors was designated “dorsal reservoir,” while the one containing ventralizing factors was termed “ventral reservoir” (Fig. 1b).
The µSCLS protocol began with microcontact printing an array of rectangular Geltrex-coated adhesive islands onto a coverslip, which guided the formation of hPSC colonies with defined rectangular geometries. After colony seeding, the microfluidic structural layer was aligned and affixed to the coverslip (Supplementary Fig. 1b). Geltrex was chosen for its enrichment in basement membrane proteins characteristic of the NT environment17,18. The positions of hPSC colonies were precisely aligned with the microfluidic channel using positional markers engraved on the device (Fig. 1b and Supplementary Fig. 1b).
Two days after initial hPSC colony formation in mTeSR medium, the microfluidic channel was filled with 100% Geltrex to embed the colonies within a three-dimensional matrix, thereby supporting epithelialization and lumen formation (Fig. 1c and Supplementary Fig. 1b). At this point, the culture medium was switched to neural induction medium (NIM; see “Methods”). Because early activation of WNT signaling promotes SC fate during neural induction30,32,34, we supplemented NIM with the WNT agonist CHIR99021 (3 µM) from day 3 to day 5 to promote SC identity specification in the hPSC colonies (Fig. 1c).
To induce DV patterning in µSCLS, opposing gradients of BMP4 (150 ng mL−1) and smoothened agonist (SAG; 250 nM), a SHH signaling agonist, were established by adding these factors to the dorsal and ventral reservoirs, respectively, starting on day 5 (Fig. 1c and Supplementary Fig. 1b). Passive diffusion of fluorescent dextran was used as a proxy to assess morphogen gradient formation, confirming that stable chemical gradients were established within approximately 48 h (Supplementary Fig. 1c–e). Our µSCLS protocol supported continuous tissue growth over time (Fig. 1d, e). By day 12, µSCLSs had acquired a thoracic SC identity, as indicated by expression of the thoracic SC marker HOXC9 and the absence of the more caudal marker HOXC10 (Fig. 1f).
We also performed temporal immunofluorescence analysis and live imaging to examine tissue dynamics during µSCLS development. Between day 2 and day 6, we observed gradual cellular reorganization, characterized by progressive alignment of the long axes of cell nuclei along the thickness of µSCLS tissues, indicative of increasing epithelialization (Supplementary Fig. 1f). During this period, µSCLS acquired neural identity, as evidenced by gradual loss of OCT4 and NANOG expression and sustained expression of SOX2 (Supplementary Fig. 1f). Development of the µSCLS progressed through a transient neuromesodermal progenitor (NMP) state, with a significant portion of cells on day 4 co-expressing SOX2 and Brachyury (BRA) (Supplementary Fig. 1g). The SOX2+BRA+ putative NMP cells were restricted to the dorsal and ventral poles of the µSCLS on day 6 and became undetectable by day 10 (Supplementary Fig. 1g).
Live imaging using a Lifeact-GFP reporter hPSC line revealed the formation of a tubular µSCLS structure enclosing a single, continuous central lumen within 2 days (Supplementary Fig. 1h and Supplementary Movie 1). Immunostaining for ZO-1, a tight junction protein and apical polarity marker, confirmed that the apical surfaces of µSCLS faced the central lumen (Supplementary Fig. 1i). In vivo, neural progenitor cells in the pseudostratified neuroepithelium of the NT undergo interkinetic nuclear migration, where mitotic cells localize at the apical surface and nuclei in S phase are displaced toward the basal side. To assess whether µSCLSs recapitulate this organization, we performed EdU incorporation assay (marking S-phase nuclei) and immunostaining for phospho-histone H3 (pH3, a mitosis marker) on day 5 and day 12. EdU-labeled nuclei were preferentially located at the basal side, whereas pH3+ mitotic cells were concentrated near the apical surface (Supplementary Fig. 1j). To further confirm their pseudostratified cytoarchitecture, we generated µSCLSs by spiking single cytoplasmic EGFP-expressing hPSCs into wild-type cells. On both day 5 and day 12, EGFP-labeled cytoplasm of individual cells was observed spanning from the apical to the basal surfaces, consistent with the formation of a pseudostratified epithelium (Supplementary Fig. 1j).
Immunofluorescence analysis of day 12 µSCLSs revealed successful DV patterning, as evidenced by the spatially organized expression of canonical DV markers PAX3, PAX6, and OLIG2 along the DV axis (Fig. 1g). PAX6 and OLIG2 were expressed in mutually exclusive domains, whereas PAX3 and PAX6 exhibited slight overlap, consistent with the dorsal intermediate regions of the SC21,35 (Fig. 1g and Supplementary Fig. 2a).
The dorsal intermediate regions within the µSCLS were further identified by the co-expression of PAX6 and PAX7, which marks the dp4–dp6 domains21,36 (Supplementary Fig. 2b). The ventral intermediate domain p2 was identified by co-expression of PAX6 and NKX6-1, along with the absence of OLIG221,37 (Supplementary Fig. 2c). Additional intermediate domains in the µSCLS were delineated through analysis of DBX1 and DBX2 expression using RNA fluorescence in situ hybridization (RNA-FISH) (Supplementary Fig. 2d, e). DBX1 was localized to a distinct band dorsal to the OLIG2 expression domain, indicative of the intermediate domains p0 and dp62,21 (Supplementary Fig. 2d). DBX2, a broader intermediate domain marker, largely overlapped with DBX1 expression (Supplementary Fig. 2e). Notably, a DBX2+DBX1− region was observed dorsal to the DBX1+ domain, suggesting the presence of the dorsal intermediate domain dp52,21 (Supplementary Fig. 2e).
The presence of RP- and FP-like regions in day 12 µSCLSs was confirmed by LMX1a expression at the dorsal pole and FOXA2 expression at the ventral pole, respectively2,38 (Fig. 1h and Supplementary Fig. 2f). Additionally, SOX10+ NCCs were observed near the dorsal pole, delaminating from the µSCLS (Fig. 1h and Supplementary Fig. 2f). The presence of FOXA2+ FP-like region at the ventral pole of the µSCLS prompted us to examine its ventral patterning. Ventral SC progenitor markers FOXA2, NKX2-2, and OLIG2, indicative of FP-, p3-, and pMN-like identities, respectively, showed a ventral-to-dorsal organization in the µSCLS (Supplementary Fig. 2g). While expression of these lineage markers was mostly exclusive, some cells co-expressed NKX2-2 and OLIG2, early emergence of which is a characteristic of human spinal cord development21,22 (Supplementary Fig. 2g). Additionally, some cells in the ventral µSCLS region co-expressed NKX2-2 and FOXA2; such cells have also been identified in e9.5 mice, corresponding to CS12 human embryos39 (Supplementary Fig. 2g).
µSCLSs exhibited high patterning efficiency across two human embryonic stem cell (hESC) lines and one human induced pluripotent stem cell (hiPSC) line, with an average efficiency of 97.9 ± 6.2% (mean ± s.d.). Patterning efficiency was defined by the spatial segregation of PAX3, OLIG2, and FOXA2 expression domains along the DV axis (Fig. 1i and Supplementary Fig. 3a–c; see “Methods”). In contrast, imposing a single gradient of either the dorsalizing signal BMP4 (150 ng mL−1) or the ventralizing cue SAG (250 nM) resulted in lower and more variable efficiencies of 77.6 ± 30.5% and 36.1 ± 37.6% (mean ± s.d.), respectively (Supplementary Fig. 3d–f).
DV patterning in µSCLSs was tunable through modulation of morphogen concentrations. Specifically, reducing BMP4 from 150 to 50 ng mL−1 led to a contraction of the dorsal PAX3 expression domain (Supplementary Fig. 3g). Similarly, reducing SAG from 250 to 125 nM resulted in shortened expression domains of the ventral markers OLIG2 and FOXA2 (Supplementary Fig. 3h, i). Interestingly, while ventral OLIG2 expression remained unaffected by BMP4 reduction, FOXA2 expression at the ventral pole of µSCLSs showed an unexpected shrinkage, suggesting cross-regulatory effects between dorsalizing and ventralizing signals (Supplementary Fig. 3i). These findings underscore the complexity of SC DV patterning under the influence of opposing morphogen gradients.
DV patterning also promoted region-specific neurogenesis in µSCLSs. By day 12, neurite-like processes were observed extending from µSCLSs (Fig. 1j), and immunostaining for βIII-TUBULIN confirmed their neuronal identity (Fig. 1k). ISL1⁺HB9− sensory neurons were identified among the dorsal NCC population, while ISL1⁺HB9⁺ motor neurons emerged at the periphery of the ventral OLIG2⁺ motor neuron progenitor domain, resembling the developing mantle layer40 (Fig. 1l). Some motor neuron precursors retained OLIG2 expression, indicating an early stage of neuronal differentiation41 (Fig. 1l).
Transcriptome analysis of µSCLS
Next, we performed single-cell RNA sequencing (scRNA-seq) on day 12 µSCLSs to assess their cellular composition. Unbiased clustering using Uniform Manifold Approximation and Projection (UMAP) identified five distinct cell clusters, annotated as “Neuroepithelium” (NE), “Neural Crest” (NC), “Dorsal Root Ganglion” (DRG), “Neuron“, and “Mesoderm”, based on lineage-specific marker expression (Fig. 2a, b and Supplementary Fig. 4a–c). The small population of Mesoderm cluster likely arose as a byproduct of early WNT activation during neural induction, which is known to transiently induce a neuromesodermal progenitor state34. The Mesoderm cluster contained only a small proportion of cells and showed high expression of sclerotome-related markers PAX1, PAX9, and NKX3-2 (Supplementary Fig. 4b)42. Immunostaining for the mesenchymal marker TWIST1 revealed the presence of these mesodermal cells, mostly located near the ventral pole of µSCLSs (Supplementary Fig. 4d). Sclerotome specification requires the ventralizing signal SHH43,44, potentially accountable for the ventral localization of TWIST1+ mesodermal cells in µSCLSs. All cell populations in µSCLSs expressed thoracic markers HOXA9, HOXB9, HOXC9, and more anterior HOX genes, whereas more caudal HOX genes were not expressed (Supplementary Fig. 4e), confirming a thoracic identity of µSCLS.
