Abstract
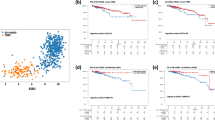
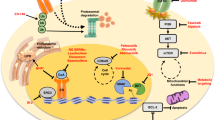
Next-generation selective estrogen receptor-α (ERα) antagonist/degraders (SERDs) are being developed for ER-positive breast cancer (ER+ BC), with intentions of improving outcomes for patients. In recent clinical trials of metastatic ER+ BC, next-generation SERDs demonstrated clinical activity, and elacestrant received an approval for advanced ESR1-mutant disease. However, responses to these drugs were highly heterogeneous: across trials and independent of ESR1 status, 30–50% of patients progressed by their first follow-up scan while other patients sustained benefit for 2 years or more. Here, we interrogate the basis for heterogeneous responses by comparing biopsies from non-responding patients (NR; progression-free survival <2 months) and responding patients (Resp; PFS ≥ 2 months) who received the next-generation SERD giredestrant. While Resp tumors maintain high dependency on ERα signaling, NR tumors exhibit loss of luminal lineage identity and by extension, ERα dependence. NR tumors instead up-regulate multiple ERα-independent proliferative pathways, such as EGFR/MAPK and Hippo/TEAD, which may represent targetable dependencies in NR disease. Modeling resistance and lineage plasticity in vitro, we find that giredestrant-resistant ER+ BC cell lines exhibit profound shifts in chromatin accessibility, with the transcription factors, FOXA1 and FOXM1, implicated in gene expression of NR-upregulated proliferative pathways.
Similar content being viewed by others
Data availability
Raw and processed files from sequencing experiments have been uploaded to the NCBI Gene Expression Omnibus under the following accession IDs: GSE295127 (RNA-seq; www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE295127), GSE295128 (ATAC-seq; www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE295128), and GSE305432 (ChIP-seq; www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE305432). Due to data privacy laws and the terms of informed consent for study GO39932, raw data from patient-derived F-ACT and RNA-seq cannot be deposited in a public repository nor made available upon request. The remaining data for Figs. 1–4 and all Supplementary Figs. have been provided as Source Data files. Source data are provided with this paper.
Code availability
Analysis code is available at https://doi.org/10.5281/zenodo.18644938. For ATAC-seq and ChIP-seq, all analysis code uses publicly available R packages. For RNA-seq, some analyses code includes custom in-house functions. The custom code was designed to analyze RNA-seq data within our institution’s cloud-based computing environment, and will not function properly outside of this environment. However, comparable analyses can be performed using publicly available R packages.
References
Sung, H. et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Toy, W. et al. ESR1 ligand-binding domain mutations in hormone-resistant breast cancer. Nat. Genet. 45, 1439–1445 (2013).
Robinson, D. R. et al. Activating ESR1 mutations in hormone-resistant metastatic breast cancer. Nat. Genet. 45, 1446–1451 (2013).
Jeselsohn, R. et al. Emergence of constitutively active estrogen receptor-α mutations in pretreated advanced estrogen receptor-positive breast cancer. Clin. Cancer Res. 20, 1757–1767 (2014).
Schiavon, G. et al. Analysis of ESR1 mutation in circulating tumor DNA demonstrates evolution during therapy for metastatic breast cancer. Sci. Transl. Med. 7, ra182 (2015).
Guan, J. et al. Therapeutic ligands antagonize estrogen receptor function by impairing its mobility. Cell 178, 949–963 e918 (2019).
van Kruchten, M. et al. Measuring residual estrogen receptor availability during fulvestrant therapy in patients with metastatic breast cancer. Cancer Discov. 5, 72–81 (2015).
Mottamal, M., Kang, B., Peng, X. & Wang, G. From pure antagonists to pure degraders of the estrogen receptor: evolving strategies for the same target. ACS Omega 6, 9334–9343 (2021).
Liang, J. et al. GDC-9545 (Giredestrant): a potent and orally bioavailable selective estrogen receptor antagonist and degrader with an exceptional preclinical profile for ER+ breast cancer. J. Med. Chem. 64, 11841–11856 (2021).
Liang, J. et al. Giredestrant counters a progesterone hypersensitivity program driven by estrogen receptor mutations in breast cancer. Sci. Transl. Med. 14, eabo5959 (2022).
