Abstract
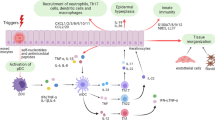
Metabolic dysregulation within the epithelial immune microenvironment (EIME) drives chronic inflammatory skin diseases like psoriasis, but the immune mechanisms and downstream consequences remain unclear. Here we perform in-depth metabolomic analysis showing that nucleotide metabolism is enhanced in psoriatic patients, with elevated adenosine levels closely correlating with disease severity. Single-cell and spatial transcriptomics analyses revealed that adenosine is primarily generated from a population of CD73high fibroblasts in psoriatic skin through enhanced metabolic processes and catalytic capability. Adenosine acts as a mediator between fibroblasts and keratinocytes, causing mitochondrial dysfunction and generating oxidative stress, resulting in the release of pro-inflammatory mediators in keratinocytes via ADORA2B. Deletion of Cd73 in fibroblasts, Adora2b in keratinocytes, or the use of pharmacological inhibitors of the pathways involved, reduces epidermal inflammation in the imiquimod- and IL-23A-induced mouse skin inflammation models. Our study thus identifies the CD73high fibroblast subsets as regulators of epithelial inflammation through metabolic microenvironment interactions with keratinocytes, providing proof of principle for therapeutic strategies targeting fibroblast-keratinocyte crosstalk in inflammatory skin diseases.
Similar content being viewed by others
Data availability
The raw data generated in this study, including spatial transcriptomics, bulk RNA-seq, and metabolomics sequencing data, have been deposited in the genome sequence archive (GSA) under accession numbers HRA011913, HRA009804, and OMIX010507. The patient-derived spatial transcriptomics data are available under restricted access due to the inclusion of potentially identifiable human genetic and clinical information. Access can be obtained by submitting a request to the corresponding author for non-commercial academic research purposes and is subject to institutional approval and a data use agreement. Responses to access requests are typically provided within 2–4 weeks. Public gene expression microarray and bulk RNA-seq datasets of psoriasis and associated cohort information were obtained from the GEO database under accession numbers GSE67785, GSE26866, GSE11903, GSE51440, GSE14905, GSE85034, GSE6710, GSE41662, GSE117239, GSE30999, GSE69967, GSE106992, GSE41663, GSE136757, GSE67853, GSE34248, GSE54456, GSE79704, GSE53431, GSE83582, and GSE52471. Single-cell RNA-sequencing data were obtained from GEO under accession number GSE173706. Source data are provided with this paper.
References
Dainichi, T. et al. The epithelial immune microenvironment (EIME) in atopic dermatitis and psoriasis. Nat. Immunol. 19, 1286–1298 (2018).
Zhou, Y. et al. The epidermal immune microenvironment plays a dominant role in psoriasis development, as revealed by mass cytometry. Cell. Mol. Immunol. 19, 1400–1413 (2022).
Griffiths, C. E. M., Armstrong, A. W., Gudjonsson, J. E. & Barker, J. Psoriasis. Lancet 397, 1301–1315 (2021).
Parisi, R., Symmons, D. P., Griffiths, C. E. & Ashcroft, D. M. Global epidemiology of psoriasis: a systematic review of incidence and prevalence. J. Investig. Dermatol. 133, 377–385 (2013).
Ma, F. et al. Single cell and spatial sequencing define processes by which keratinocytes and fibroblasts amplify inflammatory responses in psoriasis. Nat. Commun. 14, 3455 (2023).
Deng, C. C. et al. Single-cell RNA-seq reveals fibroblast heterogeneity and increased mesenchymal fibroblasts in human fibrotic skin diseases. Nat. Commun. 12, 3709 (2021).
Wang, L., Wang, B., Kou, E., Du, L. & Zhu, Y. New insight into the role of fibroblasts in the epithelial immune microenvironment in the single-cell era. Front. Immunol. 14, 1259515 (2023).
Cibrian, D., de la Fuente, H. & Sánchez-Madrid, F. Metabolic pathways that control skin homeostasis and inflammation. Trends Mol. Med. 26, 975–986 (2020).
Jiang, Y. et al. Nicotinamide metabolism face-off between macrophages and fibroblasts manipulates the microenvironment in gastric cancer. Cell Metab. 36, 1806–1822 (2024).
Notarangelo, G. et al. Oncometabolite d-2HG alters T cell metabolism to impair CD8(+) T cell function. Science 377, 1519–1529 (2022).
Chen, L. L. et al. Itaconate inhibits TET DNA dioxygenases to dampen inflammatory responses. Nat. Cell Biol. 24, 353–363 (2022).
Huang, B. & Cao, X. Metabolically targeting immunosuppression and immunoescape for future cancer immunotherapy: a narrative review. Holist. Integr. Oncol. 1, 15 (2022).
Chen, S. et al. CD73: an emerging checkpoint for cancer immunotherapy. Immunotherapy 11, 983–997 (2019).
Merleev, A. et al. Proprotein convertase subtilisin/kexin type 9 is a psoriasis-susceptibility locus that is negatively related to IL36G. JCI Insight 7, e141193 (2022).