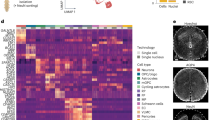
a UMAP plot of single-cell transcriptome data of day 12 µSCLS. b Dot plot showing expression of key marker genes across cell clusters. Dot sizes and colors indicate proportions of cells expressing corresponding genes and their averaged scaled values of log-transformed expression, respectively. c PCA plot of averaged gene expression profile of day 12 µSCLS and human SC across developmental stages. SC spinal cord, TH Thoracic spinal cord, BR Brachial spinal cord. d (Left) UMAP projection of integrated scRNA-seq data from day 12 µSCLS and human SC. (Right) UMAP plots of data from day 12 µSCLS and human SC, separated from the integrated UMAP plot on the left. e Heatmap showing Pearson correlation coefficients between cell types in day 12 µSCLS and human SC. Black boxes highlight the highest correlation coefficients in each row. f (Left) UMAP plot of integrated scRNA-seq data from day 12 µSCLS NE and human CS12 SC NE. (Right) UMAP projections of data from day 12 µSCLS NE and human CS12 SC NE, separated from the integrated UMAP plot on the left. g Heatmap of Pearson correlation coefficients between 13 NE subtypes from day 12 µSCLS and CS12 human SC. Black boxes highlight the highest correlation coefficients in each row. h Bar plot showing proportion of NE subtypes in day 12 µSCLS and CS12 human SC. i Heatmap of averaged expression levels of primate-specific genes in CS12 human SC and day 12 µSCLS, ordered by expression levels in CS12 human SC. Human-specific genes are marked in red. j Averaged and normalized expression levels of NKX6−2 in NE subtypes of day 12 µSCLS, CS12 human SC, and stage-matched E9.5 mouse SC. k Proportion of NE cells expressing both NKX2-2 and OLIG2 to NE cells expressing NKX2-2 or OLIG2. l Violin plots showing expression levels of indicated oligodendrocyte and neurogenic markers in day 12 µSCLS and CS12 human SC.
To determine the developmental stage represented by day 12 µSCLSs, we conducted principal component analysis (PCA) comparing the averaged gene expression profiles of µSCLSs with those of human embryonic SC tissues across developmental stages (Fig. 2c). This analysis revealed that the transcriptomic profile of day 12 µSCLSs most closely resembled that of the Carnegie stage (CS) 12 human SC (Fig. 2c).
To validate the authenticity of cell types within µSCLSs, we integrated the µSCLS transcriptomic dataset with that of human CS12 SC. The mesodermal population from the µSCLS dataset was excluded from this analysis due to its low abundance and limited relevance. Additionally, given the scarcity of DRG cells in CS12 human samples, DRG cells from CS14 human embryos were included to improve coverage in the integrated analysis. Upon integration, day 12 µSCLS clusters showed strong concordance with corresponding human SC cell types, with substantial overlap between µSCLS and CS12 human clusters (Fig. 2d). Consistently, Pearson correlation coefficients calculated for matched clusters—based on averaged gene expression - were highest between corresponding µSCLS and human SC cell types (Fig. 2e).
The NE cluster was further analyzed using a “knowledge matrix” approach, which assigns cell identities based on the expression of established molecular markers22,45,46. This analysis revealed the presence of all 13 embryonic SC NE subtypes (Fig. 2f, g and Supplementary Fig. 4f, g). Upon integration with NE cells from human CS12 SC tissues, µSCLS-derived NE cells exhibited a coherent DV organization, with each subtype aligning closely with its in vivo counterpart in the integrated UMAP plot (Fig. 2f and Supplementary Fig. 4f). Likewise, Pearson correlation coefficients calculated from average gene expression profiles were highest between most corresponding NE subtypes from µSCLS and human SC (Fig. 2g). Some discrepancies were noted among dorsal intermediate progenitors, potentially due to their low abundance in the datasets (Fig. 2g and Supplementary Fig. 4f). Marker genes used to classify NE subtypes also showed highly similar expression patterns between NE cells from µSCLS and human SC tissues (Supplementary Fig. 4f, g). When NE progenitors were grouped into RP, dorsal (dp1–3), dorsal intermediate (dp4–6), ventral intermediate (p0–2), pMN, p3, and FP domains, the resulting cellular compositions were comparable between µSCLS and human SC NE populations (Fig. 2h).
The gene expression profiles of NE subtypes within µSCLS were consistent with our immunofluorescence and RNA-FISH results (Supplementary Fig. 4g). Specifically, the RP population exhibited upregulated expression of LMX1A, while the FP population strongly expressed FOXA2 (Fig. 1h and Supplementary Fig. 4g). PAX3 was broadly expressed in dorsal progenitors, whereas PAX6 was upregulated in intermediate progenitors but absent in the pMN domain and more ventral regions (Fig. 1g and Supplementary Fig. 4g). The pMN population showed elevated OLIG2 expression but lacked PAX6 expression (Fig. 1g and Supplementary Fig. 4g). Additionally, WNT1 and SHH were expressed in the RP and FP cell populations, respectively, consistent with their established roles as signaling centers that secrete these morphogens2 (Supplementary Fig. 4g). Co-expression of PAX7 and PAX6 was observed throughout the dorsal intermediate domains (Supplementary Figs. 2b and 4g). Within the p2 domain, cells co-expressed NKX6-1 and PAX6 but did not show upregulation of OLIG2 (Supplementary Figs. 2c and 4g). DBX1 and DBX2 were co-expressed in the p0 and dp6 domains, with DBX2 additionally extending into the dp5 domain (Supplementary Figs. 2d, e and 4g).
“Knowledge matrix”-based cell identification was also applied to the Neuron population. Upon integration of the transcriptomic data from µSCLS neurons with that of human CS12 SC neurons and projection onto a UMAP plot, µSCLS neurons exhibited an overall DV organization and colocalized with their in vivo counterparts (Supplementary Fig. 5a, b). Most neuronal subtypes were identified; however, dI1 neuron was undetected in both µSCLS and human CS12 SC (Supplementary Fig. 5b). Considering that dI1 neuron has been identified in CS14 human SC, it is possible that the developmental stage of day 12 µSCLS was too early for the appearance of dI1 neurons22. Similar to human CS12 SC, µSCLS contained a higher proportion of ventral neurons (V0–V3, MN) compared to dorsal interneurons (dI1–dI6) (Supplementary Fig. 5b, c). When interneurons were grouped into dorsal (dI1–dI3), dorsal intermediate (dI4–dI6), and ventral intermediate (V0–V2) categories, Pearson correlation coefficients of averaged gene expression levels were highest between the corresponding neuron subtypes from µSCLS and human CS12 SC (Supplementary Fig. 5d). Additionally, marker genes used to classify neuron subtypes displayed similar expression patterns across matched neuron subtypes from µSCLS and human CS12 SC (Supplementary Fig. 5e).
Human-specific gene expression signatures were faithfully recapitulated in hPSC-derived µSCLS. Of 51 primate-enriched protein-coding genes expressed in the human CNS—including 10 human-specific genes—µSCLS expressed 34, with expression patterns resembling those in human CS12 SC22,47,48 (Fig. 2i). Another example is NKX6-2, which is broadly expressed across ventral domains in human CS12 SC, in contrast to its more restricted p1 domain expression in stage-matched mouse E9.5 SC22,49. This broader expression pattern of NKX6-2 was also observed in the ventral regions of µSCLS (Fig. 2j). Previous studies have shown that NKX2-2 and OLIG2 co-expressing cells arise earlier in human SC than in mouse SC21,22. Consistent with observations made using immunofluorescence, cells co-expressing NKX2-2 and OLIG2 were identified in the µSCLS transcriptomic data (Fig. 2k and Supplementary Fig. 2g). Importantly, among µSCLS cells expressing either NKX2-2 or OLIG2, 8% co-expressed both genes—higher than in mouse E9.5 SC (2%) and comparable to human CS12 SC (10%) (Fig. 2k). These co-expressing cells in µSCLS also exhibited expression patterns of oligodendrocyte markers similar to those in human CS12 SC and showed weak expression of neurogenic genes, in line with their identity as oligodendrocyte precursors21,22 (Fig. 2l).
Together, transcriptomic analyses demonstrate that µSCLS comprises all major cell types, including all 13 NE subtypes, and faithfully recapitulates the transcriptomic landscape of the human embryonic SC.
Effect of RA in SC DV patterning
RA has been reported to promote the expression of both dorsal and ventral markers, making its overall effect on SC DV patterning ambiguous14,26,27,50,51 (Fig. 3a). To investigate the effect of RA on SC DV patterning, we compared DV patterning of the µSCLS cultured with or without RA (500 nM; Fig. 3b). To minimize endogenous RA production, we replaced the basal medium with N2B27 lacking vitamin A and reduced BMP4 concentration to 50 ng mL−1. RA treatment produced an overall dorsalizing effect: the expression domain of the dorsal marker PAX3 expanded ventrally, while expression of the ventral marker OLIG2 was markedly reduced and FOXA2 expression was completely abolished (Fig. 3c, d).
a (Left) Schematic of somites flanking SC and secreting RA. (Right) Schematic of RA promoting expression of both dorsal and ventral SC markers as indicated. b Protocol for assessing the effect of RA in µSCLS DV patterning. c Representative stitched confocal micrographs of DV patterned µSCLS without (Top) or with (Bottom) uniformly supplemented RA. d Dorsal and ventral expression boundaries of PAX3, OLIG2, and FOXA2 in day 12 µSCLS treated with or without RA. nExperiment = 4, nPAX3,−RA = 23, nOLIG2,−RA = 21, nFOXA2,−RA = 21, nPAX3,+RA = 30, nOLIG2,+RA = 26, and nFOXA2,+RA = 26. e Representative stitched confocal micrographs of µSCLS patterned with ventral SAG gradient but without dorsal BMP4 gradient. Top: µSCLS without RA treatment; Bottom: µSCLS uniformly treated with RA. f Proportion of cells expressing OLIG2 or FOXA2 in µSCLS patterned with ventral SAG gradient with or without RA as indicated. nExperiment = 3, nOLIG2,−RA = 34, nFOXA2,−RA = 34, nOLIG2,+RA = 35, and nFOXA2,+RA = 35. g (Top) Representative western blot membrane showing increased phosphorylation of SMAD1/5 under RA treatment. (Bottom) Relative SMAD1/5 phosphorylation level in RA-treated samples compared to untreated controls. µSCLSs were patterned with antiparallel gradients of BMP4 and SAG. nExperiment = 4, n−RA = 5, n+RA = 5. In (d, f, g), error bars represent mean ± s.d. Two tailed t-test was used to assess statistical significance in (f). Two tailed one sample t-test with a hypothesized mean of 1 was used to assess statistical significance of SMAD1/5 phosphorylation level increase in RA-treated samples in (g), yielding a P-value of 0.4062. N.S.: p ≥ 0.05, *: p < 0.05. All scale bars, 200 µm.
RA signaling has been shown to influence rostrocaudal axial identity13,34. To exclude this possibility in RA-treated µSCLSs, we stained RA-treated or untreated µSCLSs for the thoracic marker HOXC9 and the lumbar marker HOXC10 (Supplementary Fig. 6a). Irrespective of RA treatment, µSCLSs were HOXC9+HOXC10−, supporting a stable thoracic identity. Thus, the RA-induced alterations in DV patterning of µSCLSs are unlikely to result from changes in rostrocaudal axial identity.
