Abstract
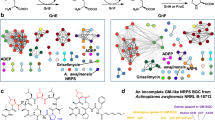
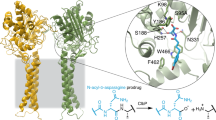
The discovery of natural products with specific modes of action from metagenomes remains challenging. Here, we present resistance-CONKAT-seq, a pipeline that links biosynthetic gene clusters (BGCs) to self-resistance genes, enabling identification of metabolites with desired molecular targets. Using clpP-directed resistance-CONKAT-seq, we identify the calprotamides, which activate native ClpP and enhance its activity. Cryo-EM and bioinformatic analyses reveal that the calprotamides’ medium-chain N-acylphenylalanine substructure is a convergently evolved ClpP-targeting motif and identify additional BGCs predicted to encode this moiety, including some with co-localized clp genes. The synthesis of structures bioinformatically inspired by two such clp-linked BGCs, desmethyl jomthonic acid C and tuscamide, reveals that both enhance ClpP activity. Extending our bioinformatically guided synthesis study to additional BGCs lacking nearby clp genes shows that ClpP activity enhancement correlated with antibacterial activity, with the strongest enhancers exhibiting narrow-spectrum antibiotic activity. These findings establish N-acylphenylalanine as a previously unrecognized but common natural motif for targeting ClpP, which should help guide the discovery of both natural and synthetic ClpP modulators for antibiotic and anticancer development. Resistance-CONKAT-seq offers a scalable method for exploring biosynthetic dark matter for metabolites with desired modes of action.
Similar content being viewed by others
Data availability
Source Data are provided in the Source Data files with this paper. Demultiplexed reads of clpP genes and ketosynthase (KS) domains are available from Zenodo [https://zenodo.org/records/18763872] (KS) and [https://zenodo.org/records/18763886] (clpP). The assembled sequencing data has been submitted to the NCBI GenBank database under accession no. PV247683. The atomic models have been deposited in the Protein Data Bank under accession codes 9P53 (calprotamide A-bound) and 9P54 (calprotamide A-unbound). The corresponding Cryo-EM maps have been deposited in the Electron Microscopy Data Bank (EMDB) under accession codes EMD-71299 (calprotamide A-bound) and EMD-71300 (calprotamide A-unbound). The ADEP-bound and apo Mtb ClpP1/P2 structures were obtained from the Protein Data Bank under accession codes 6VGN (ADEP-bound) and 6VGK (apo). Source data are provided with this paper.
References
Butler, M. S. Natural products to drugs: natural product-derived compounds in clinical trials. Nat. Prod. Rep. 25, 475–516 (2008).
Newman, D. J. & Cragg, G. M. Natural products as sources of new drugs over the last 25 years. J. Nat. Prod. 70, 461–477 (2007).
Atanasov, A. G., Zotchev, S. B., Dirsch, V. M., International Natural Product Sciences, T. & Supuran, C. T. Natural products in drug discovery: advances and opportunities. Nat. Rev. Drug Discov. 20, 200–216 (2021).
Moffat, J. G., Vincent, F., Lee, J. A., Eder, J. & Prunotto, M. Opportunities and challenges in phenotypic drug discovery: an industry perspective. Nat. Rev. Drug Discov. 16, 531–543 (2017).
Newman, D. J. & Cragg, G. M. Natural Products as Sources of New Drugs from 1981 to 2014. J. Nat. Prod. 79, 629–661 (2016).
Rutledge, P. J. & Challis, G. L. Discovery of microbial natural products by activation of silent biosynthetic gene clusters. Nat. Rev. Microbiol. 13, 509–523 (2015).
Freel, K. C., Millan-Aguinaga, N. & Jensen, P. R. Multilocus sequence typing reveals evidence of homologous recombination linked to antibiotic resistance in the genus Salinispora. Appl. Environ. Microbiol. 79, 5997–6005 (2013).
Kale, A. J., McGlinchey, R. P., Lechner, A. & Moore, B. S. Bacterial self-resistance to the natural proteasome inhibitor salinosporamide A. ACS Chem. Biol. 6, 1257–1264 (2011).
Ziemert, N. et al. Diversity and evolution of secondary metabolism in the marine actinomycete genus Salinispora. Proc. Natl. Acad. Sci. USA 111, E1130–E1139 (2014).
Yan, Y., Liu, N. & Tang, Y. Recent developments in self-resistance gene directed natural product discovery. Nat. Prod. Rep. 37, 879–892 (2020).
Alanjary, M. et al. The Antibiotic Resistant Target Seeker (ARTS), an exploration engine for antibiotic cluster prioritization and novel drug target discovery. Nucleic Acids Res 45, W42–W48 (2017).
Yan, Y. et al. Resistance-gene-directed discovery of a natural-product herbicide with a new mode of action. Nature 559, 415–418 (2018).
Fukuda, D. et al. A natural plasmid uniquely encodes two biosynthetic pathways creating a potent anti-MRSA antibiotic. PLoS One 6, e18031 (2011).
