Abstract
State-supported research funding agencies are critical to the scientific enterprise. However, it remains unclear how funding agencies cooperate with academic communities to realize common scientific goals. Here, we present a fully digital archive assembled by the National Human Genome Research Institute (NHGRI), focusing on the nascent stages of “genomics” as a scientific field and the everyday workings of the Human Genome Project and subsequent major genomics projects. We identify early events behind the conception of genome-wide association studies, clarify hitherto obscured factors around funding decisions, and how NHGRI and academics outside NHGRI ensured continuity in technical expertise across projects. The computational models we developed correctly recapitulate how academic experts and NHGRI increased adoption of genomics by jointly deciding which organisms’ genomes to sequence. Taken together, these findings reveal how a funding agency contributed to scientific innovation in a nascent field of science by repeatedly cooperating with the broader scientific community.
Similar content being viewed by others
Introduction
State-funded scientific research agencies serve as a critical means for societies to invest in science and generate scientific solutions to pressing challenges1,2,3,4. These agencies support the development of novel scientific concepts through funding5, direct employment of researchers6, and the creation of shared community resources7,8. From the case for unplanned science9 to mission-oriented state planning, research funding agencies have long been identified to foster innovation9,10,11,12,13,14. While existing scholarship has explored the roles of funding agencies15,16,17,18,19,20, the limits of available data have precluded analyses on how funding agencies organized alongside thousands of academics to realize common scientific goals, while doing so in an efficient, cost-effective and innovative manner.
We present and leverage a fully digital archive assembled by the National Human Genome Research Institute (NHGRI) to identify and characterize pivotal decisions and developments for a now ubiquitous field of science. This archive documents the development of the field of genomics across model organism sequencing, human variation research and genetic epidemiology. Unlike preceding modes of gene research, genomics focuses research on large sets of genes21,22. Today, genomics accounts for at least 20% of all biomedical research into human genes23.
Established in 1997 as one of the institutes and centers of the National Institutes of Health (NIH), NHGRI, along with the Department of Energy (DOE), led the US efforts toward the International Human Genome Project (HGP). The HGP and parallel commercial efforts, such as those conducted by Celera Genomics, cataloged the set of genes that form a human genome24,25 and enabled a wider adoption of genomics in biomedical research26. Prompted by the complex historical links between genetics and eugenics, as well as other issues such as genetic determinism and genetic reductionism, the HGP set aside funds (5% of the HGP’s total budget) to support research on ethical, legal, and social implications (ELSI)27. NHGRI continues to support ELSI research28, including the History of Genomics Program, which archived documents from the HGP and subsequent genomic projects. The NHGRI archive presently comprises 2 + million pages and continues to grow at an annual rate of 5%.
Leading scholars in genetics and the history and philosophy of biomedicine have already used the NHGRI archives for research29,30,31. However, these efforts have relied on close readings of individual documents, an approach that does not scale well for addressing questions like ours, where answers may be embedded across thousands of documents.
We therefore turned to computationally augmented mixed-methods approaches. We used and, when necessary, optimized existing open-source software to extract information without the need for a proprietary cloud infrastructure. Based on our companion research32 into the ethical use of artificial intelligence (AI) in analyzing archives with sensitive documents, we restricted our scholarly investigations to the Core Collection of the NHGRI archive. This collection, manually assembled and annotated by historians over more than 10 + years, contains a subset of documents of elevated importance to NHGRI, such as those of the HGP. Each document in the Core Collection is manually reviewed to ensure that it does not contain personal information unrelated to the institutional mission of NHGRI.
Here, we show through multiple interlinked examples how NHGRI, with academics from universities, academic institutions, and private, research-based non-profit institutions, established a foundation for what is now the influential field of genomics. Through a custom legal and computational framework for archival data, we enable internal government documents to be computationally accessible to AI and scholars at scale. In these documents, we identify how NHGRI directly responded to scientific communities by creating resources that would create an impactful technology (GWAS), how NHGRI leadership engaged with the external scientific community to resolve complex technical issues in large collaborations, and how, together, they decided upon funding proposals for the sequencing of non-human organisms.
Results
Core Collection chronicles a range of important genomics initiatives
The Core Collection consists of two primary types of digital documents, “born-physical” (e.g., physical letters, subsequently digitized) and “born-digital” (e.g., Microsoft Office files). Our data processing framework (Fig. 1A) divides the content of single files into individual documents because different physical documents were scanned together to increase throughput (“Methods”, Supplementary Fig. S1). Second, we use computer vision methods to remove handwriting, a processing step identified by our research into the ethics of AI32 (Fig. 1A). Lastly, we use machine learning (ML) approaches to extract text, mask and code personal information (Supplementary Figs. S2, S3 and Supplementary Table S1), and generate further metadata used throughout our study.
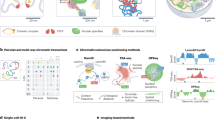
A Overview of the digitization, curation, and digital enrichment of the NHGRI archive guided by ethical considerations described in our companion manuscript32. PSS refers to page stream segmentation of collated scanned files. Founding vision refers to three themes of genomics from Collins et al., 200328. B Breakdown of the number of files, documents, and pages of archival materials inside the Core Collection. MS refers to Microsoft Office suite files. C The majority of documents whose dates were extracted from the Core Collection fall between 1988 to 2012. D The timeline of major genomics projects chronicled by the Core Collection. PAGE refers to Population Architecture Using Genomics and Epidemiology Consortium, eMERGE refers to Electronic Medical Records and Genomics Network, H3Africa refers to Human, Heredity & Health in Africa. E t-SNE projection of unsupervised representation of all text documents (dots) from the Core Collection. Colors correspond to folders assigned by the History of Genomics Program before computational analysis (Supplementary Data S1).
The Core Collection comprises 2073 born-digital files and 1039 PDF files (Fig. 1B, outer ring), corresponding to a total of 22,843 different documents (Fig. 1B, center ring) with a total of 92,519 pages of text (Fig. 1B, inner ring). More than 80% of the documents for which we could computationally extract a date (Supplementary Figs. S4–S6 and Supplementary Table S2, Supplementary Text – Robustness Analysis) originated between 1993 and 2007 (Fig. 1C), the timespan anticipated to be covered by the underlying curation efforts (Supplementary Fig. S7). The Core Collection covers the HGP and subsequent large genomics projects which were supported by NHGRI (Fig. 1D, Supplementary Fig. S7 and Supplementary Data S1). Covered projects include the International HapMap Project, which developed a genome-wide comparative map of haplotype variation; the Large-Scale Genome Sequencing and Analysis Centers (LSAC), which sequenced a significant number of non-human genomes by utilizing the knowledge and expertise gained from sequencing a human genome; the Encyclopedia of DNA Elements (ENCODE), which aimed to identify functional elements of a human genome; and modENCODE, which aimed to identify functional, and often, conserved elements of non-human genomes.
As certain projects captured in the Core Collection are scientifically and temporally close, it is unclear whether computational approaches can retrieve meaningful information to distinguish projects. Before advancing to more targeted analyses, we hence evaluated the feasibility of computational approaches. Before the start of our current study, the History of Genomics Program had manually organized files into folders, which correspond to different genomics projects (Supplementary Data S1). We could hence contrast this manual organization (Supplementary Data S1) against an unsupervised representation of text documents based on word frequencies (“Methods”, Supplementary Text – Robustness Analysis). Reassuringly, this unsupervised representation aligned with the manual organization. Documents of the HGP separated from documents of subsequent efforts, such as LSAC, ENCODE, and modENCODE (Fig. 1E and Supplementary Figs. S8–S12). This confirms that computational approaches can recover project-specific information from documents. Core Collection documents also separate from publicly facing data, such as NHGRI’s requests for grant proposals or research articles supported by NHGRI funding (Supplementary Fig. S13). This suggests that archival materials could enable distinctive insights into the workings of NHGRI.
NHGRI promoted the early development of a transformative technique
Computational analyses of publicly available bibliometric data demonstrate that NHGRI has been an unusually successful funding agency. For instance, research publications supported through NHGRI funding have received more citations than research publications of any of the other 20 core institutes of the NIH, and are more likely to be among the 5% most cited biomedical publications (Supplementary Fig. S14). Research publications supported by NHGRI received the second-highest number of citations by patents (Supplementary Fig. S14). They also rank fourth in disrupting33 the scientific literature, where disruption is quantified through the extent that NIH-supported publications uncouple references in later publications from preceding scientific literature (Supplementary Fig. S14). However, such analyses cannot resolve how NHGRI contributed to early stages of scientific innovation that precede individual research grants or publication metrics.
We thus wanted to ensure that our investigations into historical materials of NHGRI were focused on developments important to NHGRI as it supported the transition of the nascent field of genomics from the HGP to subsequent genomics efforts. Only a fraction of the documents in the Core Collection have hitherto been manually annotated with keywords (Fig. 2A). Therefore, we used entity recognition and pattern matching to extract keywords for each document in the Core Collection, covering biological phenomena, techniques, organizations and people (Fig. 2B and Supplementary Fig. S15). Among 69,895 keywords, 1246 keywords (hereto referred to as “enriched”) appeared statistically more often in documents from one of the four genomic projects that followed it. To guide interpretation, we hierarchically clustered enriched keywords (Supplementary Figs. S15–S18) according to their occurrence in each project. We highlighted some of the most frequent keywords in Fig. 2C (see Supplementary Data S2 for the full list).
