Abstract
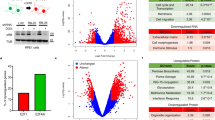
The retinoblastoma protein (RB) is a well-characterized repressor of E2F transcriptional activity that controls genes involved in cell proliferation during the G1 phase. Here, we examine the effect of RB on chromatin organization and uncover a non-canonical role for RB in which it promotes the removal of cohesin from insulators and thereby induces the expression of adjacent genes. We identify that RB colocalizes with cohesin across the human genome. During mitosis, RB facilitates the removal of cohesin from CTCF sites, with this effect persisting into early G1, impacting loop formation and extrusion at topologically associating domain (TAD) boundaries. Chromosome conformation capture assays reveal that RB reduces insulation at TAD boundaries and promotes enhancer-promoter interactions marked by histone 3 lysine 27 acetylation. In this way, RB enhances the transcription of hundreds of genes beyond the E2F transcriptional program, a finding that expands our understanding of this key tumor suppressor and cell cycle regulator.
Similar content being viewed by others
Data availability
Source data are provided as a Source Data file. The RB, E2F1, CTCF, c-Jun, H3K4me1, H3K4me3, and H3K27ac ChIP-seq from asynchronous hTERT-RPE1 cells are available at GEO under the Series GSE17603523 [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE176035]. The SMC3, acSMC3, H3K27ac, and H3K27me3 ChIP-seq, ATAC-seq, MNase-seq, and RNA-seq datasets from hTERT-RPE1 cells are available at GEO under the SuperSeries GSE246946. Individual accession codes are as follows: ATAC-seq, GSE246942; ChIP-seq, GSE246943; RNA-seq, GSE246944; and MNase-seq, GSE246945. Additionally, STAG1, STAG2, and MED15 ChIP-seq from G1-arrested hTERT-RPE1 cells and SMC3 ChIP-seq from Eg5 inhibitor-treated hTERT-RPE1 cells are available at GEO under the Series GSE315529. Single cell RNA-seq from RBWT and RBKO hTERT-RPE1 cells are available at GEO under the Series GSE315528. The Micro-C dataset is available at the EMBL-EBI ArrayExpress database under the accession code E-MTAB-13566, and the processed.mcool files are available at EMBL-EBI BioStudies database under the accession code S-BSST1221. The H3K27ac HiChIP in G1-arrested hTERT-RPE1 cells is available at GEO under the Series GSE266412. All mass spectrometry data can be accessed through the MassIVE data repository under the accession number MSV000098256 [https://massive.ucsd.edu/ProteoSAFe/dataset.jsp?task=1c31585ab6ae49a2a3599d8eec197c9d]. Source data are provided with this paper.
Code availability
Source codes utilized for the bioinformatics analyses are publicly available at our GitHub repository79 [https://github.com/hanjunlee21/Lee-et-al-2023].
References
Weinberg, R. A. The retinoblastoma protein and cell cycle control. Cell 81, 323–330 (1995).
Dyson, N. The regulation of E2F by pRB-family proteins. Genes Dev. 12, 2245–2262 (1998).
Lees, J. A., Buchkovich, K. J., Marshak, D. R., Anderson, C. W. & Harlow, E. The retinoblastoma protein is phosphorylated on multiple sites by human cdc2. EMBO J. 10, 4279–4290 (1991).
Chellappan, S. P., Hiebert, S., Mudryj, M., Horowitz, J. M. & Nevins, J. R. The E2F transcription factor is a cellular target for the RB protein. Cell 65, 1053–1061 (1991).
Chittenden, T., Livingston, D. M. & Kaelin, W. G. Jr. The T/E1A-binding domain of the retinoblastoma product can interact selectively with a sequence-specific DNA-binding protein. Cell 65, 1073–1082 (1991).
Bagchi, S., Weinmann, R. & Raychaudhuri, P. The retinoblastoma protein copurifies with E2F-I, an E1A-regulated inhibitor of the transcription factor E2F. Cell 65, 1063–1072 (1991).
Knudsen, E. S. & Knudsen, K. E. Tailoring to RB: tumour suppressor status and therapeutic response. Nat. Rev. Cancer 8, 714–724 (2008).
Rubin, S. M., Sage, J. & Skotheim, J. M. Integrating old and new paradigms of G1/S control. Mol. Cell 80, 183–192 (2020).