Bidard, F. C. et al. Elacestrant (oral selective estrogen receptor degrader) versus standard endocrine therapy for estrogen receptor-positive, human epidermal growth factor receptor 2-negative advanced breast cancer: results from the randomized phase III EMERALD trial. J. Clin. Oncol. JCO2200338 (2022).https://doi.org/10.1200/JCO.22.00338
Will, M., Liang, J., Metcalfe, C. & Chandarlapaty, S. Therapeutic resistance to anti-oestrogen therapy in breast cancer. Nat. Rev. Cancer 23, 673–685 (2023).
Oliveira, M. et al. Abstract GS3-02: GS3-02 Camizestrant, a next generation oral SERD vs fulvestrant in post-menopausal women with advanced ER-positive HER2-negative breast cancer: Results of the randomized, multi-dose Phase 2 SERENA-2 trial. Cancer Res. 83, GS3-02 (2023).
Jhaveri, K. L. et al. Phase Ia/b study of giredestrant ±palbociclib and ±luteinizing hormone-releasing hormone agonists in estrogen receptor-positive, HER2-negative, locally advanced/metastatic breast cancer. Clin. Cancer Res. (2023). https://doi.org/10.1158/1078-0432.CCR-23-1796
Shomali, M. et al. SAR439859, a novel selective estrogen receptor degrader (SERD), demonstrates effective and broad antitumor activity in wild-type and mutant ER-positive breast cancer models. Mol. Cancer Ther. 20, 250–262 (2021).
Bhagwat, S. V. et al. Imlunestrant is an oral, brain-penetrant selective estrogen receptor degrader with potent antitumor activity in ESR1 wildtype and mutant breast cancer. Cancer Res. (2024). https://doi.org/10.1158/0008-5472.CAN-24-2608
Martín, M. et al. Giredestrant for estrogen receptor-positive, HER2-negative, previously treated advanced breast cancer: results from the randomized, phase II acelERA breast cancer study. J. Clin. Oncol. JCO2301500 (2024). https://doi.org/10.1200/JCO.23.01500
Tolaney, S. M. et al. AMEERA-3: randomized phase II study of amcenestrant (oral selective estrogen receptor degrader) versus standard endocrine monotherapy in estrogen receptor-positive, human epidermal growth factor receptor 2-negative advanced breast cancer. J. Clin. Oncol. 41, 4014–4024 (2023).
Jhaveri, K. L. et al. Imlunestrant with or without abemaciclib in advanced breast cancer. N. Engl. J. Med. (2024). https://doi.org/10.1056/NEJMoa2410858
Iams, W. T. et al. Concurrent tissue and circulating tumor DNA molecular profiling to detect guideline-based targeted mutations in a multicancer cohort. JAMA Netw. Open 7, e2351700 (2024).
Chen, J. W. et al. ER+, HER2- advanced breast cancer treated with taselisib and fulvestrant: genomic landscape and associated clinical outcomes. Mol. Oncol. (2023). https://doi.org/10.1002/1878-0261.13416
Razavi, P. et al. The genomic landscape of endocrine-resistant advanced breast cancers. Cancer Cell 34, 427–438.e426 (2018).
Cristofanilli, M. et al. Fulvestrant plus palbociclib versus fulvestrant plus placebo for treatment of hormone-receptor-positive, HER2-negative metastatic breast cancer that progressed on previous endocrine therapy (PALOMA-3): final analysis of the multicentre, double-blind, phase 3 randomised controlled trial. Lancet Oncol. 17, 425–439 (2016).
Beltran, H. et al. The role of lineage plasticity in prostate cancer therapy resistance. Clin. Cancer Res. 25, 6916–6924 (2019).
Wang, Y., Ci, X., Choi, S. Y. C., Crea, F. & Lin, D. Molecular events in neuroendocrine prostate cancer development. Nat. Rev. Urol. 18, 581–596 (2021).
Liang, J. et al. ERα dysfunction caused by ESR1 mutations and therapeutic pressure promotes lineage plasticity in ER+ breast cancer. Nat. Cancer 6, 357–371 (2025).
Chan, J. M. et al. Lineage plasticity in prostate cancer depends on JAK/STAT inflammatory signaling. Science 377, 1180–1191 (2022).
Griffiths, J. I. et al. Serial single-cell genomics reveals convergent subclonal evolution of resistance as early-stage breast cancer patients progress on endocrine plus CDK4/6 therapy. Nat. Cancer 2, 658–671 (2021).
