Abstract
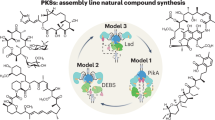
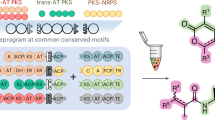
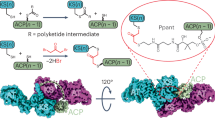
Assembly line biosynthesis creates numerous structurally diverse natural products using a common modular synthetic strategy. The collinearity between the architectures of modular polyketide synthases (PKS) and the structures of their polyketide products would seem to render these biosynthetic machineries excellent platforms for designer biosynthesis, yet reliable strategies to reprogram these assembly lines without diminishing their activities have not been identified. Here, as a best practice for PKS engineering, we demonstrate the reprogramming of the mediomycin PKS without significant loss of productivity. Using in vitro CRISPR/Cas9 gene editing followed by heterologous expression, we reconstruct an inaccessible drug lead of the fibrinogen receptor, tetrafibricin, at 82 ± 3 mg/L yield, retaining 26% productivity after five-step module editing using an evolution-supported cut site, downstream of the acyltransferase domain. A macrocyclic aminopolyol is also accessed through thioesterase swapping. These results pave the way toward the rational reprogramming of PKSs to access desired complex organic molecules.
Similar content being viewed by others
Data availability
All data supporting the findings of the study are available from the corresponding author upon request. Source data are provided with this paper.
References
Fischbach, M. A. & Walsh, C. T. Assembly-line enzymology for polyketide and nonribosomal peptide antibiotics: logic, machinery, and mechanisms. Chem. Rev. 106, 3468–3496 (2006).
Staunton, J. & Weissman, K. J. Polyketide biosynthesis: a millennium review. Nat. Prod. Rep. 18, 380–416 (2001).
Herbst, D. A., Townsend, C. A. & Maier, T. The architectures of iterative type I PKs and FAS. Nat. Prod. Rep. 35, 1046–1069 (2018).
Paiva, P. et al. Animal fatty acid synthase: a chemical nanofactory. Chem. Rev. 121, 9502–9553 (2021).
Khosla, C. Harnessing the biosynthetic potential of modular polyketide synthases. Chem. Rev. 97, 2577–2590 (1997).
Keatinge-Clay, A. T. The structures of type I polyketide synthases. Nat. Prod. Rep. 29, 1050–1073 (2012).
Katsuyama, Y. & Miyanaga, A. Recent advances in the structural biology of modular polyketide synthases and nonribosomal peptide synthetases. Curr. Opin. Chem. Biol. 71, 102223 (2022).
Grininger, M. Enzymology of assembly line synthesis by modular polyketide synthases. Nat. Chem. Biol. 19, 401–415 (2023).
Ranganathan, A. et al. Knowledge-based design of bimodular and trimodular polyketide synthases based on domain and module swaps: a route to simple statin analogues. Chem. Biol. 6, 731–741 (1999).
Menzella, H. G. et al. Combinatorial polyketide biosynthesis by de novo design and rearrangement of modular polyketide synthase genes. Nat. Biotechnol. 23, 1171–1176 (2005).
Walker, M. C. et al. Expanding the fluorine chemistry of living systems using engineered polyketide synthase pathways. Science 341, 1089–1094 (2013).
Peng, H., Ishida, K., Sugimoto, Y., Jenke-Kodama, H. & Hertweck, C. Emulating evolutionary processes to morph aureothin-type modular polyketide synthases and associated oxygenases. Nat. Commun. 10, 3918 (2019).
Kudo, K. et al. In vitro cas9-assisted editing of modular polyketide synthase genes to produce desired natural product derivatives. Nat. Commun. 11, 4022 (2020).
Su, L. et al. Engineering the stambomycin modular polyketide synthase yields 37-membered mini-stambomycins. Nat. Commun. 13, 515 (2022).
Cane, D. E., Walsh, C. T. & Khosla, C. Harnessing the biosynthetic code: combinations, permutations, and mutations. Science 282, 63–68 (1998).
Weissman, K. J. Genetic engineering of modular PKSs: from combinatorial biosynthesis to synthetic biology. Nat. Prod. Rep. 33, 203–230 (2016).
Klaus, M. & Grininger, M. Engineering strategies for rational polyketide synthase design. Nat. Prod. Rep. 35, 1070–1081 (2018).
Donadio, S., Staver, M. J., McAlpine, J. B., Swanson, S. J. & Katz, L. Modular organization of genes required for complex polyketide biosynthesis. Science 252, 675–679 (1991).
