Abstract
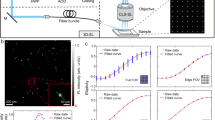
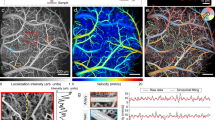
Quantitative, volumetric imaging of cerebrovascular networks and microcirculation is essential for understanding brain function. However, rapid mesoscopic 3D imaging remains challenging because of fundamental trade-offs between spatiotemporal resolution, field of view, and sensitivity to functional parameters. Here we present a mesoscopic fluorescence imaging platform featuring a double-helix phase mask for real-time, depth-resolved measurements through the intact mouse skull. The compact phase-mask design is compatible with both laser-scanning and widefield microscopy. Using multifocal laser scanning, we demonstrate real-time volumetric in vivo imaging while discriminating calvarial from cerebral vasculature across 6.6×6.6×0.8 mm3 volume. Beyond high-resolution structural imaging, perfusion time-to-peak values are extracted from the laser-scanning configuration while accurate flow velocity/direction information is provided via widefield tracking of fluorescently labeled cells. We demonstrate the platform’s capabilities by analyzing brain-layer-specific perfusion dynamics and vascular topology in glioma-bearing mouse brains, offering unprecedented views for probing cerebrovascular alterations in both physiological and pathological contexts.
Similar content being viewed by others
Data availability
The main data supporting the finding of this study are available within the main text or Supplementary Information. Representative raw imaging datasets supporting the findings of this study have been deposited in Zenodo and are publicly available at https://doi.org/10.5281/zenodo.18876905. Source data are provided with this paper.
Code availability
Localization of fluorescence emitters was performed with the open-source TrackNTrace toolbox63. RBC tracking was performed using the SimpleTracker algorithm36. ADMM reconstruction was performed with the open-source DiffuserCam code69. Pseudocode outlining the main DH-PSF image reconstruction workflow is included in the Supplementary Information (Supplementary Algorithm 1). Custom MATLAB codes for data analysis which are available for research purposes from the corresponding author upon request.
References
Zlokovic, B. V. The blood-brain barrier in health and chronic neurodegenerative disorders. Neuron 57, 178–201 (2008).
Zlokovic, B. V. Neurovascular pathways to neurodegeneration in Alzheimer’s disease and other disorders. Nat. Rev. Neurosci. 12, 723–738 (2011).
Blinder, P. et al. The cortical angiome: an interconnected vascular network with noncolumnar patterns of blood flow. Nat. Neurosci. 16, 889–897 (2013).
Montgomery, M. K. et al. Glioma-induced alterations in neuronal activity and neurovascular coupling during disease progression. Cell Rep. 31, 107500 (2020).
Huang, C. et al. Construction of a whole-brain panorama for glioma vasculature reveals tumor heterogeneity and blood-brain barrier disruption. Sci. Adv. 11, eadw8330 (2025).
Liu, J. et al. Vascular remodeling after ischemic stroke: mechanisms and therapeutic potentials. Prog. Neurobiol. 115, 138–156 (2014).
Verret, L. et al. Inhibitory interneuron deficit links altered network activity and cognitive dysfunction in Alzheimer model. Cell 149, 708–721 (2012).
Zhou, Q. et al. Cortex-wide transcranial localization microscopy with fluorescently labeled red blood cells. Nat. Commun. 15, 3526 (2024).
Shen, Z., Lu, Z., Chhatbar, P. Y., O’Herron, P. & Kara, P. An artery-specific fluorescent dye for studying neurovascular coupling. Nat. Methods 9, 273–276 (2012).
Walek, K. W. et al. Near-lifespan longitudinal tracking of brain microvascular morphology, topology, and flow in male mice. Nat. Commun. 14, 2982 (2023).
Sun, J., Kuschmierz, R., Katz, O., Koukourakis, N. & Czarske, J. W. Lensless fiber endomicroscopy in biomedicine. PhotoniX 5,18 (2024).
Helmchen, F. & Denk, W. Deep tissue two-photon microscopy. Nat. Methods 2, 932–940 (2005).
