Abstract
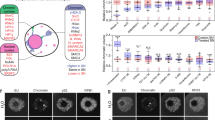
Oocyte-specific isoforms play crucial roles in oocyte maturation, while current understanding of the oocyte transcriptome is mainly focused on gene level. Here, we utilize single-cell full-length isoform sequencing to detect entire transcripts in human and mouse oocytes. Isoform diversity during oocyte maturation is systematically profiled, including 7154 and 4875 putative novel human and mouse transcripts, respectively. More than half of novel isoforms are categorized as novel-not-in-catalog (NNC) and may serve specific functions in oocytes. For example, ARHGAP18 mainly encoded by novel isoforms colocalizes with microtubules, and targeted knockdown of novel isoforms disrupts oocyte maturation. Moreover, approximately 30% of NNC isoforms are derived from transposable elements, and their incorporation within transcripts could enhance isoform stability during oocyte maturation. Altogether, our findings represent a valuable resource showcasing the complexity and diversity of RNA isoforms in oocytes, as well as transposable element co-option for novel isoform generation and isoform stability enhancement.
Similar content being viewed by others
Data availability
All the raw sequencing data generated in this study have been deposited in the Genome Sequence Archive for human (GSA-Human) and Genome Sequence Archive of China National Center for Bioinformation (Human: HRA006583, Mouse: CRA014675). The processed data including assembled transcriptomes (annotated gtf files), new splice junctions and expression tables are publicly available in the OMIX, China National Center for Bioinformation (OMIX012135 [https://ngdc.cncb.ac.cn/omix/release/OMIX012135], OMIX012137). Publicly available oocyte short-read RNA-seq datasets used in this work were retrieved from SRA (SRP011546 [https://www.ncbi.nlm.nih.gov/sra/?term=SRP011546], SRP285893, SRP361878, SRP086707, SRP189718, SRP062106). Publicly available deepCAGE and PAIso-seq datasets used in this work were retrieved from FANTOM5 [https://fantom.gsc.riken.jp/5/datafiles/latest/basic/mouse.tissue.hCAGE/, https://fantom.gsc.riken.jp/5/datafiles/reprocessed/hg38_latest/basic/human.tissue.hCAGE/], GSA-Human (HRA001288 [https://ngdc.cncb.ac.cn/gsa-human/browse/HRA001288]) and SRA under the accession number PRJNA529588 [https://www.ncbi.nlm.nih.gov/biosample?Db=biosample&DbFrom=bioproject&Cmd=Link&LinkName=bioproject_biosample&LinkReadableName=BioSample&ordinalpos=1&IdsFromResult=529588]. Publicly available multi-tissue short-read RNA-seq data used in this work was retrieved from ENCODE portal (https://www.encodeproject.org/) with the following identifiers: ENCSR278TQR, ENCSR197GCF, ENCSR197GCF, ENCSR344MQK, ENCSR029KNZ, ENCSR000AEU, ENCSR000AFB, ENCSR621PZI, ENCSR096LTX, ENCSR150QJY, ENCSR504NIU, ENCSR759TPN, ENCSR773COB. UCSC genome browser sessions have been created for the human and mouse assembled transcriptomes (https://genome.ucsc.edu/s/kratos12138/Human%20oocyte%20long%2Dread%20transcriptome; https://genome.ucsc.edu/s/kratos12138/Mouse%20oocyte%20long%2Dread%20transcriptome). Source data are provided with this paper.
Code availability
All analyses utilized publicly available algorithms and software. The code used to generate the figures has been deposited on GitHub (https://github.com/Kratos12138/Oocyte-long-read-transcriptome).
References
Cheng, S. et al. Mammalian oocytes store mRNAs in a mitochondria-associated membraneless compartment. Science 378, eabq4835 (2022).
Su, Y. Q. et al. Selective degradation of transcripts during meiotic maturation of mouse oocytes. Dev. Biol. 302, 104–117 (2007).
Wan, Y. et al. LSM14B is essential for oocyte meiotic maturation by regulating maternal mRNA storage and clearance. Nucleic Acids Res. 51, 11652–11667 (2023).
Do, D. V. et al. SRSF3 maintains transcriptome integrity in oocytes by regulation of alternative splicing and transposable elements. Cell Discov. 4, 33 (2018).
Kocabas, A. M. et al. The transcriptome of human oocytes. Proc. Natl. Acad. Sci. USA 103, 14027–14032 (2006).
Llonch, S. et al. Single human oocyte transcriptome analysis reveals distinct maturation stage-dependent pathways impacted by age. Aging Cell 20, e13360 (2021).
Pietroforte S. et al. Specific processing of meiosis-related transcript is linked to final maturation in human oocytes. Mol Hum Reprod 29, gaad021 (2023).
