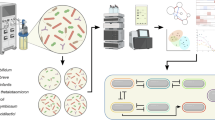
Abstract
Infant gut microbiota development involves frequent colonization by Enterobacteriaceae, particularly Escherichia coli, yet their ecological role in healthy infants is unclear. Here, we analyse longitudinal stool samples from healthy, term-born, breastfed infants (n = 41) and related mothers (n = 30) using shotgun metagenomics and novel computational approaches. Strain-resolved profiling indicates that Bifidobacterium species are frequently shared within families, whereas E. coli derive from external sources, but often persist within individuals. Despite differing ecological strategies, these genera co-exist and share evolutionary adaptations related to lactose acquisition in the infant gut. In vitro, we demonstrate that interactions between E. coli and Bifidobacterium bifidum are mutualistic in co-culture, where E. coli supplies cysteine to its auxotrophic partner, facilitating cooperative degradation of 2′-fucosyllactose, the predominant human milk oligosaccharide. In turn, the liberated monosaccharides sustain E. coli growth, highlighting a cooperative cross-feeding interaction that may contribute to regulating E. coli abundance within the infant host.
Similar content being viewed by others
Introduction
Infant health is tied to the postnatal assembly of the gut microbiota. Despite major research interest1,2,3,4, our understanding of the principles underlying microbial community assembly is still lacking. This is largely due to difficulties in identifying true microbe-microbe and host-microbe interactions in natural populations, whose dynamics arise from the interplay between deterministic factors such as diet5 and geographical location6 with countless other stochastic components.
Human breastmilk represents the primary source of nutrition during early life. Its most abundant carbohydrate is lactose, which provides nutrition for the infant, although it can also be metabolized by a wide range of gut-associated bacterial taxa7. To sustain an intestinal population of commensal microbes, lactose is additionally incorporated within greater oligosaccharide structures, collectively known as human milk oligosaccharides (HMOs). HMOs are not absorbed in the proximal gut and thus remain available to microbial strains with the enzymatic capacity to degrade them8. Together, these features impose two key ecological constraints on infant gut colonization: (i) the ability to capture and degrade HMOs, and (ii) competition for accessible mono- and disaccharides.
Bifidobacterium frequently dominates the gut microbiota of breastfed infants9. Their presence has been associated with several beneficial functions, including induction of immunological tolerance10, establishment of colonisation resistance against pathogens11, improved vaccine response during the first year of life12, and maintenance of gut-barrier integrity11. This early dominance is supported by their rich set of glycoside hydrolases enabling efficient HMO-degradation13 and direct maternal transfer14. At the same time, Escherichia coli is commonly detected in breastfed infants, typically at low relative abundances but with high prevalence across individuals10,15,16,17,18,19,20,21,22. E. coli are generally considered pathobionts23, as upon overgrowth, they can cause infectious diarrhoea, which remains a significant cause of neonatal sepsis and mortality24, particularly in low- and middle-income countries25. E. coli lacks the capacity to degrade HMOs26, but its generalist metabolic strategy and rapid turnover of mono- and disaccharides are thought to underly colonization of the intestinal ecosystem27,28,29. In breastmilk, soluble lactose is not completely absorbed in the small intestine30 and may therefore contribute to E. coli persistence. Additionally, E. coli may access simple sugars by cross-feeding interactions with extracellular degraders of HMOs, two mechanisms that are not mutually exclusive. Milk oligosaccharide cross-feeding has been reported between Bacteroides and E. coli17, but it remains unclear if Bifidobacterium could similarly provide accessible sugars to E. coli.
To investigate ecological factors associated with infant gut community composition, we performed metagenomic sequencing of stool samples from a Dutch cohort of healthy neonates (n = 41). Notably, these infants represent a selected subset of the larger LucKi Birth cohort31, chosen for their rare status of exclusive breastfeeding post-delivery before gradually transitioning to a more complex diet during the first year. This well-defined dietary trajectory provides a unique model system to investigate organizing principles of infant microbiome community dynamics. To explore it in depth, we developed a new computational pipeline - MAJIC (Mean across Jaccard index checkerboards), and profiled microdiversity of infant and maternal gut microbiota, revealing ecological aspects for the transmission of strains among individual hosts. Furthermore, without relying on annotation of genes, we identified ecological links between Bifidobacterium and Escherichia concerning their strategies to acquire lactose, supported by their consistent co-occurrence in neonates. Lastly, our experimental findings demonstrate that in co-culture, E. coli actively supplies cysteine to auxotrophic B. bifidum, facilitating the cooperative breakdown of 2’-fucosyllactose (2’FL), one of the predominant HMOs. This in vitro interaction highlights a cross-feeding mechanism that may contribute to the regulation of E. coli persistence and abundance within the infant host.