Burkhart, D. L. & Sage, J. Cellular mechanisms of tumour suppression by the retinoblastoma gene. Nat. Rev. Cancer 8, 671–682 (2008).
Witkiewicz, A. K., Kumarasamy, V., Sanidas, I. & Knudsen, E. S. Cancer cell cycle dystopia: heterogeneity, plasticity, and therapy. Trends Cancer 8, 711–725 (2022).
Knudson, A. G. Jr. Mutation and cancer: statistical study of retinoblastoma. Proc. Natl. Acad. Sci. USA 68, 820–823 (1971).
Godbout, R., Dryja, T. P., Squire, J., Gallie, B. L. & Phillips, R. A. Somatic inactivation of genes on chromosome 13 is a common event in retinoblastoma. Nature 304, 451–453 (1983).
Harbour, J. W. et al. Abnormalities in structure and expression of the human retinoblastoma gene in SCLC. Science 241, 353–357 (1988).
Ku, S. Y. et al. Rb1 and Trp53 cooperate to suppress prostate cancer lineage plasticity, metastasis, and antiandrogen resistance. Science 355, 78–83 (2017).
Mu, P. et al. SOX2 promotes lineage plasticity and antiandrogen resistance in TP53- and RB1-deficient prostate cancer. Science 355, 84–88 (2017).
Dick, F. A., Goodrich, D. W., Sage, J. & Dyson, N. J. Non-canonical functions of the RB protein in cancer. Nat. Rev. Cancer 18, 442–451 (2018).
Beltran, H. et al. Divergent clonal evolution of castration-resistant neuroendocrine prostate cancer. Nat. Med. 22, 298–305 (2016).
Gardner, E. E. et al. Lineage-specific intolerance to oncogenic drivers restricts histological transformation. Science 383, eadj1415 (2024).
Walter, D. M. et al. RB constrains lineage fidelity and multiple stages of tumour progression and metastasis. Nature 569, 423–427 (2019).
Hernando, E. et al. Rb inactivation promotes genomic instability by uncoupling cell cycle progression from mitotic control. Nature 430, 797–802 (2004).
Manning, A. L., Longworth, M. S. & Dyson, N. J. Loss of pRB causes centromere dysfunction and chromosomal instability. Genes Dev. 24, 1364–1376 (2010).
Manning, A. L. et al. Suppression of genome instability in pRB-deficient cells by enhancement of chromosome cohesion. Mol. Cell 53, 993–1004 (2014).
Sanidas, I. et al. Chromatin-bound RB targets promoters, enhancers, and CTCF-bound loci and is redistributed by cell-cycle progression. Mol. Cell 82, 3333–3349 e3339 (2022).
Sanidas, I., Lawrence, M. S. & Dyson, N. J. Patterns in the tapestry of chromatin-bound RB. Trends Cell Biol. 34, 288–298 (2024).
Krishnan, B. et al. Active RB causes visible changes in nuclear organization. J. Cell Biol. 221, e202102144 (2022).
Cheng, Y. et al. Perturb-tracing enables high-content screening of multiscale 3D genome regulators. Nat Methods 22, 950–961 (2025).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Davidson, I. F. et al. DNA loop extrusion by human cohesin. Science 366, 1338–1345 (2019).
Gabriele, M. et al. Dynamics of CTCF- and cohesin-mediated chromatin looping revealed by live-cell imaging. Science 376, 496–501 (2022).
Filippova, G. N. et al. An exceptionally conserved transcriptional repressor, CTCF, employs different combinations of zinc fingers to bind diverged promoter sequences of avian and mammalian c-myc oncogenes. Mol. Cell Biol. 16, 2802–2813 (1996).
Chakraborty, S. et al. Enhancer-promoter interactions can bypass CTCF-mediated boundaries and contribute to phenotypic robustness. Nat. Genet. 55, 280–290 (2023).
Kim, K. L. et al. Dissection of a CTCF topological boundary uncovers principles of enhancer-oncogene regulation. Mol. Cell 84, 1365–1376 e1367 (2024).
Hsieh, T. H. et al. Mapping nucleosome resolution chromosome folding in yeast by micro-C. Cell 162, 108–119 (2015).
Mumbach, M. R. et al. HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat. Methods 13, 919–922 (2016).
