Abstract
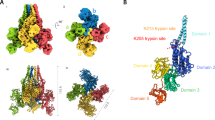
Insecticidal toxins from Bacillus thuringiensis (Bt) have been extensively and successfully used in genetically engineered crops for decades but continue to face challenges from the adaptive resistance in the insect population. The Bt binary toxin Vip1Ad1(Vip1) and Vip2Ag1(Vip2), a promising next-generation candidate gene combination for transgenic crops, have demonstrated high efficacy against the destructive coleopteran pest white grubs, however, their mode of action remains largely elusive. In this study, we report cryo-EM structures of the heptameric Vip1-pore and Vip2-bound Vip1-pore complex, capturing a series of putative assembly-related intermediates that suggest a binary toxin assembly and translocation pathway. Together with structure-guided mutagenesis, these data provide insights into a sequential assembly of binary complex and a sequence-independent translocation mechanism. Proof-of-principle experiments showed successful delivery of a desired protein cargo into host cells based on the mini-Vip2-Vip1 pore system, paving the way for developing much needed extracellular pesticidal protein delivery platforms. These findings not only clarify the assembly and translocation mechanism of the binary insecticidal toxin pair but also offer an excellent alternative model to investigate human-pathogenic pore-forming toxins because of its similarity and biosafety.
Similar content being viewed by others
Data availability
Cryo-EM maps were deposited to the Electron Microscopy Data Bank (https://www.ebi.ac.uk/emdb/) under accession codes EMD-65010 (Vip1 heptameric pore), EMD-65009 (Vip2 bound heptameric pore complex), EMD-65007 (1:5 (Vip2:Vip1) complex). The corresponding coordinates were deposited to the Protein Data Bank (https://www.rcsb.org/) under accession codes 9VEK (Vip1 heptameric pore), 9VEJ (Vip2 bound heptameric pore complex), 9VEG (1:5 (Vip2:Vip1) complex). All other published PDB codes cited in this paper are 6SMS, 7YVQ and 1QS1. All data generated or analyzed during this study are included in this article. Source data are provided as a Source Data file. Source data are provided with this paper.
References
Pardo-López, L., Soberón, M. & Bravo, A. Bacillus thuringiensis insecticidal three-domain Cry toxins: mode of action, insect resistance and consequences for crop protection. FEMS Microbiol. Rev. 37, 3–22 (2013).
Chakroun, M., Banyuls, N., Bel, Y., Escriche, B. & Ferré, J. Bacterial vegetative insecticidal proteins (Vip) from entomopathogenic bacteria. Microbiol. Mol. Biol. Rev. 80, 329–350 (2016).
Li, J., Carroll, J. & Ellar, D. J. Crystal structure of insecticidal δ-endotoxin from Bacillus thuringiensis at 2.5 Å resolution. Nature 353, 815–821 (1991).
Badran, A. H. et al. Continuous evolution of Bacillus thuringiensis toxins overcomes insect resistance. Nature 533, 58–63 (2016).
Soberón, M. et al. Engineering modified bt toxins to counter insect resistance. Science 318, 1640–1642 (2007).
Ye, Z. et al. Too much of a good thing: Lessons from compromised rootworm Bt maize in the US Corn Belt. Science 387, 984–989 (2025).
Warren, G.W. Advances in Insect Control: the Role of Transgenic Plants (eds. Carozzi, N.B. & Koziel, M.G.), 109–121 (CRC Press, 1997).
Syed, T. et al. Current insights on vegetative insecticidal proteins (Vip) as next generation pest killers. Toxins 12, 522 (2020).
Han, S., Craig, J. A., Putnam, C. D., Carozzi, N. B. & Tainer, J. A. Evolution and mechanism from structures of an ADP-ribosylating toxin and NAD complex. Nat. Struct. Biol. 6, 932–936 (1999).
Crickmore, N. et al. Revision of the nomenclature for the Bacillus thuringiensis pesticidal crystal proteins. Microbiol. Mol. Biol. Rev. 62, 807–813 (1998).
Bi, Y. et al. Genomic sequencing identifies novel Bacillus thuringiensis Vip1/Vip2 binary and Cry8 toxins that have high toxicity to Scarabaeoidea larvae. Appl. Microbiol. Biotechnol. 99, 753–760 (2015).
Suzuki, T. et al. Editorial: plant-pest interactions volume III: coleoptera and lepidoptera. Front. Plant Sci. 12, 730290 (2021).
Gibert, M. et al. Differential requirement for the translocation of clostridial binary toxins: Iota toxin requires a membrane potential gradient. FEBS Lett. 581, 1287–1296 (2007).
Knapp, O., Benz, R. & Popoff, M. R. Pore-forming activity of clostridial binary toxins. Biochim. Biophys. Acta (BBA) Biomembr. 1858, 512–525 (2016).
Barbut, F. et al. Clinical features of Clostridium difficile-associated diarrhoea due to binary toxin (actin-specific ADP-ribosyltransferase)-producing strains. J. Med. Microbiol. 54, 181–185 (2005).
