Abstract
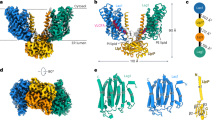
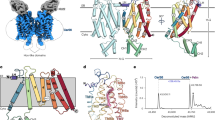
Ceramide synthases (CerSs) are crucial enzymes in sphingolipid metabolism and have shown therapeutic potential for treating various metabolic disorders. However, their regulatory mechanisms remain poorly understood. In this study, we report the cryo-electron microscopy structure of a yeast CerS (yCerS), composed of a catalytic Lac1 subunit and a regulatory Lip1 subunit, organized into a higher-order 4:4 assembly. This assembly is formed by dimerization of two 2:2 Lac1-Lip1 subcomplexes via an interface primarily involving the Lac1 subunit. Notably, within this interface, the C-terminal transmembrane helix (TM8) of Lac1 adopts a dramatically twisted conformation and engages in extensive interactions with TMs 6/7/8 of the adjacent Lac1 subunit. This structural rearrangement sterically occludes the catalytic chamber and blocks acyl-CoA substrate entry. Functional assays further demonstrate that, although structurally reminiscent of an autoinhibitory conformation, this interface is required for the regulation of ceramide output and cellular adaption during perturbation of complex sphingolipid biosynthesis. Together, our findings uncover a complex oligomerization-mediated regulatory mechanism in yCerS, advancing the mechanistic understanding of ceramide synthesis control and highlighting the nuanced role of oligomerization in modulating CerS activity.
Similar content being viewed by others
Data availability
The EM density maps and atomic coordinates for the 4:4 Lac1-Lip1 complex have been deposited in the EMDB under accession code EMD-65613 and in the PDB under the accession code 9W3Z, respectively. PDB codes of previously published structures used in this study are 8Y2N, 8IZD, and 8QZ6. All data needed to evaluate the conclusions in the paper are present in the paper and/or the Supplementary Information. Source Data are provided as a Source Data file. Source data are provided with this paper.
References
Hannun, Y. A. & Obeid, L. M. Many ceramides. J. Biol. Chem. 286, 27855–27862 (2011).
Obeid, L. M., Linardic, C. M., Karolak, L. A. & Hannun, Y. A. Programmed cell death induced by ceramide. Science 259, 1769–1771 (1993).
Hannun, Y. A. & Obeid, L. M. Principles of bioactive lipid signalling: lessons from sphingolipids. Nat. Rev. Mol. Cell Biol. 9, 139–150 (2008).
Gault, C. R., Obeid, L. M. & Hannun, Y. A. An overview of sphingolipid metabolism: from synthesis to breakdown. Adv. Exp. Med. Biol. 688, 1–23 (2010).
Choi, M. J. & Maibach, H. I. Role of ceramides in barrier function of healthy and diseased skin. Am. J. Clin. Dermatol. 6, 215–223 (2005).
Maceyka, M. & Spiegel, S. Sphingolipid metabolites in inflammatory disease. Nature 510, 58–67 (2014).
Hannun, Y. A. & Obeid, L. M. Sphingolipids and their metabolism in physiology and disease. Nat. Rev. Mol. Cell Biol. 19, 175–191 (2018).
Summers, S. A., Chaurasia, B. & Holland, W. L. Metabolic messengers: ceramides. Nat. Metab. 1, 1051–1058 (2019).
Hla, T. & Dannenberg, A. J. Sphingolipid signaling in metabolic disorders. Cell Metab. 16, 420–434 (2012).
Chaurasia, B. & Summers, S. A. Ceramides in metabolism: key lipotoxic players. Annu. Rev. Physiol. 83, 303–330 (2021).
Holland, W. L. et al. Inhibition of ceramide synthesis ameliorates glucocorticoid-, saturated-fat-, and obesity-induced insulin resistance. Cell Metab. 5, 167–179 (2007).
Turpin, S. M. et al. Obesity-induced CerS6-dependent C16:0 ceramide production promotes weight gain and glucose intolerance. Cell Metab. 20, 678–686 (2014).
Summers, S. A. Could ceramides become the new cholesterol? Cell Metab. 27, 276–280 (2018).
Apostolopoulou, M. et al. Specific hepatic sphingolipids relate to insulin resistance, oxidative stress, and inflammation in nonalcoholic steatohepatitis. Diab. Care 41, 1235–1243 (2018).
Choi, R. H., Tatum, S. M., Symons, J. D., Summers, S. A. & Holland, W. L. Ceramides and other sphingolipids as drivers of cardiovascular disease. Nat. Rev. Cardiol. 18, 701–711 (2021).
Hajduch, E., Lachkar, F., Ferré, P. & Foufelle, F. Roles of ceramides in non-alcoholic fatty liver disease. J. Clin. Med. 10, 792 (2021).
Ogretmen, B. & Hannun, Y. A. Biologically active sphingolipids in cancer pathogenesis and treatment. Nat. Rev. Cancer 4, 604–616 (2004).
Furuya, H., Shimizu, Y. & Kawamori, T. Sphingolipids in cancer. Cancer Metastasis Rev. 30, 567–576 (2011).