A Documents in the Core Collection have been annotated with keywords by the History of Genomics Program, but the scale of the collection left gaps in annotation, even after one decade of manual annotation work. B Entity recognition and pattern matching fill this gap by computationally generating relevant keywords on biological phenomena, techniques, organizations, and individuals. We use two-sided Fisher’s Exact Test with Bonferroni correction to detect keywords that appear more in projects that follow the HGP (we call this “enriched” henceforth). C Hierarchical clustering of 1246 keywords from panel B by the share of documents in each project containing the keyword, normalized across projects by a standard Z score. See Data S2 for the list of keywords and Fig. S17 for the entire row-wide dendrograms. D Swarm plot shows that genomic techniques (dots) enriched among the four genomics projects that followed the HGP already occurred in documents before the start of these projects. One such technique is genome-wide association studies (red dot). E Bibliometric analysis of investigative techniques (n = 2601; defined using Medical Subject Headings) in the biomedical literature. (top) shows the share of publications that are among the 5% most cited publications. (bottom) shows the share of new genes introduced to the biomedical literature according to investigative techniques used in these initial publications. The red vertical line indicates GWAS. F Timeline of the development of GWAS. The histogram shows the occurrence of GWAS in the Core Collection. The dashed lines indicate key publications: Risch and Merikangas demonstrated the mathematical feasibility of GWAS33, the start of the International HapMap Project34, and – what have been independently considered35,36,37 to be – the first GWAS article38, and the first large-scale GWAS39.
A first insight was that most keywords are enriched only for one project (Supplementary Fig. S18). This demonstrates that parallel genomics projects differed. A second insight was that project-specific keywords reassuringly match prior understanding of early genomics research. For example, single nucleotide polymorphism (SNP), a term for sites or base pairs of the genome that vary among individuals, is prominent within HapMap documents (Fig. 2C). This was to be expected as HapMap aimed to uncover recurring patterns of SNPs (co-inherited through haplotypes)34. Equally reassuring, animal models featured prominently in documents of LSAC, a program that leveraged sequencing capacity to analyze the genomes of a variety of organisms (Fig. 2C). The Department of Energy (DOE) is prominent in HGP documents (Fig. 2C), consistent with HGP’s origins inside the DOE’s Office of Science35. Aligning with oral histories, which identified former NHGRI Deputy Director Elke Jordan (1988–2002) as one of NHGRI’s most significant but least publicly acknowledged leaders of early genomics efforts35, we found frequent mentions of Dr. Jordan in documents from HGP and the LSAC (Fig. 2C and Supplementary Fig. S19). Overall, the statistically identified keywords align with known insights into the history of genomics and highlight important differences among projects.
To guide our analyses toward early phases of scientific innovation, we triangulated the first occurrence of techniques that were enriched among the four genomics projects that followed the HGP. Of a total of 21 genomic techniques, 20 have occurred prior to the start of the relevant projects (Fig. 2D). To understand the specifics of NHGRI’s role, we focused on the most frequently occurring technique, genome-wide association studies (GWAS). This technique searches for small variations in the genomes of large populations to identify gene variants that are associated with phenotypes, such as the occurrence of a specific disease36. Between 2005, the year of publication of the first research article using GWAS36, and 2020, a total of 30,174 research articles mentioned GWAS. These articles were more likely to receive more citations than articles mentioning most other techniques (top 0.6% of all 2845 techniques, Fig. 2E). Demonstrating impact beyond citations, studies mentioning GWAS introduced the greatest number of previously uncharacterized genes into the scientific literature (Fig. 2E).
The earliest mention of GWAS in the Core Collection is in the agenda for a March 1997 NHGRI workshop about human variation. This workshop was prompted by a perspective article published in September 1996 in Science, which outlined GWAS’ mathematical feasibility, but remarked that too few single- nucleotide polymorphisms were known to implement GWAS37. To answer this call to the community, NHGRI leadership invited a group of external academic experts to a workshop where they discussed the path to develop GWAS (Fig. 2F and Supplementary Figs. S20, 21). The following year, NHGRI leadership allocated infrastructure and resources that could support the future development of GWAS38. NHGRI supported and co-led the International HapMap Project, which started in 2002. The HapMap project identified millions of co-inherited single-nucleotide polymorphisms, which were indeed used by the initial GWAS39,40. Jointly, these observations – which have been absent from public histories of GWAS41,42 – demonstrate that NHGRI and academics cooperated toward the shared goal of developing an innovative technique, GWAS, with NHGRI contributing by coordinating actions toward a resource beyond the capacity of typical research laboratories or biomedical investigators.
NHGRI and a small group of academic experts provided continuity between HGP and HapMap
To understand how NHGRI cooperated with academics during the everyday operations of individual genomics projects, we turned to email communications. From a total of 4,111 email documents, we extracted 47,406 sender–recipient pairs (“Methods”, Supplementary Fig. S23, S24 and Supplementary Table S3 and Supplementary Data S3). Since email domains did not always differentiate NHGRI from other NIH institutes (e.g., @nih.gov), we did not distinguish between NHGRI staff and staff from other NIH centers and institutes in our analysis.
We represented these email communications as a directed network with arcs pointing from senders to recipients and arcs weighted by the number of emails (Fig. 3A). Because the emails archived for the LSAC program, ENCODE, and modENCODE projects constituted less than 0.5% of the emails in the Core Collection, we focused on the HGP and HapMap projects. Emails from NIH staff to other NIH staff made up 36.1% of all email conversations, NIH staff to academics 17.8% and academics to NIH staff 7.6%. 47% of identifiable NIH staff were involved in communications regarding the HGP and HapMap projects, compared to only 21% of academics (P < 0.001, two-sided Fisher’s Exact Test) (Fig. 3B), suggesting a higher rate of turnover among academics during the transition from HGP to HapMap.
A Directed, weighted network of 47,046 email conversations found in 4111 scanned emails. Every node represents an email recipient or sender. Edges are colored by the same manual folder-level organization as Fig. 1E. Parentheses are percentages of edges. B Overlap of nodes between HGP and HapMap email conversations for NIH staff and academics. C Communities detected among sender-recipient pairs from the International HapMap Project using nested stochastic block models45. The color of the nodes indicates individuals’ affiliation with NIH, academia, or others. Others are commercial, non-NIH governmental and non-academic institutions. D As C, but colored by the number of emails the individual sent and received. E As C but colored by membership in the Kitchen Cabinet or membership in the Steering Committee (but not the Kitchen Cabinet). The rest of the HapMap are individuals outside these two groups. F Sankey diagram of individuals inside HGP and HapMap email communication. Blocks are partitions of nodes and were assigned by the stochastic block model. G Triad analysis of brokerage roles46 among three individuals connected by directed emails (two sender-recipient pairs). The top scheme illustrates triad architecture around middle individuals (n) with white/gray/black illustrating membership in different groups (Kitchen Cabinet, Steering Committee, and the rest of HapMap). The box-and-whisker plot shows a fraction of triads within a group; whiskers bound the min–max values, the bounds of the box represent lower and upper quartiles, and the central value contains the median value. All comparisons across three groups by a two-sided Mann-Whitney Test resulted in P < 0.001. The U statistics, listed in the order of Kitchen Cabinet vs. Steering Committee and Steering Committee vs. the rest of HapMap, are 254.0, 665.0, 20.0, 99.0, 51.0, 114.5, 262.0, 684.0, 24.0, and 572.0.
We anticipated that email networks could recover latent organizational structures as they provide a proxy for human behavior and communication43,44. Using a stochastic block modeling approach (“Methods”, Supplementary Table S4), we found that most participants in HGP communication do not group into one single community (Supplementary Fig. S24). Of the communities detected, one comprised primarily of NIH staff (Supplementary Fig. S24). Unlike the HGP, large portions of email communication in the HapMap (40.0%) occurred between two major communities that together accounted for 13% of participants: one comprising primarily of academics and the other consisting of both NIH staff and academics (Figs. 3C, D). The former includes members of HapMap’s International Steering Committee, whereas the latter includes a previously not publicly acknowledged group, which a member of NHGRI leadership sometimes referred to as a “Kitchen Cabinet” (Fig. 3E).
The Kitchen Cabinet consisted of five members of NIH leadership (including Francis Collins), two members of other staff, and four academics. Nine members of the Kitchen Cabinet already appeared in the communications of the HGP. Of these, six were members of NIH, one an academic who served on the National Advisory Council for Human Genome Research (NACHGR) during HGP, and two were academics who founded one of the five core sequencing centers from HGP (so called “G5” in Supplementary Fig. S7). NACHGR was established in 1990 by the Public Health Services Act (42 U.S.C. 284a) to advise, consult, and make recommendations about activities regarding NHGRI. Hence, the Kitchen Cabinet may have been critical to providing continuity between the HGP and HapMap (Fig. 3F). To better describe the latent role of the Kitchen Cabinet, we implemented Gould and Fernandez’s45 brokerage typology, which categorizes how information flows within a network. Members of the Kitchen Cabinet, including Francis Collins, most frequently acted as consultants or liaisons (Fig. 3G and Supplementary Figs. S25, S26).
To identify specific examples of how Kitchen Cabinet members acted as consultants or liaisons, we extracted meeting agendas of the Kitchen Cabinet and of the subsequent meetings of the Steering Committee meetings. A comparison of the two sets of agendas revealed that items in the Kitchen Cabinet meetings mostly concerned technical matters, and that those items were also included in the subsequent Steering Committee meetings (Supplementary Table S5, Reviewer Materials). This suggests that the primary function of the Kitchen Cabinet was to undertake preparation work on complex issues ahead of the Steering Committee meetings. Overall, our data-driven analysis of the email communication of HapMap demonstrates that NHGRI leadership closely worked with academic experts and ensured the continuity of tacit technical and organizational knowledge between the HGP and HapMap.
How academics and NHGRI jointly decided which organisms’ genomes to sequence
LSAC’s sequence target selection effort was one of several major genomics projects, along with HapMap and the ENCODE Project, that followed the HGP (Fig. 1D) and utilized “lessons learned” from that effort. While HapMap was organized around the creation of a single resource, leveraging an emerging technology – scalable, accurate genotyping – a critical main objective of the LSAC was the creation of genome sequences of non-human organisms. Innovation in the LSAC, therefore, in the context of comparative model organism sequencing, originated from applying existing genome sequencing technology to novel domains of biomedical research. Accordingly, NHGRI needed to include the input from many scientific communities in deciding which organisms to sequence. This raises the question of whether computational approaches can retrospectively recreate these decisions and find correlates of funding outcomes.
