Abstract
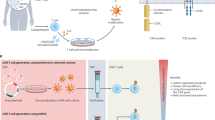
Despite striking efficacy against hematologic malignancies, the cost and complexity of CAR T manufacturing present significant barriers to broader patient access. Beyond manufacturing challenges, ex vivo expansion of T cells may be detrimental to their function and persistence. Thus, delivery of CARs to reprogram host cells in vivo would represent a significant advance towards a readily available therapy, but has been limited by low efficiency, low specificity, and immunogenicity of viral vectors. Here, we describe the design of pseudotyped lentiviral vectors (LV) with superior functionality and high target specificity. We show that LV pseudotyped with chimeric envelope glycoproteins from dolphin morbillivirus (DMV) can be engineered to selectively infect human T cells and evade neutralizing antibody responses in measles-vaccinated human serum. We further demonstrate that camelid-derived nanobodies are a superior retargeting domain, overcoming limitations inherent to the use of single-chain variable fragment antibodies. Using a chimeric DMV-pseudotyped virus targeting the CD7 receptor, we demonstrate efficient and highly specific infection of T cells both in vitro and in vivo, generating functional CAR T cells and inducing therapeutic efficacy in a preclinical B cell lymphoma model.
Similar content being viewed by others
Data availability
Data supporting the findings of this work are available within the paper and the Source Data file. The immunofluorescence data generated in this study have been deposited in the BioImage Archive under accession code S-BIAD2901. Source data are provided with this paper.
References
Merten, O.-W., Hebben, M. & Bovolenta, C. Production of lentiviral vectors. Mol. Ther. Methods Clin. Dev. 3, 16017 (2016).
Raguram, A., Banskota, S. & Liu, D. R. Therapeutic in vivo delivery of gene editing agents. Cell 185, 2806–2827 (2022).
Milone, M. C. & O’Doherty, U. Clinical use of lentiviral vectors. Leukemia 32, 1529–1541 (2018).
Sweeney, N. P. & Vink, C. A. The impact of lentiviral vector genome size and producer cell genomic to gag-pol mRNA ratios on packaging efficiency and titre. Mol. Ther. Methods Clin. Dev. 21, 574–584 (2021).
Abordo-Adesida, E. et al. Stability of lentiviral vector-mediated transgene expression in the brain in the presence of systemic antivector immune responses. Hum. Gene Ther. 16, 741–751 (2005).
Baekelandt, V., Eggermont, K., Michiels, M., Nuttin, B. & Debyser, Z. Optimized lentiviral vector production and purification procedure prevents immune response after transduction of mouse brain. Gene Ther. 10, 1933–1940 (2003).
Argaw, T. et al. In vivo targeting of lentiviral vectors pseudotyped with the Tupaia paramyxovirus H glycoprotein bearing a cell-specific ligand. Mol. Ther. Methods Clin. Dev. 21, 670–680 (2021).
Frank, A. M. & Buchholz, C. J. Surface-engineered lentiviral vectors for selective gene transfer into subtypes of lymphocytes. Mol. Ther. Methods Clin. Dev. 12, 19–31 (2019).
Buchholz, C. J., Friedel, T. & Büning, H. Surface-engineered viral vectors for selective and cell type-specific gene delivery. Trends Biotechnol. 33, 777–790 (2015).
Zhou, Q. et al. T-cell receptor gene transfer exclusively to human CD8(+) cells enhances tumor cell killing. Blood 120, 4334–4342 (2012).
Zhou, Q. et al. Exclusive transduction of human CD4+ T cells upon systemic delivery of CD4-targeted lentiviral vectors. J. Immunol. 195, 2493–2501 (2015).
Verhoeyen, E. et al. IL-7 surface-engineered lentiviral vectors promote survival and efficient gene transfer in resting primary T lymphocytes. Blood 101, 2167–2174 (2003).
Maurice, M. et al. Efficient gene transfer into human primary blood lymphocytes by surface-engineered lentiviral vectors that display a T cell-activating polypeptide. Blood 99, 2342–2350 (2002).
Yang, L., Bailey, L., Baltimore, D. & Wang, P. Targeting lentiviral vectors to specific cell types in vivo. Proc. Natl. Acad. Sci. USA 103, 11479–11484 (2006).
Funke, S. et al. Pseudotyping lentiviral vectors with the wild-type measles virus glycoproteins improves titer and selectivity. Gene Ther. 16, 700–705 (2009).