Hanker, A. B., Sudhan, D. R. & Arteaga, C. L. Overcoming endocrine resistance in breast cancer. Cancer Cell 37, 496–513 (2020).
Keenan, A. B. et al. ChEA3: transcription factor enrichment analysis by orthogonal omics integration. Nucleic Acids Res. 47, W212–W224 (2019).
Sanders, D. A., Ross-Innes, C. S., Beraldi, D., Carroll, J. S. & Balasubramanian, S. Genome-wide mapping of FOXM1 binding reveals co-binding with estrogen receptor alpha in breast cancer cells. Genome Biol. 14, R6 (2013).
Millour, J. et al. FOXM1 is a transcriptional target of ERalpha and has a critical role in breast cancer endocrine sensitivity and resistance. Oncogene 29, 2983–2995 (2010).
Arceci, A. et al. FOXM1 deubiquitination by USP21 regulates cell cycle progression and paclitaxel sensitivity in basal-like breast cancer. Cell Rep. 26, 3076–3086.e3076 (2019).
Lu, X. F. et al. FoxM1 is a promising candidate target in the treatment of breast cancer. Oncotarget 9, 842–852 (2018).
Kaminska, K. et al. Distinct mechanisms of resistance to fulvestrant treatment dictate level of ER independence and selective response to CDK inhibitors in metastatic breast cancer. Breast Cancer Res. 23, 26 (2021).
Yan, Z. et al. ATP6AP1 promotes cell proliferation and tamoxifen resistance in luminal breast cancer by inducing autophagy. Cell Death Dis. 16, 201 (2025).
Joseph, J. D. et al. The selective estrogen receptor downregulator GDC-0810 is efficacious in diverse models of ER+ breast cancer. Elife 5, e15828 (2016).
Hurtado, A., Holmes, K. A., Ross-Innes, C. S., Schmidt, D. & Carroll, J. S. FOXA1 is a key determinant of estrogen receptor function and endocrine response. Nat. Genet 43, 27–33 (2011).
Glont, S. E., Chernukhin, I. & Carroll, J. S. Comprehensive Genomic Analysis Reveals that the Pioneering Function of FOXA1 Is Independent of Hormonal Signaling. Cell Rep. 26, 2558–2565.e2553 (2019).
Wang, Y., Ung, M. H., Xia, T., Cheng, W. & Cheng, C. Cancer cell line specific co-factors modulate the FOXM1 cistrome. Oncotarget 8, 76498–76515 (2017).
Hafner, M., Niepel, M., Chung, M. & Sorger, P. K. Growth rate inhibition metrics correct for confounders in measuring sensitivity to cancer drugs. Nat. Methods 13, 521–527 (2016).
Guan, J. et al. Giredestrant immobilizes the Estrogen Receptor to potently suppress ER-active breast cancers. San. Antonio Breast Cancer Symp. PS 11, 1–12 (2025).
Hagenbeek, T. J. et al. An allosteric pan-TEAD inhibitor blocks oncogenic YAP/TAZ signaling and overcomes KRAS G12C inhibitor resistance. Nat. Cancer 4, 812–828 (2023).
Dardenne, E. et al. N-Myc Induces an EZH2-mediated transcriptional program driving neuroendocrine prostate cancer. Cancer Cell 30, 563–577 (2016).
Gianni, C. et al. Cell-Free DNA fragmentomics: a promising biomarker for diagnosis, prognosis and prediction of response in breast cancer. Int. J. Mol. Sci. 23 (2022). https://doi.org/10.3390/ijms232214197
Qi, T. et al. Cell-free DNA fragmentomics: the novel promising biomarker. Int. J. Mol. Sci. 24 (2023). https://doi.org/10.3390/ijms24021503
Clark, T. A. et al. Analytical validation of a hybrid capture-based next-generation sequencing clinical assay for genomic profiling of cell-free circulating tumor DNA. J. Mol. Diagn. 20, 686–702 (2018).
Wu, T. D. & Nacu, S. Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 26, 873–881 (2010).
Wu, T. D., Reeder, J., Lawrence, M., Becker, G. & Brauer, M. J. GMAP and GSNAP for genomic sequence alignment: enhancements to speed, accuracy, and functionality. Methods Mol. Biol. 1418, 283–334 (2016).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Wagle, M. C. et al. A transcriptional MAPK Pathway Activity Score (MPAS) is a clinically relevant biomarker in multiple cancer types. NPJ Precis. Oncol. 2, 7 (2018).