Before investigating the decisions themselves, we first reconstructed the decision-making process through manual inspection of archived materials (Fig. 4A). Using the LSAC, NHGRI publicly called for proposals on which organisms to sequence. Proposals, which originated from both scientific laboratories outside of NHGRI and from NHGRI working groups (e.g., annotating the human genome working group, comparative genome evolution working group), comprised of both NHGRI staff and academic experts, laid out the rationale for the sequencing of a particular organism. These proposals were reviewed by a committee comprised of NHGRI staff and two members from NACHGR. The committee’s recommendations were later submitted to the entire NACHGR, which formally approved or rejected proposals. The production of genome sequences of approved organisms was delegated to sequencing centers established during the HGP. To ensure data completeness, we compared genome sequences deposited at NCBI’s BioProject to NACHGR decisions in our data and found coverage of 97.4% of the organisms supported by the LSAC in 10 years following the HGP (Supplementary Fig. S27).
A Graphical overview of the decision-making process behind non-human organism sequencing within the LSAC. B Proposal submissions colored by approval (red for rejected, blue for approved) and ordered by animal phyla. We indicate statistical overrepresentation of NHGRI submissions compared to the taxa diversity (Supplementary Fig. S28). Statistical significance according to the two-sided Fisher’s Exact test is displayed by * only for those below P < 0.05. C Receiver operating characteristic curve (ROC) of machine learning models trained on 100 Monte Carlo cross-validation sets of 13 features. Legend shows mean (solid line) and +/− 1 standard deviation (shaded area) of the area under the ROC (AUROC). D SHAP analysis for two features, the community size (measured in log10 publications) and lexical diversity. The scatter points are SHAP values, with positive SHAP values indicating an association with approval. The line indicates a rolling average of 1,000 values. Bars are a histogram of feature values. E The box-and-whisker plot represents the share of publications that use genomics approaches for approved (n = 159) and rejected (n = 88) model organisms; whiskers bound the min–max values, the bounds of the box represent lower and upper quartiles, and the central value contains the median value. Two-sided Mann-Whitney U test between approved and rejected groups showed significant differences for 1-5 years prior to council decision (U = 5791.5, P = 0.023), 1-5 years after the council decision (U = 8154.0, P = 0.031), and 6–10 years after the council decision (U = 8220.5, P = 0.023).
The original call for proposals emphasized “improvement of human health” and “informing human biology” (Supplementary Note). Consistent with such a scope, we encountered among submitted proposals an overrepresentation of Chordata, the phylum of homo sapiens (Fig. 4B and Supplementary Fig. S28). This is also consistent with a recent systematic review of publicly available animal genomes46 and our observation of the term phylogeny being overrepresented in LSAC documents (Fig. 2C).
To uncover some of the reasons behind NACHGR’s decisions beyond Chordata, we developed a machine learning model to recapitulate these decisions (Data S5). We surmised that decisions could have been influenced by features that capture biological phenomena (e.g., genome size), organizational aspects (e.g., if multiple organisms are bundled in the same proposal), reputation (e.g., H-index of proposal authors), or linguistics (e.g., argumentativeness). Given the prevalence of categorical data, we used CatBoost47 as our machine learning approach. Given the historical nature of the LSAC and thus the absence of independent data to evaluate the performance of our models, we applied Monte Carlo cross-validation, which trains models on random subsets of NACHGR’s decisions and evaluates models on decisions excluded from the training process. Demonstrating that this machine learning approach accurately recapitulates NACHGR’s decisions, the models yielded an average area under the receiver operating characteristic (AUROC) of 0.94 + /− 0.02 (Fig. 4C). Models trained on all features performed significantly better than models trained only on subsets of features. This suggests that no single feature captures the full decision-making process at NHGRI (Fig. 4C).
An analysis of the features’ SHAP values48 showed that all features contributed to the final models. The largest contribution stemmed from the size of the research community, with a higher number of publications on individual organisms being associated with approval. The second and third largest contributions stemmed from the proposals’ lexical diversity and argumentativeness, with intermediate levels of both seeming most associated with approval (Fig. 4C and Supplementary Figs. S29, S30). Reassuringly, approval was not associated with the origins of the proposals, whether they originated from an NHGRI internal working group or from external academic communities.
Studies of scientific innovation implicitly surmise that novelty alone is not sufficient for innovation – innovation also requires a measurable impact on the conduit of science6,49. While it has been hypothesized that the choice of an organism for sequencing changed how subsequent research on these organisms was done50,51,52,53, this hypothesis could not be tested before due to the lack of a counterfactual. In contrast, the Core Collection enabled us to build such a counterfactual using the set of organisms that were considered for sequencing but were rejected. We thus scanned the published biomedical literature to determine whether studies on specific organisms utilized genomic approaches. Indeed, after NACHGR’s approval, publications on approved organisms (n = 159) saw a higher rate of use of genomic approaches (Fig. 4E, P = 0.031, median of 0.15 and 0.10, respectively) during the 5 years after the decision. Simultaneously, there was no statistically significant difference on the overall amount of research done on individual organisms after NACHGR’s decisions on approved (N = 159) and rejected (N = 88) projects (Supplementary Fig. S31). This suggests that NACHGR’s decisions align with the conduit of science and that the availability of a genome sequence accelerated the adoption of genomic approaches in organismal communities by existing researchers.
Overall, our analysis of the entire decision-making process behind the LSAC’s sequence target section process and its impact on research surrounding specific organisms demonstrates that it is possible to trace innovation in a nascent field of inquiry back to the intricate cooperation patterns between a funding agency and groups of external grantees, as well as other advisors. While existing insights reveal the end outcome of decisions to fund sequencing of organisms46, the application of computational approaches on archived materials enabled us to quantitatively recapitulate how these decisions were made (Fig. 4C).
Discussion
While funding agencies are expected to contribute to innovation (Supplementary Fig. S14), our manuscript shows how a funding agency supported nascent stages of one of the most successful fields of contemporary science. A repeating theme in our analysis is that the NHGRI staff were substantially involved in a variety of genomics projects and convened with academic experts to support the creation of shared data resources, including information on haplotype structure and comparative organismal sequences. At first glance, the institutional behavior of NHGRI appears collaborative. However, anecdotes published elsewhere show that multiple prominent academics perceived NHGRI’s involvement as restricting academics’ ability to pursue investigator-initiated research54,55. Likewise, NHGRI staff who served during the HGP described interactions with academics as cooperative to emphasize NHGRI’s responsibility toward the projects rather than individual investigators35. In line with such a view, we see that NHGRI repeatedly identified and amplified voices within the scientific community that aligned with NHGRI’s mission.
Some level of close cooperation between NHGRI and academics is by design: For instance, NHGRI’s advisory council, NACHGR, is mandated to recruit from not only leading representatives of the health and scientific disciplines, but also of public policy, behavioral and sociological research, and economics56. In addition, one of the formal responsibilities of NACHGR is to hold ad-hoc meetings and working groups, which often brought together NACHGR members, intramural researchers, and external academics35.
The lack of strict separation among academia, the outside grantee community, and NHGRI may be conceptualized as funding agencies being boundary organizations4,5, bodies jointly created by both science and non-science stakeholders to advance mutual interests and resolve conflicts arising from inherent differences57,58,59. Further supporting this view, we found that NHGRI staff, including the top leadership, consistently exhibited behaviors of both scientists (e.g., honing technical details) and administrators (e.g., organizing new groups and the holistic evaluation of projects). This interaction promoted long-term relationships (e.g., sequencing centers from HGP) and stabilized the boundary (e.g., degree of cooperation).
The fully digital Core Collection of the NHGRI archive enabled us to retrospectively identify important organizational and conceptual continuities between the HGP and subsequent genomic projects. Our findings complicate any discussion of “post genomics” that also presents the HGP as a monolithic “project,” largely separated from later genomics efforts35,60. We suspect that these superficially conflicting perspectives may have been shaped by the selective availability of public data. As our study demonstrates, important innovations and structures for fostering that innovation61, were already established ahead of the awarding of grants or the publication of research results and thus might not be visible using externally available sources like funding announcements or citations. This further confirms previous scholarship12,14 that innovation in funding agencies take time and programmatic efforts, connecting back to the mission-driven NIH served to be generative, not incidental, of innovation. Motivated by the Common Disease – Common Variant hypothesis62,63, NHGRI leadership identified GWAS as the technically feasible64 direction towards studying genetic drivers of common disease and thus programmatically created resources such as HapMap that later enabled adoption of GWAS (Fig. 3). Similarly, NHGRI decisions accelerated the adoption of genomics in existing model organism communities (Fig. 4E).
Since our conclusions are based on documents archived by one collection of documents, it is unclear to what extent they generalize to the entirety of NHGRI or other research funding agencies or scientific fields. Arguing for generalizability, one must highlight that, at sufficient granularity, the Core Collection of the NHGRI archive covers a variety of major initiatives across many domains of genomics that brought numerous distinct scientific communities into the process. Our findings on NHGRI and early genomics also complement recent sociological research into the National Cancer Institute, and suggest that findings of funding agencies as builders of techno-capacity are part of a more generalizable pattern12,14,65,66. Similarly, that – beyond individual funding agencies – staff of funding agencies acts as research administrators and scientists13. However, one can also argue that innovation in the field of genomics is one of those special cases where generalizability is secondary to importance, where the case of genomics also illustrates radical novelty. Genomics has been one of the most transformative research areas in the recent history of science, whether evaluated by its current prevalence (Fig. 4), the number of citations received23 or the inclusion of new genes into the scientific literature (Fig. 2E). It also led to an increase of team science in biology23 and policies around the open sharing of data67. Scientific endeavors of equivalent importance seem few and far in-between. It may be then no surprise that – when evaluated through citation-based impact metrics – research supported by NHGRI has been a notable success of US science (Supplementary Fig. S14).