Results
E. coli and Bifidobacterium co-occur consistently during early-life
Although the order of colonizing bacteria is well described4,32,33, the mechanisms driving assembly of the early-life gut microbiome remain poorly understood. Here, we employed shotgun metagenomics to sequence 78 longitudinal stool samples from 41 healthy neonates. These included an early time-point (n = 18) at 2 months post delivery, during which all infants were exclusively breastfed. Additional samples were collected at 6 months post delivery (n = 25), a time-point that commonly marks the onset of weaning34, and a late time-point at 11 months post delivery (n = 35), by which complex foods were introduced. In comparison to the broader LucKi Birth cohort31, where 74.1% of infants are no longer exclusively breastfed by 6 months, our cohort represents a rare subset. Additionally, to evaluate potentially relevant components of vertical transfer between mother and infant we sequenced 30 stool samples of respective mothers at 2 weeks post delivery (Fig. 1A and Supplementary Data 1).
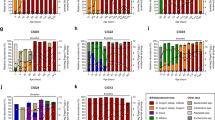
A Overview of study-design. Samples were obtained from babies at two-, six-, and eleven months post-delivery (B-02, B-06, B-11), and one maternal sample was obtained two weeks post-delivery (M). B Microbial richness measured by Shannon diversity (Two-sided unpaired t-test with FDR correction). C Principal component analysis (PCA) of relative species counts over Hellinger distance. D Scatterplot highlighting prevalence (percentage of samples in which species are present) and high abundance (percentage of samples in which species represent >20% of the microbial community) for Bifidobacterium and Escherichia coli in maternal and infant samples. E Bar plot displaying the distribution of microbial colonization patterns across sample categories. Bars represent the percentage of samples (y-axis) classified according to different combinations of abundances between Bifidobacterium (including B. adolescentis, B. breve, B. bifidum, and B. longum subsp. longum) and E. coli. F Differences in mean Jaccard dissimilarity ∆µi = µi+ - µi-, wherein negative values indicate that microbiota are more restrictive when focal species is missing [σi- (µi-)], and conversely more facilitating when ∆µi > 0, therefore focal species being present [σi+ (µi+)]. It was identified via comparisons to shuffled data whether focal species (σi) is state-associated (X) or state-dissociated (O), and colour indicates age-groupings. On the x-axis, Bifidobacterium are highlighted in red and E. coli in black. ⊗ in violin plots indicate group median.
Overall, alpha diversity was similar at 2 and 6 months post-delivery (Shannon, t.test, p.adj = 0.061) and increased significantly thereafter (Shannon, t.test, p.adj <0.001, respectively; Figs. 1B and S1A). For between-sample comparisons, species present in <10% of samples at a minimum relative abundance of 0.1% within each age group were excluded to improve statistical robustness and reduce noise. Principal component analysis (PCA) indicated that infant gut microbiotas were most dissimilar to each other at 2 months post delivery, gradually becoming more similar to same-aged infants and to maternal microbiotas over time (Fig. 1C). This trend was more pronounced between infants and their own mothers compared to unrelated mothers (Fig. S1B). Microbiome composition varied significantly with age (PERMANOVA, p.adj <0.05), except between 2 and 6 months post delivery (PERMANOVA, p.adj = 0.731), highlighting the selective pressure exerted by a breastmilk-based diet. Bifidobacterium breve was on average the most abundant species in infants, followed by Bifidobacterium bifidum, and Bifidobacterium longum subspecies longum. As long as breastmilk was supplied, E. coli occurred at low relative abundance, but consistently (74% and 96% prevalence at 2 and 6 months post delivery), alongside initially dominant Bifidobacterium. In maternal samples, Bifidobacterium adolescentis was the most abundant bifidobacterial species on average (Figs. 1D and S1C, and Supplementary Data 2). To assess the distribution of taxa across infants, we calculated a “cross-sample Shannon diversity” index (CSI) for each species, quantifying the evenness of the species’ abundance across all samples. Prevalent Bifidobacterium species, including B. bifidum (CSI = 3.8), B. longum subsp. longum (CSI = 3.8), B. breve (CSI = 3.5), and B. adolescentis (CSI = 3.4), ranked within the top 5% of this distribution ( > 95th percentile), indicating broad and even prevalence across the cohort. In contrast, other Bifidobacterium species (e.g., B. animalis, B. catenulatum) exhibited markedly lower CSI, reflecting restricted occurrence and uneven abundance patterns.
