Abstract
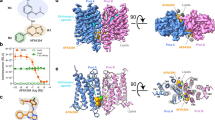
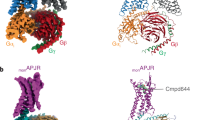
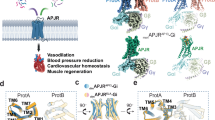
G-protein-coupled receptors (GPCRs) are significant signal transducers that exist as monomers and in multiple oligomeric forms. However, molecular mechanism driving their dynamic interconversion to regulate intricate signaling in class A GPCRs remains elusive, compounding our understanding of their related pathophysiological functions. Here, we present a set of 12 assemblies of the apelin receptor (APLNR), including dimeric apo state, agonistic small molecule- or nanobody-bound state of monomeric and dimeric APLNR with and without G-proteins, providing a detailed dynamic view of the monomer-dimer transition. High-resolution cryo-EM structures reveal that different ligands induce varying degrees of pre-dissociation of dimers in the absence of G-protein, with G-protein coupling facilitating the transition from dimeric to monomeric receptor. These insights enhance our understanding of the dynamic regulation of class A GPCRs between monomeric and dimeric forms and advance the rational drug design strategies aimed at selectively modulating of APLNR signaling.
Similar content being viewed by others
Data availability
The cryo-EM density map generated in this study of the apo APLNR dimer, JN241-9–dimerLBS1a, JN241-9–dimerLBS1b, JN241-9–dimerLBS1c, AMG986–dimerLBS2a, AMG986–dimerLBS2b, JN241-9–monomer–Gi1, JN241-9–dimer–Gi1, AMG986–monomer–Gi1, AMG986–dimerACT2a–Gi1 and AMG986–dimerACT2b–Gi1 have been deposited in the Electron Microscopy Data Bank (EMDB) under accession code EMD-63304 (apo APLNR dimer), EMD-63305 (JN241-9–dimerLBS1a), EMD-63307 (JN241-9–dimerLBS1b), EMD-63308 (JN241-9–dimerLBS1c), EMD-63309 (AMG986–dimerLBS2a), EMD-63310 (AMG986–dimerLBS2b), EMD-63312 (JN241-9–monomer–Gi1), EMD-63313 (JN241-9–dimer–Gi1), EMD-63314 (AMG986–monomer–Gi1), EMD-65254 (AMG986–dimerACT2a–Gi1), EMD-65255 (AMG986–dimerACT2b–Gi1), and model coordinates have been deposited in the Protein Data Bank (PDB) under accession number 9LQT (apo APLNR dimer), 9LQU (JN241-9–dimerLBS1a), 9LQW (JN241-9–dimerLBS1b), 9LQX (JN241-9–dimerLBS1c), 9LQY (AMG986–dimerLBS2a), 9LQZ (AMG986–dimerLBS2b), 9LR1 (JN241-9–monomer–Gi1), 9LR2 (JN241-9–dimer–Gi1), 9LR3 (AMG986–monomer–Gi1), 9VPM (AMG986–dimerACT2a–Gi1), 9VPN (AMG986–dimerACT2b–Gi1), respectively. All other data generated in this study are provided in the Supplementary Information and Source data files. Source data are provided with this paper.
References
Ginés, S. et al. Dopamine D1 and adenosine A1 receptors form functionally interacting heteromeric complexes. Proc. Natl. Acad. Sci. USA 97, 8606–8611 (2000).
Albizu, L. et al. Time-resolved FRET between GPCR ligands reveals oligomers in native tissues. Nat. Chem. Biol. 6, 587–594 (2010).
Angers, S. et al. Detection of beta 2-adrenergic receptor dimerization in living cells using bioluminescence resonance energy transfer (BRET). Proc. Natl. Acad. Sci. USA 97, 3684–3689 (2000).
Cai, X., Bai, B., Zhang, R., Wang, C. & Chen, J. Apelin receptor homodimer-oligomers revealed by single-molecule imaging and novel G protein-dependent signaling. Sci. Rep. 7, 40335 (2017).
Liu, J. et al. Biased signaling due to oligomerization of the G protein-coupled platelet-activating factor receptor. Nat. Commun. 13, 6365 (2022).
Fricke, F., Beaudouin, J., Eils, R. & Heilemann, M. One, two or three? Probing the stoichiometry of membrane proteins by single-molecule localization microscopy. Sci. Rep. 5, 14072 (2015).
Prinster, S. C., Hague, C. & Hall, R. A. Heterodimerization of g protein-coupled receptors: specificity and functional significance. Pharmacol. Rev. 57, 289–298 (2005).