Horton, N. G. et al. In vivo three-photon microscopy of subcortical structures within an intact mouse brain. Nat. Photonics 7, 205–209 (2013).
Chen, Z., Zhou, Q., Robin, J. & Razansky, D. Widefield fluorescence localization microscopy for transcranial imaging of cortical perfusion with capillary resolution. Opt. Lett. 45, 3470–3473 (2020).
Gluck, C. et al. Pia-FLOW: Deciphering hemodynamic maps of the pial vascular connectome and its response to arterial occlusion. Proc. Natl. Acad. Sci. USA. 121, e2402624121 (2024).
Errico, C. et al. Transcranial functional ultrasound imaging of the brain using microbubble-enhanced ultrasensitive Doppler. Neuroimage 124, 752–761 (2016).
Sauvage, J. et al. 4D functional imaging of the rat brain using a large aperture row-column array. IEEE Trans. Med. Imaging 39, 1884–1893 (2020).
Jensen, J. A. et al. Anatomic and functional imaging using row–column arrays. IEEE Trans. Ultrason. Ferroelectr. Freq. Control 69, 2722–2738 (2022).
Heiles, B. et al. Nonlinear sound-sheet microscopy: Imaging opaque organs at the capillary and cellular scale. Science 388, eads1325 (2025).
Deán-Ben, X. L. et al. Deep optoacoustic localization microangiography of ischemic stroke in mice. Nat. Commun. 14, 3584 (2023).
Pavani, S. R. et al. Three-dimensional, single-molecule fluorescence imaging beyond the diffraction limit by using a double-helix point spread function. Proc. Natl. Acad. Sci. USA. 106, 2995–2999 (2009).
Xu, K., Babcock, H. P. & Zhuang, X. Dual-objective STORM reveals three-dimensional filament organization in the actin cytoskeleton. Nat. Methods 9, 185–188 (2012).
Gahlmann, A. et al. Quantitative multicolor subdiffraction imaging of bacterial protein ultrastructures in three dimensions. Nano Lett. 13, 987–993 (2013).
Jia, S., Vaughan, J. C. & Zhuang, X. Isotropic 3D super-resolution imaging with a self-bending point spread function. Nat. Photonics 8, 302–306 (2014).
Thompson, M. A., Lew, M. D., Badieirostami, M. & Moerner, W. E. Localizing and tracking single nanoscale emitters in three dimensions with high spatiotemporal resolution using a double-helix point spread function. Nano Lett. 10, 211–218 (2010).
Li, H. et al. Particles 3D tracking with large axial depth by using the 2π-DH-PSF. Opt. Lett. 46, 5088–5091 (2021).
Pavani, S. R. P. & Piestun, R. High-efficiency rotating point spread functions. Opt. Express 16, 3484–3489 (2008).
Chen, Z. et al. High-speed large-field multifocal illumination fluorescence microscopy. Laser Photonics Rev. 14, 1900070 (2019).
Zhou, Q. et al. Three-dimensional wide-field fluorescence microscopy for transcranial mapping of cortical microcirculation. Nat. Commun. 13, 7969 (2022).
York, A. G. et al. Resolution doubling in live, multicellular organisms via multifocal structured illumination microscopy. Nat. Methods 9, 749–754 (2012).
Ingaramo, M. et al. Two-photon excitation improves multifocal structured illumination microscopy in thick scattering tissue. Proc. Natl. Acad. Sci. 111, 5254–5259 (2014).
Shechtman, Y., Weiss, L. E., Backer, A. S., Sahl, S. J. & Moerner, W. E. Precise three-dimensional scan-free multiple-particle tracking over large axial ranges with tetrapod point spread functions. Nano Lett. 15, 4194–4199 (2015).
Siemons, M. E., Kapitein, L. C. & Stallinga, S. Axial accuracy in localization microscopy with 3D point spread function engineering. Opt. Express 30, 28290–28300 (2022).
Herisson, F. et al. Direct vascular channels connect skull bone marrow and the brain surface enabling myeloid cell migration. Nat. Neurosci. 21, 1209–1217 (2018).