The dorsalizing effect of RA contrasts with existing in vitro SC DV patterning models and motor neuron differentiation protocols, where RA has been reported to promote ventral marker expression, including FOXA2 and OLIG226,27,50. Importantly, those studies were conducted under uniform culture conditions containing only ventralizing cues. To test the hypothesis that RA and BMP4 may interact synergistically, we repeated the experiment in the absence of a dorsal-ventral BMP4 gradient. Under these conditions, RA did not affect the proportion of FOXA2⁺ or OLIG2⁺ cells, suggesting a crosstalk between RA and BMP signaling pathways (Fig. 3e, f). Further supporting this interaction, µSCLS patterned with antiparallel BMP4 and SAG gradients exhibited a 67.0% increase in SMAD1/5 phosphorylation upon RA treatment (Fig. 3g). To examine whether SHH signaling is also required for RA-mediated dorsalization of µSCLSs, we repeated the experiment in the absence of the ventral-dorsal SAG gradient. Even without SAG, RA successfully dorsalized µSCLSs, as the expression domain of the dorsal marker PAX3 expanded, whereas the expression domain of the intermediate marker PAX6 was reduced (Supplementary Fig. 6b, c). Together, these data suggest that SHH signaling is not required for RA to exert a dorsalizing effect on DV patterning of µSCLSs.
To investigate the mechanism underlying RA-mediated enhancement of BMP signaling, we re-analyzed data from a mouse ESC-based motor neuron differentiation study that included retinoic acid receptor (RAR) chromatin immunoprecipitation sequencing (ChIP-seq) for RA-treated and untreated samples52. This analysis revealed 16 RAR binding sites located near genes involved in BMP signaling regulation, with 5 sites specific to RA-untreated cells and 8 unique to RA-treated cells (Supplementary Table 1). These findings suggest that RA may enhance SMAD1/5 phosphorylation by modulating RAR binding to regulatory elements of BMP pathway-associated genes.
To identify which of the 16 genes were differentially expressed in µSCLSs in response to RA, we quantified mRNA levels of each gene in RA-treated and untreated µSCLSs using quantitative polymerase chain reaction (qPCR). Four of the 16 genes—GDF3, SKI, SKIL, and SFRP4—were significantly upregulated following RA treatment (Supplementary Fig. 6d). To test whether these genes mediated the dorsalizing effect of RA, we individually knocked down each gene using small interfering RNA (siRNA) and compared DV patterning in RA-treated and untreated µSCLSs. Among the siRNAs tested, only knockdown (KD) of GDF3 resulted in a significant reduction of target mRNA levels (Supplementary Table 7). We therefore focused on examining DV patterning in RA-treated and untreated µSCLSs under GDF3-KD. Notably, in GDF3-KD µSCLSs, RA treatment no longer altered the expression domain of the ventral marker OLIG2 (Supplementary Fig. 6e, f). In contrast, RA treatment still reduced FOXA2 expression, suggesting that RA-mediated DV patterning involves domain-specific downstream mediators (Supplementary Fig. 6e, g). To further investigate the epistatic relationship among RA signaling, BMP signaling, and GDF3, we performed qPCR under conditions with or without RA and with or without a dorsal-ventral BMP4 gradient. GDF3 expression was significantly upregulated by RA treatment but was unaffected by BMP4, supporting a model in which RA directly induces expression of GDF3, which in turn directly or indirectly modulates BMP signaling to regulate DV patterning.
NCC migration
To track the delamination and migration of NCCs, we developed a SOX10::T2A-Cre lineage-tracing hESC line. A T2A-Cre cassette was knocked in downstream of the endogenous SOX10 locus in a hESC line harboring a Cre-inducible ZsGreen cassette at the AAVS1 locus (Fig. 4a). Live imaging from day 6 to day 8 of µSCLS derived from this lineage-tracing line revealed gradual delamination of NCCs from the dorsal side, with ZsGreen expression initiating after delamination (Supplementary Movie 2). This observation aligns with in vivo murine studies reporting that SOX10 expression in NCCs begins post-delamination15,19. Prolonged culture showed some NCCs detached from dorsal NCC clusters, became completely embedded in the surrounding matrix, and migrated ventrally (Fig. 4b, c, Supplementary Fig. 7a, and Supplementary Movie 3). Tracking individual NCCs demonstrated directional migration toward the ventral side (Fig. 4d), supported by a polar histogram of migratory trajectories skewed ventrally (Fig. 4e). Compared to a random walk simulation, migrating NCCs exhibited reduced circular standard deviation and increased directionality, consistent with directed migration (Fig. 4f). Their mean square displacement exhibited a parabolic trend, further characteristic of directional migration (Fig. 4g).
a Schematic of lineage tracing hESC line. b Protocol for continuous development. c (Top) Representative micrograph showing day 15 µSCLS generated using SOX10::T2A-Cre lineage tracer. (Bottom) Zoom-in views highlight ventral migration at indicated time points. d Micrographs of representative NCC migration trajectories. e Polar histogram of NCC migratory directions. nmigratory_tracks = 602. f Bar plots comparing circular standard deviation (Left) and directionality (Right) between NCCs and simulated random walk. nExperiment = 5 and nSimulation = 5. g Mean square displacement (MSD) of NCCs and random walk simulations over time. nExperiment = 5 and nSimulation = 5. h Representative micrographs showing distribution of SOX10+ non-DRG NCCs and SIX1 + DRG cells in day 18 µSCLS, untreated with AMD3100. i Schematic of criteria for dorsal versus ventral location quantification. j Bar plot comparing number of ventrally located non-DRG NCCs and DRG cells. nExperiment = 4 and nSample = 14. k Representative confocal micrographs showing negative staining for ASCL1 in ventrally located NCCs. Dotted line encircles ventrally located ZsGreen+ NCCs. l Feature plots of day 12 µSCLS showing expression of CXCL12 in the FP and CXCR4 in both NC and DRG clusters. m Representative micrographs showing altered distribution of non-DRG NCCs and DRG cells in AMD3100-treated day 18 µSCLS. n Bar plot comparing ventrally located non-DRG NCCs in control vs. AMD-treated samples. nExperiment = 4, nCtrl_sample = 14, and nAMD_sample = 14. o, p Violin plots showing DV positional distribution of non-DRG NCCs (o) and DRG cells (p) in control vs. AMD-treated conditions. nExperiment = 4, nCtrl_non_DRG_NCCs = 95,867, nAMD_non_DRG_NCCs = 102,218, nCtrl_DRG_cells = 38,377, and nAMD_DRG_cells = 43,180. q, r Bar plots comparing dorsally located non-DRG NCCs (q) and DRG cells (r) between conditions. nExperiment = 4, nCtrl_sample = 14, and nAMD_sample = 14. s Schematic showing effect of CXCR4 inhibition in µSCLS. Error bars represent mean ± s.d. in (f, j, n, q, r). Two tailed t-test was used to assess statistical significance in (f, q, r), yielding P-values of 0.03856 (f, left), 0.00173 (f, right), 0.02966 (q), and 0.01159 (r). Two-sided zero-inflated rank test was used in (j, n), yielding P-values of 0.00188 (j) and 0.04686 (n). *: p < 0.05, **: p < 0.01. Scale bars, 500 µm (c, Top), 100 µm (c, Bottom), 50 µm (d), 200 µm (h, k, m).
The earliest fate bifurcation of trunk NCCs occurs between the sensory and sympathetic lineages. Sensory neuron precursors arrest migration adjacent to the NT to form the DRG, whereas sympathetic neuron precursors migrate further ventrally toward the dorsal aorta15,16,17,18. Immunofluorescence analysis revealed that expression of SIX1, a DRG marker, was mutually exclusive with SOX10 (Fig. 4h). SIX1+ DRG cells were intermingled with SOX10+ NCCs in the dorsal region, whereas no DRG cells were observed among the more ventrally located, sparsely distributed NCCs, presumed to be migrating ventrally (Fig. 4h). For quantification, we defined cells located above the dorsal tip of the µSCLS as “dorsally located,” and those below the midpoint as “ventrally located” (Fig. 4i). On average, the number of ventrally located SOX10+ non-DRG NCCs was 6.79 ± 9.74 per device, while DRG cells were rarely found ventrally (0.14 ± 0.36) (Fig. 4j). These ventrally migrating non-DRG NCCs, suspected to be sympathetic neuron precursors, were examined for ASCL1 expression by immunostaining but were negative (Fig. 4k), possibly due to a lack of maturation cues from the dorsal aorta17.
To determine the timing of NCC lineage specification, µSCLSs derived from the SOX10::T2A-Cre lineage tracer were immunostained for SOX10 and SIX1 at multiple time points (Supplementary Fig. 7b). SOX10 expression was detected on day 9 in delaminated NCCs, whereas ZsGreen expression appeared with a one-day delay, becoming detectable on day 10. SIX1 expression was observed in day 11 µSCLS, suggesting that NCC specification toward the DRG lineage occurs after delamination. ZsGreen expression in SIX1+ DRG cells further indicates that these cells were originated from a multipotent SOX10+ NCC pool. This observation aligns with previous studies showing that delaminating NCCs are multipotent progenitors and that fate bifurcation occurs during migration15.
Both sympathetic and sensory NCC lineages are reportedly responsive to chemoattraction by CXCL1219,20. Consistent with this, transcriptomic analysis of day 12 µSCLS revealed expression of CXCR4, the receptor for CXCL12, in both NC and DRG clusters (Fig. 4l). Additionally, CXCL12 was expressed in the FP, potentially driving ventral NCC migration (Fig. 4l). Chemical inhibition of CXCR4 with AMD3100 (200 nM), a CXCR4 antagonist, resulted in a dorsal shift in the localization of both non-DRG NCCs and DRG cells (Fig. 4m). Specifically, the number of ventrally located non-DRG NCCs was significantly reduced from 6.79 ± 9.74 to 1.54 ± 4.96 per device (Fig. 4n). Distribution analyses confirmed a dorsal shift for both populations (Fig. 4o–r). The proportion of dorsally located non-DRG NCCs increased from 40.70 ± 24.92% to 64.71 ± 29.95% (Fig. 4q), and that of DRG cells increased from 38.16 ± 27.14% to 69.24 ± 32.96% (Fig. 4r). These results are consistent with in vivo findings demonstrating CXCL12 as a chemoattractant for both sympathetic and sensory precursors19,20. Together, these findings show that µSCLS faithfully recapitulates the chemoattraction-based, fate-dependent migration behavior of NCCs.
Discussion
Unlike existing SC patterning models that rely on uniform culture medium conditions26,27,28,29, the µSCLS employs bioengineered, controllable morphogen gradients to achieve robust and complete DV patterning. This includes the specification of both the most dorsal (delaminating NCCs and RP) and ventral (FP) cell types in the SC. Recent studies have demonstrated that antiparallel gradients, generated through microfluidics or artificial signaling centers, can also produce DV patterned SC models30,32,33. However, artificial signaling centers require genetic modifications to alter the dose or composition of secreted molecules, and precise spatiotemporal control over the resulting morphogen gradients remains challenging33. Moreover, the genetically modified cells that serve as artificial signaling centers often integrate into the SC-like structure, reducing experimental control and physiological relevance33. Similarly, current microfluidic platforms for modeling SC DV patterning obstruct the ventral migratory path of NCCs, preventing faithful recapitulation of NCC migration and lineage development32.