Jordan, B. A., Trapaidze, N., Gomes, I., Nivarthi, R. & Devi, L. A. Oligomerization of opioid receptors with beta 2-adrenergic receptors: a role in trafficking and mitogen-activated protein kinase activation. Proc. Natl. Acad. Sci. USA 98, 343–348 (2001).
Bellot, M. et al. Dual agonist occupancy of AT1-R-α2C-AR heterodimers results in atypical Gs-PKA signaling. Nat. Chem. Biol. 11, 271–279 (2015).
Callén, L. et al. Cannabinoid receptors CB1 and CB2 form functional heteromers in brain. J. Biol. Chem. 287, 20851–20865 (2012).
Yue, Y. et al. Structural insight into apelin receptor-G protein stoichiometry. Nat. Struct. Mol. Biol. 29, 688–697 (2022).
Yue, Y. et al. Structural insights into the regulation of monomeric and dimeric apelin receptor. Nat. Commun. 16, 310 (2025).
Cox, C. M., D’Agostino, S. L., Miller, M. K., Heimark, R. L. & Krieg, P. A. Apelin, the ligand for the endothelial G-protein-coupled receptor, APJ, is a potent angiogenic factor required for normal vascular development of the frog embryo. Dev. Biol. 296, 177–189 (2006).
Szokodi, I. et al. Apelin, the novel endogenous ligand of the orphan receptor APJ, regulates cardiac contractility. Circ. Res. 91, 434–440 (2002).
Rossin, D. et al. APJ as promising therapeutic target of peptide analogues in myocardial infarction- and hypertension-induced heart failure. Pharmaceutics 15, https://doi.org/10.3390/pharmaceutics15051408 (2023).
Chapman, F. A. et al. The therapeutic potential of apelin in kidney disease. Nat. Rev. Nephrol. 17, 840–853 (2021).
Vinel, C. et al. The exerkine apelin reverses age-associated sarcopenia. Nat. Med. 24, 1360–1371 (2018).
Enoki, Y. et al. The G protein-coupled receptor ligand apelin-13 ameliorates skeletal muscle atrophy induced by chronic kidney disease. J. Cachexia Sarcopenia Muscle 14, 553–564 (2023).
Wang, W. W. et al. Structure-based design of non-hypertrophic apelin receptor modulator. Cell 187, 1460–1475.e1420 (2024).
Scimia, M. C. et al. APJ acts as a dual receptor in cardiac hypertrophy. Nature 488, 394–398 (2012).
Brame, A. L. et al. Design, characterization, and first-in-human study of the vascular actions of a novel biased apelin receptor agonist. Hypertension 65, 834–840 (2015).
Read, C. et al. Cardiac action of the first G protein biased small molecule apelin agonist. Biochem. Pharmacol. 116, 63–72 (2016).
Yeganeh-Hajahmadi, M., Moosavi-Saeed, Y. & Rostamzadeh, F. Apelin receptor dimerization and oligomerization. Curr. Mol. Pharmacol. 17, e180823219999 (2024).
Hu, S. et al. Apelin receptor dimer: classification, future prospects, and pathophysiological perspectives. Biochim. Biophys. Acta Mol. Basis Dis. 1870, 167257 (2024).
Cai, X. et al. Apelin receptor homodimerisation inhibits hippocampal neuronal autophagy via G protein-dependent signalling in vascular dementia. Mol. Neurobiol. https://doi.org/10.1007/s12035-024-04383-2 (2024).
Cai, X., Wang, D., Zhang, R., Chen, Y. & Chen, J. The transmembrane domains of GPCR dimers as targets for drug development. Drug Discov. Today 28, 103419 (2023).
Trivedi, A. et al. A Phase I, open-label, single-dose study to evaluate the pharmacokinetics, safety, and tolerability of AMG 986 in healthy Japanese subjects. Drugs R&D 22, 141–146 (2022).
Winkle, P. et al. A first-in-human study of AMG 986, a novel apelin receptor agonist, in healthy subjects and heart failure patients. Cardiovasc. Drugs Ther. 37, 743–755 (2023).
Ason, B. et al. Cardiovascular response to small-molecule APJ activation. JCI Insight 5, https://doi.org/10.1172/jci.insight.132898 (2020).
Ma, Y. et al. Structure-guided discovery of a single-domain antibody agonist against human apelin receptor. Sci. Adv. 6, eaax7379 (2020).