Lévy, C. et al. Measles virus envelope pseudotyped lentiviral vectors transduce quiescent human HSCs at an efficiency without precedent. Blood Adv. 1, 2088–2104 (2017).
Anliker, B. et al. Specific gene transfer to neurons, endothelial cells and hematopoietic progenitors with lentiviral vectors. Nat. Methods 7, 929–935 (2010).
Hashiguchi, T. et al. Structure of the measles virus hemagglutinin bound to its cellular receptor SLAM. Nat. Struct. Mol. Biol. 18, 135–141 (2011).
Navaratnarajah, C. K., Leonard, V. H. J. & Cattaneo, R. Measles virus glycoprotein complex assembly, receptor attachment, and cell entry. Curr. Top. Microbiol Immunol. 329, 59–76 (2009).
Takeda, M., Tahara, M., Nagata, N. & Seki, F. Wild-Type Measles Virus is Intrinsically Dual-Tropic. Front Microbiol 2, 279 (2011).
Lin, L.-T. & Richardson, C. D. The host cell receptors for measles virus and their interaction with the viral Hemagglutinin (H) Protein. Viruses 8, (2016).
White, J. M., Delos, S. E., Brecher, M. & Schornberg, K. Structures and mechanisms of viral membrane fusion proteins: multiple variations on a common theme. Crit. Rev. Biochem. Mol. Biol. 43, 189–219 (2008).
Duvergé, A. & Negroni, M. Pseudotyping lentiviral vectors: when the clothes make the Virus. Viruses 12, (2020).
Ou, W. et al. Specific targeting of human interleukin (IL)−13 receptor α2-positive cells with lentiviral vectors displaying IL-13. Hum. Gene Ther. Methods 23, 137–147 (2012).
Frank, A. M. et al. CD8-specific designed ankyrin repeat proteins improve selective gene delivery into human and primate T lymphocytes. Hum. Gene Ther. 31, 679–691 (2020).
Münch, R. C. et al. DARPins: an efficient targeting domain for lentiviral vectors. Mol. Ther. 19, 686–693 (2011).
Lonez, C. & Breman, E. Allogeneic CAR-T therapy technologies: Has the promise been met? Cells 13, 146 (2024).
Friedel, T. et al. Receptor-targeted lentiviral vectors are exceptionally sensitive toward the biophysical properties of the displayed single-chain Fv. Protein Eng. Des. Sel. 28, 93–106 (2015).
Lech, P. J. et al. Antibody neutralization of retargeted measles viruses. Virology 454-455, 237–246 (2014).
Bah, E. S., Nace, R. A., Peng, K. W., Muñoz-Alía, M. Á & Russell, S. J. Retargeted and stealth-modified oncolytic measles viruses for systemic cancer therapy in measles immune patients. Mol. Cancer Ther. 19, 2057–2067 (2020).
Marino, M. P. et al. A scalable method to concentrate lentiviral vectors pseudotyped with measles virus glycoproteins. Gene Ther. 22, 280–285 (2015).
Bieback, K. et al. Hemagglutinin protein of wild-type measles virus activates toll-like receptor 2 signaling. J. Virol. 76, 8729–8736 (2002).
Rall, G. F. et al. A transgenic mouse model for measles virus infection of the brain. Proc. Natl. Acad. Sci. USA 94, 4659–4663 (1997).
Chiu, M. L., Goulet, D. R., Teplyakov, A. & Gilliland, G. L. Antibody structure and function: The basis for engineering therapeutics. Antibodies 8, 55 (2019).
Sun, W. et al. A combined strategy improves the solubility of aggregation-prone single-chain variable fragment antibodies. Protein Expr. Purif. 83, 21–29 (2012).
Völkel, T., Müller, R. & Kontermann, R. E. Isolation of endothelial cell-specific human antibodies from a novel fully synthetic scFv library. Biochem. Biophys. Res. Commun. 317, 515–521 (2004).
Ahmad, Z. A. et al. scFv antibody: principles and clinical application. Clin. Dev. Immunol. 2012, 980250 (2012).
Goyvaerts, C. et al. Development of the nanobody display technology to target lentiviral vectors to antigen-presenting cells. Gene Ther. 19, 1133–1140 (2012).
Strebinger, D. et al. Cell type-specific delivery by modular envelope design. Nat. Commun. 14, 5141 (2023).
Dobson, C. S. et al. Antigen identification and high-throughput interaction mapping by reprogramming viral entry. Nat. Methods 19, 449–460 (2022).