Sanidas, I. et al. A code of mono-phosphorylation modulates the function of RB. Mol. Cell 73, 985–1000 e1006 (2019).
Narasimha, A. M. et al. Cyclin D activates the Rb tumor suppressor by mono-phosphorylation. eLife 3, e02872 (2014).
Nicolay, B. N. et al. Proteomic analysis of pRb loss highlights a signature of decreased mitochondrial oxidative phosphorylation. Genes Dev. 29, 1875–1889 (2015).
Cremer, T. & Cremer, C. Chromosome territories, nuclear architecture and gene regulation in mammalian cells. Nat. Rev. Genet. 2, 292–301 (2001).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Schwarzer, W. et al. Two independent modes of chromatin organization revealed by cohesin removal. Nature 551, 51–56 (2017).
Vian, L. et al. The energetics and physiological impact of cohesin extrusion. Cell 173, 1165–1178 e1120 (2018).
Naumova, N. et al. Organization of the mitotic chromosome. Science 342, 948–953 (2013).
Haarhuis, J. H. I. et al. The cohesin release factor WAPL restricts chromatin loop extension. Cell 169, 693–707 e614 (2017).
Crane, E. et al. Condensin-driven remodelling of X chromosome topology during dosage compensation. Nature 523, 240–244 (2015).
Giorgetti, L. et al. Structural organization of the inactive X chromosome in the mouse. Nature 535, 575–579 (2016).
Munoz, S., Minamino, M., Casas-Delucchi, C. S., Patel, H. & Uhlmann, F. A role for chromatin remodeling in cohesin loading onto chromosomes. Mol. Cell 74, 664–673 e665 (2019).
Losada, A., Hirano, M. & Hirano, T. Identification of Xenopus SMC protein complexes required for sister chromatid cohesion. Genes Dev. 12, 1986–1997 (1998).
Waizenegger, I. C., Hauf, S., Meinke, A. & Peters, J. M. Two distinct pathways remove mammalian cohesin from chromosome arms in prophase and from centromeres in anaphase. Cell 103, 399–410 (2000).
Agarwal, H., Reisser, M., Wortmann, C. & Gebhardt, J. C. M. Direct observation of cell-cycle-dependent interactions between CTCF and chromatin. Biophys. J. 112, 2051–2055 (2017).
Tanaka, T., Cosma, M. P., Wirth, K. & Nasmyth, K. Identification of cohesin association sites at centromeres and along chromosome arms. Cell 98, 847–858 (1999).
Bell, A. C., West, A. G. & Felsenfeld, G. The protein CTCF is required for the enhancer blocking activity of vertebrate insulators. Cell 98, 387–396 (1999).
Pearson, J. D. et al. Binary pan-cancer classes with distinct vulnerabilities defined by pro- or anti-cancer YAP/TEAD activity. Cancer Cell 39, 1115–1134 e1112 (2021).
Zacksenhaus, E. et al. pRb controls proliferation, differentiation, and death of skeletal muscle cells and other lineages during embryogenesis. Genes Dev. 10, 3051–3064 (1996).
Li, F. Q., Coonrod, A. & Horwitz, M. Selection of a dominant negative retinoblastoma protein (RB) inhibiting satellite myoblast differentiation implies an indirect interaction between MyoD and RB. Mol. Cell Biol. 20, 5129–5139 (2000).
Zappia, M. P., Rogers, A., Islam, A. & Frolov, M. V. Rbf activates the myogenic transcriptional program to promote skeletal muscle differentiation. Cell Rep. 26, 702–719 e706 (2019).
Muller, G. A. et al. The CHR promoter element controls cell cycle-dependent gene transcription and binds the DREAM and MMB complexes. Nucleic Acids Res. 40, 1561–1578 (2012).
Ferrari, R. et al. Adenovirus small E1A employs the lysine acetylases p300/CBP and tumor suppressor Rb to repress select host genes and promote productive virus infection. Cell Host Microbe 16, 663–676 (2014).
Kareta, M. S. et al. Inhibition of pluripotency networks by the Rb tumor suppressor restricts reprogramming and tumorigenesis. Cell Stem Cell 16, 39–50 (2015).
Gonzalez-Vasconcellos, I. et al. The Rb1 tumour suppressor gene modifies telomeric chromatin architecture by regulating TERRA expression. Sci. Rep. 7, 42056 (2017).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320 e324 (2017).