Ballesteros, J. A. & Weinstein, H. Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations in G protein-coupled receptors. In Methods in Neurosciences (ed Sealfon, S. C.) 366–428 (Academic Press, 1995).
McCorvy, J. D. et al. Structural determinants of 5-HT(2B) receptor activation and biased agonism. Nat. Struct. Mol. Biol. 25, 787–796 (2018).
Liu, W. et al. Serial femtosecond crystallography of G protein-coupled receptors. Science 342, 1521–1524 (2013).
Wang, W., Qiao, Y. & Li, Z. New insights into modes of GPCR activation. Trends Pharmacol. Sci. 39, 367–386 (2018).
Nemoto, W. & Toh, H. Prediction of interfaces for oligomerizations of G-protein coupled receptors. Proteins 58, 644–660 (2005).
Moreno, J. L., Holloway, T. & González-Maeso, J. G protein-coupled receptor heterocomplexes in neuropsychiatric disorders. Prog. Mol. Biol. Transl. Sci. 117, 187–205 (2013).
Guidolin, D., Agnati, L. F., Marcoli, M., Borroto-Escuela, D. O. & Fuxe, K. G-protein-coupled receptor type A heteromers as an emerging therapeutic target. Expert Opin. Ther. Targets 19, 265–283 (2015).
Guixà-González, R. et al. Membrane omega-3 fatty acids modulate the oligomerisation kinetics of adenosine A2A and dopamine D2 receptors. Sci. Rep. 6, 19839 (2016).
Asico, L. D., Rozyyev, S., Crusan, A. M., Jose, P. A. & Villar, V. A. M. Elucidating the role of lipid rafts on G protein-coupled receptor function in the mouse kidney: an in vivo approach. Methods Mol. Biol. 2187, 187–206 (2021).
Chini, B. & Parenti, M. G-protein coupled receptors in lipid rafts and caveolae: How, when and why do they go there?. J. Mol. Endocrinol. 32, 325–338 (2004).
Jakubík, J. & El-Fakahany, E. E. Allosteric modulation of GPCRs of Class A by cholesterol. Int. J. Mol. Sci. 22, https://doi.org/10.3390/ijms22041953 (2021).
Duncan, A. L., Song, W. & Sansom, M. S. P. Lipid-dependent regulation of ion channels and G protein-coupled receptors: insights from structures and simulations. Annu. Rev. Pharmacol. Toxicol. 60, 31–50 (2020).
Villar, V. A., Cuevas, S., Zheng, X. & Jose, P. A. Localization and signaling of GPCRs in lipid rafts. Methods Cell Biol. 132, 3–23 (2016).
Legler, D. F. et al. Modulation of chemokine receptor function by cholesterol: new prospects for pharmacological intervention. Mol. Pharmacol. 91, 331–338 (2017).
Mao, C. et al. Cryo-EM structures of inactive and active GABA(B) receptor. Cell Res. 30, 564–573 (2020).
Ma, S. et al. Molecular basis for hormone recognition and activation of corticotropin-releasing factor receptors. Mol. Cell 77, 669–680.e664 (2020).
Bader, J. E. & Beck-Sickinger, A. G. Fluorescence resonance energy transfer to study receptor dimerization in living cells. Methods Mol. Biol. 259, 335–352 (2004).
Oliveira, P. A. et al. Angiotensin II type 1/adenosine A (2A) receptor oligomers: a novel target for tardive dyskinesia. Sci. Rep. 7, 1857 (2017).
Mellado, M. et al. Chemokine receptor homo- or heterodimerization activates distinct signaling pathways. EMBO J. 20, 2497–2507 (2001).
Hu, J. et al. Novel structural and functional insights into M3 muscarinic receptor dimer/oligomer formation. J. Biol. Chem. 288, 34777–34790 (2013).
Harikumar, K. G. et al. Glucagon-like peptide-1 receptor dimerization differentially regulates agonist signaling but does not affect small molecule allostery. Proc. Natl. Acad. Sci. USA 109, 18607–18612 (2012).
Pioszak, A. A., Harikumar, K. G., Parker, N. R., Miller, L. J. & Xu, H. E. Dimeric arrangement of the parathyroid hormone receptor and a structural mechanism for ligand-induced dissociation. J. Biol. Chem. 285, 12435–12444 (2010).
Ke, R., Lok, S. I. S., Singh, K., Chow, B. K. C. & Lee, L. T. O. GIP receptor suppresses PAC1receptor-mediated neuronal differentiation via formation of a receptor heterocomplex. J. Neurochem. 157, 1850–1860 (2021).