Steffensky, M., Muhlenweg, A., Wang, Z. X., Li, S. M. & Heide, L. Identification of the novobiocin biosynthetic gene cluster of Streptomyces spheroides NCIB 11891. Antimicrob. Agents Chemother. 44, 1214–1222 (2000).
Panter, F., Krug, D., Baumann, S. & Muller, R. Self-resistance guided genome mining uncovers new topoisomerase inhibitors from myxobacteria. Chem. Sci. 9, 4898–4908 (2018).
Burian, J. et al. Bioactive molecules unearthed by terabase-scale long-read sequencing of a soil metagenome. Nat. Biotechnol. https://doi.org/10.1038/s41587-025-02810-w (2025).
Libis, V. et al. Uncovering the biosynthetic potential of rare metagenomic DNA using co-occurrence network analysis of targeted sequences. Nat. Commun. 10, 3848 (2019).
Libis, V. et al. Multiplexed mobilization and expression of biosynthetic gene clusters. Nat. Commun. 13, 5256 (2022).
Aljghami, M. E., Barghash, M. M., Majaesic, E., Bhandari, V. & Houry, W. A. Cellular functions of the ClpP protease impacting bacterial virulence. Front. Mol. Biosci. 9, 1054408 (2022).
Bhandari, V. et al. The role of ClpP protease in bacterial pathogenesis and human diseases. ACS Chem. Biol. 13, 1413–1425 (2018).
Deepa, S. S. et al. Down-regulation of the mitochondrial matrix peptidase ClpP in muscle cells causes mitochondrial dysfunction and decreases cell proliferation. Free Radic. Biol. Med. 91, 281–292 (2016).
Amor, A. J., Schmitz, K. R., Baker, T. A. & Sauer, R. T. Roles of the ClpX IGF loops in ClpP association, dissociation, and protein degradation. Protein Sci. 28, 756–765 (2019).
Mabanglo, M. F. & Houry, W. A. Recent structural insights into the mechanism of ClpP protease regulation by AAA+ chaperones and small molecules. J. Biol. Chem. 298, 101781 (2022).
Weinhäupl, K. et al. Activation mechanism and structural assembly of the Mycobacterium tuberculosis ClpP1P2 protease and its associated ATPases. Preprint at https://www.biorxiv.org/content/10.1101/2025.02.28.640722v1 (2025).
Brady, S. F. Construction of soil environmental DNA cosmid libraries and screening for clones that produce biologically active small molecules. Nat. Protoc. 2, 1297–1305 (2007).
Kim, S. H. et al. Atolypenes, Tricyclic Bacterial Sesterterpenes Discovered Using a Multiplexed In Vitro Cas9-TAR Gene Cluster Refactoring Approach. ACS Synth. Biol. 8, 109–118 (2019).
Zhao, C. et al. Oxazolomycin biosynthesis in Streptomyces albus JA3453 featuring an “acyltransferase-less” type I polyketide synthase that incorporates two distinct extender units. J. Biol. Chem. 285, 20097–20108 (2010).
Benaroudj, N., Raynal, B., Miot, M. & Ortiz-Lombardia, M. Assembly and proteolytic processing of mycobacterial ClpP1 and ClpP2. BMC Biochem. 12, 61 (2011).
Vahidi, S. et al. An allosteric switch regulates Mycobacterium tuberculosis ClpP1P2 protease function as established by cryo-EM and methyl-TROSY NMR. Proc. Natl. Acad. Sci. USA 117, 5895–5906 (2020).
Akopian, T. et al. The active ClpP protease from M. tuberculosis is a complex composed of a heptameric ClpP1 and a ClpP2 ring. EMBO J. 31, 1529–1541 (2012).
Li, M. et al. Structure and Functional Properties of the Active Form of the Proteolytic Complex, ClpP1P2, from Mycobacterium tuberculosis. J. Biol. Chem. 291, 7465–7476 (2016).
Famulla, K. et al. Acyldepsipeptide antibiotics kill mycobacteria by preventing the physiological functions of the ClpP1P2 protease. Mol. Microbiol. 101, 194–209 (2016).
Li, L. et al. Biosynthetic Interrogation of Soil Metagenomes Reveals Metamarin, an Uncommon Cyclomarin Congener with Activity against Mycobacterium tuberculosis. J. Nat. Prod. 84, 1056–1066 (2021).
Vasudevan, D., Rao, S. P. & Noble, C. G. Structural basis of mycobacterial inhibition by cyclomarin A. J. Biol. Chem. 288, 30883–30891 (2013).
Brotz-Oesterhelt, H. et al. Dysregulation of bacterial proteolytic machinery by a new class of antibiotics. Nat. Med. 11, 1082–1087 (2005).
Lee, B. G. et al. Structures of ClpP in complex with acyldepsipeptide antibiotics reveal its activation mechanism. Nat. Struct. Mol. Biol. 17, 471–478 (2010).