Morad, S. A. F. & Cabot, M. C. Ceramide-orchestrated signalling in cancer cells. Nat. Rev. Cancer 13, 51–65 (2013).
Kuo, A. & Hla, T. Regulation of cellular and systemic sphingolipid homeostasis. Nat. Rev. Mol. Cell Biol. 25, 802–821 (2024).
Guillas, I. et al. C26-CoA-dependent ceramide synthesis of Saccharomyces cerevisiae is operated by Lag1p and Lac1p. EMBO J. 20, 2655–2665 (2001).
Schorling, S., Vallée, B., Barz, W. P., Riezman, H. & Oesterhelt, D. Lag1p and Lac1p are essential for the acyl-CoA-dependent ceramide synthase reaction in Saccharomyces cerevisiae. Mol. Biol. cell 12, 3417–3427 (2001).
Funato, K. & Riezman, H. Vesicular and nonvesicular transport of ceramide from ER to the Golgi apparatus in yeast. J. Cell Biol. 155, 949–960 (2001).
Kumagai, K. et al. CERT mediates intermembrane transfer of various molecular species of ceramides. J. Biol. Chem. 280, 6488–6495 (2005).
Limar, S. et al. Yeast Svf1 binds ceramides and contributes to sphingolipid metabolism at the ER cis-Golgi interface. J. Cell Biol. 222, e202109162 (2023).
Kitatani, K., Idkowiak-Baldys, J. & Hannun, Y. A. The sphingolipid salvage pathway in ceramide metabolism and signaling. Cell. Signal. 20, 1010–1018 (2008).
Mullen, T. D., Hannun, Y. A. & Obeid, L. M. Ceramide synthases at the centre of sphingolipid metabolism and biology. Biochem. J. 441, 789–802 (2012).
Park, J.-W., Park, W.-J. & Futerman, A. H. Ceramide synthases as potential targets for therapeutic intervention in human diseases. Biochim. Biophys. Acta (BBA) - Mol. Cell Biol. Lipids 1841, 671–681 (2014).
Raichur, S. Ceramide synthases are attractive drug targets for treating metabolic diseases. Front. Endocrinol. 11, 483 (2020).
Brachtendorf, S., El-Hindi, K. & Grösch, S. Ceramide synthases in cancer therapy and chemoresistance. Prog. Lipid Res. 74, 160–185 (2019).
Wang, E., Norred, W. P., Bacon, C. W., Riley, R. T. & Merrill, A. H. Inhibition of sphingolipid biosynthesis by fumonisins. Implications for diseases associated with Fusarium moniliforme. J. Biol. Chem. 266, 14486–14490 (1991).
MANDALA, S. M. et al. The discovery of australifungin, a novel inhibitor of sphinganine N-acyltransferase from Sporormiella australis. producing organism, fermentation, isolation, and biological activity. J. Antibiot. 48, 349–356 (1995).
Munshi, M. A. et al. The role of ceramide synthases in the pathogenicity of Cryptococcus neoformans. Cell Rep. 22, 1392–1400 (2018).
Vallée, B. & Riezman, H. Lip1p: a novel subunit of acyl-CoA ceramide synthase. EMBO J. 24, 730–741 (2005).
Venkataraman, K. et al. Upstream of growth and differentiation factor 1 (uog1), a mammalian homolog of the yeast longevity assurance gene 1 (LAG1), regulates N-stearoyl-sphinganine (C18-(Dihydro)ceramide) synthesis in a fumonisin B1-independent manner in mammalian cells. J. Biol. Chem. 277, 35642–35649 (2002).
Guillas, I. et al. Human homologues of LAG1 reconstitute acyl-CoA-dependent ceramide synthesis in yeast. J. Biol. Chem. 278, 37083–37091 (2003).
Riebeling, C., Allegood, J. C., Wang, E., Merrill, A. H. & Futerman, A. H. Two mammalian longevity assurance gene (LAG1) family members, trh1 and trh4, regulate dihydroceramide synthesis using different fatty acyl-CoA donors. J. Biol. Chem. 278, 43452–43459 (2003).
Mizutani, Y., Kihara, A. & Igarashi, Y. Mammalian Lass6 and its related family members regulate synthesis of specific ceramides. Biochem. J. 390, 263–271 (2005).
Mizutani, Y., Kihara, A. & Igarashi, Y. LASS3 (longevity assurance homologue 3) is a mainly testis-specific (dihydro)ceramide synthase with relatively broad substrate specificity. Biochem. J. 398, 531–538 (2006).
Laviad, E. L. et al. Characterization of ceramide synthase 2: tissue distribution, substrate specificity, and inhibition by sphingosine 1-phosphate. J. Biol. Chem. 283, 5677–5684 (2008).
Levy, M. & Futerman, A. H. Mammalian ceramide synthases. IUBMB Life 62, 347–356 (2010).
Stiban, J., Tidhar, R. & Futerman, A. H. Ceramide synthases: roles in cell physiology and signaling. Adv. Exp. Med. Biol. 688, 60–71 (2010).