Hamilton, J. R. et al. In vivo human T cell engineering with enveloped delivery vehicles. Nat. Biotechnol. https://doi.org/10.1038/s41587-023-02085-z (2024).
Nikolic, J. et al. Structural basis for the recognition of LDL-receptor family members by VSV glycoprotein. Nat. Commun. 9, 1029 (2018).
Sun, X., Yau, V. K., Briggs, B. J. & Whittaker, G. R. Role of clathrin-mediated endocytosis during vesicular stomatitis virus entry into host cells. Virology 338, 53–60 (2005).
Morizono, K. et al. The soluble serum protein Gas6 bridges virion envelope phosphatidylserine to the TAM receptor tyrosine kinase Axl to mediate viral entry. Cell Host Microbe 9, 286–298 (2011).
Sommermeyer, D. et al. Chimeric antigen receptor-modified T cells derived from defined CD8+ and CD4+ subsets confer superior antitumor reactivity in vivo. Leukemia 30, 492–500 (2016).
Russell, S. J. et al. Remission of disseminated cancer after systemic oncolytic virotherapy. Mayo Clin. Proc. 89, 926–933 (2014).
Dispenzieri, A. et al. Phase I trial of systemic administration of Edmonston strain of measles virus genetically engineered to express the sodium iodide symporter in patients with recurrent or refractory multiple myeloma. Leukemia 31, 2791–2798 (2017).
Lévy, C. et al. Lentiviral vectors displaying modified measles virus gp overcome pre-existing immunity in in vivo-like transduction of human T and B cells. Mol. Ther. 20, 1699–1712 (2012).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Bouche, F. B., Ertl, O. T. & Muller, C. P. Neutralizing B cell response in measles. Viral Immunol. 15, 451–471 (2002).
de Swart, R. L., Yüksel, S., Langerijs, C. N., Muller, C. P. & Osterhaus, A. D. M. E. Depletion of measles virus glycoprotein-specific antibodies from human sera reveals genotype-specific neutralizing antibodies. J. Gen. Virol. 90, 2982–2989 (2009).
de Swart, R. L., Yüksel, S. & Osterhaus, A. D. M. E. Relative contributions of measles virus hemagglutinin- and fusion protein-specific serum antibodies to virus neutralization. J. Virol. 79, 11547–11551 (2005).
Bender, R. R. et al. Receptor-targeted Nipah virus glycoproteins improve cell-type selective gene delivery and reveal a preference for membrane-proximal cell attachment. PLoS Pathog. 12, e1005641 (2016).
Enkirch, T. et al. Targeted lentiviral vectors pseudotyped with the Tupaia paramyxovirus glycoproteins. Gene Ther. 20, 16–23 (2013).
Engeland, C. E. et al. A Tupaia paramyxovirus vector system for targeting and transgene expression. J. Gen. Virol. 98, 2248–2257 (2017).
Plattet, P. et al. Conserved leucine residue in the head region of morbillivirus fusion protein regulates the large conformational change during fusion activity. Biochemistry 48, 9112–9121 (2009).
von Messling, V. & Cattaneo, R. Amino-terminal precursor sequence modulates canine distemper virus fusion protein function. J. Virol. 76, 4172–4180 (2002).
von Messling, V., Milosevic, D., Devaux, P. & Cattaneo, R. Canine distemper virus and measles virus fusion glycoprotein trimers: partial membrane-proximal ectodomain cleavage enhances function. J. Virol. 78, 7894–7903 (2004).
Funke, S. et al. Targeted cell entry of lentiviral vectors. Mol. Ther. 16, 1427–1436 (2008).
Muñoz-Alía, M. A. & Russell, S. J. Probing Morbillivirus Antisera neutralization using functional chimerism between measles virus and canine distemper virus envelope glycoproteins. Viruses 11, 688 (2019).
Corey Elizabeth, A. & Iorio Ronald, M. Mutations in the stalk of the measles virus hemagglutinin protein decrease fusion but do not interfere with virus-specific interaction with the homologous fusion protein. J. Virol. 81, 9900–9910 (2007).
Rust, B. J. et al. Envelope-specific adaptive immunity following transplantation of hematopoietic stem cells modified with VSV-G Lentivirus. Mol. Ther. Methods Clin. Dev. 19, 438–446 (2020).
DePolo, N. J. et al. VSV-G pseudotyped lentiviral vector particles produced in human cells are inactivated by human serum. Mol. Ther. 2, 218–222 (2000).