Given our anticipation of a near future where AI-enabled archives become commonplace, we feel responsible to set a good reference on how to balance accuracy with completeness, and openness with the autonomy and rights of individuals whose activities are represented in archived documents. Concerning the former, we believe that the expert-annotated and human-reviewed Core Collection enabled us to quantify the accuracy of our processes, thus building confidence in the appropriateness of our approach (Fig. 1E and Supplementary Figs. S2–S13). Concerning the latter, we undertook a careful study of the ethics of using AI on the NHGRI archive32. We consciously chose to refrain from technically feasible but ethically uncertain analyses. These and other decisions were part of a multi-layered approach that included the encoding of sensitive information, secure local cyber-infrastructure, manual and computational screening of documents, and a legal framework for the exchange of data. This approach is the basis of a consortium we recently established with the responsibility to create training opportunities, maintain and improve our tools, and enable data sharing so that other historical archives are enabled to preserve and study their domain data (See Code availability). As part of this consortium, the extension of the legal and computational framework presented in the current manuscript towards the entire NHGRI archive is ongoing.
Although our algorithms to mask people and sensitive information work well (Supplementary Fig. S2 and Supplementary Table S1), we advocate that data made accessible to AI-enabled tools be “coded” rather than “deidentified” to ensure accountability from data stewards and to reduce the risk of harm to individuals while simultaneously enabling scholarly insight32. Importantly, we contend this approach represents a new posture towards potentially sensitive data in archival domains, which acknowledges the difficulty of predicting potential future risks (e.g., the capability of AI and other such tools to reidentify). Inversely, we underscore that prioritizing ethics over technical feasibility will enable more studies involving partnerships between governmental agencies and academic scholars.
Methods
Ethics approval
Upon submission to the Northwestern University Institutional Review Board, our study (STU00218837) was determined to be exempt. Prior to approval, we evaluated potential risks associated with applying computational methods to the Core Collection, as described in an accompanying manuscript32. Three key measures were implemented to minimize these risks: (1) preserving data ownership with NHGRI; (2) creating a digital version of the archive in which potentially sensitive information was removed, encrypted, or not made publicly available; and (3) conducting analyses within an isolated, encrypted, access-controlled computational environment (i.e., a dedicated workstation).
Page stream segmentation
To enable the economic digitization of physical documents, most documents in the Core Collection were digitized through a high-throughput scanning process using a Panasonic High-Speed scanner at 300 DPI resolution that collates multiple documents into a single PDF file. Page stream segmentation involves recovering the individual document boundaries from such compilations. This process was essential as it enabled the metadata assignment to each respective logical document. We created a custom interface using LabelStudio v1.8.0 to manually label all 79,743 pages from 1,039 PDFs in 80 person-hours (16 pages per minute). We assessed the consistency of this splitting approach by comparing documents that are resolved from PDFs with the same content by hashing. From such duplicated PDFs (N = 134) that independently underwent this manual segmentation, we report a 100% consistency. We followed the rules shown in Supplementary Fig. S1 to guide segmentation.
Handwriting removal
We found 43% of scanned Core Collection pages to contain handwritten marks from crossed-out lines and circled words to handwritten comments in the margins and fully handwritten documents. Unlike traditional printed text, handwriting uses a more informal, personal language. As discussed in our accompanying manuscript on the ethics of our data processing32, we determined that, because of its more personal nature of communication and lack of optical character recognition (OCR) support for individual handwriting styles, handwriting requires separate handling from printed text. Moreover, we found that even formal data, such as Social Security Numbers, credit card details, and phone numbers, are often recorded in archive documents in handwritten form.
Handwriting presents an additional layer of encoded information on top of the printed content, but it is also an ‘artifact’ that obstructs traditional OCR engines. To deconvolute printed text from handwritten data present on the same page and often with mixed information, we used a U-Net based68 segmentation model. We trained the model using a corpus of synthetic documents that included both handwritten and printed information with labeled positions.
For building the synthetic corpus, we sourced printed materials from Industrial Documents Library69 files with no handwriting data, training manuals from the web, forms70,71 and selected printed documents from the Core Collection. We sourced handwritten marks from HASYv272 and Google’s “The Quick! Draw!” Dataset73, handwritten digit data from UCI74, and handwritten text from the IAM-database75. To replicate how humans may circle, underline, and write near existing words, we identified locations of random words in the printed background using the Tesseract v4.1.1 OCR engine. We then, at random, underlined, crossed out, or circled these words with synthetic handwritten marks to mimic those found in the Core Collection. In addition, we randomly placed generated handwritten words and sentences with varying font sizes, orientations, and structures. For form layouts, we placed handwritten characters in the logically appropriate regions, using both cursive and block letters.
The developed model operates on images resized to 2,048 ×2,048 pixels. It uses a convolutional network structure with symmetric downscaling and upscaling paths connected by bottleneck layers, allowing precise localization and context integration. The network architecture includes standard components such as convolutional layers, max pooling, and up-convolutional layers. We use the negative of the Sørensen-Dice coefficient76 where \({p}_{i}\) is the predicted value of class i and \({g}_{i}\) is the ground truth of class i, and \({{\mathscr{L}}}\) as the loss function, to enhance the boundary delineation between different image regions.
We trained the model on a Quadro RTX-8000 GPU using curriculum learning77 first with 512 by 512 pixels, then 1024 by 1024, and finally 2048 by 2048 pixels. We used the TensorFlow v2.8.0 framework, the Adam optimizer, and specific callbacks for early stopping. For the 300 synthetic handwritten documents of the test set, we achieved an average F-1 score of the redacted handwriting of 0.961 (0.018).
Text extraction
We classified PDFs in the Core Collection based on their origin: those with a digital origin contain embedded text, whereas those with a physical origin were ingested through scanning and contain embedded images. For PDFs with a digital origin, we used Apache Tika v2.6.0 to extract the text with default parameters. For scanned PDFs, we first removed handwritten marks and then used Tesseract v4.1.178 to apply OCR with automatic page segmentation mode to the resulting images. For Microsoft Office files, we used Apache Tika’s Microsoft Parser modules in the default setting to extract the content in raw text and the associated metadata.
Entity recognition and disambiguation
To protect the privacy of personally identifiable information (PII) captured in the archive and attribute metadata in archive documents, we detected in the raw text the following categories of data: name, email address, physical address, identifiers (e.g., numerical combinations that could be social security numbers, phone numbers, or credit card information), and organizations. To accomplish this goal, we fine-tuned a RoBERTa79 model using spaCy v3.7.2, a popular natural language processing library. We used LabelStudio v1.8.0 to annotate 600 pages from the Core Collection, which could then be used to train the model. Precision, recall, and F-1 scores on the categories listed above on a holdout validation set of 100 pages from the Core Collection exceed 0.86, 0.88, and 0.87, as shown in Supplementary Fig. S2 and Supplementary Table S1.
Because RoBERTa can only consider 512 tokens as the context window, we divided all texts by a stride of 300 tokens with a 50-token window to ensure that long texts are accommodated during entity recognition.
While entity recognition detected names of individuals in the corpus, linking such mentions to unique individuals required separate tasks, called entity disambiguation. For this section, we linked and disambiguated the names of individuals. Entity disambiguation approaches commonly draw upon existing knowledge bases around individuals. However, no such resource existed for individuals captured by the Core Collection. Therefore, we created a method inspired by the “Seed + expand” procedure80 to overcome an incomplete landscape of individuals or the lack of metadata information about each possible individual. We also opted for explicitly stated rules and string patterns to make the procedure interpretable and easily adaptable for other use cases.
Specifically, we first manually seeded our entity list by collecting the names of NHGRI staff, National Advisory Council on Human Genome Research (NACHGR) members, public officials (politicians), and prominent HGP scholars. Next, we created aliases for each of these individuals, including different combinations of their first, middle, and last names. We normalized all names by making the text lower-case, removing common prefixes and suffixes to names (e.g., Dr., Miss, Ph.D.), and non-alphabetic characters except for a comma in a name, which we considered to be an inverted name (e.g., Smith, Jane).
For every detected individual in the corpus, we matched their names against the aliases using a Weighted Ratio metric of the Levenshtein distance calculated using the Python library TheFuzz v0.22.1 (curve B in Supplementary Fig. S3). Empirically, we found that 92% of the Weighted Ratio match names close to their intended individual as some mentions of names included OCR artifacts, such as incorrectly recognized characters, superscripts, and subscripts.
Based on the number of individuals matched to the aliases, we concluded that our initial list of individuals is not comprehensive enough. Therefore, we extended it by grouping unmatched names by considering all of the mentions to be an alias of the same name if they fall at or above 92% Weighted Ratio score (curve C in Supplementary Fig. S3). We then created aliases for newly added individuals and matched them against the corpus once more. This procedure yielded 260,099 matched names in the corpus to disambiguate to 39,252 individuals.
We noted that the top 500 disambiguated individuals make up 50.7% of the mentions, suggesting a power-law relationship of individuals captured in organizations.
Masking and coding personal information
Coded data is one of the three categories of data defined in the Institutional Review Board guidelines. Coded data differs from deidentified data in that indirect identifiers or codes replace the 18 PII direct identifiers defined by the Health Insurance Portability and Accountability Act (HIPPA). Declaring all data in the Core Collection as deidentified would eliminate data stewardship and remove any restrictions on its use. Therefore, we created coded datasets and removed potentially sensitive information (e.g., names and addresses). By declaring this data as coded, we would enable data stewardship and avoid falsely declaring the data deidentified. Specifically, we replaced the mentions of names of individuals by the disambiguated identifiers if disambiguation was possible or by a generic category label (e.g., PERSON) if disambiguation was not possible (e.g., only having a first name). Similarly, we encoded physical locations, email addresses, and phone numbers by their non-identifying category labels. The model discussed under the above subsection Entity recognition and disambiguation was used to find mentions of names, locations, and email addresses. To further decrease false negative incidents of sensitive identifiers, we used a regular expression approach to match for numeric patterns commonly associated with social security and credit card numbers.
Date extraction
We inferred the date of a Microsoft Office document using “creation” and “modified” dates reported in its metadata. This metadata was extracted during the text extraction step using Apache Tika. Some documents showed inaccurate creation or late modification dates caused by template reuse or file conversion during preservation (resulting in a spurious later modification date). To account for such inaccuracies, we used the median of all valid dates as the document’s inferred date.