Cugurra, A. et al. Skull and vertebral bone marrow are myeloid cell reservoirs for the meninges and CNS parenchyma. Science 373, eabf7844 (2021).
Tinevez, J.-Y. simpletracker. A simple object tracking plug-in for ImageJ. https://github.com/tinevez/simpletracker (2019).
Glandorf, L. et al. Bessel beam optical coherence microscopy enables multiscale assessment of cerebrovascular network morphology and function. Light. Sci. Appl. 13, 307 (2024).
Glandorf, L. et al. In vivo network-level cerebrovascular mapping reveals the impact of flow topology on capillary stalls after stroke. bioRxiv. 2025.2007.2028.667165 (2025).
Leitgeb, R. A., Werkmeister, R. M., Blatter, C. & Schmetterer, L. Doppler optical coherence tomography. Prog. Retin. Eye Res. 41, 26–43 (2014).
Bouvy, W. H. et al. Assessment of blood flow velocity and pulsatility in cerebral perforating arteries with 7-T quantitative flow MRI. NMR Biomed. 29, 1295–1304 (2016).
Tang, J., Kilic, K., Szabo, T. L. & Boas, D. A. Improved color doppler for cerebral blood flow axial velocity imaging. IEEE Trans. Med. Imaging 40, 758–764 (2021).
Miller, D. R. et al. Multi-modal microvascular cerebral blood flow velocity mapping with 14T single-vessel MRI and optical microscopy in the mouse brain. bioRxiv 2023, 566001 (2023).
Hardee, M. E. & Zagzag, D. Mechanisms of glioma-associated neovascularization. Am. J. Pathol. 181, 1126–1141 (2012).
Louis, D. N. et al. The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol. 131, 803–820 (2016).
Gratton, J.-P. et al. Selective inhibition of tumor microvascular permeability by cavtratin blocks tumor progression in mice. Cancer Cell 4, 31–39 (2003).
Lugano, R., Ramachandran, M. & Dimberg, A. Tumor angiogenesis: causes, consequences, challenges and opportunities. Cell. Mol. Life Sci. 77, 1745–1770 (2020).
Chen, Z., Han, F., Du, Y., Shi, H. & Zhou, W. Hypoxic microenvironment in cancer: molecular mechanisms and therapeutic interventions. Signal Transduct. Target. Ther. 8, 70 (2023).
Secomb, T. W., Dewhirst, M. W. & Pries, A. R. Structural adaptation of normal and tumour vascular networks. Basic Clin. Pharmacol. Toxicol. 110, 63–69 (2012).
Kane, K., Edwards, D. & Chen, J. The influence of endothelial metabolic reprogramming on the tumor microenvironment. Oncogene 44, 51–63 (2025).
Fukumura, D. & Jain, R. K. Tumor microvasculature and microenvironment: targets for anti-angiogenesis and normalization. Microvasc.Res. 74, 72–84 (2007).
Fukumura, D. A. I., Duda, D. G., Munn, L. L. & Jain, R. K. Tumor microvasculature and microenvironment: novel insights through intravital imaging in pre-clinical models. Microcirculation 17, 206–225 (2010).
Nagy, J. A., Chang, S. H., Dvorak, A. M. & Dvorak, H. F. Why are tumour blood vessels abnormal and why is it important to know? Br. J. Cancer 100, 865–869 (2009).
Sofroniew, N. J., Flickinger, D., King, J. & Svoboda, K. A large field of view two-photon mesoscope with subcellular resolution for in vivo imaging. eLife 5, e14472 (2016).
Chen, C. et al. Study of neurovascular coupling by using mesoscopic and microscopic imaging. iScience 24, 103176 (2021).
Dencks, S. & Schmitz, G. Ultrasound localization microscopy. Z. Med. Phys. 33, 292–308 (2023).
Bureau, F. et al. Ultrasound matrix imaging for 3D transcranial in vivo localization microscopy. Sci. Adv. 11, eadt9778 (2025).