Baldauf, H.-M. et al. SAMHD1 restricts HIV-1 infection in resting CD4(+) T cells. Nat. Med. 18, 1682–1687 (2012).
Doitsh, G. et al. Abortive HIV infection mediates CD4 T cell depletion and inflammation in human lymphoid tissue. Cell 143, 789–801 (2010).
Zack, J. A. et al. HIV-1 entry into quiescent primary lymphocytes: molecular analysis reveals a labile, latent viral structure. Cell 61, 213–222 (1990).
Finkelshtein, D., Werman, A., Novick, D., Barak, S. & Rubinstein, M. LDL receptor and its family members serve as the cellular receptors for vesicular stomatitis virus. Proc. Natl. Acad. Sci. USA 110, 7306–7311 (2013).
Amirache, F. et al. Mystery solved: VSV-G-LVs do not allow efficient gene transfer into unstimulated T cells, B cells, and HSCs because they lack the LDL receptor. Blood 123, 1422–1424 (2014).
Zhang, D. K. Y. et al. Enhancing CAR-T cell functionality in a patient-specific manner. Nat. Commun. 14, 1–13 (2023).
Mescher, M. F. et al. Signals required for programming effector and memory development by CD8+ T cells. Immunol. Rev. 211, 81–92 (2006).
Melenhorst, J. J. et al. Decade-long leukaemia remissions with persistence of CD4+ CAR T cells. Nature 602, 503–509 (2022).
Liu, E. et al. Use of CAR-transduced natural killer cells in CD19-positive lymphoid tumors. N. Engl. J. Med. 382, 545–553 (2020).
Marin, D. et al. Safety, efficacy and determinants of response of allogeneic CD19-specific CAR-NK cells in CD19+ B cell tumors: a phase 1/2 trial. Nat. Med. 30, 772–784 (2024).
Chambers, B. J., Salcedo, M. & Ljunggren, H. G. Triggering of natural killer cells by the costimulatory molecule CD80 (B7-1). Immunity 5, 311–317 (1996).
van Til, N. P. et al. Kupffer cells and not liver sinusoidal endothelial cells prevent lentiviral transduction of hepatocytes. Mol. Ther. 11, 26–34 (2005).
Milani, M. et al. Phagocytosis-shielded lentiviral vectors improve liver gene therapy in nonhuman primates. Sci. Transl. Med. 11, eaav7325 (2019).
Morizono, K. et al. Redirecting lentiviral vectors pseudotyped with Sindbis virus-derived envelope proteins to DC-SIGN by modification of N-linked glycans of envelope proteins. J. Virol. 84, 6923–6934 (2010).
Ziegler, L. et al. Targeting lentiviral vectors to antigen-specific immunoglobulins. Hum. Gene Ther. 19, 861–872 (2008).
Zhang, X.-Y. et al. Cell-specific targeting of lentiviral vectors mediated by fusion proteins derived from Sindbis virus, vesicular stomatitis virus, or avian sarcoma/leukosis virus. Retrovirology 7, 3 (2010).
Aires da Silva, F., Costa, M. J. L., Corte-Real, S. & Goncalves, J. Cell type-specific targeting with Sindbis pseudotyped lentiviral vectors displaying anti-CCR5 single-chain antibodies. Hum. Gene Ther. 16, 223–234 (2005).
Muñoz-Alía, M. Á, Nace, R. A., Zhang, L. & Russell, S. J. Serotypic evolution of measles virus is constrained by multiple co-dominant B cell epitopes on its surface glycoproteins. Cell Rep. Med. 2, 100225 (2021).
Aditi & Shariff, M. Nipah virus infection: A review. Epidemiol. Infect. 147, e95 (2019).
Pallister, J. A. et al. Vaccination of ferrets with a recombinant G glycoprotein subunit vaccine provides protection against Nipah virus disease for over 12 months. Virol. J. 10, 237 (2013).
Nieba, L., Honegger, A., Krebber, C. & Plückthun, A. Disrupting the hydrophobic patches at the antibody variable/constant domain interface: improved in vivo folding and physical characterization of an engineered scFv fragment. Protein Eng. 10, 435–444 (1997).
Xu, J. et al. Nanobodies from camelid mice and llamas neutralize SARS-CoV-2 variants. Nature 595, 278–282 (2021).
Ruffin, N. et al. Low SAMHD1 expression following T-cell activation and proliferation renders CD4+ T cells susceptible to HIV-1. AIDS 29, 519–530 (2015).