Furthermore, the metadata for digitally and physically born PDF documents, specifically the creation and modified dates, were not consistent due to the archival process. Therefore, to identify the dates of PDF documents, we compiled a set of common date and time patterns as regular expressions listed in Supplementary Table S2. Next, we detected all valid date-like strings using these regular expression patterns in a case-insensitive manner from the first two pages of the document. Then, we used DateParser v1.2.0 to parse the detected date strings. For both PDFs and MS documents, we inferred the document date as the median of all detected dates and accepted this inference if all detected dates fell within a range of 30 days. Based on the curation of the corpus, we did not consider any dates before 1980 or after 2020. Additional robustness analysis is described in Supplementary Text (section Robustness Analysis).
Projection of document similarity
We visualized the scope of the Core Collection (Fig. 1E) by training a Doc2Vec81 model with extracted text from all handwriting-removed born-physical PDFs, born-digital PDFs, MS Word and MS PowerPoint documents. We removed blank page documents, filtered out non-alphanumeric characters and make every token lowercase. We excluded MS Excel documents from this projection because they may contain too many numerical values. In total, we considered 22,070 documents from the Core Collection.
We used 100-dimensional vectors and trained the model for 50 epochs and a minimum token count of 2. Following a common procedure in the field82, we calculated pairwise document similarity using the cosine similarity of the two vectors representing the documents. Repeating this calculation for all pairs of documents, we obtained a 22,070 × 22,070 matrix.
Next, we used the openTSNE v1.0.083 implementation of t-distributed Stochastic Neighbor Embedding (t-SNE)84 to project the 100-dimensional document vectors onto two dimensions. We followed a procedure specially developed for projecting large document corpora85. Specifically, we calculated uniform affinities on an approximate k-nearest neighbors’ graph with neighbor count k of 10. Then, we optimized for 250 iterations of early exaggeration annealing and then 2000 subsequent optimization iterations. We used default parameters for the principal component analysis initialization and a momentum value of 0.8. We highlighted projects that were NHGRI-led from the Human Genome Project to the end of 2015 and represented in the Core Collection. Additional robustness analysis is described in Supplementary Text (section Robustness Analysis).
Keyword extraction
We assigned relevant computationally generated keywords by using encoded individuals, the detected organizations and matched keywords from the National Library of Medicine’s MeSH headings. The individuals and organizations are direct outputs from Entity recognition and disambiguation. We used MeSH tree branch E’s “Analytical, Diagnostic and Therapeutic Techniques, and Equipment” tree to be the technique keywords and G’s “Phenomena and Processes” to be the phenomena keywords. We applied case-insensitive string matching to their headings and aliases listed in MeSH.
Keyword enrichment analysis
To determine which concepts and ideas may have transitioned from the HGP to subsequent large projects, we calculated the enrichment or depletion of keywords that describe each document. We assigned the document’s project association by its folder curation. We identified entities that remained statistically significant when comparing frequencies of those contained documents associated with the HGP and those contained in documents associated with each of the four large subsequent projects: Large-scale genome sequencing and analysis. Centers (LSAC), International HapMap Project, ENCODE, and model organisms ENCODE. We identified the keywords that appeared frequently enough to statistically detect a meaningful difference in the projects. To do so, we calculated the power of the two-sided Fisher’s Exact Test with a Bonferroni correction by simulation using statmod v.1.5.0 in R. We then used sciPy v1.10.1 Fisher’s Exact implementation with a Bonferroni correction using statsmodels v0.14.0 to find statistical significance in terms that pass the power threshold. We then hierarchically clustered the share of the documents with each of the statistically significant terms using Seaborn v0.12.2 clustermap with a Ward variance minimization algorithm86,87. Statistical significance was defined to be P < 0.05 / n where n is the number of keywords. After multiple hypothesis correction, the \(\alpha\) levels were 1.625e-6 (HGP vs. LSAC), 1.649e-6 (HGP vs. HapMap), 2.425e-6 (HGP vs. modENCODE), and 2.366e-6 (HGP vs. ENCODE).
Terms highlighted in the main figure were manually chosen, without blinding, to illustrate the scope of information provided in the full clustermap provided in the supplementary materials.
GWAS investigation
We identified every publication associated with a MeSH term from tree branch E’s “Analytical, Diagnostic, and Therapeutic Techniques, and Equipment” from 2006 to 2016, as the first GWAS paper36 was published in 2005. We then used NIH’s iCite88 to calculate the publication rank by the number of citations per year. We only considered papers with at least one citation. We then determined, for all papers associated with each MeSH term, the proportion of high-impact papers that we defined to be in the top 5% rank by publication year. For GWAS, we track the MeSH term “D055106:Genome-wide association study”89.
To determine how previously uncharacterized genes were introduced to the biomedical literature, we used PubTator on MEDLINE to detect human genes mentioned in the title or abstract of a publication. Then, we declared the first year a gene is mentioned in the literature to be the year of characterization and look for new genes that get introduced after the draft sequence of a human genome is released in 2000. We restricted our time window from 2006 to 2016 and calculated, on average, the proportion of new genes to all genes mentioned for each publication associated with a technique from MeSH.
We considered the publication having mentioned GWAS if the title, abstract, author-submitted keywords, or MeSH terms contained one of the following phrases in a case-insensitive manner: genome-wide association study, genome wide association study, genome-wide association studies, genome wide association studies, gwas, gwa study, gwa studies, whole genome association study, whole genome association analysis, genome-wide association analysis, genome wide association analysis, genome-wide association analyses, genome wide association analyses, genome wide association scan, genome-wide association scan.
Identification of emails
We used the DiT90 model fine-tuned on the Ryerson Vision Lab Complex Document Information Processing (RVL-CDIP) dataset91. We only inferred the document type scanned for scanned images of the first page of the document from which handwritten marks had been removed. We assumed that the rest of the document pages share the same label as the first page. We verified the appropriate fit of the pre-trained model on the Core Collection by randomly sampling 100 pages and manually annotating their classes (Supplementary Fig. S22).
Reconstructing the email network of NHGRI
Email communication networks can help reveal collaboration dynamics and participant centrality in large teams and organizations92. To create an email communication network, one needs information on the sender, the email recipients, the timestamp, and the header. We fine-tuned a LayoutLMv393 model on 150 documents from the Core Collection that had been categorized as emails, which we manually annotated using LabelStudio v1.8.0. Specifically, we fine-tuned 100 examples for each of the 10 Monte Carlo cross-validated datasets evaluated on sets of 25 examples each and chose the model with the highest overall F-1 score.
Our email recognition model considered both the image and text modalities during model training to take advantage of the layout of email correspondence. Therefore, the LayoutLMv3 tokenizer automatically ran OCR on the image regions to align both modalities. To accommodate this tokenizer, we encoded the individuals after the email recognition model using the same disambiguation scheme above. The top 500 individuals make up 89.2% of all email pairs (Supplementary Fig. S23). We only kept the emails with a valid date (n = 4111).
To ensure correct affiliation, we restricted the network to the top 500 individuals, whose affiliations we assigned by either manually inspecting and assigning them or by taking the domain detected from the email address (if available) and inferring the institution. For individuals who may have changed affiliations (notably between NIH and external academia), we note separately to ensure that they do not get accounted for in the analysis.
We assigned the affiliation as NIH if an email address was from any of the 27 NIH Institutes or Centers. We defined external academia as universities, academic institutions, and private, research-based non-profit institutions. The list of external academic institutions considered in the disambiguated, top 500 people network is included in the Supplementary Data S4.
We represent the email communication network as a directed graph, where arcs point from the sender to the recipients. This graph comprises 500 nodes and 44,975 arcs. We used Gephi v0.10.1 with a ForceAtlas2 algorithm94 to visualize the network with a 0.01 strength parameter, a scaling parameter of 3, and a LinLog transformation of the attraction force95. We used NetworkX v3.3 to calculate in- and out-degrees and prepare the network for Gephi. We assigned projects inside the email network by the folders which the scanned email page originated from. We removed self-looped edges from the network.
More institutions may have participated in NHGRI activities; these only reflect the email network from the Core Collection in the top 500 individuals. In the disambiguated email network of the top 500 individuals, 75.3% of all identified individuals were affiliated with the NIH or external academic institutions. Other individuals in the network are notably affiliated with commercial partners, non-NIH government agencies, associations, and staffing/consulting firms.
Finding communities in emails
To avoid potentially spurious random artifacts that many modularity-based methods face96, we employed hierarchical stochastic block models (hSBM) suited for weighted, directed networks to detect communities using Graph-tool v2.5997. Specifically, to accommodate edge weights, we used the variation of SBM from Piexoto98, which treats edge weights as additional covariates that can guide the final node partition. We used a nested microcanonical degree-corrected SBM99,100, as this approach is less likely to underfit with large networks. For the microcanonical distribution, we considered the four available through Graph-tool: exponential, geometric, Poisson, and binomial. We chose a continuous exponential distribution as the model for the weights as it led to the smallest minimum description length of SBM (Supplementary Table S4). We used simulated annealing with an inverse temperature range of 1 to 10 and 1000 steps to find the optimal parameters for hSBM. Because of its stochastic nature, there may have been more than one possible node partition of hSBM that leads to similar posterior probabilities. Therefore, for every optimization to find the best model fit, we looped this approach 10 times and chose the partition with the smallest minimum description length.
We considered an individual a Kitchen Cabinet member or a HapMap steering committee member by inspecting two archived lists of individuals found inside the Core Collection. We use Floweaver v2.0.0 to create Sankey diagrams of HapMap individuals.
Brokerage roles in email network
Brokerage is the process of how an actor in a network connects otherwise disconnected actors or groups and is a key part of social network analysis. Many argue that brokers play elevated roles in networks45,101,102,103, either from access to new information or control over information flow. Inspired by how brokerage roles may describe social networks from various domains104,105,106, we used the brokerage definitions from Gould and Fernandez45 to describe the NHGRI email network.
Edge weights present a challenge when finding local brokerage measures, as it is unclear which edges should be considered for analysis. A popular approach is dichotomization of the network, where a certain threshold is applied to determine which edges to preserve for brokerage analysis107.