Nonetheless, the µSCLS employs microfluidic gradient designs similar to those utilized in other microfluidic platforms32 for modeling SC DV patterning. However, this study extends beyond existing approaches by uncovering a previously unrecognized crosstalk between RA and BMP signaling and by enabling direct visualization and mechanistic dissection of lineage-specific ventral migration of NCCs, as well as divergent sensory vs. sympathetic fate trajectories. More specifically, the µSCLS successfully recapitulates the chemoattraction-based, fate-dependent ventral migration behavior of NCCs. Transcriptomic analysis confirmed that the µSCLS was fully patterned, containing all 13 NE subtypes with strong transcriptomic similarity to their in vivo counterparts. Additionally, the hPSC-derived µSCLS exhibited human-specific transcriptional signatures, including the expression of primate- and human-specific genes, a distinctive NKX6-2 expression pattern, and the early emergence of oligodendrocyte precursors - together supporting the human relevance of this model.
We further leveraged the µSCLS to address discrepancies between in vivo and in vitro studies regarding the role of RA. Although RA has been reported to exert a dorsalizing effect in vivo12,14, several in vitro studies have described the opposite26,27,50. In the µSCLS, RA exhibited a strong dorsalizing effect, consistent with in vivo observations. This effect was dependent on the presence of BMP4, suggesting a synergistic interaction between the two signaling pathways. These findings may explain conflicting in vitro reports, as prior studies administered only ventralizing cues.
To identify genes mediating the interaction between RA and BMP signaling, we interrogated an external dataset and identified 16 BMP pathway-related genes that show differential RAR binding in the presence of RA. Among these, four genes—GDF3, SKI, SKIL, and SFRP4—were upregulated in µSCLSs. GDF3-KD abolished the effect of RA on expression of the pMN marker OLIG2, confirming the involvement of GDF3 in RA-induced dorsalization. In contrast, expression of the FP marker FOXA2 remained sensitive to RA, suggesting domain-specific regulation. Consistent with this idea, differences in chromatin accessibility among neural progenitor domains have been reported53, supporting the existence of domain-specific downstream targets of RA signaling. GDF3 is a member of the TGF-β superfamily and can function as a BMP inhibitor54,55; thus, it is not immediately apparent how GDF3 mediates the dorsalizing effect of RA. Further analysis using single-cell multiome tools may identify additional domain-specific RA targets and clarify the mechanism by which GDF3 contributes to RA-induced dorsalization in µSCLSs.
The µSCLS also provides a unique platform for investigating fate-dependent NCC migration. NCCs in the µSCLS migrated ventrally via chemotaxis mediated by CXCL12-CXCR4 signaling. This migration was fate-dependent, as only non-DRG NCCs migrated ventrally. Although the ventrally migrating population is suspected to include sympathetic neuron precursors, their precise identity remains unknown. Importantly, DRG cells clearly ceased migration despite responding to CXCL12, suggesting a distinct regulatory mechanism for migration arrest. As the mechanism underlying DRG migration cessation remains elusive18,20,56, the µSCLS represents a powerful system to study fate-dependent NCC migration, particularly the regulation of DRG cell arrest.
Some limitations of the µSCLS should be acknowledged. First, the morphogen gradients in the µSCLS system were assessed to be linear, whereas in vivo morphogen signals are more complex and challenging to quantify. Beyond passive diffusion, the regulation of in vivo morphogen gradients involves a range of coordinated mechanisms, including receptor-mediated uptake and feedback regulation. If the morphogen gradients in the µSCLS were indeed linear, this may have contributed to variability and less organized patterning compared to the embryonic SC, which are thought to involve exponential morphogen gradients10. Moreover, the absence of certain signaling cues—such as specific BMP isoforms or RA—could affect patterning fidelity. For example, distinct BMP ligands have been shown to promote specific dorsal progenitor identities57, and their absence may underlie the weaker correlation observed for some dorsal intermediate subtypes. Similarly, external guidance cues for NCC migration, such as signals from surrounding mesenchyme and somites18,20,58,59, were absent. This could explain variability in ventrally located NCCs across samples. Nevertheless, these limitations are likely addressable given the flexibility of the µSCLS platform. Additional signaling factors, including specific BMP ligands and chemoattractants, can readily be applied in gradients to refine DV patterning or enhance NCC migration fidelity.
In summary, the µSCLS represents a robust, controllable, scalable, and human-relevant in vitro model of SC development that recapitulates full DV patterning. Its high level of controllability and compatibility with live imaging and genetic lineage tracing make it a powerful tool for investigating fundamental aspects of SC development, physiology, and disease. These include morphogen signal decoding, progenitor fate specification, developmental robustness and precision, and cell-cell interactions.
Methods
Ethical statement
All experimental procedures used in this work with hPSCs were approved by the Human Pluripotent Stem Cell Research Oversight Committee at the University of Michigan. The research conforms to the 2021 Guidelines for Stem Cell Research and Clinical Translation recommended by the International Society for Stem Cell Research.
Cell line
hPSC lines used in this study were H9 hESC (WA09, WiCell; NIH registration number: 0062), H1 hESC (WA01, WiCell; NIH registration number: 0043), and 1196a (human induced pluripotent stem cell) lines from the University of Michigan Pluripotent Stem Cell Core. Additionally, a Lifeact-GFP H2B-mCherry WIBR3 hESC line60, a gift from O. Reiner, a cytoplasmic EGFP-expressing H9 hESC line, and a SOX10::T2A-Cre lineage tracer H9 hESC line were used. hPSC lines used in this study were all validated by original sources and in-house by immunofluorescence staining for pluripotency markers along with successful differentiation to three germ layers. All hPSC lines were maintained in a feeder-free culture system for at least ten passages and were authenticated as karyotypically normal by Cell Line Genetics. All hPSC lines were not contamination by mycoplasma (LookOut Mycoplasma PCR Detection kit; Sigma-Aldrich).
Cell culture
All hPSC lines were maintained under standard feeder-free conditions using mTeSR medium (STEMCELL Technologies). Cells were cultured on tissue culture plates coated with dehydrogenase-elevating virus (LDEV)-free, hESC-qualified, reduced growth factor basement membrane matrix Geltrex (Thermo Fisher Scientific). Cultures were visually inspected at each passage for the presence of mesenchymal-like cells, which indicate spontaneous differentiation. All hPSCs were used between passages 60 and 80.
Cytoplasmic EGFP-expressing hESC line
To generate cytoplasmic EGFP-expressing hESC line, the lentiviral vector containing EGFP61 was co-transfected with pMDLg/pRRE, pRSV-Rev, and pMD2.G (Addgene IDs: 12251, 12253, and 12259) into 293 T cells (Takara Bio) using CalFectin (SignaGen Laboratories). 10 μM Forskolin (Selleck Chemicals) was added after 24 h of transfection. On the third day following transfection, the supernatant was collected, concentrated using Lenti-X Concentrator (Takara Bio), and used for the transduction of an H9 ES cell line. Then, EGFP-positive cells were sorted by flow cytometry with Bigfoot spectral cell sorter (Thermo Fisher Scientific).
SOX10::T2A-Cre lineage tracer
The Sox10::T2A-Cre lineage tracer hESC line was generated in the background of the previously published ZsGreen conditional reporter ES cell line32. Briefly, T2A-Cre was inserted before the stop codon of the SOX10 locus by using CRISPR-Cas9-mediated genome editing. The donor plasmid containing the left and right arms in the SOX10 locus, T2A-Cre, PGK-Hydromycin as well as gRNA plasmids targeting the SOX10 locus was generated as shown in our previous publication32. The gRNA sequence was: 5′ cactgtcccggccctaaagg 3′.
The donor and gRNA plasmids were transfected into the ZsGreen conditional reporter H9 hESCs and positive clones were selected in the presence of hygromycin (50 µg mL−1) for 7 days. Individual ES cell clones were genotyped and checked for a successful knock-in.
Device fabrication
The microfluidic device consisted of a polydimethylsiloxane (PDMS) structural layer attached to a coverslip. To fabricate PDMS structural layer, PDMS curing agent and base polymer (Sylgard 184; Dow Corning) were mixed at a ratio of 1:10. The mixed PDMS prepolymer was cast onto a microfabricated silicon mold and baked at 110 °C for 1 h. Harris Uni-Core punch tool (6 mm diameter, Ted Pella) was used to punch medium reservoirs in the PDMS structural layer. In parallel, PDMS stamps with rectangular micropatterns were fabricated by casting PDMS prepolymer (with the ratio of curing agent to base polymer of 1:20) onto a microfabricated silicon mold. This was baked at 110 °C for 1 h and 30 min. PDMS stamps were then peeled off, sterilized using 100% ethanol, and coated with 1% (v/v) Geltrex solution at 4 °C overnight. The next day, glass coverslips (Thermo Fisher Scientific) were cleaned by first sonicating in 2% (v/v) Hellmanex III (Hellma USA, 9-307-011-4-507) for 30 min, rinsing 3 times in deionized water, and then sonicating in 100% ethanol for 30 min. The cleaned coverslips were blow-dried and treated with ultraviolet ozone (Ozone cleaner; Jelight) for 7 min. The Geltrex-coated PDMS stamps were blow-dried and placed in conformal contact with ultraviolet ozone-treated coverslips to transfer Geltrex adhesive patterns onto the coverslips. Stamps were peeled off and PDMS structural layers were then attached to coverslips. Alignment of microcontact-printed Geltrex islands in the patterning region was guided by position markers engraved on the structural layer and was done using a desktop aligner62.
Generation of µSCLS
On day 0, colonies of human PS cells in tissue culture plates were dissociated using Accutase (Sigma-Aldrich) at 37 °C for 8 min before being suspended in DMEM/F12 (Gibco) as single cells. Cells were centrifuged and resuspended at a concentration of 20 × 106 cells per mL in mTeSR containing the ROCK inhibitor Y27632 (10 μM, Tocris) to prevent dissociation-induced apoptosis of hPSCs63. 7 μL of the hPSC suspension was injected into the microfluidic channel through openings to the reservoirs. The microfluidic device was incubated for 45 min to allow hPSCs to attach to the Geltrex adhesive islands. Unattached cells were flushed out by adding fresh mTeSR containing 10 μM Y27632 to one of the reservoirs. The resulting monolayer hPSC colonies were cultured one day more in mTeSR without Y27632. On day 2, mTeSR medium was aspirated from the microfluidic channel and 7 µL of 100% Geltrex was injected into the channel. The device was incubated for 10 min for gelation, establishing a 3D culture environment. Neural induction medium (NIM) was added to both reservoirs after gelation. NIM consisted of basal medium and two SMAD inhibitors for neural induction: TGF-β inhibitor SB431542 (10 μM, StemCell Technologies) and BMP inhibitor LDN193189 (500 nM, StemCell Technologies). The basal medium, N2B27, comprised 1:1 mixture of DMEM/F12 (Gibco) and neurobasal medium (Gibco), 1% N2 supplement (Gibco), 2% B-27 supplement (with vitamin A, Gibco), Glutamax (2 mM, Gibco), 1% non-essential amino acids (Gibco), 100 µM 2-mercaptoethanol (Gibco), and 1% antibiotic–antimycotic (Gibco). From day 3 to day 5, CHIR99021 (CHIR, 3 μM, StemCell Technologies) was further added to the NIM. From day 5 to day 12, antiparallel signaling gradients were imposed by adding basal medium supplemented with BMP4 (150 ng mL−1, R&D Systems) to the dorsal reservoir and basal medium supplemented with SAG (250 nM, StemCell Technologies) to the ventral reservoir. For prolonged culture from day 12 to day 18, both reservoirs were filled with basal media supplemented with neurotrophic factors: BDNF (10 ng mL−1; StemCell Technologies), GDNF (10 ng mL−1; StemCell Technologies), CNTF (10 ng mL−1; StemCell Technologies), IGF-1 (10 ng mL−1; PeproTech), cAMP (1 μM; Sigma-Aldrich), and AA (0.2 μg mL−1; Sigma-Aldrich). In all cases, culture media were replenished daily.