Zhang, L. et al. Characterization of giant modular PKSs provides insight into genetic mechanism for structural diversification of aminopolyol polyketides. Angew. Chem. Int. Ed. 56, 1740–1745 (2017).
Grote, M. et al. Identification of crucial bottlenecks in engineered polyketide biosynthesis. Org. Biomol. Chem. 17, 6374–6385 (2019).
Hirsch, M., Fitzgerald, B. J. & Keatinge-Clay, A. T. How cis-acyltransferase assembly-line ketosynthases gatekeep for processed polyketide intermediates. ACS Chem. Biol. 16, 2515–2526 (2021).
Ray, K. A., Saif, N. & Keatinge-Clay, A. T. Modular polyketide synthase ketosynthases collaborate with upstream dehydratases to install double bonds. Chem. Commun. 60, 8712–8715 (2024).
Klaus, M. et al. Protein-protein interactions, not substrate recognition, dominate the turnover of chimeric assembly line polyketide synthases. J. Biol. Chem. 291, 16404–16415 (2016).
Klaus, M., Buyachuihan, L. & Grininger, M. Ketosynthase domain constrains the design of polyketide synthases. ACS Chem. Biol. 15, 2422–2432 (2020).
Mabesoone, M. F. J. et al. Evolution-guided engineering of trans-acyltransferase polyketide synthases. Science 383, 1312–1317 (2024).
Yi, D., Wakeel, M. A. & Agarwal, V. Gatekeeping activity of collinear ketosynthase domains limits product diversity for engineered type I polyketide synthases. Biochemistry 63, 2240–2244 (2024).
Yuzawa, S. et al. Comprehensive in vitro analysis of acyltransferase domain exchanges in modular polyketide synthases and its application for short-chain ketone production. ACS Synth. Biol. 6, 139–147 (2017).
Medema, M. H., Cimermancic, P., Sali, A., Takano, E. & Fischbach, M. A. A systematic computational analysis of biosynthetic gene cluster evolution: Lessons for engineering biosynthesis. PLoS Comput. Biol. 10, e1004016 (2014).
Bozhuyuk, K. A. J. et al. De novo design and engineering of non-ribosomal peptide synthetases. Nat. Chem. 10, 275–281 (2018).
Tang, Y., Kim, C. Y., Mathews, I. I., Cane, D. E. & Khosla, C. The 2.7-angstrom crystal structure of a 194-kDa homodimeric fragment of the 6-deoxyerythronolide B synthase. Proc. Natl. Acad. Sci. USA. 103, 11124–11129 (2006).
Cogan, D. P. et al. Mapping the catalytic conformations of an assembly-line polyketide synthase module. Science 374, 729–734 (2021).
Bagde, S. R., Mathews, I. I., Fromme, J. C. & Kim, C. Y. Modular polyketide synthase contains two reaction chambers that operate asynchronously. Science 374, 723–729 (2021).
Kamiyama, T. et al. Tetrafibricin, a novel fibrinogen receptor antagonist. Ii. Structural elucidation. J. Antibiot. 46, 1047–1054 (1993).
Kamiyama, T. et al. Tetrafibricin, a novel fibrinogen receptor antagonist. I. Taxonomy, fermentation, isolation, characterization and biological activities. J. Antibiot. 46, 1039–1046 (1993).
Friedrich, R. M. & Friestad, G. K. Inspirations from tetrafibricin and related polyketides: New methods and strategies for 1,5-polyol synthesis. Nat. Prod. Rep. 37, 1229–1261 (2020).
Sun, F., Xu, S., Jiang, F. & Liu, W. Genomic-driven discovery of an amidinohydrolase involved in the biosynthesis of mediomycin A. Appl. Microbiol. Biotechnol. 102, 2225–2234 (2018).
Ridley, C. P., Lee, H. Y. & Khosla, C. Evolution of polyketide synthases in bacteria. Proc. Natl. Acad. Sci. USA. 105, 4595–4600 (2008).
Gokhale, R. S., Hunziker, D., Cane, D. E. & Khosla, C. Mechanism and specificity of the terminal thioesterase domain from the erythromycin polyketide synthase. Chem. Biol. 6, 117–125 (1999).
Miyazawa, T., Hirsch, M., Zhang, Z. & Keatinge-Clay, A. T. An in vitro platform for engineering and harnessing modular polyketide synthases. Nat. Commun. 11, 80 (2020).
Ray, K. A. et al. Assessing and harnessing updated polyketide synthase modules through combinatorial engineering. Nat. Commun. 15, 6485 (2024).
Yuzawa, S. et al. Short-chain ketone production by engineered polyketide synthases in streptomyces albus. Nat. Commun. 9, 4569 (2018).