Identifying the appropriate thresholding value is a challenge, as local brokerage roles can be plagued with arbitrary network structures leftover from thresholding107. Therefore, we used an established systematic approach provided by Serrano et al.108 which extracts edges from a weighted network by filtering to retain key ties and retain connectivity of the network. We present in Supplementary Fig. S25 a range of significance levels with the number of nodes and edges retained in the HapMap email network, as well as the corresponding brokerage role descriptions. We only considered nodes with more than 10 qualifying triad edges to ensure that no one brokerage role dominates in proportion.
Modeling council decisions
To find council decisions on sequencing proposals, we identified an internal project tracking sheet inside the Core Collection that determines project proposals at Council meetings from May 2001 to June 2009. We modeled a total of 325 sequencing decisions. These originated from 113 proposals, both internal and external, for the sequencing of 249 organisms. Internal proposals included proposals from NHGRI working groups, such as the “annotating the human genome” working group that identified organisms in critical phylogenetic positions that helped elucidate human evolution. Each organism may be proposed for additional sequencing, either to resubmit a previously unapproved project or to increase the coverage from the draft assembly. We removed all projects with missing Council decisions from the tracking sheet and render deferred decisions as non-approval equivalent. We also removed decisions whose sequencing proposal was not found in the Core Collection. These decisions are represented in Data S5.
To verify that the sequencing target documents inside the Core Collection are representative of the sequencing efforts considered during this time, we compared the organisms documented inside the Core Collection to the entries in NCBI BioProject. We specifically look at NHGRI’s Large-scale sequencing program for organismal sequencing109 umbrellas and the Fungal Genome Initiative110 umbrellas. We only considered NCBI BioProject entries before 2010 to match the council decisions available within the Core Collection.
To determine optimal hyperparameters, we used a Monte Carlo cross-validation of 100 sets with grid search for CatBoost47 and trained on the following 13 properties, grouped in 4 categories:
-
1.
Biological: Genome size (in log10 of megabases), phyla (categorical)
-
2.
Linguistic: argumentativeness (a ratio from 0 to 1), lexical diversity (a ratio from 0 to 1)
-
3.
Project: multi-organism proposal (boolean), number of female proposal writers, team size, time since organism’s first submission (in years), number of support letters, internal or external origin (boolean)
-
4.
Reputation: maximum H-index of proposal authors, in-degree centrality rank of authors in the NHGRI email network, organism community size (in log10 of publication count)
We only considered the main text of the proposal for linguistic features by manually removing the appendices and references at the end of the proposal. Since the absence of a support letter in the Core Collection indicates that no support letter had been attached to the proposal, we manually inspected the appendices of proposals or attachments to extract encoded identifiers of individuals who wrote letters of support for the proposal. Notably, we found that only 30% of the proposals considered had letters of support.
To find individuals who authored the whitepapers or proposals, we either detected names from the first page of the whitepaper, manually compiled authors if the first page did not detect any names or inferred the authors from major steering committees that wrote the proposals, such as the Fungal Genome Initiative steering committee. We determined an individual’s gender by using get_gender from the Python package gender-guesser.detector v0.4.0, which allows “guessing” gender using an individual’s given (first) name. We reassigned “mostly male” and “mostly female” as their respective gender labels and maintained “unknown” for those whose gender could not be conclusively inferred.
We calculated H-index111 directly using OpenAlex112. We considered works published up to a year before the year of consideration by the Council.
We identified sentences with arguments by detecting lexicons associated with argumentation113, specifically those in priority, difficulty, necessity, causation, contrast, and emphasis categories listed by MPQA. We normalized the number of argumentative lexicons by the word count of the proposal, for which we use spaCy’s word tokenizer.
We calculated lexical diversity by measuring the type-token ratio, which is the ratio of unique words to total words, using the Python library LexicalRichness v0.5.1.
The genome size of the organism was directly compiled from the internal project tracking sheet, as the understanding of an estimated genome size may be different from its actual size. Any missing genome sizes were encoded as null to avoid introducing ex-post bias.
We calculated the community size of an organism by the log10 of the number of publications about the organism in the decade before the year of the council decision. We determined whether the publication was about a specific organism by looking for mentions of the taxon of the organism in the title, abstract, author-submitted keywords or MeSH term headings.
We used the shapr package v1.0.0 in R that implements SHAP (SHapley Additive explanation)48,114,115, a feature attribution framework based on the Shapley values116 algorithm. We specifically used the conditional inference tree correction, which accommodates both categorical and numeric variables while avoiding the feature independence assumption that many other implementations of SHAP have48. This is the main limitation behind 13 properties, as the sheer number of permutations of features would become computationally infeasible.
Definition of genomics
We considered a publication to be associated with genomics if the title, abstract, author-submitted keywords, or MeSH terms contained one of the following phrases in a case-insensitive manner. We used the list of terms we assembled in Stoeger et al.23:
chip seq, chip sequencing, chip-chip, chip-seq, chip-sequencing, chromatin immunoprecipitation followed by sequencing, clinicogenomic, clinicogenomics, clip seq, clip seq, clip sequencing, clip sequencing, clip-seq, clip-seq, clip-sequencing, clip-sequencing, epigenome, epigenomes, epigenomic, epigenomics, eqtl, eqtls, exome, expression quantitative trait loci, ewas, genome, genome-scale, genome-wide, genomes, genomic, genomics, glycome, glycomes, glycomics, glycoproteome, glycoproteomes, glycoproteomic, glycoproteomics, gwas, high-throughput nucleotide sequencing, hits seq, hits sequencing, hits-seq, hits-sequencing, in situ proximity ligation, in-situ proximity ligation, interactome, interactomes, interactomic, interactomics, metabolome, metabolomes, metabolomic, metabolomics, metagenome, metagenomes, metagenomics, microarray, microarrays, multi-ome, multi-omes, multi-omic, multi-omics, multiome, multiomes, multiomic, multiomics, next generation sequencing, next generation-sequencing, next-generation sequencing, next-generation-sequencing, ngs, nutrigenome, nutrigenomes, nutrigenomic, nutrigenomics, oligonucleotide array sequence analysis, omics, onco-genome, onco-genomes, onco-genomics, onco-genomics, onco-proteogenome, onco-proteogenomes, onco-proteogenomic, onco-proteogenomics, oncogenome, oncogenomes, oncogenomics, oncogenomics, oncoproteogenome, oncoproteogenomes, oncoproteogenomic, oncoproteogenomics, par-clip, pharmacogenome, pharmacogenomes, pharmacogenomic, pharmacogenomics, phenome, phenomes, phenomic, phenomics, phosphoproteome, phosphoproteomes, phosphoproteomic, phosphorpoteomics, protein array, protein array analysis, protein interaction map, protein interaction mapping, protein interaction maps, protein interaction network, protein interaction networks, protein–protein interaction map, protein–protein interaction mapping, protein–protein interaction maps, protein–protein interaction network, protein–protein interaction networks, proteogenome, proteogenomes, proteogenomic, proteogenomics, proteome, proteomes, proteomic, proteomics, radiogenome, radiogenomes, radiogenomic, radiogenomics, rna seq, rna sequencing, rna-seq, rna-sequencing, rnaseq, toxicogenome, toxicogenomes, toxicogenomic, toxicogenomics, transcriptome, transcriptomes, transcriptomic, transcriptomics, wes, wgs, whole-exome, whole-genome.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The Core Collection and meta-information extracted during this study are available under an Information Transfer Agreement (ITA) for research purposes. Because the collection may contain personally identifiable information (PII), the dataset cannot be made publicly available. Access can be obtained by contacting NHGRI’s History of Genomics Program by emailing NHGRIHistory@nih.gov. The ITA is a two-party agreement between the National Human Genome Research Institute (NHGRI) and the researcher’s sponsoring institution, typically where that researcher is employed and conducting the relevant research. The ITA outlines roles and responsibilities for each institution and sets out specific conditions for access. It underscores, most importantly, that the sponsoring research organization is receiving data and/or metadata that may contain personally identifiable information (PII). As part of the agreement, the researcher and sponsoring organization requesting the data must clearly identify their purpose for using this dataset (e.g., to ensure reproducibility) and present a summary of their core research objectives with the data. If the researcher is utilizing AI/ML tools in any way or manner, those must be specified explicitly as part of the application for data access. The research is only allowed to use the data for the purposes outlined in the ITA. Data access requests are reviewed by NHGRI to confirm that the stated research use is consistent with the terms of the ITA and that the requesting researcher acknowledges receipt of materials that may contain PII. The expected time frame for the ratification of the ITA is three weeks. Data access is granted for three years and may be extended. The researcher and/or the researcher’s sponsoring institution may also terminate the agreement at any time. Amendments or ongoing concerns over the agreement or uses of the data can be requested by the research organization or by the NHGRI. Copies of the ITA can be requested at the above address with no obligation. For all bibliometric analyses, PubMed annual baseline files (containing assigned MeSH terms) were downloaded on 2023-01-14 https://ftp.ncbi.nlm.nih.gov/pubmed/baseline/ and parsed through PubMed Parser 0.3117. MeSH definitions were downloaded from https://nlmpubs.nlm.nih.gov/projects/mesh/ in ASCII Format on 2020-12-08. NCBI taxonomy identifiers were downloaded from https://ftp.ncbi.nlm.nih.gov/pub/taxonomy/taxdmp.zip on 2021-11-08. NCBI BioProject was downloaded from https://ftp.ncbi.nlm.nih.gov/bioproject on 2021-04-01. iCite version 32 was downloaded from https://nih.figshare.com/collections/iCite_Database_Snapshots_NIH_Open_Citation_Collection_/4586573/32 NCBI Gene Info was downloaded from https://ftp.ncbi.nlm.nih.gov/gene/DATA/GENE_INFO/ on 2022-07-12. PubTator was downloaded from https://ftp.ncbi.nlm.nih.gov/pub/lu/PubTatorCentral/ on 2021-11-09. We excluded records where a gene name had been mapped to multiple NCBI Entrez Gene Identifiers by PubTator. OpenAlex, used to calculate H-indices directly, was downloaded on 2024-03-26. The datasets from NCBI are information that is created by or for the US government on this site and is within the public domain. OpenAlex is CC0. We refer to the individual data sources for clarification on copyright.