Confirmation of neural induction
To confirm neural induction in Supplementary Fig. 1f, µSCLS were stained for OCT4, NANOG, and SOX2 at different time points. A µSCLS was categorized as expressing OCT4 if at least one cell in the tissue exhibited nuclear localization of OCT4. Similar criterion was applied for NANOG and SOX2.
Confirmation of pseudostratified epithelium
To assess the structure of the neuroepithelium, cytoplasmic EGFP-expressing H9 hESCs were mixed into wild type H9 hESCs with a ratio of 1–200 before injecting into the device. Then, all steps were carried out identical to the standard µSCLS generation protocol.
µSCLS patterning efficiency
Quantification of µSCLS patterning efficiency was based on marker expressions. A µSCLS was deemed patterned if dorsal marker PAX3 was expressed in the dorsal side without a separate expression domain further ventral to the main expression domain. Similarly, for samples stained for ventral markers OLIG2 or FOXA2, similar criterion was applied to assess patterning. The success rate was quantified as the ratio of number of patterned colonies to the number of total colonies.
Microscopy
Whole-mount immunostained µSCLS were imaged using an Olympus DSUIX81 spinning-disc confocal microscope, Nikon X1 Yokogawa spinning-disc confocal microscope, or a Nikon A1R confocal microscope. An array of partially overlapping images (15% overlap) were taken to cover the entire µSCLS area. Images taken with Olympus DSUIX81 were first corrected for background and shade using the Fiji (ver. 2.16.0.) plugin BaSiC64 and then stitched using the Fiji plugin MIST65. Images taken with Nikon A1R were first denoised using the “denoise AI” functionality in Nikon NIS-Elements platform. Then, the denoised images were stitched using the “Stitch Image” functionality in Nikon NIS-Elements platform. For 3D reconstruction, z stack images were acquired with a slice thickness of 1 μm. Low-magnification brightfield images were acquired using Olympus CKX53 inverted microscope. Live imaging was conducted using an inverted epifluorescence microscope (Zeiss Axio Observer Z1; Carl Zeiss MicroImaging) enclosed in an environmental incubator (XL S1 incubator, Carl Zeiss MicroImaging) that maintained the cell culture conditions at 37 °C and 5% CO2. Images taken using Zeiss Axio Observer Z1 were stitched using the Grid/Collection stitching plugin in Fiji.
Dextran diffusion assay
Morphogen diffusion in the microfluidic channel was characterized by using Texas Red-labelled dextran (70 kDa, with a hydrodynamic radius of 5.8 nm, Invitrogen) as proxy. From day 5 to day 8 of µSCLS culture, 10 µM Texas Red-labelled dextran was supplemented to dorsal culture medium (basal medium supplemented with BMP4). Fluorescence images covering the whole microfluidic channel were taken at different time points using an inverted epifluorescence microscope (Zeiss Axio Observer Z1; Carl Zeiss MicroImaging). Images were stitched using the Grid/Collection stitching plugin in Fiji 2.16.0. Average fluorescence intensities across the width of each patterning region were obtained using Plot Profile tool in Fiji 2.16.0.
Patterning through single gradient
To assess the patterning efficiencies of µSCLS when patterned through BMP4 gradient alone in Supplementary Fig. 3d, e, 150 ng mL−1 BMP4 was supplemented to the dorsal reservoir from day 5 to 12. The ventral reservoir was filled with basal medium without SAG supplementation. Similarly, to assess the patterning efficiency through SAG gradient alone in Supplementary Fig. 3d, f, 250 nM SAG was supplemented to the ventral reservoir from day 5 to 12 while the dorsal reservoir was filled with basal medium without BMP4 supplementation.
Signaling molecule titration assay
To demonstrate controllability of patterning through titration of signaling molecule concentrations in Supplementary Fig. 3g–i, three conditions were used: Standard, Less SAG, and Less BMP4 condition. For standard condition, from day 5 to 12, 150 ng mL−1 BMP4 was supplemented to the dorsal reservoir and 250 nM SAG was supplemented to the ventral reservoir. For Less SAG condition, the SAG concentration was decreased to 125 nM. For Less BMP4 condition, BMP4 concentration was decreased to 50 ng mL−1.
RA treatment assay
To observe the effect of RA in µSCLS DV patterning in Fig. 3c, d, g, RA (500 nM, STEMCELL Technologies) was added to both dorsal and ventral reservoirs from day 5 to 12, along with 50 ng mL−1 BMP4 in the dorsal reservoir and 250 nM SAG in the ventral reservoir. To compare DV patterning in the absence of BMP4 gradient, from day 5 to 12, the dorsal reservoir was filled with basal media, and the ventral reservoir was filled with basal media supplemented with 250 nM SAG. 500 nM RA was further supplemented to both dorsal and ventral reservoirs from day 5 to 12. To compare DV patterning in the absence of SAG gradient in Supplementary Fig. 6b, c, from day 5 to 12, the ventral reservoir was filled with basal media and the dorsal reservoir was filled with basal media supplemented with 50 ng mL−1 BMP4. Throughout the µSCLS process, the basal medium was switched to N2B27 without vitamin A, which is identical to the standard N2B27 except for the use of B27 without vitamin A (Gibco).
CXCR4 inhibition assay
To inhibit CXCR4, AMD3100 (200 nM, Selleckchem) was supplemented into both reservoirs together with BDNF (10 ng mL−1; R&D Systems), GDNF (10 ng mL−1; PeproTech), CNTF (10 ng mL−1; PeproTech), IGF-1 (10 ng mL−1; PeproTech), cAMP (1 μM; Sigma-Aldrich), and AA (0.2 μg mL−1; Sigma-Aldrich) from day 12 to day 18.
Whole-mount immunofluorescence staining of µSCLS
First, µSCLS were fixed in 4% paraformaldehyde (PFA; buffered in 1× PBS) at room temperature for 1 h 30 min. Then, µSCLS were exposed by manually detaching PDMS structural layers from glass coverslips. Thinly sliced PDMS strips were placed between the PDMS structural layer and glass coverslip to provide space for solutions. The µSCLSs were washed 3 times in PBS for 5 min each. µSCLSs were permeabilized in 0.1% SDS solution (SDS dissolved in PBS) at room temperature for 3 h. After washing 3 times for 5 min in PBS, µSCLS were blocked in 4% donkey serum (Sigma-Aldrich) in PBS at 4 °C for 24 h followed by incubation with primary antibody solutions at 4 °C for another 24 h. µSCLS were washed in PBS 3 times for 5 min and labelled with donkey-raised secondary antibodies (1:400 dilution, Fisher Scientific) at 4 °C for 24 h. Both primary and secondary antibodies were dissolved in 4% donkey serum with 0.2% NaN3. 4′,6-diamidino-2-phenylindole (DAPI, Thermo Fisher Scientific) was used to counterstaining cell nuclei. Antibodies used are listed in Supplementary Table 4.
Whole-mount RNA fluorescence in situ hybridization (RNA-FISH)
Similar to the whole-mount immunofluorescence staining protocol, the µSCLS was first fixed in 4% paraformaldehyde (PFA; buffered in 1× PBS) at room temperature for 1 h 30 min. Then, µSCLS was exposed by manually detaching PDMS structural layers from glass coverslips. Thinly sliced PDMS strips were placed between the PDMS structural layer and glass coverslip to provide space for solutions. The µSCLS was washed 3 times in PBS for 5 min each. µSCLS was permeabilized in methanol in −20 °C for 30 min and in room temperature for 5 min. The µSCLS was washed once with PBSTRG and once with PBSTR. PBSTR consists of PBS with 0.1% Tween-20 and 0.1 U μL−1 Protector RNase Inhibitor (Cat. No. 3335399001; MilliporeSigma), and PBSTRG consists of PBSTR with 0.1 M Glycine. Then, the µSCLS was incubated in probes (Supplementary Table 6; Integrated DNA Technologies) diluted to 4 µM in HCRTM Probe Hybridization Buffer (v3.0; Molecular Instruments) overnight in 37 °C. The µSCLS was washed 4 times, 5 min each, with HCRTM Probe Wash Buffer (v3.0; Molecular Instruments) preheated to 37 °C. The µSCLS was further washed 2 times for 5 min each with 5× SSCT, which consists of 5× sodium chloride sodium citrate buffer with 0.1% Tween-20. Then, the µSCLS was incubated in HCRTM Amplifier Buffer (v3.0; Molecular Instruments) in room temperature for 30 min. In the meantime, HCRTM Amplifiers (v3.0; Molecular Instruments) were snap-cooled for formation of hairpin structure by heating them up to 95 °C for 90 s and cooling down to 12 °C for 30 min. The snap-cooled HCRTM Amplifiers were mixed in HCRTM Amplifier Buffer at 1/100 (v/v) dilution right before use. The µSCLS was incubated in the HCRTM Amplifiers mixed in the HCRTM Amplifier Buffer for 90 min in room temperature, avoiding light. Then, the µSCLS was washed with 5XSSCT 3 times, 5 min each. Nuclei of µSCLS was stained with 4′,6-diamidino-2-phenylindole (DAPI, Thermo Fisher Scientific) by incubating it in DAPI diluted in 5XSSCT at a ratio of 1/400 (v/v) overnight at 4 °C, avoiding light. The µSCLS was finally washed with 5XSSCT 3 times, 5 min each, before proceeding to tissue clearing.
Tissue clearing
For downstream microscopy after whole-mount immunofluorescence staining or whole-mount RNA-FISH, µSCLS were optically cleared by incubating in a refractive index (RI)-matching solution. The solution comprised of 6.3 mL ddH2O, 9.2 ml OptiPrep Density Gradient medium (MilliporeSigma), 4 × g N-methyl-d-glucamine (MilliporeSigma), and 5 × g diatrizoic acid (MilliporeSigma)66. 10 µL of RI-matching solution was applied onto the µSCLS, and PDMS structure layer was attached back to the glass coverslip. The re-assembled sample was incubated in RT for at least 30 min before imaging.