Chen, W., You, J., Gu, X., Du, C. & Pan, Y. High-speed swept source optical coherence Doppler tomography for deep brain microvascular imaging. Sci. Rep. 6, 38786 (2016).
Merkle, C. W., Zhu, J., Bernucci, M. T. & Srinivasan, V. J. Dynamic contrast optical coherence tomography reveals laminar microvascular hemodynamics in the mouse neocortex in vivo. NeuroImage 202, 116067 (2019).
Veldman, M. B. et al. Brainwide genetic sparse cell labeling to illuminate the morphology of neurons and glia with cre-dependent MORF mice. Neuron 108, 111–127.e116 (2020).
Chen, X., Wang, X., Huang, F. & Ma, D. Multicolor single-molecule localization microscopy: review and prospect. PhotoniX 5, 29 (2024).
Chen, X. et al. Self-supervised denoising for multimodal structured illumination microscopy enables long-term super-resolution live-cell imaging. PhotoniX 5, 4 (2024).
Shen, B. et al. Surmounting photon limits and motion artifacts for biological dynamics imaging via dual-perspective self-supervised learning. PhotoniX 5, 1 (2024).
Stein, S. C. & Thiart, J. TrackNTrace: a simple and extendable open-source framework for developing single-molecule localization and tracking algorithms. Sci. Rep. 6, 37947 (2016).
Boyd, S., Parikh, N., Chu, E., Peleato, B. & Eckstein, J. Distributed optimization and statistical learning via the alternating direction method of multipliers. Found. Trends® Mach. Learn. 3, 1–122 (2011).
Drees, D., Scherzinger, A., Hägerling, R., Kiefer, F. & Jiang, X. Scalable robust graph and feature extraction for arbitrary vessel networks in large volumetric datasets. BMC Bioinform. 22, 346 (2021).
Zhang, Z. et al. Imaging volumetric dynamics at high speed in mouse and zebrafish brain with confocal light field microscopy. Nat. Biotechnol. 39, 74–83 (2021).
Huisjes, R. et al. Squeezing for life - properties of red blood cell deformability. Front. Physiol. 9, 656 (2018).
Leal, J. K. F., Adjobo-Hermans, M. J. W. & Bosman, G. Red blood cell homeostasis: mechanisms and effects of microvesicle generation in health and disease. Front. Physiol. 9, 703 (2018).
Antipa, N. et al. DiffuserCam: lensless single-exposure 3D imaging. Optica 5, 1–9 (2018).
Acknowledgements
D.R. acknowledges grant support from the Swiss National Science Foundation (10.006.824 and 10.003.762). Z.C. acknowledges support from the Fundamental Research Funds for the Central Universities in China (13702150142) and the National Natural Science Foundation of China (62575214). W.W. acknowledges support from the National Natural Science Foundation of China (52275527).
Author information
Authors and Affiliations
Contributions
B.Z., Q.Z. and Z.C. conceived the experimental design. B.Z. and Q.Z. carried out the experiments. B.Z. and Q.Z. conducted data analysis. L.G., E.J. and D.S. carried out the segmentation and visualization of vessels. M.R. and T.J. assisted with the animal experiments. L.T. performed RBC staining. C.G. and B.W. contributed to the interpretation of the results. X.C. contributed to the optimization of the ADMM algorithm. S.G., X.D., Q.F., H.A. and W.H. contributed to the design and optimization of the phase mask. Y.C., X.L.D-B. and S.L. established the tumor model and contributed to the interpretation of the results. W.W., X.D., Z.C., D.R. and Q.Z. supervised the work. D.R. acquired funding for the project. B.Z. and Q.Z. drafted the manuscript. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Xiaochuan Zhang, Robert Prevedel and the other anonymous reviewer for their contribution to the peer review of this work. [A peer review file is available].
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, B., Guo, S., Tang, L. et al. Double-helix optical point spread function enables real-time mesoscopic 3D functional microangiography in the living mouse brain and skull. Nat Commun (2026). https://doi.org/10.1038/s41467-026-71746-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-71746-9