Abstract
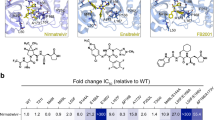
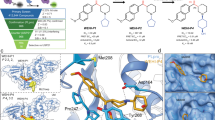
It remains elusive to design peptidomimetic inhibitors of SARS-CoV-2 main protease (Mpro) refractory to multiple deficiencies of Paxlovid (ritonavir-boosted nirmatrelvir), pertaining mainly to E166X mutations-conferred drug resistance and inherent pharmacokinetic limitations to nirmatrelvir. We identify via virtual screening an iso-quinoline P1 moiety in place of the traditional γ-lactam and design iso-quinoline-containing inhibitors with high affinity for Mpro and its nirmatrelvir-resistant E166X mutants. Further optimization at P4 cultivates distinctive peptidomimetic inhibitors with drastically improved pharmacokinetic properties and significantly enhanced antiviral efficacy independent of ritonavir. Two such inhibitors, FD3-32 and FD3-36, also potent against SARS-CoV-1 and MERS-CoV Mpro, are more effective as a monotherapy regimen than Paxlovid in reducing viral loads in vivo and protecting infected male mice from acute lung injury. Here, we report the discovery of next-generation SARS-CoV-2 Mpro inhibitors that overcome the deficiencies of Paxlovid, promising efficacious antivirals critical for mitigating the current and future pandemics of coronaviruses.
Similar content being viewed by others
Data availability
The PDB accession number for coordinates and structure factors of SARS-CoV-2 Mpro in complex with FD6-31 is 9L13. All other data are available in the main text or the supplementary materials. The initial coordinates and final outputs of MD simulations involved in this study have been deposited in the Zenodo database (https://doi.org/10.5281/zenodo.16948252). Resources, reagents and materials generated in this study are available from the Lead Contact, W.L. (luwuyuan@fudan.edu.cn), upon request. Source data are provided with this paper as Source Data files. Source data are provided in this paper.
Code availability
Input scripts for MD simulations in this study are available in the Zenodo database https://doi.org/10.5281/zenodo.16948252. The codes used for analysis and visualization of chemical space are publicly available on GitHub at https://github.com/LogenLuo/Noncanonical_P1_screening and deposited in Zenodo https://doi.org/10.5281/zenodo.18823333.
References
Drosten, C. et al. Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N. Engl. J. Med. 348, 1967–1976 (2003).
Zaki, A. M., van Boheemen, S., Bestebroer, T. M., Osterhaus, A. D. & Fouchier, R. A. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N. Engl. J. Med. 367, 1814–1820 (2012).
Zhou, P. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270–273 (2020).
Li, G., Hilgenfeld, R., Whitley, R. & De Clercq, E. Therapeutic strategies for COVID-19: progress and lessons learned. Nat. Rev. Drug Discov. 22, 449–475 (2023).
Davis, H. E., McCorkell, L., Vogel, J. M. & Topol, E. J. Long COVID: major findings, mechanisms and recommendations. Nat. Rev. Microbiol. 21, 133–146 (2023).
Wu, F. et al. A new coronavirus associated with human respiratory disease in China. Nature 579, 265–269 (2020).
Anand, K. et al. Structure of coronavirus main proteinase reveals combination of a chymotrypsin fold with an extra alpha-helical domain. EMBO J. 21, 3213–3224 (2002).
Zhao, Y. et al. Structural basis for replicase polyprotein cleavage and substrate specificity of main protease from SARS-CoV-2. Proc. Natl. Acad. Sci. USA 119, e2117142119 (2022).
Hegyi, A. & Ziebuhr, J. Conservation of substrate specificities among coronavirus main proteases. J. Gen. Virol. 83, 595–599 (2002).
Anand, K., Ziebuhr, J., Wadhwani, P., Mesters, J. R. & Hilgenfeld, R. Coronavirus main proteinase (3CLpro) structure: basis for design of anti-SARS drugs. Science 300, 1763–1767 (2003).
Song, L. et al. Medicinal chemistry strategies towards the development of non-covalent SARS-CoV-2 Mpro inhibitors. Acta Pharm. Sin. B 14, 87–109 (2024).
Hu, Q. et al. The SARS-CoV-2 main protease (Mpro): Structure, function, and emerging therapies for COVID-19. MedComm 3, e151 (2022). 2020.
Owen, D. R. et al. An oral SARS-CoV-2 Mpro inhibitor clinical candidate for the treatment of COVID-19. Science 374, 1586–1593 (2021).