EdU cell-proliferacion assay
Cell proliferation was assessed using the Click-iT EdU Alexa Fluor 488 Imaging Kit (Thermo Fisher Scientific), following the manufacturer’s protocol. On days 5 and 12, PDMS structural layers were manually detached from the glass coverslips, exposing the µSCLSs that remained on the PDMS surfaces. The µSCLSs were then incubated for 1 h in basal medium containing 20 μM EdU, followed by fixation, permeabilization, and incubation with the Click-iT reaction cocktail for 30 min. Afterward, samples were blocked with 4% donkey serum in PBS and incubated with primary antibodies. DAPI was used to counterstain cell nuclei. EdU-labeled proliferating cells were visualized using confocal microscopy.
Fluorescence intensity maps
To generate heatmaps in Fig. 1g and Supplementary Fig. 3g–i, a custom MATLAB (R2024b) script was developed to process fluorescence images of lineage markers and generate fluorescence intensity maps. First, stitched images of µSCLS were manually rotated to align the dorsoventral (DV) axis vertically. Binary masks were then generated from DAPI images by selecting pixels with intensities above 20% of the maximum signal, effectively outlining the µSCLS contours. Pixel intensities outside the DAPI mask were automatically set to zero. The remaining pixel intensities were normalized to the maximum intensity within each image to produce normalized fluorescence intensity maps. Normalized signal values were then averaged across the thickness of the tissue to generate a one-dimensional intensity profile along its length. PAX6 staining images had high cytoplasmic signals at the ventral tip. This led to misleading expression profiles with high expression at the ventral end, where the thickness of the tissues taper off. Thus, normalized PAX6 staining signals were summed up, instead of being averaged, across the thickness of the tissue. Because the tissue lengths varied across samples, each profile was interpolated to 1000 equidistant points using the MATLAB function interp1. Each interpolated intensity profile was subsequently normalized by its minimum and maximum values. Finally, normalized profiles from different tissues were averaged to produce an average normalized intensity heatmap.
To generate heatmaps in Supplementary Fig. 1j showing cell cycle markers along the apical-basal axis, confocal micrograph of individual µSCLS was cut in half along the lumen. The lumen side was marked as apical and the exterior as basal. Fluorescence intensity maps were generated similarly and was averaged over the length of the tissue. The resultant expression profile along the apical-basal axis was interpolated to 1000 equidistant points using the MATLAB function interp1. Each interpolated intensity profiles were normalized by its minimum and maximum values and averaged across different tissues.
Boundary determination
To determine the expression domain boundaries, the intensity profile of each marker obtained as described in “Fluorescence intensity maps” section was first filtered using a moving average filter with a window size of 50 to smooth out local fluctuations. Then, the point where the expression value becomes 50% of the maximum was determined as the boundary of expression domain.
scRNA-seq
µSCLS was dissociated into single cells by manually detaching the PDMS structural layers and incubating the released µSCLS in Accutase for 2 h. The resulting cell suspension was collected into PBS containing 0.5% BSA and centrifuged at 200 × g for 5 min. Cell pellet was resuspended in PBS with 0.5% BSA and filtered through a 40 μm cell strainer (pluriSelect USA, 43-50040-51) to remove debris and aggregates. Within 1 h of dissociation, cells were loaded into a 10× Genomics Chromium system (10× Genomics). 10× Genomics v.3 libraries were prepared following manufacturer’s instructions. Then, libraries were sequenced using paired-end sequencing with a minimum coverage of 20,000 raw reads per cell using an Illumina NovaSeqXPlus. scRNA-seq data were aligned and quantified using Cell Ranger Single-Cell software suite (v.7.1.0, 10× Genomics) against Homo sapiens (human) genome assembly GRCh38 (version refdata-gex-GRCh38-2020-A, 10× Genomics).
Analysis of scRNA-seq data
scRNA-seq data analysis were conducted using the R (ver. 4.2.2.) package Seurat (v.5.1.0, https://satijalab.org/seurat/)67. Unless noted otherwise, default setups in Seurat were used. Briefly, scRNA-seq dataset was filtered based on the total number of genes detected and the percentage of mitochondrial genes. Raw counts were normalized by the total counts, multiplied by 10,000, and log-transformed. To regress out cell cycle information, cell cycle scores were first calculated using CellCycle Scoring function. Then, the cell cycle scores were regressed out during the data-scaling process through ScaleData. For dimensional reduction, top 2000 variable genes were first identified using FindVariableFeatures. Identified variable genes were used for PCA analysis using RunPCA. The principal components were used for embedding the data into low-dimensional space with uniform manifold approximation and projection algorithm using RunUMAP. Neighboring cells were identified in the principal component space using FindNeighbors and clusters were identified by performing a shared nearest neighbor modularity optimization-based clustering algorithm using FindClusters. Differentially expressed genes (DEGs) in each cluster were identified using FindAllMarkers. Dot plots in Fig. 2b and Supplementary Figs. 4g and 5e were generated using DotPlot. Heatmap in Supplementary Fig. 4a was generated using DoHeatmap. Feature plots in Fig. 4l and Supplementary Fig. 4c were generated using FeaturePlot. Heatmap in Supplementary Fig. 4b was generated by first averaging the scaled expression levels of each gene in each cluster and using ggplot in conjunction with geom_tile of the R package ggplot2. Heatmap in Supplementary Fig. 4e was generated by first averaging the mRNA counts of each HOX gene in each cluster and using pheatmap function of the R package pheatmap.
Comparison with human embryo data
To deduce the developmental stage of µSCLS, PCA analysis was done on average counts of each transcript. Then, Euclidean distances were calculated in the principal component space between µSCLS and human SCs from different developmental stages22. To compare the transcriptome of µSCLS and stage-matched human SC in Fig. 2d, the development systems “Neural crest progenitor”, “Neuron”, and “Progenitor” were isolated from CS12 SC dataset. “Peripheral neuron”, corresponding to the DRG population, was obtained from CS14 SC dataset because of its scarcity in the CS12 SC dataset. For comparison of NE clusters in Fig. 2f, g and Supplementary Fig. 4f, g, clusters “RP”, “dP1”, “dP2”, “dP3”, “dP4”, “dP5”, “dP6”, “p0”, “p1”, “p2”, “pMN”, “p3”, “FP” were isolated from human CS12 SC dataset. For comparison of neuron clusters in Supplementary Fig. 5, “Neuron” cluster was isolated from human CS12 SC dataset. The datasets were integrated using a reciprocal PCA approach based on 30 dimensions and k.anchor = 20 through IntegrateLayers. Annotations were based on their original annotation before integration. Pearson’s correlations between cell clusters in Fig. 2e, g and Supplementary Fig. 5d were calculated using the cor function in the R package Stats based on their integrated principal components averaged over each gene in each cluster. Red and gray feature plots marking subtype localization in UMAP plots in Supplementary Figs. 4f and 5b were generated using FeaturePlot. Color-graded feature plots showing average expression of marker genes in Supplementary Figs. 4f and 5b were generated by first averaging the scaled expression of marker genes used to identify each subtype (Supplementary Table 2) and using FeaturePlot. Heatmap in Fig. 2i was generated by averaging the scaled expression levels of each gene and using ggplot in conjunction with geom_tile of the R package ggplot2. Violin plots in Fig. 2l were generated by first isolating NE cells co-expressing OLIG2 and NKX2.2, determined by a threshold of one UMI count, and using VlnPlot.
NE and neuron subtype partitioning
Partitioning of NE and neuron population into subtypes was done as previously reported22,45,46. First, expression levels of marker genes were binarized with a threshold of one UMI count. Then, Euclidean distance from a cell to each ideal subtype expressing all the marker genes was calculated in the binary space. The cell was labeled as the closest subtype. Markers used are listed in Supplementary Table 2.
Re-analysis of mouse data
Raw count matrices and metadata of scRNA-seq data from mouse embryos at E9.5 were re-analyzed for Fig. 2j, k. NE subtypes “RP“, “dP1”, “dP2”, “dP3”, “dP4”, “dP5”, “dP6”, “p0”, “p1”, “p2”, “pMN”, “p3”, “FP” were isolated from mouse E9.5 SC dataset46. In Fig. 2j, raw counts were averaged within each NE subtypes and normalized by maximum mean expression. In Fig. 2k, expression of NKX2.2 and OLIG2 were determined by a threshold of one UMI count.
Re-analysis of Chip seq
Table of aligned peaks from retinoic acid receptor chromatin immunoprecipitation sequencing experiment done on motor neurons differentiated from mouse embryonic stem cells with or without retinoic acid was reanalyzed52. Among the list of genes closest to peaks, BMP signaling pathway regulatory genes were identified. List of BMP signaling pathway regulatory genes was found from the Molecular Signature Database (MSigDB)68,69,70 (Supplementary Table 3).
Tissue area and length
To measure projected area and length of tissues in Fig. 1e, brightfield images of µSCLS at different days were first rotated so that the DV axis was aligned with the vertical axis. Then, tissues were manually contoured using freehand selection tool in Fiji 2.16.0. The height and area in pixels from Fiji measurements were converted to length and projected area in µm and µm2.
Cell number and position measurement
To specify position and number of NCCs in Fig. 4, or that of TWIST1+ mesodermal cells in Supplementary Fig. 4d, the fluorescent images were first binarized using a threshold. Then, adjoining cells were separated using watershed process in Fiji 2.16.0. The locations of separated cells were identified using analyze particle functionality in Fiji 2.16.0. Due to the high cell density in the NE, proportion of cells in Fig. 3f was estimated using proportion of pixel numbers. First, DAPI image was binarized by using Otsu’s thresholding. The positive pixels were counted and further used as a mask for fluorescence images of OLIG2 and FOXA2. After DAPI masking, OLIG2 and FOXA2 images were binarized by using Otsu’s thresholding and positive pixels were counted. Proportion of cells positive for OLIG2 was estimated by the proportion of OLIG2-positive pixels with to the number of DAPI-positive pixels.
Nuclei long axis angle quantification
To measure the angle of long axes of nuclei from DAPI images in Supplementary Fig. 1f, the DAPI images were first binarized by setting a threshold. Then, adjoining nuclei were separated using watershed process in Fiji 2.16.0. Each nucleus was fitted to an ellipse using analyze particle functionality in Fiji 2.16.0. The long axis of the ellipse was used to estimate the long axis of the nucleus.
Western blot
First, µSCLSs were dissociated into single cells. To do so, PDMS structural layer was manually detached from glass coverslips to release µSCLS, and released µSCLSs were incubated with Accutase for 20 min. Then, pellets were centrifuged and lysed for 10 min using ice cold radioimmunoprecipitation assay (RIPA) buffer supplemented with 1× protease and phosphatase inhibitor cocktail (Thermo Scientific). The lysates were vortexed every 2 min during lysis. The lysates were then centrifuged at maximum speed for 15 min. Total protein concentrations were measured using BCA Protein Assay Kit following manufacturer’s instructions (Thermo Scientific). 20 µg of total protein was loaded in each pocket of SDS-polyacrylamide gel electrophoresis gel (BioRad) along with Precision Plus Protein Dual Color Standards (BioRad). The gel was then transferred onto a polyvinylidene difluoride membrane. The membrane was blocked in 5% BSA (w/v) dissolved in TBS with 0.1% Tween (v/v) (TBS-T) for 2 h, washed three times in TBS-T for 5 min, and incubated with primary antibody in 5% BSA in TBS-T at 4 °C overnight on a rotator. On the next day, the membrane was washed three times in TBS-T for 5 min and incubated with a horseradish peroxidase (HRP) antibody in 5% BSA in TBS-T at 4 °C for 2 h on a rotator. The membrane was then washed three times in PBS-T for 5 min. Chemiluminescence signal was detected using ECL Western Blotting Detection Reagent (Cat. No. 32106; Thermo Scientific). Antibodies used are listed in Supplementary Table 4. Uncropped, unprocessed scan of representative blot is presented in Supplementary Fig. 8.