Chellapandi, P. & Prisilla, A. Structure, function and evolution of Clostridium botulinum C2 and C3 toxins: insight to poultry and veterinary vaccines. Curr. Protein Pept. Sci. 18, 412–424 (2017).
Nagahama, M., Kobayashi, K. & Takehara, M. Cathepsin release from lysosomes promotes endocytosis of Clostridium perfringens iota-toxin. Toxins 13, 721 (2021).
Papatheodorou, P. et al. Identification of the Cellular Receptor of Clostridium spiroforme Toxin. Infect. Immun. 80, 1418–1423 (2012).
Hardenbrook, N. J. et al. Atomic structures of anthrax toxin protective antigen channels bound to partially unfolded lethal and edema factors. Nat. Commun. 11, 840 (2020).
Jucovic, M., Walters, F. S., Warren, G. W., Palekar, N. V. & Chen, J. S. From enzyme to zymogen: engineering Vip2, an ADP-ribosyltransferase from Bacillus cereus, for conditional toxicity. Protein Eng., Des. Sel. 21, 631–638 (2008).
Wynia-Smith, S. L., Brown, M. J., Chirichella, G., Kemalyan, G. & Krantz, B. A. Electrostatic ratchet in the protective antigen channel promotes anthrax toxin translocation. J. Biol. Chem. 287, 43753–43764 (2012).
Yamada, T. et al. Cryo-EM structures reveal translocational unfolding in the clostridial binary iota toxin complex. Nat. Struct. Mol. Biol. 27, 288–296 (2020).
Sheedlo, M. J., Anderson, D. M., Thomas, A. K. & Lacy, D. B. Structural elucidation of the Clostridioides difficile transferase toxin reveals a single-site binding mode for the enzyme. Proc. Natl. Acad. Sci. 117, 201920555 (2020).
Kawamoto, A. et al. Cryo-EM structures of the translocational binary toxin complex CDTa-bound CDTb-pore from Clostridioides difficile. Nat. Commun. 13, 6119 (2022).
Krantz, B. et al. A phenylalanine clamp catalyzes protein translocation through the anthrax toxin pore. Science 309, 777–781 (2005).
Wang, K. et al. Lignocellulose degradation in Protaetia brevitarsis larvae digestive tract: refining on a tightly designed microbial fermentation production line. Microbiome 10, 90 (2022).
Jiang, J., Pentelute, B. L., Collier, R. J. & Zhou, Z. H. Atomic structure of anthrax protective antigen pore elucidates toxin translocation. Nature 521, 545–549 (2015).
Barth, H., Aktories, K., Popoff, M. R. & Stiles, B. G. Binary bacterial toxins: biochemistry, biology, and applications of common Clostridium and Bacillus proteins. Microbiol Mol. Biol. Rev. 68, 373–402 (2004).
Knapp, O. et al. Residues involved in the pore-forming activity of the Clostridium perfringens iota toxin. Cell Microbiol 17, 288–302 (2015).
Larburu, N., Adams, C. J., Chen, C.-S., Nowak, P. R. & Ali, M. M. U. Mechanism of Hsp70 specialized interactions in protein translocation and the unfolded protein response. Open Biol. 10, 200089 (2020).
Schopf, F. H., Biebl, M. M. & Buchner, J. The HSP90 chaperone machinery. Nat. Rev. Mol. Cell Biol. 18, 345–360 (2017).
Kato, Y. et al. HSP70 and HSP90 differentially regulate translocation of extracellular antigen to the cytosol for cross-presentation. Autoimmune Dis. 2012, 745962 (2012).
Geng, J. et al. Bacillus thuringiensis Vip1 functions as a receptor of Vip2 toxin for binary insecticidal activity against Holotrichia parallela. Toxins 11, 440 (2019).
Thoren, K. L. & Krantz, B. A. The unfolding story of anthrax toxin translocation. Mol. Microbiol. 80, 588–595 (2011).
Feld, G. K. et al. Structural basis for the unfolding of anthrax lethal factor by protective antigen oligomers. Nat. Struct. Mol. Biol. 17, 1383–1390 (2010).
Brown, M. J., Thoren, K. L. & Krantz, B. A. Role of the α clamp in the protein translocation mechanism of anthrax toxin. J. Mol. Biol. 427, 3340–3349 (2015).
Collier, R. J. Membrane translocation by anthrax toxin. Mol. Asp. Med. 30, 413–422 (2009).
Krantz, B. A., Finkelstein, A. & Collier, R. J. Protein translocation through the anthrax toxin transmembrane pore is driven by a proton gradient. J. Mol. Biol. 355, 968–979 (2006).
Brown, M. J., Thoren, K. L. & Krantz, B. A. Charge requirements for proton gradient-driven translocation of anthrax toxin*. J. Biol. Chem. 286, 23189–23199 (2011).
Das, D. & Krantz, B. A. Peptide- and proton-driven allosteric clamps catalyze anthrax toxin translocation across membranes. Proc. Natl. Acad. Sci. 113, 9611–9616 (2016).