Zhang, J., Bista, R., Miyazawa, T. & Keatinge-Clay, A. T. Boosting titers of engineered triketide and tetraketide synthases to record levels through t7 promoter tuning. Metab. Eng. 78, 93–98 (2023).
Miyazawa, T., Fitzgerald, B. J. & Keatinge-Clay, A. T. Preparative production of an enantiomeric pair by engineered polyketide synthases. Chem. Commun. (Camb.) 57, 8762–8765 (2021).
Hagen, A. et al. Engineering a polyketide synthase for in vitro production of adipic acid. ACS Synth. Biol. 5, 21–27 (2016).
Huang, Z. et al. Plug-and-play engineering of modular polyketide synthases. Nat. Chem. Biol. 21, 1361–1367 (2025).
Englund, E. et al. Nature communicationsbiosensor guided polyketide synthases engineering for optimization of domain exchange boundaries. Nat. Commun. 14, 4871 (2023).
Itoh, H. & Inoue, M. Comprehensive structure-activity relationship studies of macrocyclic natural products enabled by their total syntheses. Chem. Rev. 119, 10002–10031 (2019).
Buyachuihan, L., Stegemann, F. & Grininger, M. How acyl carrier proteins (acps) direct fatty acid and polyketide biosynthesis. Angew. Chem. Int. Ed. 63, e202312476 (2024).
Tran, L., Broadhurst, R. W., Tosin, M., Cavalli, A. & Weissman, K. J. Insights into protein-protein and enzyme-substrate interactions in modular polyketide synthases. Chem. Biol. 17, 705–716 (2010).
Cortes, J. et al. Repositioning of a domain in a modular polyketide synthase to promote specific chain cleavage. Science 268, 1487–1489 (1995).
Kao, C. M., Luo, G. L., Katz, L., Cane, D. E. & Khosla, C. Manipulation of macrolide ring size by directed mutagenesis of a modular polyketide synthase. J. Am. Chem. Soc. 117, 9105–9106 (1995).
Menzella, H. G., Carney, J. R. & Santi, D. V. Rational design and assembly of synthetic trimodular polyketide synthases. Chem. Biol. 14, 143–151 (2007).
Hansen, D. A., Koch, A. A. & Sherman, D. H. Identification of a thioesterase bottleneck in the pikromycin pathway through full-module processing of unnatural pentaketides. J. Am. Chem. Soc. 139, 13450–13455 (2017).
Kudo, K. et al. Capability of a large bacterial artificial chromosome clone harboring multiple biosynthetic gene clusters for the production of diverse compounds. J. Antibiot. 77, 288–298 (2024).
Kudo, K., Nishimura, T., Miyako, K., Suenaga, H. & Shin-Ya, K. A new cannabigerolic acid derivative and its unprenylated precursor produced through the reconstitution of cannabinoid biosynthesis in streptomyces. J. Antibiot. 78, 126–130 (2025).
Katoh, K. & Standley, D. M. Mafft multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Price, M. N., Dehal, P. S. & Arkin, A. P. Fasttree 2–approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using clustal omega. Mol. Syst. Biol. 7, 539 (2011).
Acknowledgements
This work was supported by Japan Agency for Medical Research and Development (AMED) grant JP19ae0101045 (KS), Japan Society for the Promotion of Science (JSPS) KAKENHI grant JP23H05474 (KS), JSPS KAKENHI grant JP23H04569 (KK), and NIH grant GM145992 (ATK). We also thanks Drs. Toshio Nagashima, Huiping Zhang, and Yoshitaka Ishii (Riken) for measurement of NMR spectra at RIKEN Yokohama.
Author information
Authors and Affiliations
Contributions
All authors contributed to the writing of the paper. Conceptualization: K.K., T.H., T.A., L.Z., A.T.K., I.A., K.S. Methodology: K.K., T.H., L.Z., K.S. Investigation: K.K., T.H., T.A., T.N., J.H., I.K, N.K. Visualization: K.K., T.A., L.Z., T.N., N.K. Funding acquisition: K.K., K.S. Project administration: K.S. Supervision: H.S., I.A., K.S. Writing – original draft: K.K., L.Z. Writing – review & editing: K.K., T.A., L.Z., A.T.K., K.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Kira Weissman, Martin Grininger and the other anonymous reviewer for their contribution to the peer review of this work. [A peer review file is available].
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kudo, K., Hashimoto, T., Awakawa, T. et al. Skeletal editing via multi-step engineering of a modular polyketide synthase. Nat Commun (2026). https://doi.org/10.1038/s41467-026-71501-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-71501-0