Unoh, Y. et al. Discovery of S-217622, a noncovalent oral SARS-CoV-2 3CL protease inhibitor clinical candidate for treating COVID-19. J. Med. Chem. 65, 6499–6512 (2022).
Tiew, K. C. et al. Design, synthesis, and evaluation of inhibitors of Norwalk virus 3C protease. Bioorg. Med. Chem. Lett. 21, 5315–5319 (2011).
Kim, Y. et al. Broad-spectrum antivirals against 3C or 3C-like proteases of picornaviruses, noroviruses, and coronaviruses. J. Virol. 86, 11754–11762 (2012).
Ghosh, A. K. et al. Exploration of P1 and P4 modifications of nirmatrelvir: Design, synthesis, biological evaluation, and X-ray structural studies of SARS-CoV-2 Mpro inhibitors. Eur. J. Med. Chem. 267, 116132 (2024).
Tran, N. et al. The H163A mutation unravels an oxidized conformation of the SARS-CoV-2 main protease. Nat. Commun. 14, 5625 (2023).
Duan, Y. et al. Molecular mechanisms of SARS-CoV-2 resistance to nirmatrelvir. Nature 622, 376–382 (2023).
Kovalevsky, A. et al. Effects of SARS-CoV-2 main protease mutations at positions L50, E166, and L167 rendering resistance to covalent and noncovalent inhibitors. J. Med. Chem. 67, 18478–18490 (2024).
Ip, J. D. et al. Global prevalence of SARS-CoV-2 3CL protease mutations associated with nirmatrelvir or ensitrelvir resistance. EBioMedicine 91, 104559 (2023).
Iketani, S. et al. Functional map of SARS-CoV-2 3CL protease reveals tolerant and immutable sites. Cell Host Microbe 30, 1354–1362 (2022).
Kiso, M. et al. In vitro and in vivo characterization of SARS-CoV-2 resistance to ensitrelvir. Nat. Commun. 14, 4231 (2023).
Kiso, M. et al. In vitro and in vivo characterization of SARS-CoV-2 strains resistant to nirmatrelvir. Nat. Commun. 14, 3952 (2023).
Iketani, S. et al. Multiple pathways for SARS-CoV-2 resistance to nirmatrelvir. Nature 613, 558–564 (2023).
Doi, A., Ota, M., Saito, M. & Matsuyama, S. Sporadic occurrence of ensitrelvir-resistant SARS-CoV-2, Japan. Emerg. Infect. Dis. 30, 1289–1291 (2024).
Esler, M. A. et al. Structural basis for varying drug resistance of SARS-CoV-2 Mpro E166 variants. mBio 16, e0262424 (2025).
Pfizer. Pfizer 2024 Annual Review https://annualreview.pfizer.com/ (2024).
Kempf, D. J. et al. ABT-538 is a potent inhibitor of human immunodeficiency virus protease and has high oral bioavailability in humans. Proc. Natl. Acad. Sci. USA 92, 2484–2488 (1995).
Danner, S. A. et al. A short-term study of the safety, pharmacokinetics, and efficacy of ritonavir, an inhibitor of HIV-1 protease. N. Engl. J. Med. 333, 1528–1533 (1995).
Marzolini, C. et al. Recommendations for the management of drug-drug interactions Between the COVID-19 antiviral nirmatrelvir/Ritonavir (Paxlovid) and Comedications. Clin. Pharm. Ther. 112, 1191–1200 (2022).
Girardin, F., Manuel, O., Marzolini, C. & Buclin, T. Evaluating the risk of drug-drug interactions with pharmacokinetic boosters: the case of ritonavir-enhanced nirmatrelvir to prevent severe COVID-19. Clin. Microbiol. Infect. 28, 1044–1046 (2022).
Ross, S. B. et al. Drug interactions with nirmatrelvir-ritonavir in older adults using multiple medications. JAMA Netw. Open 5, e2220184 (2022).
Han, Y. et al. Potential drug interaction after withdrawal of nirmatrelvir-ritonavir in hospitalized patients with COVID-19 infection. J. Glob. Antimicrob. Resist. 42, 151–153 (2025).
Marzolini, C. et al. Prescribing nirmatrelvir-ritonavir: how to recognize and manage drug-drug interactions. Ann. Intern. Med. 175, 744–746 (2022).
Allerton, C. M. N. et al. A second-generation oral SARS-CoV-2 main protease inhibitor clinical candidate for the treatment of COVID-19. J. Med. Chem. 67, 13550–13571 (2024).