Leuber, M., Orlik, F., Schiffler, B., Sickmann, A. & Benz, R. Vegetative insecticidal protein (Vip1Ac) of Bacillus thuringiensis HD201: evidence for oligomer and channel formation. Biochemistry 45, 283–288 (2006).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014).
Ashkenazy, H. et al. ConSurf 2016: an improved methodology to estimate and visualize evolutionary conservation in macromolecules. Nucleic Acids Res. 44, W344–W350 (2016).
Yariv, B. et al. Using evolutionary data to make sense of macromolecules with a “face-lifted” ConSurf. Protein Sci. 32, e4582 (2023).
Crickmore, N. et al. BPPRC. (Bacterial Pesticidal Protein Resource Center, https://www.bpprc.org).
Ritter, A. T. et al. ESCRT-mediated membrane repair protects tumor-derived cells against T cell attack. Science 376, 377–382 (2022).
Schrank, B. R. et al. An antibody–toxin conjugate targeting CD47 linked to the bacterial toxin listeriolysin O for cancer immunotherapy. Nat. Cancer 6, 511–527 (2025).
Woida, P. J. & Satchell, K. J. F. The Vibrio cholerae MARTX toxin silences the inflammatory response to cytoskeletal damage before inducing actin cytoskeleton collapse. Sci. Signal. 13, eaaw9447 (2020).
Koo, S., Cheley, S. & Bayley, H. Redirecting Pore Assembly of Staphylococcal α-Hemolysin by Protein Engineering. ACS Cent. Sci. 5, 629–639 (2019).
Hermansen, S., Linke, D. & Leo, J.C. Chapter Four - Transmembrane β-barrel proteins of bacteria: From structure to function. in Advances in Protein Chemistry and Structural Biology, Vol. 128 (ed. Donev, R.) 113–161 (Academic Press, 2022).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr, D: Biol. Crystallogr. 66, 486–501 (2010).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Afonine, P.V., Poon, B.K., Read, R.J., Sobolev, O.V. & Adams, P.D. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D Struct. Biol. 74, 531–544 (2018).
Hink, W. F. Established insect cell line from the cabbage looper, Trichoplusia ni. Nature 226, 466–467 (1970).
Mi, L. et al. Enterobacter strain IPPBiotE33 displays a synergistic effect with Bacillus thuringiensis Bt185. Int. J. Mol. Sci. 24, 14193 (2023).
Zhao, H., Brautigam, C. A., Ghirlando, R. & Schuck, P. Overview of current methods in sedimentation velocity and sedimentation equilibrium analytical ultracentrifugation. Curr. Protoc. Protein Sci. 71, 20.12.1–20.12.49 (2013).
Schuck, P., Perugini, M. A., Gonzales, N. R., Howlett, G. J. & Schubert, D. Size-distribution analysis of proteins by analytical ultracentrifugation: strategies and application to model systems. Biophys. J. 82, 1096–1111 (2002).
Wang, Z. et al. JAK/STAT signaling regulated intestinal regeneration defends insect pests against pore-forming toxins produced by Bacillus thuringiensis. PLoS Pathog. 20, e1011823 (2024).
Jiang, S. et al. Generic Diagramming Platform (GDP): a comprehensive database of high-quality biomedical graphics. Nucleic Acids Res. 53, D1670–D1676 (2025).
Acknowledgements
We would like to thank Ms. Danyang Li and Xiangning Li from the Core Facility of Wuhan. University for their assistance with the cryo-EM data collection, thank Prof. Changyou Li and Dr. Guiling Zheng from Qingdao Agricultural University to supply H.parallela HP9-2 Cells, thank Prof. Qing Yang, Prof. Changlong Shu and Dr. Lili Geng from Institute of Plant Protection, Chinese Academy of Agricultural Sciences for their technical guidance about the insecticidal bioassay. We thank Dr. Qing Chang for providing AUC support at Biomolecule Preparation and Characterization Platform of the Technology Center for Protein Research, Tsinghua University. We thank Derek Duong, Dr. Linfeng Gao, and Dr. Baohua Chen for improving the language and style of the manuscript. This work is supported by grants from Natural Science Foundation of Wuhan (grant number 2024040801020228) for Z.L., Provincial Natural Science Foundation of China (grant number 2024AFB026) for Z.L., a startup fund from Wuhan University awarded to P.C., and Innovation Program of Chinese Academy of Agricultural Sciences (CAAS-CSCB-202401 and CAAS-ZDRW202304 to Z.W.).
Author information
Authors and Affiliations
Contributions
Conceptualization: T.Z. and P.C. Methodology: T.Z., J.R., L.L., Z.L., and Z.W. Investigation: T.Z., J.R., L.L., Z.W., and X.H. Writing: T.Z., Z.W., and P.C. Supervision: J.Z. and P.C. Funding acquisition: Z.L., Z.W., and P.C.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Nicholas Waterfield and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhao, T., Wang, Z., Ren, J. et al. Structural basis for the assembly and translocation of the Vip1-Vip2 insecticidal toxin from Bacillus thuringiensis. Nat Commun (2026). https://doi.org/10.1038/s41467-026-71211-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-71211-7