Code availability
Code of this study is available under https://github.com/born-physical-studied-digitally/nhgri_archive (with persistent118 https://doi.org/10.5281/zenodo.17204492). For adoption on custom infrastructure or archives, we recommend usage of the latest versions of our software, which is actively maintained and expanded by Born Physical / Studied Digitally (https://studieddigitally.org), an NSF Consortium on Cyberinfrastructure for Sustained Scientific Innovation.
References
Braun, D. Who governs intermediary agencies? principal-agent relations in research policy-making. J. Public Policy 13, 135–162 (1993).
Guston, D. H. Principal-agent theory and the structure of science policy. Sci. Public Policy 23, 229–240 (1996).
Van der Meulen, B. Science policies as principal–agent games: Institutionalization and path dependency in the relation between government and science. Res. Policy 27, 397–414 (1998).
Braun, D. The role of funding agencies in the cognitive development of science. Res. Policy 27, 807–821 (1998).
Guston, D. H. Stabilizing the boundary between US politics and science: The role of the Office of Technology Transfer as a boundary organization. Soc. Stud. Sci. 29, 87–111 (1999).
Aviles, N. B. An Ungovernable Foe: Science and Policy Innovation in the U.S. National Cancer Institute. (Columbia University Press, 2023).
Dunham, I. et al. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Towns, J. et al. XSEDE: Accelerating scientific discovery. Comput. Sci. Eng. 16, 62–74 (2014).
Bush, V. Science the Endless Frontier. (United States Government Printing Office, 1945).
Sampat, B. N. Mission-oriented biomedical research at the NIH. Res. Policy 41, 1729–1741 (2012).
Yerger, R. H. in Moonshots and the New Industrial Policy: Questioning the Mission Economy (eds Magnus H, Christian S, & Mikael S.) 109–123 (Springer Nature Switzerland, 2024).
Scheffler, R. W. Managing the future: the Special Virus Leukemia Program and the acceleration of biomedical research. Stud. Hist. Philos. Biol. Biomed. Sci. 48, 231–249 (2014). Pt B.
Aviles, N. B. Situated practice and the emergence of ethical research: HPV vaccine development and organizational cultures of translation at the National Cancer Institute. Sci. Technol. Hum. Val. 43, 810–833 (2018).
Scheffler, R. W. & Aviles, N. B. State planning, cancer vaccine infrastructure, and the origins of the oncogene theory. Soc. Stud. Sci. 52, 174–198 (2022).
Jacob, B. A. & Lefgren, L. The impact of research grant funding on scientific productivity. J. Public Econ. 95, 1168–1177 (2011).
Packalen, M. & Bhattacharya, J. NIH funding and the pursuit of edge science. Proc. Natl. Acad. Sci. USA 117, 12011–12016 (2020).
Fortin, J.-M. & Currie, D. J. Big science vs. little science: how scientific impact scales with funding. PloS ONE 8, e65263 (2013).
Liu, W. Accuracy of funding information in Scopus: a comparative case study. Scientometrics 124, 803–811 (2020).
Shin, H., Kim, K. & Kogler, D. F. Scientific collaboration, research funding, and novelty in scientific knowledge. PloS ONE 17, https://doi.org/10.1371/journal.pone.0271678 (2022).
Fortunato, S. et al. Science of science. Science 359, eaao0185 (2018).
Kuska, B. B. eer Bethesda, and biology: how “Genomics” came into being. JNCI: J. Natl. Cancer Inst. 90, 93–93 (1998).
Hieter, P. & Boguski, M. Functional genomics: it’s all how you read it. Science 278, 601–602 (1997).
Stoeger, T. & Nunes Amaral, L. A. The characteristics of early-stage research into human genes are substantially different from subsequent research. PLoS Biol. 20, e3001520 (2022).
Venter, J. C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Green, E. D. et al. Strategic vision for improving human health at The Forefront of Genomics. Nature 586, 683–692 (2020).
Langfelder, E. J. & Juengst, E. T. Ethical, legal, and social implications (Elsi) program - National Center for Human Genome Research, National Institutes of Health. Polit. Life Sci. 12, 273–275 (1993).
Collins, F. S., Green, E. D., Guttmacher, A. E., Guyer, M. S. & Institute, N. H. G. R. A vision for the future of genomics research. Nature 422, 835–847 (2003).
De Chadarevian, S. Heredity under the Microscope: Chromosomes and the Study of the Human Genome. (University of Chicago Press, 2020).
Tabery, J. Tyranny of the Gene: Personalized Medicine and its Threat to Public Health. (Knopf, 2023).
García-Sancho, M. & Lowe, J. A History of Genomics Across Species, Communities and Projects. (Springer Nature, 2023).
Hosseini, M. et al. Ethical considerations in utilizing artificial intelligence for analyzing the NHGRI’s History of Genomics and Human Genome Project archives. J. EScience Librariansh. 13, https://doi.org/10.7191/jeslib.811 (2024).
Wu, L., Wang, D. & Evans, J. A. Large teams develop and small teams disrupt science and technology. Nature 566, 378–382 (2019).
Gibbs, R. A. et al. The International HapMap Project. Nature 426, 789–796 (2003).
Perspectives on the Human Genome Project and Genomics. (University of Minnesota Press, 2026).
DeWan, A. et al. Promoter polymorphism in wet age-related macular degeneration. Science 314, 989–992 (2006).
Risch, N. & Merikangas, K. The future of genetic studies of complex human diseases. Science 273, 1516–1517 (1996).
Collins, F. S. et al. New goals for the U.S. human genome project: 1998-2003. Science 282, 682–689 (1998).
Klein, R. J. et al. Complement factor H polymorphism in age-related macular degeneration. Science 308, 385–389 (2005).
Wellcome Trust Case Control, C Genome-wide association study of 14,000 cases of seven common diseases and 3000 shared controls. Nature 447, 661–678 (2007).
Abdellaoui, A., Yengo, L., Verweij, K. J. H. & Visscher, P. M. 15 years of GWAS discovery: Realizing the promise. Am. J. Hum. Genet. 110, 179–194 (2023).
Loos, R. J. F. 15 years of genome-wide association studies and no signs of slowing down. Nat. Commun. 11, 5900 (2020).
Malmgren, R. D., Stouffer, D. B., Motter, A. E. & Amaral, L. A. N. A Poissonian explanation for heavy tails in e-mail communication. Proc. Natl. Acad. Sci. USA 105, 18153–18158 (2008).
Barabási, A.-L. The origin of bursts and heavy tails in human dynamics. Nature 435, 207–211 (2005).
Gould, R. V. & Fernandez, R. M. Structures of Mediation: A Formal Approach to Brokerage in Transaction Networks. Sociol. Methodol. 19, 89–126 (1989).
Hotaling, S., Kelley, J. L. & Frandsen, P. B. Toward a genome sequence for every animal: Where are we now. Proc. Natl. Acad. Sci. USA 118, e2109019118 (2021).
Dorogush, A.V., Ershov, V. & Gulin, A. 2018.CatBoost: gradient boosting with categorical features support. Preprint at https://doi.org/10.48550/arXiv.1810.11363 (2018).
Aas, K., Jullum, M. & Løland, A. Explaining individual predictions when features are dependent: More accurate approximations to Shapley values. Artif. Intell. 298, 103502 (2021).
Xu, F., Wu, L. & Evans, J. Flat teams drive scientific innovation. Proc. Natl. Acad. Sci. USA 119, e2200927119 (2022).
Ellegren, H. Genome sequencing and population genomics in non-model organisms. Trends Ecol. Evol. 29, 51–63 (2014).
Aitman, T. J. et al. The future of model organisms in human disease research. Nat. Rev. Genet. 12, 575–582 (2011).
Müller, B. & Grossniklaus, U. Model organisms — A historical perspective. J. Proteom. 73, 2054–2063 (2010).
Ankeny, R. A. Model organisms as models: Understanding the ‘lingua franca’of the human genome project. Philos. Sci. 68, S251–S261 (2001).
Ratti, E. The end of ‘small biology’? Some thoughts about biomedicine and big science. Big Data Soc. 3, 1–6 (2016).
Utz, Z. “The Human Genome Project is simply a bad idea”, https://www.genome.gov/virtual-exhibits/human-genome-project-is-simply-a-bad-idea (2024).
Charter for the National Advisory Council for Human Genome Research, https://www.genome.gov/sites/default/files/media/files/2024-02/NACHGR_approved_charter_2022_2024.pdf (2024).
Gieryn, T. F. Boundary-work and the demarcation of science from non-science - strains and interests in professional ideologies of scientists. Am. Socio. Rev. 48, 781–795 (1983).
Guston, D. H. Boundary organizations in environmental policy and science: an introduction. Sci. Technol. Hum. Values 26, 399–408 (2001).
Epstein, S. Impure Science: AIDS, Activism, and the Politics of Knowledge. (Univ of California Press, 1996).
Postgenomics: Perspectives on Biology after the Genome. (Duke University Press, 2015).
Simon, H. A. Administrative Behavior. (Simon and Schuster, 2013).
Reich, D. E. & Lander, E. S. On the allelic spectrum of human disease. Trends Genet. 17, 502–510 (2001).
Lander, E. S. The new genomics: global views of biology. Science 274, 536–539 (1996).
Neel, J. in The Genetics of Diabetes Mellitus 1-11 (Springer, 1976).
Studer, K. E. & Chubin, D. E. The Cancer Mission: Social Contexts of Biomedical Research. (Sage, 1980).
Scheffler, R. W. A Contagious Cause: The American Hunt for Cancer Viruses and the Rise of Molecular Medicine. (University of Chicago Press, 2019).
Dyke, S. O. M. & Hubbard, T. J. P. Developing and implementing an institute-wide data sharing policy. Genome Med. 3, https://doi.org/10.1186/gm276 (2011).