Talevich, E., Shain, A. H., Botton, T. & Bastian, B. C. CNVkit: genome-wide copy number detection and visualization from targeted DNA sequencing. PLoS Comput. Biol. 12, e1004873 (2016).
Chakravarty, D. et al. OncoKB: a precision oncology knowledge base. JCO Precis. Oncol. 2017 (2017). https://doi.org/10.1200/PO.17.00011
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Consortium, E. P. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
McLeay, R. C. & Bailey, T. L. Motif enrichment analysis: a unified framework and an evaluation on ChIP data. BMC Bioinforma. 11, 165 (2010).
Kulakovskiy, I. V. et al. HOCOMOCO: towards a complete collection of transcription factor binding models for human and mouse via large-scale ChIP-Seq analysis. Nucleic Acids Res. 46, D252–D259 (2018).
Acknowledgements
We would like to thank Ann E. Collier, Alejandro Chibly, and Xiaosai Yao for helpful discussions; Meritxell Bellet, Sherene Loi, Aditya Bardia, Valentina Boni, Joohyuk Sohn, Tomas G. Neilan, Rafael Villanueva-Vázquez, Peter Kabos, Laura García-Estévez, Elena López-Miranda, Jose A. Pérez-Fidalgo, and Jose M. Pérez-García for providing biopsies in the clinical trial (NCT03332797); and patients for their participation in the clinical trial. This study was supported by funding from Roche/Genentech.
Author information
Authors and Affiliations
Contributions
J.L. designed/conducted the biomarker analyses and wrote the manuscript. J.L. & C.M. designed the experiments. K.H., C.O., & J.Guan performed the experiments. V.K. and B.D. performed the ChIP sequencing, M.S. and Y.L. performed the RNA and ATAC sequencing, J.Giltnane performed pathology review. H.M.M., J.A., C.-W.C., M.R.G., J.E.-W., P.P.-M., K.L.J, N.C.T, & E.L. conducted the clinical study. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare the following competing interests: excepting K.L. Jhaveri, N.C. Turner, and E. Lim, all authors are Genentech/Roche employees and own shares of Roche. C.M. is a named co-inventor on patent 11081236 entitled “Diagnostic and therapeutic methods for the treatment of breast cancer”. K.L. Jhaveri has a consultant/advisory board role in Novartis, Pfizer, BMS, Jounce Therapeutics, Taiho Oncology, Genentech, Inc., AbbVie, Eisai, AstraZeneca, Blueprint Medicine, Daiichi Sankyo, Sun Pharma Advanced Research Company Ltd, Gilead, Seattle Genetics, Olema Pharmaceuticals, Menarini/Stemline, and Lilly/Loxo Oncology; and has received research funding from Novartis, Clovis Oncology, Genentech, Inc., AstraZeneca, ADC Therapeutics, Novita Pharmaceuticals, Debio Pharmaceuticals, Pfizer, Lilly Pharmaceuticals/Loxo Oncology, Zymeworks, Gilead, Puma Biotechnology, Context Therapeutics, and Merck Pharmaceuticals. N.C. Turner has received advisory board honoraria from AstraZeneca, Lilly, Pfizer, Roche–Genentech, Novartis, GlaxoSmithKline, Repare Therapeutics, Relay Therapeutics, Zentalis, Gilead, Inivata, Guardant, and Exact Sciences; and research funding from AstraZeneca, Pfizer, Roche–Genentech, Merck Sharpe & Dohme, Guardant Health, Invitae, Inivata, Personalis, and Natera. E. Lim reports research support from Ellipses, Novartis, and Pfizer; speaker compensation from AstraZeneca, Gilead, Novartis, Lilly, Pfizer and Roche; meeting and/or travel support from AstraZeneca, Novartis, and Gilead; advisory board compensation from AstraZeneca, Gilead, Lilly, MSD, Novartis, Pfizer and Roche; and leadership or fiduciary roles in the Breast Cancer Trials and University of New South Wales.
Peer review
Peer review information
Nature Communications thanks Victor Jin, who co-reviewed with Lavanya Choppavarapu, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liang, J., Ong, C., Heslop, K. et al. Loss of luminal lineage drives resistance to next-generation ERα antagonists in pretreated ER+ HER2− locally-advanced or metastatic breast cancer. Nat Commun (2026). https://doi.org/10.1038/s41467-026-71233-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-71233-1