Hu, W. et al. Single-cell transcriptome and translatome dual-omics reveals potential mechanisms of human oocyte maturation. Nat. Commun. 13, 5114 (2022).
Zhang, Y. et al. Transcriptome landscape of human folliculogenesis reveals oocyte and granulosa cell interactions. Mol. Cell 72, 1021–1034 e1024 (2018).
He, Y. et al. Pervasive 3’-UTR isoform switches during mouse oocyte maturation. Front. Mol. Biosci. 8, 727614 (2021).
Chalupnikova, K., Solc, P., Sulimenko, V., Sedlacek, R. & Svoboda, P. An oocyte-specific ELAVL2 isoform is a translational repressor ablated from meiotically competent antral oocytes. Cell Cycle 13, 1187–1200 (2014).
Li, X., Li, Y., Liu, C., Jin, M. & Lu, B. Oocyte-specific expression of mouse MEX3C652AA in the ovary and its potential role in regulating maternal Fos mRNA. Biol. Reprod. 94, 115 (2016).
Miao, B. et al. Tissue-specific usage of transposable element-derived promoters in mouse development. Genome Biol. 21, 255 (2020).
Fueyo, R., Judd, J., Feschotte, C. & Wysocka, J. Roles of transposable elements in the regulation of mammalian transcription. Nat. Rev. Mol. Cell Biol. 23, 481–497 (2022).
Georgiou, I. et al. Retrotransposon RNA expression and evidence for retrotransposition events in human oocytes. Hum. Mol. Genet. 18, 1221–1228 (2009).
Franke, V. et al. Long terminal repeats power evolution of genes and gene expression programs in mammalian oocytes and zygotes. Genome Res. 27, 1384–1394 (2017).
Modzelewski, A. J. et al. A mouse-specific retrotransposon drives a conserved Cdk2ap1 isoform essential for development. Cell 184, 5541–5558 e5522 (2021).
Peaston, A. E. et al. Retrotransposons regulate host genes in mouse oocytes and preimplantation embryos. Dev. Cell 7, 597–606 (2004).
Flemr, M. et al. A retrotransposon-driven dicer isoform directs endogenous small interfering RNA production in mouse oocytes. Cell 155, 807–816 (2013).
Treangen, T. J. & Salzberg, S. L. Repetitive DNA and next-generation sequencing: computational challenges and solutions. Nat. Rev. Genet. 13, 36–46 (2011).
Amarasinghe, S. L. et al. Opportunities and challenges in long-read sequencing data analysis. Genome Biol. 21, 30 (2020).
Uapinyoying, P. et al. A long-read RNA-seq approach to identify novel transcripts of very large genes. Genome Res. 30, 885–897 (2020).
Berrens, R. V. et al. Locus-specific expression of transposable elements in single cells with CELLO-seq. Nat. Biotechnol. 40, 546–554 (2022).
Fan, X. et al. Single-cell RNA-seq analysis of mouse preimplantation embryos by third-generation sequencing. PLoS Biol. 18, e3001017 (2020).
Glinos, D. A. et al. Transcriptome variation in human tissues revealed by long-read sequencing. Nature 608, 353–359 (2022).
Qiao, Y. et al. High-resolution annotation of the mouse preimplantation embryo transcriptome using long-read sequencing. Nat. Commun. 11, 2653 (2020).
Sun, Y. H. et al. Single-molecule long-read sequencing reveals a conserved intact long RNA profile in sperm. Nat. Commun. 12, 1361 (2021).
Torre, D. et al. Isoform-resolved transcriptome of the human preimplantation embryo. Nat. Commun. 14, 6902 (2023).
Sang, Q. et al. Homozygous mutations in WEE2 cause fertilization failure and female infertility. Am. J. Hum. Genet. 102, 649–657 (2018).
Blengini C. S. et al. AURKA controls oocyte spindle assembly checkpoint and chromosome alignment by HEC1 phosphorylation. Life Sci. Alliance 8, e202403146 (2025).
Blengini, C. S., Vaskovicova, M., Schier, J., Drutovic, D. & Schindler, K. Spatio-temporal requirements of Aurora kinase A in mouse oocyte meiotic spindle building. iScience 27, 110451 (2024).
Liu, Y., Nie, H., Liu, H. & Lu, F. Poly(A) inclusive RNA isoform sequencing (PAIso-seq) reveals wide-spread non-adenosine residues within RNA poly(A) tails. Nat. Commun. 10, 5292 (2019).
Zhao, Z. H. et al. RNA-Seq transcriptome reveals different molecular responses during human and mouse oocyte maturation and fertilization. BMC Genomics 21, 475 (2020).
Myers, K. N. et al. The bornavirus-derived human protein EBLN1 promotes efficient cell cycle transit, microtubule organisation and genome stability. Sci. Rep. 6, 35548 (2016).