Ronneberger, O., Fischer, P. & Brox, T. Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015 234-241 (Springer International Publishing, Cham, 2015).
Lewis, D. et al. in Proceedings of the 29th annual international ACM SIGIR conference on Research and development in information retrieval. 665–666 (Association for Computing Machinery).
Sarkar, M., Aggarwal, M., Jain, A., Gupta, H. & Krishnamurthy, B. European Conference on Computer Vision (Springer International Publishing).
Aggarwal, M., Sarkar, M., Gupta, H. & Krishnamurthy, B. in Proceedings of the IEEE/CVF Winter Conference on Applications of Computer Vision. 2075-2084. https://link.springer.com/chapter/10.1007/978-3-030-58604-1_39
Thoma, M. The HASYv2 dataset. Preprint at https://doi.org/10.48550/arXiv.1701.08380 (2017).
Google. The quick, draw! dataset, https://github.com/googlecreativelab/quickdraw-dataset (2017).
Communication, S. R. C. O. S. O. (UC Irvine Machine Learning Repository, 2008).
Marti, U. V. & Bunke, H. The IAM-database: an English sentence database for offline handwriting recognition. Int. J. Doc. Anal. Recognit. 5, 39–46 (2002).
Dice, L. R. Measures of the amount of ecologic association between species. Ecology 26, 297–302 (1945).
Bengio, Y., Louradour, J., Collobert, R. & Weston, J. in Proceedings of the 26th Annual International Conference on Machine Learning 41–48 (Association for Computing Machinery, Montreal, Quebec, Canada, 2009).
Smith, R. in Ninth International Conference on Document Analysis and Recognition (ICDAR 2007). 629-633.
Liu, Y. et al. Roberta: A robustly optimized bert pretraining approach. Preprint at https://doi.org/10.48550/arXiv.1907.11692 (2019).
Reijnhoudt, L., Costas, R., Noyons, E., Börner, K. & Scharnhorst, A. ‘Seed + expand’: a general methodology for detecting publication oeuvres of individual researchers. Scientometrics 101, 1403–1417 (2014).
Le, Q. & Mikolov, T. in International Conference on Machine Learning. 1188-1196 (PMLR).
Mikolov, T., Chen, K., Corrado, G. & Dean, J. Efficient estimation of word representations in vector space. Preprint at https://doi.org/10.48550/arXiv.1301.3781 (2013).
Poličar, P. G., Stražar, M. & Zupan, B. openTSNE: a modular Python library for t-SNE dimensionality reduction and embedding.https://doi.org/10.1101/731877 (2019).
Van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
González-Márquez, R., Berens, P. & Kobak, D. ICLR Workshop on Geometrical and Topological Representation Learning. https://www.springer.com/journal/44007/updates/19962244 (2022).
Müllner, D. fastcluster: Fast hierarchical, agglomerative clustering routines for R and Python. J. Stat. Softw. 53, 1–18 (2013).
Ward, J. H. Jr Hierarchical grouping to optimize an objective function. J. Am. Stat. Assoc. 58, 236–244 (1963).
ICite, H., Ian, B. & Santangelo, G. iCite Database snapshots (NIH open citation collection). https://doi.org/10.35092/yhjc.c.4586573 (2022).
D055106
Li, J. et al. in Proceeding id="MPS_d1e2376">of the 30th ACM International Conference on Multimedia 3530–3539 (Association for Computing Machinery, Lisbon, Portugal, 2022).
Harley, A. W., Ufkes, A. & Derpanis, K. G. in 2015 13th International Conference on Document Analysis and Recognition (ICDAR). 991-995 (IEEE).
Guimerà, R., Danon, L., Díaz-Guilera, A., Giralt, F. & Arenas, A. Self-similar community structure in a network of human interactions. Phys. Rev. E 68, 065103 (2003).
Huang, Y., Lv, T., Cui, L., Lu, Y. & Wei, F. in Proceedings of the 30th ACM International Conference on Multimedia 4083–4091 (Association for Computing Machinery, Lisboa, Portugal, 2022).
Jacomy, M., Venturini, T., Heymann, S. & Bastian, M. ForceAtlas2, a continuous graph layout algorithm for handy network visualization designed for the Gephi software. PloS ONE 9, e98679 (2014).
Noack, A. Unified quality measures for clusterings, layouts, and orderings of graphs, and their application as software design criteria, BTU Cottbus-Senftenberg, (2007).
Guimerà, R., Sales-Pardo, M. & Amaral, L. A. N. Modularity from fluctuations in random graphs and complex networks. Phys. Rev. E 70, 025101 (2004).
Peixoto, T. P. The graph-tool python library. Figshare https://doi.org/10.6084/m9.figshare.1164194 (2014).
Peixoto, T. P. Nonparametric weighted stochastic block models. Phys. Rev. E 97, 012306 (2018).
Peixoto, T. P. Nonparametric Bayesian inference of the microcanonical stochastic block model. Phys. Rev. E 95, 012317 (2017).
Peixoto, T. P. Hierarchical block structures and high-resolution model selection in large networks. Phys. Rev. X 4, 011047 (2014).
Stovel, K. & Shaw, L. Brokerage. Annu. Rev. Sociol. 38, 139–158 (2012).
Burt, R. S. Brokerage and Closure: An Introduction to Social Capital. (OUP Oxford, 2007).
Burt, R. S. Structural Holes: The Social Structure of Competition. (Harvard University Press, 1992).
Kirkels, Y. & Duysters, G. Brokerage in SME networks. Res. Policy 39, 375–385 (2010).
Gray, B. Enhancing transdisciplinary research through collaborative leadership. Am. J. Prevent. Med. 35, S124–S132 (2008).
Täube, V. G. Measuring the social capital of brokerage roles. Connections 26, 29–52 (2004).
Neal, Z. The backbone of bipartite projections: Inferring relationships from co-authorship, co-sponsorship, co-attendance and other co-behaviors. Soc. Netw. 39, 84–97 (2014).
Serrano, M. Á, Boguñá, M. & Vespignani, A. Extracting the multiscale backbone of complex weighted networks. Proc. Natl. Acad. Sci. USA 106, 6483–6488 (2009).
PRJNA167910.
PRJNA176382.
Hirsch, J. E. An index to quantify an individual’s scientific research output. Proc. Natl. Acad. Sci. USA 102, 16569–16572 (2005).
Priem, J.P., Piwowar, H. & Orr, R. OpenAlex: A fully-open index of scholarly works, authors, venues, institutions, and concepts. Preprint https://doi.org/10.48550/arXiv.2205.01833 (2022).
Somasundaran, S., Ruppenhofer, J. & Wiebe, J. in Proceedings of the 8th SIGdial Workshop on Discourse and Dialogue. (2007).
Lundberg, S. M. & Lee, S.-I. A unified approach to interpreting model predictions. In NIPS'17: Proceedings of the 31st International Conference on Neural Information Processing Systems (2017).
JullumM.OlsenL.HB.LachmannJ.RedelmeierA. 2025.shapr: Explaining machine learning models with conditional shapley values in R and Python. Preprint at https://doi.org/10.48550/arXiv.2504.01842 (2025).
Štrumbelj, E. & Kononenko, I. Explaining prediction models and individual predictions with feature contributions. Knowl. Inf. Syst. 41, 647–665 (2014).
Achakulvisut, T., Acuna, D. E. & Kording, K. Pubmed parser: a python parser for pubmed open-access XML subset and MEDLINE XML dataset XML dataset. J. Open Source Softw. 5, 1979 (2020).
Hong, S., Tejedor, H. & Stoeger, T. born-Phys.-Stud.-digitally/nhgri_archive https://doi.org/10.5281/zenodo.17204492 (2025).
Acknowledgements
KH is supported by NIH UM1TR005121, U24LM013751 and 1OT2DB000013-01. LANA is supported by NSF 2410335. MH is supported by UM1TR005121. T.S. is supported by R00AG068544 and NSF 2410335. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. This research (L.A.N.A., T.S.) was supported in part by grants from the NSF (DMS-2235451) and Simons Foundation (MP-TMPS-00005320) to the NSF-Simons National Institute for Theory and Mathematics in Biology (NITMB). Z.U., K.A.W., S.A.B., and C.R.D. are paid employees of NHGRI. S.H. and T.S. are unpaid Special Volunteers with NHGRI’s History of Genomics Program and the Office of Genomic Data Science. The authors declare to have used ChatGPT on the text of the manuscript to ensure the correct grammar of some sentences (i.e., “Ensure correct grammar of: [sentence(s)]”). We deeply thank Dr. Gerard Bouffard for having triggered this project by introducing the teams of Northwestern and NIH-NHGRI to each other after spotting mutual interests and complementary expertise. We also thank NVIDIA for providing in-equipment grant to support this research without the need for cloud computing. We thank Dr. Elke Jordan, Dr. Valentin Danchev, and the Office of Communications at NHGRI for their feedback, support, and consultation for this project.
Author information
Authors and Affiliations
Contributions
L.A.N.A., C.R.D., and T.S. conceptualized the study. Z.U., K.W., and C.R.D. curated the data. K.H., K.W., S.A.B., L.A.N.A., C.R.D., and T.S. acquired necessary funding. S.S.H., L.A.N.A., C.R.D., and T.S. led the investigation. S.S.H., M.H., C.Z., L.A.N.A., and T.S. created the methodology. K.W., S.A.B., L.A.N.A., C.R.D., and T.S. administered the project. S.S.H., C.Z., H.T.N., L.A.N.A., and T.S. created the software and code for the analysis. S.S.H. and T.S. created the visualizations in the project. S.S.H., L.A.N.A., C.R.D., and T.S. wrote the original draft of the publication. S.S.H., M.H., C.Z., K.H., L.A.N.A., C.R.D., and T.S. revised and edited the draft.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Alexander Gates, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hong, S.S., Utz, Z., Hosseini, M. et al. A digital archive reveals how a funding agency cooperated with academics to support the nascent field of genomics. Nat Commun 17, 3621 (2026). https://doi.org/10.1038/s41467-026-71700-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-71700-9