Chen, X. et al. Preclinical evaluation of the SARS-CoV-2 M(pro) inhibitor RAY1216 shows improved pharmacokinetics compared with nirmatrelvir. Nat. Microbiol. 9, 1075–1088 (2024).
Hert, J. et al. Comparison of topological descriptors for similarity-based virtual screening using multiple bioactive reference structures. Org. Biomol. Chem. 2, 3256–3266 (2004).
Ma, C. et al. Discovery of Di- and trihaloacetamides as covalent SARS-CoV-2 main protease inhibitors with high target specificity. J. Am. Chem. Soc. 143, 20697–20709 (2021).
Drayman, N. et al. Masitinib is a broad coronavirus 3CL inhibitor that blocks replication of SARS-CoV-2. Science 373, 931–936 (2021).
Gao, S. et al. Discovery and crystallographic studies of trisubstituted piperazine derivatives as non-covalent SARS-CoV-2 main protease inhibitors with high target specificity and low toxicity. J. Med. Chem. 65, 13343–13364 (2022).
Boby, M. L. et al. Open science discovery of potent noncovalent SARS-CoV-2 main protease inhibitors. Science 382, eabo7201 (2023).
Robertson, A. J., Ying, J. & Bax, A. NMR Observation of sulfhydryl signals in SARS-CoV-2 main protease aids structural studies. Chembiochem 23, e202200471 (2022).
Altincekic, N. et al. Targeting the main protease (Mpro, nsp5) by growth of fragment scaffolds exploiting structure-based methodologies. ACS Chem. Biol. 19, 563–574 (2024).
Kao, R. Y. et al. Characterization of SARS-CoV main protease and identification of biologically active small molecule inhibitors using a continuous fluorescence-based assay. FEBS Lett. 576, 325–330 (2004).
Jin, Z. et al. Structure of M(pro) from SARS-CoV-2 and discovery of its inhibitors. Nature 582, 289–293 (2020).
Wilkesman, J. Cysteine protease zymography: brief review. Methods Mol. Biol. 1626, 25–31 (2017).
Zhong, B. et al. Oridonin Inhibits SARS-CoV-2 by targeting its 3C-like protease. Small Sci. 2, 2100124 (2022).
Craven, G. B. et al. Mutant-selective AKT inhibition through lysine targeting and neo-zinc chelation. Nature 637, 205–214 (2025).
Meyer, C. et al. Small-molecule inhibition of SARS-CoV-2 NSP14 RNA cap methyltransferase. Nature 637, 1178–1185 (2025).
Boras, B. et al. Preclinical characterization of an intravenous coronavirus 3CL protease inhibitor for the potential treatment of COVID19. Nat. Commun. 12, 6055 (2021).
Ullrich, S. & Nitsche, C. The SARS-CoV-2 main protease as drug target. Bioorg. Med. Chem. Lett. 30, 127377 (2020).
Jiang, X. et al. Structure-based development and preclinical evaluation of the SARS-CoV-2 3C-like protease inhibitor simnotrelvir. Nat. Commun. 14, 6463 (2023).
Zhu, Y. et al. In vitro selection and analysis of SARS-CoV-2 nirmatrelvir resistance mutations contributing to clinical virus resistance surveillance. Sci. Adv. 10, eadl4013 (2024).
Moghadasi, S. A. et al. Transmissible SARS-CoV-2 variants with resistance to clinical protease inhibitors. Sci. Adv. 9, eade8778 (2023).
Heilmann, E. et al. SARS-CoV-2 3CLpro mutations selected in a VSV-based system confer resistance to nirmatrelvir, ensitrelvir, and GC376. Sci. Transl. Med. 15, eabq7360 (2023).
Abdelnabi, R. et al. Nirmatrelvir-resistant SARS-CoV-2 is efficiently transmitted in female Syrian hamsters and retains partial susceptibility to treatment. Nat. Commun. 14, 2124 (2023).
Hu, Y. et al. Naturally occurring mutations of SARS-CoV-2 main protease confer drug resistance to nirmatrelvir. ACS Cent. Sci. 9, 1658–1669 (2023).
Sasaki, M. et al. S-217622, a SARS-CoV-2 main protease inhibitor, decreases viral load and ameliorates COVID-19 severity in hamsters. Sci. Transl. Med. 15, eabq4064 (2023).
Estudante, M., Morais, J. G., Soveral, G. & Benet, L. Z. Intestinal drug transporters: an overview. Adv. Drug Deliv. Rev. 65, 1340–1356 (2013).