Wu, Y. et al. Spatiotemporal immune landscape of colorectal cancer liver metastasis at single-cell level. Cancer Discov. 12, 134–153 (2022).
Mayr, B. & Montminy, M. Transcriptional regulation by the phosphorylation-dependent factor CREB. Nat. Rev. Mol. Cell Biol. 2, 599–609 (2001).
Liu, H. et al. Beneficial role of erythrocyte adenosine A2B receptor-mediated AMP-activated protein kinase activation in high-altitude hypoxia. Circulation 134, 405–421 (2016).
Liu, H. et al. ADORA1 Inhibition Promotes Tumor Immune Evasion by Regulating the ATF3-PD-L1 Axis. Cancer Cell 37, 324–339.e328 (2020).
Cheng, Z. et al. Foxo1 integrates insulin signaling with mitochondrial function in the liver. Nat. Med. 15, 1307–1311 (2009).
Converso, D. P. et al. HO-1 is located in liver mitochondria and modulates mitochondrial heme content and metabolism. FASEB J. 20, 1236–1238 (2006).
Taillé, C. et al. Induction of heme oxygenase-1 inhibits NADPH oxidase activity by down-regulating cytochrome b558 expression via the reduction of heme availability. J. Biol. Chem. 279, 28681–28688 (2004).
González-Halphen, D., Lindorfer, M. A. & Capaldi, R. A. Subunit arrangement in beef heart complex III. Biochemistry 27, 7021–7031 (1988).
Lynch, M. D. & Watt, F. M. Fibroblast heterogeneity: implications for human disease. J. Clin. Investig. 128, 26–35 (2018).
Gurtner, G. C., Werner, S., Barrandon, Y. & Longaker, M. T. Wound repair and regeneration. Nature 453, 314–321 (2008).
desJardins-Park, H. E., Foster, D. S. & Longaker, M. T. Fibroblasts and wound healing: an update. Regener. Med. 13, 491–495 (2018).
Xu, Z. et al. Anatomically distinct fibroblast subsets determine skin autoimmune patterns. Nature 601, 118–124 (2022).
Cai, X. et al. Tenascin C(+) papillary fibroblasts facilitate neuro-immune interaction in a mouse model of psoriasis. Nat. Commun. 14, 2004 (2023).
Aghajanian, H. et al. Targeting cardiac fibrosis with engineered T cells. Nature 573, 430–433 (2019).
Andrés, R. M. et al. Adenosine A(2A) and A(2B) receptors differentially modulate keratinocyte proliferation: possible deregulation in psoriatic epidermis. J. Investig. Dermatol. 137, 123–131 (2017).
Pasquini, S., Contri, C., Borea, P. A., Vincenzi, F. & Varani, K. Adenosine and inflammation: here, there and everywhere. Int. J. Mol. Sci. 22 (2021).
Novitskiy, S. V. et al. Adenosine receptors in regulation of dendritic cell differentiation and function. Blood 112, 1822–1831 (2008).
Karmouty-Quintana, H. et al. The A2B adenosine receptor modulates pulmonary hypertension associated with interstitial lung disease. FASEB J. 26, 2546–2557 (2012).
Zaynagetdinov, R. et al. Attenuation of chronic pulmonary inflammation in A2B adenosine receptor knockout mice. Am. J. Respir. Cell Mol. Biol. 42, 564–571 (2010).
De Palma, G., Caiazzo, E., Ialenti, A. & Cicala, C. P38 experimental evidence of ecto-5′-nucleotidase (CD73) modulation in psoriasis-like inflammation. Br. J. Dermatol. (2024).
Vijayan, D., Young, A., Teng, M. W. L. & Smyth, M. J. Targeting immunosuppressive adenosine in cancer. Nat. Rev. Cancer 17, 709–724 (2017).
Bach, N., Winzer, R., Tolosa, E., Fiedler, W. & Brauneck, F. The clinical significance of CD73 in cancer. Int. J. Mol. Sci. 24, 11759 (2023).
Young, A. et al. Co-inhibition of CD73 and A2AR adenosine signaling improves anti-tumor immune responses. Cancer Cell 30, 391–403 (2016).
Schön, M. P., Schön, M. & Klotz, K. N. The small antitumoral immune response modifier imiquimod interacts with adenosine receptor signaling in a TLR7- and TLR8-independent fashion. J. Investig. Dermatol. 126, 1338–1347 (2006).
Villarreal-Martinez, A. et al. Mitochondrial dysfunction: the pathological link between psoriasis and insulin resistance? J. Eur. Acad. Dermatol. Venereol. 37, 340–347 (2023).
Dobrică, E. C., Cozma, M. A., Găman, M. A., Voiculescu, V. M. & Găman, A. M. The involvement of oxidative stress in psoriasis: a systematic review. Antioxidants 11, 282 (2022).
Bansal, S., Biswas, G. & Avadhani, N. G. Mitochondria-targeted heme oxygenase-1 induces oxidative stress and mitochondrial dysfunction in macrophages, kidney fibroblasts and in chronic alcohol hepatotoxicity. Redox Biol. 2, 273–283 (2014).