Nielsen, S. J. et al. Rb targets histone H3 methylation and HP1 to promoters. Nature 412, 561–565 (2001).
Siddiqui, H., Fox, S. R., Gunawardena, R. W. & Knudsen, E. S. Loss of RB compromises specific heterochromatin modifications and modulates HP1alpha dynamics. J. Cell Physiol. 211, 131–137 (2007).
Markey, M. P. et al. Loss of the retinoblastoma tumor suppressor: differential action on transcriptional programs related to cell cycle control and immune function. Oncogene 26, 6307–6318 (2007).
Lu, X., Keo, V., Cheng, I. et al. NKX2-1 drives neuroendocrine transdifferentiation of prostate cancer via epigenetic and 3D chromatin remodeling. Nat Genet 57, 1966–1980 (2025).
Freeburg, N. F., Peterson, N., Ruiz, D. A., Gladstein, A. C. & Feldser, D. M. Metastatic competency and tumor spheroid formation are independent cell states governed by RB in lung adenocarcinoma. Cancer Res. Commun. 3, 1992–2002 (2023).
Thangavel, C. et al. RB loss promotes prostate cancer metastasis. Cancer Res. 77, 982–995 (2017).
Gibcus, J. H. et al. A pathway for mitotic chromosome formation. Science 359 https://doi.org/10.1126/science.aao6135 (2018).
Longworth, M. S., Herr, A., Ji, J. Y. & Dyson, N. J. RBF1 promotes chromatin condensation through a conserved interaction with the Condensin II protein dCAP-D3. Genes Dev. 22, 1011–1024 (2008).
Coschi, C. H. et al. Haploinsufficiency of an RB-E2F1-Condensin II complex leads to aberrant replication and aneuploidy. Cancer Discov. 4, 840–853 (2014).
Marshall, A. E., Ishak, C. A. & Dick, F. A. An RB-condensin II complex mediates long-range chromosome interactions and influences expression at divergently paired genes. Mol. Cell Biol. 40 https://doi.org/10.1128/MCB.00452-19 (2020).
Edwards, A. & Haas, W. Multiplexed quantitative proteomics for high-throughput comprehensive proteome comparisons of human cell lines. Methods Mol. Biol. 1394, 1–13 (2016).
Ting, L., Rad, R., Gygi, S. P. & Haas, W. MS3 eliminates ratio distortion in isobaric multiplexed quantitative proteomics. Nat. Methods 8, 937–940 (2011).
Elias, J. E. & Gygi, S. P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Lee, H. RB loss modulates chromatin organization by regulating cohesin-dependent loops and enhancer-promoter interactions. Zenodo https://doi.org/10.5281/zenodo.10062483 (2026).
Knight, P. A. & Ruiz, D. A fast algorithm for matrix balancing. IMA J. Numer. Anal. 33, 1029–1047 (2012).
Acknowledgements
We thank Dr. Nicholas J. Dyson for his unwavering support and critical reading of the manuscript. We also thank Dr. Miguel N. Rivera and Dr. Christopher J. Ott for their helpful discussions and feedback. This study was funded by National Institutes of Health grant R01CA236538 to Nicholas J. Dyson and Ioannis Sanidas and by Rullo Family Innovation Award to Michael S. Lawrence.
Author information
Authors and Affiliations
Contributions
Conceptualization: H.L., M.S.L., and I.Sanidas. Methodology: H.L., I.-M.G., M.S.L., and I.Sanidas. Investigation: H.L., I.-M.G., C.G.M., B.K., S.A., I.Salinas, R.M., and I.Sanidas. Visualization: H.L. Funding acquisition: M.S.L. and I.Sanidas. Project administration: M.S.-F., W.H., M.S.L., and I.Sanidas. Supervision: M.S.L. and I.Sanidas. Writing – original draft: H.L. Writing – review & editing: H.L., I.-M.G., M.S.L., and I.Sanidas.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Rolf Ohlsson and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lee, H., Gkotinakou, IM., McGrath, C.G. et al. RB loss modulates chromatin organization by regulating cohesin-dependent loops and enhancer-promoter interactions. Nat Commun (2026). https://doi.org/10.1038/s41467-026-71655-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-71655-x