Luttens, A. et al. Ultralarge virtual screening identifies SARS-CoV-2 main protease inhibitors with broad-spectrum activity against coronaviruses. J. Am. Chem. Soc. 144, 2905–2920 (2022).
Zhang, Y., Huo, M., Zhou, J. & Xie, S. PKSolver: An add-in program for pharmacokinetic and pharmacodynamic data analysis in Microsoft Excel. Comput. Methods Prog. Biomed. 99, 306–314 (2010).
Wang, H. et al. Macrophage ferroportin serves as a therapeutic target against bacteria-induced acute lung injury by promoting barrier restoration. iScience 25, 105698 (2022).
Jakalian, A., Jack, D. B. & Bayly, C. I. Fast, efficient generation of high-quality atomic charges. AM1-BCC model: II. Parameterization and validation. J. Comput. Chem. 23, 1623–1641 (2002).
Wang, J., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. Development and testing of a general amber force field. J. Comput. Chem. 25, 1157–1174 (2004).
Tian, C. et al. ff19SB: Amino-acid-specific protein backbone parameters trained against quantum mechanics energy surfaces in solution. J. Chem. Theory Comput. 16, 528–552 (2020).
Sun, H., Li, Y., Tian, S., Xu, L. & Hou, T. Assessing the performance of MM/PBSA and MM/GBSA methods. 4. Accuracies of MM/PBSA and MM/GBSA methodologies evaluated by various simulation protocols using PDBbind data set. Phys. Chem. Chem. Phys. 16, 16719–16729 (2014).
Wong, F. et al. Discovery of a structural class of antibiotics with explainable deep learning. Nature 626, 177–185 (2024).
Brinson, R. G. et al. Enabling adoption of 2D-NMR for the higher order structure assessment of monoclonal antibody therapeutics. MAbs 11, 94–105 (2019).
Williamson, M. P. Using chemical shift perturbation to characterise ligand binding. Prog. Nucl. Magn. Reson Spectrosc. 73, 1–16 (2013).
Acknowledgements
We thank Dr. Lingyun Yang in the NMR platform of iHuman Institute, ShanghaiTech University, for his excellent technical assistance on 2D NMR experiments; Prof. Youhua Xie in Shanghai Medical College, Fudan University, for his technical support on virological assays. We also thank the Center for Molecular Modeling and Design, School of Pharmacy, Fudan University, for support of computational resources. We thank the staff of the BL10U2 beamline at Shanghai Synchrotron Radiation Facility (SSRF), Shanghai, China, for assistance during data collection. This work was supported by grants from the Shanghai Municipal Science and Technology Major Project (ZD2021CY001 to F.W.), the National Natural Science Foundation of China (82030062 to W.L. and 82400683 to J.Z.), the Young Elite Scientists Sponsorship Program by CAST-Doctoral Student Special Plan (G.L.), the China Postdoctoral Science Foundation (2023M740661 to J.Z.), and the National Key Research and Development Program of China (2021YFC2300703 to L.L.).
Author information
Authors and Affiliations
Contributions
W.L., G.L., X.G., and F.W. conceived, supervised, and/or designed the research; G.L. and W.L. designed the inhibitors; G.L., L.F., and J.J.Z. performed or supervised chemical synthesis, purification and structural characterizations; G.W. and M.L. conducted all protein expression and purification in this study; G.L. and G.W. performed 2D-NMR experiments and analyzed spectra; G.L. performed enzymatic inhibition assays and TSA assays; B.Z. and G.W. contributed to crystallization and diffraction data collection. B.Z. determined and analyzed the crystal structure; J.Z. performed in vitro cytotoxicity assays; W.X. and L.L. supervised virological assays in this study; C. L., W.X., G.L., Z.W., J.R., G.W., J.Z., and Y.Y. performed both in vitro and in vivo antiviral assays, as well as related characterizations; G.L. and W.L. wrote the manuscript with assistance of G.W., B.Z., and L.F.
Corresponding authors
Ethics declarations
Competing interests
W.L., G.L., and X.G. are inventors in a patent application (CN202410660959.1; PCT/CN2025/096704) related to compounds in this study. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Asim K. Debnath, Jun Zhang and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Luo, G., Wang, G., Liao, C. et al. Next-generation inhibitors of SARS-CoV-2 Mpro overcome the deficiencies of Paxlovid. Nat Commun (2026). https://doi.org/10.1038/s41467-026-71436-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-71436-6