Ayer, A., Zarjou, A., Agarwal, A. & Stocker, R. Heme oxygenases in cardiovascular health and disease. Physiol. Rev. 96, 1449–1508 (2016).
Yang, Y. et al. Targeting ferroptosis suppresses osteocyte glucolipotoxicity and alleviates diabetic osteoporosis. Bone Res. 10, 26 (2022).
Chang, L. C. et al. Heme oxygenase-1 mediates BAY 11-7085 induced ferroptosis. Cancer Lett. 416, 124–137 (2018).
Jais, A. et al. Heme oxygenase-1 drives metaflammation and insulin resistance in mouse and man. Cell 158, 25–40 (2014).
Kimura, S. et al. Increasing heme oxygenase-1-expressing macrophages indicates a tendency of poor prognosis in advanced colorectal cancer. Digestion 101, 401–410 (2020).
Chau, L. Y. Heme oxygenase-1: emerging target of cancer therapy. J. Biomed. Sci. 22, 22 (2015).
Mitterstiller, A. M. et al. Heme oxygenase 1 controls early innate immune response of macrophages to Salmonella Typhimurium infection. Cell. Microbiol. 18, 1374–1389 (2016).
Scharn, C. R. et al. Heme oxygenase-1 regulates inflammation and mycobacterial survival in human macrophages during Mycobacterium tuberculosis infection. J. Immunol. 196, 4641–4649 (2016).
Carasi, P. et al. Heme-oxygenase-1 expression contributes to the immunoregulation induced by fasciola hepatica and promotes infection. Front. Immunol. 8, 883 (2017).
Gorg, B. et al. O-GlcNAcylation-dependent upregulation of HO1 triggers ammonia-induced oxidative stress and senescence in hepatic encephalopathy. J. Hepatol. 71, 930–941 (2019).
Wen, J. et al. Impaired erectile function in CD73-deficient mice with reduced endogenous penile adenosine production. J. Sex. Med. 8, 2172–2180 (2011).
Liu, Y. et al. Profiling with senescence-associated secretory phenotype score identifies GDC-0879 as a small molecule sensitizing glioblastoma to anti-PD1. Cell Death Dis. 16, 602 (2025).
Lu, H. et al. Novel gene regulatory networks identified in response to nitro-conjugated linoleic acid in human endothelial cells. Physiol. Genomics 51, 224–233 (2019).
Huang, X. et al. Fast, long-term, super-resolution imaging with Hessian structured illumination microscopy. Nat. Biotechnol. 36, 451–459 (2018).
Zhao, W. et al. Sparse deconvolution improves the resolution of live-cell super-resolution fluorescence microscopy. Nat. Biotechnol. 40, 606–617 (2022).
Kardon, J. R. et al. Mitochondrial ClpX activates a key enzyme for heme biosynthesis and erythropoiesis. Cell 161, 858–867 (2015).
Vandekeere, S. et al. Serine synthesis via PHGDH is essential for heme production in endothelial cells. Cell Metab. 28, 573–587.e513 (2018).
Acknowledgements
This work was supported by the Key Program of National Natural Science Foundation of China (No. U22A20329 to H.L.), Fundamental and Interdisciplinary Disciplines Breakthrough Plan of the Ministry of Education of China (JYB2025XDXM603 to X.C.), the Natural Science Foundation of China for outstanding Young Scholars (No. 82022060 to H.L.), Key Program of National Natural Science Foundation of China (No.82130090 to X.C.), Science Found for Creative Research Groups of the National Natural Science Foundation of China (No. 82221002 to X.C.), National Natural Science Foundation of China (No. 82203931 and No. 82572076 to Y.T., No. 82504290 to J.G.), China Postdoctoral Science Foundation (No. BX20220355 to Y.T., No. 2023M733957 to Y.T., No. BX20250226 to J.G., and No. 2024M763721 to J.G.), The Excellent Youth Project of Hunan Provincial Natural Science Foundation (No. 2024JJ4094 to Y.T.), as well as The Scientific Research Program of FuRong Laboratory (No. 2023SK2095 to H.L., No. 2024PT5104 to Y.Z., 2025PT5010 to Y.T).
Author information
Authors and Affiliations
Contributions
Y.T., J.G., J.S., X.Z., G.Z., X.C. and H.L. developed the concepts and discussed experiments. Y.T. and J.G. performed the experiments and wrote the manuscript. J.S. and G.Z. performed bioinformatics analysis. Y.T., J.G., J.S., X.Z., L.Y., Y.Z., J.Z., C.X., X.X., Y.X. and H.B. analysed and discussed the data. X.Y., G.Z., and Y.Z. verified statistical methods. J.Z., C.X. and X.X. identified clinical samples. P.L. provided the normal foreskin tissues donated by circumcision patients. X.C. and H.L. revised the manuscript and supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Maria Cavinato, Daniela Sauma and the other, anonymous, reviewers for their contribution to the peer review of this work. A.peer review file is available
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tian, Y., Guo, J., Sun, J. et al. CD73high fibroblasts orchestrate keratinocyte inflammation in the psoriasis-associated epithelial immune microenvironment. Nat Commun (2026). https://doi.org/10.1038/s41467-026-71323-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-71323-0