Garcia-Salcedo, R. et al. Characterization of the Jomthonic Acids Biosynthesis Pathway and Isolation of Novel Analogues in Streptomyces caniferus GUA-06-05-006A. Mar. Drugs 16, 259 (2018).
Igarashi, Y. et al. Jomthonic acid, a modified amino acid from a soil-derived Streptomyces. J. Nat. Prod. 75, 986–990 (2012).
Carney, D. W. et al. A simple fragment of cyclic acyldepsipeptides is necessary and sufficient for ClpP activation and antibacterial activity. Chembiochem. 15, 2216–2220 (2014).
Alm, R. A. & Lahiri, S. D. Narrow-Spectrum Antibacterial Agents-Benefits and Challenges. Antibiotics 9, 418 (2020).
Gigante, V. et al. Multi-year analysis of the global preclinical antibacterial pipeline: trends and gaps. Antimicrob. Agents Chemother. 68, e0053524 (2024).
Kan, J. et al. Oxydifficidin, a potent Neisseria gonorrhoeae antibiotic due to DedA-assisted uptake and ribosomal protein RplL sensitivity. Elife 13, e99281 (2025).
Leimer, N. et al. A selective antibiotic for Lyme disease. Cell 184, 5405–5418.e5416 (2021).
Qian, Y. et al. A Potent and Narrow-Spectrum Antibacterial against Clostridioides difficile Infection. J. Med. Chem. 66, 13891–13899 (2023).
Baker, T. A. & Sauer, R. T. ATP-dependent proteases of bacteria: recognition logic and operating principles. Trends Biochem Sci. 31, 647–653 (2006).
Kim, G. L., Akoolo, L. & Parker, D. The ClpXP Protease Contributes to Staphylococcus aureus Pneumonia. J. Infect. Dis. 222, 1400–1404 (2020).
Mawla, G. D. et al. ClpP1P2 peptidase activity promotes biofilm formation in Pseudomonas aeruginosa. Mol. Microbiol. 115, 1094–1109 (2021).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks - PubMed. Genome Res. 13, 2498–2504 (2003).
Frey, S. & Gorlich, D. A new set of highly efficient, tag-cleaving proteases for purifying recombinant proteins. J. Chromatogr. A 1337, 95–105 (2014).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Punjani, A. Algorithmic Advances in Single Particle Cryo-EM Data Processing Using CryoSPARC. Microsc. Microanal. 26, 2322–2323 (2020).
Rubinstein, J. L. & Brubaker, M. A. Alignment of cryo-EM movies of individual particles by optimization of image translations. J. Struct. Biol. 192, 188–195 (2015).
Zivanov, J., Nakane, T. & Scheres, S. H. Estimation of high-order aberrations and anisotropic magnification from cryo-EM data sets in RELION-3.1. IUCrJ 7, 253–267 (2020).
Zivanov, J., Nakane, T. & Scheres, S. H. A Bayesian approach to beam-induced motion correction in cryo-EM single-particle analysis. IUCrJ 6, 5–17 (2019).
Pettersen, E. F. et al. UCSF Chimera-a visualization system for exploratory research and analysis. J. Comput Chem. 25, 1605–1612 (2004).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr. D. Struct. Biol. 74, 519–530 (2018).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D. Struct. Biol. 74, 531–544 (2018).
Pettersen, E. F. et al. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Steinegger, M. & Soding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
Paysan-Lafosse, T. et al. The Pfam protein families database: embracing AI/ML. Nucleic Acids Res. 53, D523–D534 (2025).
Klimke, W. et al. The National Center for Biotechnology Information’s Protein Clusters Database. Nucleic Acids Res. 37, D216–D223 (2009).
Capra, N. et al. Adapting an acyl CoA ligase from Metallosphaera sedula for lactam formation by structure-guided protein engineering. Front. Catal. 4, 1360129 (2024).
Acknowledgements
We are grateful to Yiming Niu at The Rockefeller University for providing detailed guidance on cryo-EM data analysis. We thank Mark Ebrahim, Johanna Sotiris, Honkit Ng and the Evelyn Gruss Lipper Cryo-Electron Microscopy Resource Center for cryo-EM sample preparation and data collection support. This work was supported by National Institutes of Health grant R35GM122559 (SFB). K.S. is supported by NIH T32 GM136640-Tan to K.S.
Author information
Authors and Affiliations
Contributions
S.F.B. conceptualized and supervised the study. J.K. and K.S. designed and performed most experiments and data analysis. A.M. performed the HRMS and NMR data analysis. Y.H. performed the bioinformatics analysis. M.A.T. performed the molecular cloning. J.B. contributed to the design of molecular cloning. C.P. assisted with the large-scale extraction. R.E.B. synthesized tuscamide. A.B. synthesized the jomthonic acid analog. S.F.B., J.K. and K.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Yi Yu and Jingdan Liang for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kan, J., Spotton, K., Morales-Amador, A. et al. Mode of action guided metagenomic natural product discovery reveals convergent evolution of a ClpP-targeting motif. Nat Commun (2026). https://doi.org/10.1038/s41467-026-71586-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-71586-7