He, P. et al. Knock-down of endogenous bornavirus-like nucleoprotein 1 inhibits cell growth and induces apoptosis in human oligodendroglia cells. Int. J. Mol. Sci. 17, 435 (2016).
Lovelace, M. D. et al. The RhoGAP protein ARHGAP18/SENEX localizes to microtubules and regulates their stability in endothelial cells. Mol. Biol. Cell 28, 1066–1078 (2017).
Maeda, M. et al. ARHGAP18, a GTPase-activating protein for RhoA, controls cell shape, spreading, and motility. Mol. Biol. Cell 22, 3840–3852 (2011).
Zou, Z. et al. Translatome and transcriptome co-profiling reveals a role of TPRXs in human zygotic genome activation. Science 378, abo7923 (2022).
Xiong, Z. et al. Ultrasensitive Ribo-seq reveals translational landscapes during mammalian oocyte-to-embryo transition and pre-implantation development. Nat. Cell Biol. 24, 968–980 (2022).
Sadato, D. et al. Eukaryotic translation initiation factor 3 (eIF3) subunit e is essential for embryonic development and cell proliferation. FEBS Open Bio 8, 1188–1201 (2018).
Baralle, F. E. & Giudice, J. Alternative splicing as a regulator of development and tissue identity. Nat. Rev. Mol. Cell Biol. 18, 437–451 (2017).
Wang, H. H. et al. Rab3A, Rab27A, and Rab35 regulate different events during mouse oocyte meiotic maturation and activation. Histochem. Cell Biol. 145, 647–657 (2016).
Senft, A. D. & Macfarlan, T. S. Transposable elements shape the evolution of mammalian development. Nat. Rev. Genet. 22, 691–711 (2021).
Hashimoto, K. et al. Embryonic LTR retrotransposons supply promoter modules to somatic tissues. Genome Res. 31, 1983–1993 (2021).
Chen F. et al. Ribonucleic acid export 1 is a kinetochore-associated protein that participates in chromosome alignment in mouse oocytes. Int. J. Mol. Sci. 22, 4841 (2021).
Xu, X. L. et al. The microtubule-associated protein ASPM regulates spindle assembly and meiotic progression in mouse oocytes. PLoS ONE 7, e49303 (2012).
Wang, H. et al. hnRNP A1 antagonizes cellular senescence and senescence-associated secretory phenotype via regulation of SIRT1 mRNA stability. Aging Cell 15, 1063–1073 (2016).
Leung, S. K. et al. Full-length transcript sequencing of human and mouse cerebral cortex identifies widespread isoform diversity and alternative splicing. Cell Rep. 37, 110022 (2021).
Liao, Y. et al. High-throughput and high-sensitivity full-length single-cell RNA-seq analysis on third-generation sequencing platform. Cell Discov. 9, 5 (2023).
Veiga, D. F. T. et al. A comprehensive long-read isoform analysis platform and sequencing resource for breast cancer. Sci. Adv. 8, eabg6711 (2022).
Li, R., Qiao, J., Wang, L., Zhen, X. & Lu, Y. Serum progesterone concentration on day of HCG administration and IVF outcome. Reprod. Biomed. Online 16, 627–631 (2008).
Liu, T. et al. Lipid metabolism was associated with oocyte in vitro maturation in women with polycystic ovarian syndrome undergoing unstimulated natural cycle. Front. Cell Dev. Biol. 9, 719173 (2021).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Davidson, N. M. et al. JAFFAL: detecting fusion genes with long-read transcriptome sequencing. Genome Biol. 23, 10 (2022).
Chen, Y. et al. Gene fusion detection and characterization in long-read cancer transcriptome sequencing data with FusionSeeker. Cancer Res. 83, 28–33 (2023).
Liu, Q. et al. LongGF: computational algorithm and software tool for fast and accurate detection of gene fusions by long-read transcriptome sequencing. BMC Genomics 21, 793 (2020).
Wang, L. et al. CPAT: Coding-Potential Assessment Tool using an alignment-free logistic regression model. Nucleic Acids Res. 41, e74 (2013).
Pertea G., Pertea M. GFF utilities: GffRead and GffCompare. F1000Res 9, ISCB Comm J-304 (2020).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Consortium, F. et al. A promoter-level mammalian expression atlas. Nature 507, 462–470 (2014).
Pardo-Palacios, F. J. et al. SQANTI3: curation of long-read transcriptomes for accurate identification of known and novel isoforms. Nat. Methods 21, 793–797 (2024).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Wang, L., Wang, S. & Li, W. RSeQC: quality control of RNA-seq experiments. Bioinformatics 28, 2184–2185 (2012).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Patowary, A. et al. Developmental isoform diversity in the human neocortex informs neuropsychiatric risk mechanisms. Science 384, eadh7688 (2024).