Uchida, N. et al. TRIM5α variation affects Lentiviral transduction efficiency for long-term hematopoietic repopulating cells in a rhesus stem cell transplantation model. Blood 118, 2056–2056 (2011).
Arcangeli, S. et al. CAR T cell manufacturing from naive/stem memory T lymphocytes enhances antitumor responses while curtailing cytokine release syndrome. J. Clin. Invest. 132, e150807 (2022).
Han, J., Tam, K., Tam, C., Hollis, R. P. & Kohn, D. B. Improved lentiviral vector titers from a multi-gene knockout packaging line. Mol. Ther. Oncol. 23, 582–592 (2021).
Dautzenberg, I. J. C., Rabelink, M. J. W. E. & Hoeben, R. C. The stability of envelope-pseudotyped lentiviral vectors. Gene Ther. 28, 89–104 (2021).
Moreira, A. S. et al. Implementation of novel affinity ligand for Lentiviral vector purification. Int. J. Mol. Sci. 24, 3354 (2023).
Annoni, A., Gregori, S., Naldini, L. & Cantore, A. Modulation of immune responses in lentiviral vector-mediated gene transfer. Cell. Immunol. 342, 103802 (2019).
Arduini, A., Katiyar, H. & Liang, C. Progress in pseudotyping Lentiviral vectors towards cell-specific gene delivery in vivo. Viruses 17, 802 (2025).
Shirley, J. L., de Jong, Y. P., Terhorst, C. & Herzog, R. W. Immune responses to viral gene therapy vectors. Mol. Ther. 28, 709–722 (2020).
Weidner, T. et al. Genetic in vivo engineering of human T lymphocytes in mouse models. Nat. Protoc. 16, 3210–3240 (2021).
Bankhead, P. et al. QuPath: Open source software for digital pathology image analysis. Sci. Rep. 7, 1–7 (2017).
Goldsborough, T. et al. A novel channel invariant architecture for the segmentation of cells and nuclei in multiplexed images using InstanSeg. bioRxiv https://doi.org/10.1101/2024.09.04.611150 (2024).
Acknowledgements
This work was supported by the Merkin Institute for Transformative Technologies in Healthcare and Calico Life Sciences. This work was supported in part by funding from Calico Life Sciences, LLC. We thank Branka Horvat and the Measles Virus Biobank for the generous gift of the CL55 and Y503 antibodies, INSERM, France, Measles Virus Biobank https://ciri.ens-lyon.fr/teams/IbIV/measles-biobank. We thank Natan Pirete and the Broad Institute Flow Cytometry Core for their help, training, and assistance in our flow cytometry experiments, and the Broad Institute mouse facilities for providing training and assistance with in vivo experiments. We would like to thank Professor Fritz Melcher from the German Rheumatology Research Center Leibniz-Institute for providing the FGK4.5 hybridoma, Dr. Hasaya Akiba for providing the RMT3-23 hybridoma, and Dr. Yoji Murata for providing the MY-1 and Miap301 hybridomas.
Author information
Authors and Affiliations
Contributions
K.S.M., K.B.Y., and R.T.M. conceived the project. K.E.I., K.S.M., and P.M.A. designed and conducted the majority of experiments, analyzed experimental data, and wrote the manuscript. O.I.A., A.J.M., S.K.L.R., A.R., and N.H.K. contributed to mouse and viral experiments. C.C. conducted preliminary experiments. J.M.C. conducted liver histology and analysis and contributed to manuscript preparation. K.B.Y. and R.T.M. supervised the work and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
R.T.M., K.B.Y., K.S.M., K.E.I., and P.M.A. are inventors on patent applications filed by the Broad Institute relating to retargeted retroviral vector technologies and the use of morbillivirus-derived and VHH-targeted vectors described in this work (U.S. Patent Application Publications US 2024/0294942 A1 and US 2024/0307555 A1). R.T.M. has received speaking or consulting fees from Bristol Myers Squibb, Gilead Sciences, Kumquat Biosciences, Immunai Therapeutics, and BioNTech and has equity ownership in OncoRev, LLC, and Jumble Therapeutics. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Johannes Heidbuechel and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ibrahim, K.E., Mears, K.S., Allen, P.M. et al. In vivo CAR T cell generation using retargeted and functionalized lentiviral vectors with reduced immunogenicity. Nat Commun (2026). https://doi.org/10.1038/s41467-026-71395-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-71395-y