Quantitative polymerase chain reaction (qPCR)
To quantify gene expression in Supplementary Fig. 6d, h, we performed qPCR. First, µSCLS were lysed using Lysis Buffer RLT from RNeasy Mini Kit (Cat. No. 74104; Qiagen). Three tissues in one device were lysed together. Then, the lysates were homogenized using QIAshredder (Cat. No. 79654; Qiagen). RNA was extracted from the homogenized lysates using RNeasy Mini Kit (Cat. No. 74104; Qiagen) following manufacturer’s instructions. The RNA concentrations of resultant solutions were measured using a spectrophotometer (NanoDropTM One/OneC; Thermo Scientific). Then, the solutions were diluted in nuclease-free water to equalize the RNA concentrations across samples within the same batch. Complementary DNA (cDNA) was synthesized from the RNA solutions in a thermal cycler (T100; Bio-Rad Laboratories, Inc.) using iScriptTM cDNA Synthesis Kit (Cat. No. 1708890; Bio-Rad Laboratories, Inc.) following manufacturer’s instructions. Then, qPCR was performed on CFX Connect Real-Time PCR Detection System (Bio-Rad Laboratories, Inc.) by loading mixtures of the cDNA solutions, primers (Supplementary Table 5; Integrated DNA Technologies), and iTaq Universal SYBR Green Supermix (Cat. No. 1725121; Bio-Rad Laboratories, Inc.). The ratio and running conditions were set according to manufacturer’s instructions of iTaq Universal SYBR Green Supermix (Cat. No. 1725121; Bio-Rad Laboratories, Inc.). qPCR measurement was repeated two times for technical repeat. The quantification cycle (Cq) values were used for quantifying the initial mRNA concentrations. First, the Cq values from two technical repeats were averaged. Then, the initial mRNA amount was calculated as \({2}^{-{C}_{q}}\). Then, the mRNA amount was normalized by the geometric mean of two housekeeping genes, GAPDH and ACTB. The mRNA expression levels normalized by housekeeping genes were further normalized by expression levels in hESCs.
Knock down assay using small interfering RNA (siRNA)
Differentially expressed genes identified in Supplementary Fig. 6d were knocked down using siRNA (Supplementary Fig. 6e–g and Supplementary Table 7). Pre-designed siRNAs were purchased from Integrated DNA Technologies. Three siRNAs were provided per gene. Individual siRNAs were diluted in Nuclease-Free Duplex Buffer (Integrated DNA Technologies) to a concentration of 10 µM. Equal volumes of diluted siRNAs targeting the same gene were mixed. The siRNA mixture was further combined with LipofectamineTM RNAiMAX Transfection Reagent (Cat. No. 13778030; Thermo Fisher Scientific) and Opti-MEMTM (Cat. No. 31985062; Thermo Fisher Scientific) at a volumetric ration of 1:2:50. The mixture was incubated at room temperature for 5 min before being added to culture media, adjusting the final concentration of siRNA in media to be 30 nM. Note that the 30 nM was the total concentration of three different siRNAs targeting the same gene. siRNAs were added to culture media on days 5, 7, 9, and 11.
Dorsally and ventrally located cells
Dorsally located and ventrally located cells in Fig. 4 were determined by the cells’ DV location relative to the NE. Location of the dorsal and ventral poles of three tissues in one device were averaged to produce an averaged dorsal pole and an averaged ventral pole. The DV location of averaged ventral pole was defined as 0 and that of the averaged dorsal pole was defined as 1. Then, the locations of individual cells were projected onto the DV axis defined by averaged dorsal and ventral poles. Cells with DV positions of greater than 1 were defined as dorsally located and cells with DV position of less than 0.5 were defined as ventrally located.
Migration tracking
To track the migration of NCCs, µSCLSs were fabricated with SOX10::T2A-Cre lineage reporter hESC line. On day 12 and onward, the samples were observed daily for emergence of cells expressing ZsGreen with flattened morphology, indicating migrating NCC. When such cell was identified, the sample was imaged every 20 min using an inverted epifluorescence microscope (Zeiss Axio Observer Z1; Carl Zeiss MicroImaging) enclosed in an environmental incubator (XL S1 incubator, Carl Zeiss MicroImaging), maintaining the cell culture conditions at 37 °C and 5% CO2. Then, the migration of individual NCCs were tracked using the Fiji plugin TrackMate. A LoG detector was used on the ZsGreen images with estimated object diameter of 25 µm and quality threshold of 0.05 to locate the NCCs. Then, migration was tracked using a simple LAP tracker with linking max distance of 64 pixels, gap-closing max distance of 64 pixels, and gap-closing max frame gap of 2. The polar histogram in Fig. 4e was generated by first calculating the migratory direction. Migratory direction of each cell was represented by the angle between the displacement vector connecting the start and end point of a NCC and the DV axis. Then, the angles of all NCCs were pooled into a polar histogram using a MATLAB function polarhistogram.
Mean square displacement
Mean square displacement at each timepoint t in Fig. 4g was calculated as below:
Where \(N(t)\) is the number of cells tracked until time t, \({{{\bf{x}}}}^{\left(i\right)}\left(t\right)\) is the position vector of cell i at time t, and \({{{\bf{x}}}}^{\left(i\right)}\left(0\right)\) is the position vector of cell i at time 0.
Random walk simulation
A 2-dimensional random walk simulation was performed using a custom MATLAB script. In the simulation, a particle randomly took a step in either of 4 orthogonal directions in the x-y plane, namely, −x, +x, −y, and +y directions. The number of particles were set to 120, which was the average number of tracks of migrating NCCs in each experimental sample. To determine the step size, the mean square displacement (MSD) of NCCs were fitted to a parabolic curve with the constant term set to 071:
Then, considering that images were taken every 1/3 h, step size was determined as below to match the diffusion coefficient observed in migrating NCCs:
The simulation was performed 5 times.
Quantification of directionality and circular standard deviation
Directionality was quantified by dividing the distance between initial and final position of a cell by the total length of the cell’s migratory path. To calculate the circular standard deviation of migratory directions72, First, the mean resultant length \(\bar{R}\) was calculated as below:
Here,
Then, the circular standard deviation was calculated as:
Statistical analysis
All experiments were conducted in at least three biological replicates, except scRNA-seq assays, which were performed in one independent experiment. Sample sizes are indicated in the figure legends. For comparisons between two datasets, P values were calculated using two-sided Student’s t-test. For analysis of significance of SMAD1/5 complex phosphorylation level increase in Fig. 3g, a one-sample t-test with a hypothesized mean of 1. For comparison of zero-inflated data that does not follow normal distribution in Fig. 4j, n, a zero-inflated rank test was used73. For comparison of gene expression levels across more than two conditions in Supplementary Fig. 6h, a one-way ANOVA followed by a post-hoc Tukey’s HSD test were performed. No statistical methods were used to predetermine sample size. Samples were randomly allocated to different experimental groups. Investigators were not blinded to allocation during experiments and outcome assessment. All statistical analyses were performed using R 4.4.0.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data supporting findings of this study are available within the article and its Supplementary Information files and from the corresponding authors upon request. The scRNA-seq data generated in this study have been deposited in the Gene Expression Omnibus under accession number GSE300459. All source data for graphs included in the paper are available in the online version of the paper. Source data are provided with this paper.
Code availability
R, Python, and MATLAB scripts used in this work are available from the lead contact upon reasonable request.
References
Gouti, M., Metzis, V. & Briscoe, J. The route to spinal cord cell types: a tale of signals and switches. Trends Genet. 31, 282–289 (2015).
Sagner, A. & Briscoe, J. Establishing neuronal diversity in the spinal cord: a time and a place. Development 146, dev182154 (2019).
Andrews, M. G., Kong, J., Novitch, B. G. & Butler, S. J. New perspectives on the mechanisms establishing the dorsal-ventral axis of the spinal cord. Curr. Top. Dev. Biol. 132, 417 (2019).
Newbern, J. M. Molecular control of the neural crest and peripheral nervous system development. Curr. Top. Dev. Biol. 111, 201 (2015).
Barth, K. A. et al. Bmp activity establishes a gradient of positional information throughout the entire neural plate. Development 126, 4977–4987 (1999).
Yu, W., McDonnell, K., Taketo, M. M. & Bai, C. B. Wnt signaling determines ventral spinal cord cell fates in a time-dependent manner. Development 135, 3687–3696 (2008).
Liem, K. F., Tremml, G., Roelink, H. & Jessell, T. M. Dorsal differentiation of neural plate cells induced by BMP-mediated signals from epidermal ectoderm. Cell 82, 969–979 (1995).
Zagorski, M. et al. Decoding of position in the developing neural tube from antiparallel morphogen gradients. Science 356, 1379–1383 (2017).
Briscoe, J. & Small, S. Morphogen rules: design principles of gradient-mediated embryo patterning. Development 142, 3996–4009 (2015).
Chamberlain, C. E., Jeong, J., Guo, C., Allen, B. L. & McMahon, A. P. Notochord-derived Shh concentrates in close association with the apically positioned basal body in neural target cells and forms a dynamic gradient during neural patterning. Development 135, 1097–1106 (2008).
Rekler, D., Ofek, S., Kagan, S., Friedlander, G. & Kalcheim, C. Retinoic acid, an essential component of the roof plate organizer, promotes the spatio-temporal segregation of dorsal neural fates. Development https://doi.org/10.1242/dev.202973 (2024).
Novitch, B. G., Wichterle, H., Jessell, T. M. & Sockanathan, S. A requirement for retinoic acid-mediated transcriptional activation in ventral neural patterning and motor neuron specification. Neuron 40, 81–95 (2003).
Diez del Corral, R. & Morales, A. V. Retinoic acid signaling during early spinal cord development. J. Dev. Biol. 2, 174–197 (2014).
Wilson, L., Gale, E., Chambers, D. & Maden, M. Retinoic acid and the control of dorsoventral patterning in the avian spinal cord. Dev. Biol. 269, 433–446 (2004).
Soldatov, R. et al. Spatiotemporal structure of cell fate decisions in murine neural crest. Science 364, eaas9536 (2019).
Furlan, A. et al. Multipotent peripheral glial cells generate neuroendocrine cells of the adrenal medulla. Science 357, eaal3753 (2017).
Morikawa, Y. et al. BMP signaling regulates sympathetic nervous system development through Smad4-dependent and -independent pathways. Development 136, 3575 (2009).