Li, Y., Rao, X., Mattox, W. W., Amos, C. I. & Liu, B. RNA-seq analysis of differential splice junction usage and intron retentions by DEXSeq. PLoS ONE 10, e0136653 (2015).
Vitting-Seerup, K. & Sandelin, A. IsoformSwitchAnalyzeR: analysis of changes in genome-wide patterns of alternative splicing and its functional consequences. Bioinformatics 35, 4469–4471 (2019).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Xiao, Z. et al. De novo annotation and characterization of the translatome with ribosome profiling data. Nucleic Acids Res. 46, e61 (2018).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinform. 10, 421 (2009).
Duffy, E. E. et al. Developmental dynamics of RNA translation in the human brain. Nat. Neurosci. 25, 1353–1365 (2022).
Sha, Q. Q. et al. Dynamics and clinical relevance of maternal mRNA clearance during the oocyte-to-embryo transition in humans. Nat. Commun. 11, 4917 (2020).
Babarinde, I. A. et al. Transposable element sequence fragments incorporated into coding and noncoding transcripts modulate the transcriptome of human pluripotent stem cells. Nucleic Acids Res. 49, 9132–9153 (2021).
Lizio, M. et al. Gateways to the FANTOM5 promoter level mammalian expression atlas. Genome Biol. 16, 22 (2015).
Liu, Y. et al. Remodeling of maternal mRNA through poly(A) tail orchestrates human oocyte-to-embryo transition. Nat. Struct. Mol. Biol. 30, 200–215 (2023).
Sloan, C. A. et al. ENCODE data at the ENCODE portal. Nucleic Acids Res. 44, D726–D732 (2016).
Mukherjee, A. B., Chan, M., Waite, R., Metzger, M. I. & Yaffee, S. J. Inhibition of RNA synthesis by acetyl salicylate and actinomycin D during early development in the mouse. Pediatr. Res. 9, 652–657 (1975).
Huarte, J., Belin, D. & Vassalli, J. D. Plasminogen activator in mouse and rat oocytes: induction during meiotic maturation. Cell 43, 551–558 (1985).
Gill, A., Jamnongjit, M. & Hammes, S. R. Androgens promote maturation and signaling in mouse oocytes independent of transcription: a release of inhibition model for mammalian oocyte meiosis. Mol. Endocrinol. 18, 97–104 (2004).
Chousal, J. et al. Chromatin modification and global transcriptional silencing in the oocyte mediated by the mRNA decay activator ZFP36L2. Dev. Cell 44, 392–402 e397 (2018).
Li, Y. et al. 2’-O-methylation at internal sites on mRNA promotes mRNA stability. Mol. Cell 84, 2320–2336 e2326 (2024).
Acknowledgements
We thank the generous donors whose contributions have enabled this research. We thank all the staff in the Center for Reproductive Medicine of Peking University Third Hospital. This project is funded by National Natural Science Foundation of China (82288102 to L.Y., 82125013 to L.Y., 82201838 to P.Y., 82522039 to P.Y.), National Key Research and Development Program (2022YFC2702200 to P.Y., 2023YFA1800300 to L.Y., 2024YFA1802100 to J.Q.), Peking University Third Hospital Fund for Interdisciplinary Research (BYSYJC2023001 to P.Y.), Clinical Medicine Plus X - Young Scholars Project, Peking University, the Fundamental Research Funds for the Central Universities (PKU2024LCXQ005 to P.Y.), and CAMS Innovation Fund for Medical Sciences (2019-I2M-5-001 to J.Q.).
Author information
Authors and Affiliations
Contributions
J.Q., P.Y., L.Y., and Q.L. supervised the project. Y.W., W.W., Yujun Liu, Q.L., P.Y., L.Y., and J.Q. conceived and formulated the concept. Y.W. and Yujun Liu collected the samples and performed all experiments including library construction, single-cell sequencing, siRNA knock down, RNA stability assay, RT-qPCR, western blotting, and immunofluorescence under the supervision of P.Y., L.Y., and J.Q. W.W. and Y.H. conducted bioinformatics analyses and visualization on full-length isoform sequencing and next generation sequencing data under the supervision of P.Y. M.Y., N.W., X.W., L.D., Y.K. and Ying Lian assisted with samples collection and siRNA injection. Y.X., Z.D., and L.C. assisted with ESCs and GCs collection. Y.W., W.W., Yujun Liu, Y.H., and H.S. drafted the initial manuscript. All authors discussed the results and reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, Y., Wang, W., Liu, Y. et al. Single-oocyte full-length isoform sequencing unveils the impact of transposable elements on RNA diversity and stability. Nat Commun (2026). https://doi.org/10.1038/s41467-026-71425-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-71425-9