Cingolani, F., Futerman, A. H. & Casas, J. Ceramide synthases in biomedical research. Chem. Phys. Lipids 197, 25–32 (2016).
Winter, E. & Ponting, C. P. TRAM, LAG1 and CLN8: members of a novel family of lipid-sensing domains? Trends Biochem. Sci. 27, 381–383 (2002).
Spassieva, S. et al. Necessary role for the Lag1p Motif in (Dihydro)ceramide synthase activity. J. Biol. Chem. 281, 33931–33938 (2006).
Mesicek, J. et al. Ceramide synthases 2, 5, and 6 confer distinct roles in radiation-induced apoptosis in HeLa cells. Cell. Signal. 22, 1300–1307 (2010).
Laviad, E. L., Kelly, S., Merrill, A. H. & Futerman, A. H. Modulation of ceramide synthase activity via dimerization. J. Biol. Chem. 287, 21025–21033 (2012).
Kim, J. L., Ben-Dor, S., Rosenfeld-Gur, E. & Futerman, A. H. A novel C-terminal DxRSDxE motif in ceramide synthases involved in dimer formation. J. Biol. Chem. 298, 101517 (2022).
Muir, A., Ramachandran, S., Roelants, F. M., Timmons, G. & Thorner, J. TORC2-dependent protein kinase Ypk1 phosphorylates ceramide synthase to stimulate synthesis of complex sphingolipids. eLife 3, e03779 (2014).
Kobayashi, S. D. & Nagiec, M. M. Ceramide/long-chain base phosphate rheostat in Saccharomyces cerevisiae: regulation of ceramide synthesis by Elo3p and Cka2p. Eukaryot. Cell 2, 284–294 (2003).
Fresques, T. et al. Regulation of ceramide synthase by casein kinase 2-dependent phosphorylation in Saccharomyces cerevisiae. J. Biol. Chem. 290, 1395–1403 (2015).
Sassa, T., Hirayama, T. & Kihara, A. Enzyme activities of the ceramide synthases CERS2–6 Are regulated by phosphorylation in the C-terminal region. J. Biol. Chem. 291, 7477–7487 (2016).
Xie, T. et al. Structure and mechanism of a eukaryotic ceramide synthase complex. EMBO J. 42, e114889 (2023).
Zhang, Z., Fang, Q., Xie, T. & Gong, X. Mechanism of ceramide synthase inhibition by fumonisin B(1). Structure 32, 1419–1428.e1414 (2024).
Pascoa, T. C. et al. Structural basis of the mechanism and inhibition of a human ceramide synthase. Nat. Struct. Mol. Biol. 32, 431–440 (2025).
Schäfer, J. H. et al. Structure of the yeast ceramide synthase. Nat. Struct. Mol. Biol. 32, 441–449 (2025).
Tani, M. & Kuge, O. Defect of synthesis of very long-chain fatty acids confers resistance to growth inhibition by inositol phosphorylceramide synthase repression in yeast Saccharomyces cerevisiae. J. Biochem. 148, 565–571 (2010).
Schäfer, J. H. et al. Structure of the ceramide-bound SPOTS complex. Nat. Commun. 14, 6196 (2023).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7, e42166 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Chen, S. X. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013).
Pettersen, E. F. et al. UCSF Chimera-a visualization system for exploratory research and analysis. J. Comput Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. 66, 213–221 (2010).
Williams, C. J. et al. MolProbity: more and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
DeLano, W. L. The PyMOL molecular graphics system on World Wide Web http://www.pymol.org. (2002).
Guan, X. L. & Wenk, M. R. Mass spectrometry-based profiling of phospholipids and sphingolipids in extracts from Saccharomyces cerevisiae. Yeast 23, 465–477 (2006).
Shui, G. et al. Toward one step analysis of cellular lipidomes using liquid chromatography coupled with mass spectrometry: application to Saccharomyces cerevisiae and Schizosaccharomyces pombe lipidomics. Mol. Biosyst. 6, 1008–1017 (2010).
Liu, S. Y. et al. The ER calcium channel Csg2 integrates sphingolipid metabolism with autophagy. Nat. Commun. 14, 3725 (2023).
Acknowledgements
We thank the Cryo-EM Facility of the Southern University of Science and Technology (SUSTech) for providing the facility support. This work was supported by the National Natural Science Foundation of China (32401001) and the Shenzhen Science and Technology Program (JCYJ20240813094404006, KQTD20190929173906742, ZDSYS20230626091659010).
Author information
Authors and Affiliations
Contributions
X.G. conceived and supervised the project. X.G., Q.F. and T.X. designed the experiments. Q.F. and T.X. conducted the experiments. C.Y. and N.Y. contributed to the lipidomics analysis. All authors contributed to the data analysis. X.G. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jan-Hannes Schaefer and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fang, Q., Yang, C., Yao, N. et al. Structural and functional dissection of a higher-order oligomerization interface in yeast ceramide synthase. Nat Commun (2026). https://doi.org/10.1038/s41467-026-71272-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-71272-8