Saito, D., Takase, Y., Murai, H. & Takahashi, Y. The dorsal aorta initiates a molecular cascade that instructs sympatho-adrenal specification. Science 336, 1578–1581 (2012).
Belmadani, A. et al. The chemokine stromal cell-derived factor-1 regulates the migration of sensory neuron progenitors. J. Neurosci. 25, 3995 (2005).
Kasemeier-Kulesa, J. C., McLennan, R., Romine, M. H., Kulesa, P. M. & Lefcort, F. CXCR4 controls ventral migration of sympathetic precursor cells. J. Neurosci. 30, 13078–13088 (2010).
Marklund, U. et al. Detailed expression analysis of regulatory genes in the early developing human neural tube. Stem Cells Dev. 23, 5–15 (2014).
Rayon, T., Maizels, R. J., Barrington, C. & Briscoe, J. Single-cell transcriptome profiling of the human developing spinal cord reveals a conserved genetic programme with human-specific features. Development 148, dev199711 (2021).
Curtis, E. et al. A first-in-human, phase I study of neural stem cell transplantation for chronic spinal cord injury. Cell Stem Cell 22, 941–950.e6 (2018).
Martin, J. R. et al. Long-term clinical and safety outcomes from a single-site phase 1 study of neural stem cell transplantation for chronic thoracic spinal cord injury. Cell Rep. Med. 5, 101841 (2024).
Bydon, M. et al. Intrathecal delivery of adipose-derived mesenchymal stem cells in traumatic spinal cord injury: Phase I trial. Nat. Commun. 15, 1–11 (2024).
Zheng, Y. et al. Dorsal-ventral patterned neural cyst from human pluripotent stem cells in a neurogenic niche. Sci. Adv. 5, 5933–5944 (2019).
Meinhardt, A. et al. 3D reconstitution of the patterned neural tube from embryonic stem cells. Stem Cell Rep. 3, 987–999 (2014).
Ranga, A. et al. Neural tube morphogenesis in synthetic 3D microenvironments. Proc. Natl. Acad. Sci. Usa. 113, E6831–E6839 (2016).
Ogura, T., Sakaguchi, H., Miyamoto, S. & Takahashi, J. Three-dimensional induction of dorsal, intermediate and ventral spinal cord tissues from human pluripotent stem cells. Development 145, dev162214 (2018).
Rifes, P. et al. Modeling neural tube development by differentiation of human embryonic stem cells in a microfluidic WNT gradient. Nat. Biotechnol. 38, 1265–1273 (2020).
Demers, C. J. et al. Development-on-chip: In vitro neural tube patterning with a microfluidic device. Dev. (Camb.) 143, 1884–1892 (2016).
Xue, X. et al. A patterned human neural tube model using microfluidic gradients. Nature 628, 391–399 (2024).
Luo, T. et al. Establishing dorsal-ventral patterning in human neural tube organoids with synthetic organizers. Cell Stem Cell 0, 1–16 (2025).
Lippmann, E. S. et al. Deterministic HOX patterning in human pluripotent stem cell-derived neuroectoderm. Stem Cell Rep. 4, 632–644 (2015).
Buckley, D. M., Burroughs-Garcia, J., Lewandoski, M. & Waters, S. T. Characterization of the Gbx1−/− mouse mutant: a requirement for Gbx1 in normal locomotion and sensorimotor circuit development. PLoS ONE 8, e56214 (2013).
Rondon, R. et al. Dual transcriptional activities of PAX3 and PAX7 spatially encode spinal cell fates through distinct gene networks. PLoS Biol. 23, e3003448 (2025).
Briscoe, J., Pierani, A., Jessell, T. M. & Ericson, J. A Qhomeodomain protein code specifies progenitor cell identity and neuronal fate in the ventral neural tube. Cell 101, 435–445 (2000).
Jessell, T. M. Neuronal specification in the spinal cord: inductive signals and transcriptional codes. Nat. Rev. Genet. 1, 20–29 (2000).
Lek, M. et al. A homeodomain feedback circuit underlies step-function interpretation of a Shh morphogen gradient during ventral neural patterning. Development 137, 4051–4060 (2010).
Schoenwolf, G. C., Bleyl, S. B., Brauer, P. R., Francis-West, P. H. Larsen’s Human Embryology (Elsevier, Philadelphia, 2021).
Lee, S. K., Lee, B., Ruiz, E. C. & Pfaff, S. L. Olig2 and Ngn2 function in opposition to modulate gene expression in motor neuron progenitor cells. Genes Dev. 19, 282 (2005).
Tani, S., Chung, U. il, Ohba, S. & Hojo, H. Understanding paraxial mesoderm development and sclerotome specification for skeletal repair. Exp. Mol. Med. 52, 1166–1177 (2020).
McMahon, J. A. et al. Noggin-mediated antagonism of BMP signaling is required for growth and patterning of the neural tube and somite. Genes Dev. 12, 1438–1452 (1998).
Cairns, D. M., Sato, M. E., Lee, P. G., Lassar, A. B. & Zeng, L. A gradient of Shh establishes mutually repressing somitic cell fates induced by Nkx3.2 and Pax3. Dev. Biol. 323, 152–165 (2008).
Delile, J. et al. Single cell transcriptomics reveals spatial and temporal dynamics of gene expression in the developing mouse spinal cord. Development 146, dev173807 (2019).
Xu, Y. et al. A single-cell transcriptome atlas profiles early organogenesis in human embryos. Nat. Cell Biol. 25, 604–615 (2023).
Florio, M. et al. Evolution and cell-type specificity of human-specific genes preferentially expressed in progenitors of fetal neocortex. Elife 7, e32332 (2018).
Liu, J. et al. The primate-specific gene TMEM14B marks outer radial glia cells and promotes cortical expansion and folding. Cell Stem Cell 21, 635–649.e8 (2017).
Vallstedt, A. et al. Different levels of repressor activity assign redundant and specific roles to Nkx6 genes in motor neuron and interneuron specification. Neuron 31, 743–755 (2001).
Calder, E. L. et al. Retinoic acid-mediated regulation of GLI3 enables efficient motoneuron derivation from human ESCs in the absence of extrinsic SHH activation. J. Neurosci. 35, 11462 (2015).
Gupta, S. et al. Deriving dorsal spinal sensory interneurons from human pluripotent stem cells. Stem Cell Rep. 10, 390 (2018).
Mahony, S. et al. Ligand-dependent dynamics of retinoic acid receptor binding during early neurogenesis. Genome Biol. 12, 1–15 (2011).
Delás, M. J. et al. Developmental cell fate choice in neural tube progenitors employs two distinct cis-regulatory strategies. Dev. Cell https://doi.org/10.1016/J.DEVCEL.2022.11.016 (2022).
Levine, A. J. & Brivanlou, A. H. GDF3, a BMP inhibitor, regulates cell fate in stem cells and early embryos. Development 133, 209–216 (2006).
Levine, A. J., Levine, Z. J. & Brivanlou, A. H. GDF3 is a BMP inhibitor that can activate Nodal signaling only at very high doses. Dev. Biol. 325, 43 (2008).
Lee, V. M. et al. Molecular events controlling cessation of trunk neural crest migration and onset of differentiation. Front. Cell Dev. Biol. 8, 506999 (2020).
Andrews, M. G. et al. BMPs direct sensory interneuron identity in the developing spinal cord using signal-specific not morphogenic activities. Elife 6, e30647 (2017).
Kalcheim, C. & Teillet, M. A. Consequences of somite manipulation on the pattern of dorsal root ganglion development. Development 106, 85–93 (1989).
Nitzan, E. & Kalcheim, C. Neural crest and somitic mesoderm as paradigms to investigate cell fate decisions during development. Dev. Growth Differ. 55, 60–78 (2013).
Karzbrun, E., Kshirsagar, A., Cohen, S. R., Hanna, J. H. & Reiner, O. Human brain organoids on a chip reveal the physics of folding. Nat. Phys. 14, 515–522 (2018).
Shibata, S. Modeling embryo-endometrial interface recapitulating human embryo implantation. Sci. Adv. 10, eadi4819 (2024).
Li, X. et al. Desktop aligner for fabrication of multilayer microfluidic devices. Rev. Sci. Instrum. 86, 75008 (2015).
Watanabe, K. et al. A ROCK inhibitor permits survival of dissociated human embryonic stem cells. Nat. Biotechnol. 25, 681–686 (2007).
Peng, T. et al. A BaSiC tool for background and shading correction of optical microscopy images. Nat. Commun. 8, 14836 (2017).
Chalfoun, J. et al. MIST: accurate and scalable microscopy image stitching tool with stage modeling and error minimization. Sci. Rep. 7, 4988 (2017).
Serra, D. et al. Self-organization and symmetry breaking in intestinal organoid development. Nature 569, 66–72 (2019).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2023).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Liberzon, A. et al. The molecular signatures database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417 (2015).
Codling, E. A., Plank, M. J. & Benhamou, S. Random walk models in biology. J. R. Soc. Interface 5, 813–834 (2008).
Mardia, K. V. & Jupp, P. E. Directional Statistics 1–432 https://doi.org/10.1002/9780470316979. (2008).
Wang, W., Chen, E. & Li, H. Truncated rank-based tests for two-part models with excessive zeros and applications to microbiome data. Ann. Appl. Stat. 17, 1663–1680 (2021).
Acknowledgements
We thank Dr. O. Reiner for providing us the Lifeact-GFP H2B-mCherry WIBR3 hESC line. This study is supported by the Michigan-Cambridge Collaboration Initiative, University of Michigan Mcubed Fund, 21st Century Jobs Trust Fund received through the Michigan Strategic Fund from the State of Michigan (Grant CASE-315037), University of Michigan Mid-career Biosciences Faculty Achievement Recognition Award, National Science Foundation of the United States (CBET 1901718, PFI 2213845, and EFMA 2422149), National Institutes of Health of the United States (R21 NS127983, R01 GM143297, and R01 NS129850), and US-Israel Binational Science Foundation (US-Israel BSF 2023009). We acknowledge the Michigan Medicine Microscopy Core for training and support in microscopy imaging, the Michigan Advanced Genomics Core for scRNA-seq service, and the Michigan Lurie Nanofabrication Facility for support in microfabrication.
Author information
Authors and Affiliations
Contributions
J.B. and J.F. conceived and initiated this project; J.B. designed, performed, and quantified most experiments, including scRNA-seq data analysis and interpretation; Y.S.K. generated SOX10::T2A-Cre lineage tracing hESC line; F.C., C.G., and S.S. helped repeat experiments; N.K. generated cytoplasmic EGFP-expressing hESC line; Z.Z. helped with CHIP-seq data analysis; A.T. quantified µSCLS growth; X.X. developed MATLAB scripts for image processing; D.H.N designed probes for RNA-FISH; P.L. helped with data interpretation and experimental designs; J.B. and J.F. wrote manuscript. J.F. supervised the study. All authors edited and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Yuchuan Miao and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bok, J., Kim, Y.S., Cheng, F. et al. A controllable human spinal cord model with full dorsoventral patterning. Nat Commun 17, 4539 (2026). https://doi.org/10.1038/s41467-026-71162-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-71162-z