Calebiro, D. et al. Single-molecule analysis of fluorescently labeled G-protein-coupled receptors reveals complexes with distinct dynamics and organization. Proc. Natl. Acad. Sci. USA 110, 743–748 (2013).
Hern, J. A. et al. Formation and dissociation of M1 muscarinic receptor dimers seen by total internal reflection fluorescence imaging of single molecules. Proc. Natl. Acad. Sci. USA 107, 2693–2698 (2010).
Kasai, R. S. et al. Full characterization of GPCR monomer-dimer dynamic equilibrium by single molecule imaging. J. Cell Biol. 192, 463–480 (2011).
Tabor, A. et al. Visualization and ligand-induced modulation of dopamine receptor dimerization at the single molecule level. Sci. Rep. 6, 33233 (2016).
Masullo, L. A.-O. et al. Spatial and stoichiometric in situ analysis of biomolecular oligomerization at single-protein resolution. Nat. Commun. 16, 4202 (2025).
Walker, G. A.-O. et al. Oligomeric organization of membrane proteins from native membranes at nanoscale spatial and single-molecule resolution. Nat. Nanotechnol. 19, 85–94 (2024).
Liang, Y. L. et al. Dominant negative G proteins enhance formation and purification of agonist-GPCR-G protein complexes for structure determination. ACS Pharmacol. Transl. Sci. 1, 12–20 (2018).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Kimanius, D., Dong, L., Sharov, G., Nakane, T. & Scheres, S. H. W. New tools for automated cryo-EM single-particle analysis in RELION-4.0. Biochem. J. 478, 4169–4185 (2021).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Pettersen, E. F. et al. UCSF Chimera-a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Acknowledgements
The authors thank the Cryo-Electron Microscopy Center of Liangzhu laboratory for help with cryo-EM data collection of apo, AMG986-, JN241-9-bound APLNR, and AMG986-, JN241-9-bound APLNR–Gi1–scFv16 complexes. This project was supported by National Natural Science Foundation of China grant (32430051, 92353303, 32141004, 81922071 to Y.Z., 32330049 to Q.S., and 32400575 to W.W.); the “Pioneer” and “Leading Goose” R&D Program of Zhejiang (2024C03147 to Y.Z.); the Key R&D Projects of Zhejiang Province (2021C03039 to Y.Z.); The STI2030-Major Projects (2022ZD0205400 to Q.S.), the China Postdoctoral Science Foundation (2024T170783 to W.W.); Postdoctoral Fellowship Program of CPSF (GZC20232326 to W.W.); Zhejiang Provincial Postdoctoral Research Project (ZJ2024043 to W.W.); Hubei Province Key Laboratory of Ischemic Cardiovascular Disease Open Fund Project (SZ202405 to W.W.); Y.Z. is also supported by the Fundamental Research Funds for the Central Universities and Peak Discipline Cultivation Program of Zhejiang University School of Basic Medical Sciences.
Author information
Authors and Affiliations
Contributions
Y.Z. initiated the study, conceived and supervised the whole project; Y.Z., Q.S., S.-Y.J., W.-W.W., and Y.Y. participated in the data analysis and interpretation. S.-Y.J. and W.-W.W. designed the constructs of APLNR and expressed the proteins; S.-Y.J., W.-W.W., and P.X. purified these protein complexes; Y.Y. generated APLNR mutants for the cell-based G-protein activity assays and β-arrestin recruitment assays. Y.Y. generated APLNR mutants for the cell-based cAMP accumulation assays. X.Z. and D.-D.S. evaluated the samples by negative-stain EM; S.-K.Z. collected the cryo-EM data, and S.-Y.J. performed cryo-EM data processing. S.-Y.J. and Y.Y. performed model building; Y.Y. and J.Z. performed the cellular functional assays. S.-Y.J. and W.-W.W. performed structural analysis supervised by Y.Z.; W.-W.W. and S.-Y.J. prepared the figures; S.-Y.J., W.-W.W., Q.S., and Y.Z. wrote the manuscript; Y.Z., Q.S., K.X., and C.M. provided important discussions and essential revisions. S.-Y.J., W.-W.W., and Y.Y. provided Figs. 1–4, Supplementary Figs. 1–8, Supplementary Tables 1 and 2 and Supplementary Note 1.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Moritz Bünemann and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ji, SY., Wang, WW., Yang, Y. et al. Dynamic monomer-dimer transition in ligand-induced apelin receptor activation. Nat Commun (2026). https://doi.org/10.1038/s41467-026-71325-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-71325-y