In the majority of cases, E. coli co-occurred with either a single dominant Bifidobacterium species or multiple highly abundant Bifidobacterium species during the initial 6 months post delivery (Figs. 1E and S1D, E). However, at 2 months postpartum, the relative abundance of E. coli was notably higher when only one single Bifidobacterium species was highly abundant (relative abundance > 20%) compared to when multiple Bifidobacterium species were highly abundant. But this difference did not remain statistically significant after correction for multiple testing (t-test, p = 0.043, p.adj=0.14; Fig. S1F). In rare cases where E. coli was present without a highly abundant Bifidobacterium, the following alternative species were so instead: at 2 months post delivery, Clostridium perfringens (45%-abundance) and Bifidobacterium pseudocatenulatum (53%), once each. At 6 months post delivery, Ruminococcus gnavus (42%-abundance), Bacteroides fragilis (31%-abundance), B. pseudocatenulatum (29%), and Bacteroides clarus (42%-abundance) in four separate cases (Fig. S1G). Notably, all of these species are evidentially capable of degrading HMOs35,36,37, while E. coli lack this ability38.
Together, our findings show that E. coli and Bifidobacterium co-exist in the gut of most breastfed infants. However, E. coli is not restricted to communities dominated by extracellular HMO degraders (e.g., B. bifidum), but also appear alongside species that primarily import HMOs for intracellular degradation (e.g., B. longum subsp. longum, B. breve)39 (Fig. S1D). This implies that residual lactose may be sufficient to sustain their co-existence.
Mean across Jaccard index checkerboards (MAJIC)
Before focusing specifically on interactions between Bifidobacterium and E. coli, we first aimed to capture the broader landscape of putative interactions within the infant gut microbiota on the species level. To this end, we developed a computational pipeline - MAJIC - to systematically assess the extent to which focal microbes may influence gut ecology. Our approach involves comparing microbial communities in which focal species (i) is present (σi+) to those where it is absent (σi-). To ensure robust statistical testing, we generate 10,000 random σi+ and σi- datasets by shuffling the presence/absence labels of focal species (i) each time. This randomization scheme accounts for differences in sample sizes and abundance distributions of individual species. For each focal species, we calculate the mean Jaccard dissimilarity of samples in σi+ (µi+), and in σi- (µi-), as well as the difference in mean Jaccard dissimilarity (∆µi = µi+-µi-), comparing these values to the random distributions generated from shuffled data. A negative ∆µi (<0) indicates that microbiotas are more constrained in the presence of the focal species (since µi+ <µi-), whereas a positive ∆µi (>0) indicates greater community variability when the focal species is present. Furthermore, we define a species as ‘state-associated’ when the distribution of its μᵢ⁺ values, computed from observed data, differs significantly from that obtained by shuffled inputs (ks.test, p < 0.05). Conversely, “state-dissociated” species show no significant difference (p ≥ 0.05), suggesting no strong association with specific community states (Fig. S2A).
At 2 months post delivery, infant microbiotas demonstrated a high degree of compositional flexibility, which was gradually lost over time. By 11 months, and in maternal microbiotas, communities became more compositionally constrained, with an increased prevalence of “state-associated” focal species, compared to earlier time points (Fig. S2B). We identified R. gnavus as the most facilitating species in maternal microbiota (∆µ = 0.12), and Bacteroides stercoris as the most restrictive species in infant microbiota 11 months post delivery (∆µ = −0.25), suggesting critical roles in niche occupation for these putative keystone species. Interestingly, highly abundant bifidobacteria (B. breve, B. bifidum, and B. longum subsp. longum and B. adolescentis) were classified as “state-dissociated” at 2 months post delivery, suggesting no strong association with specific community states. This suggests that early post-delivery, despite their recognised importance for infant development and gut microbiome assembly, highly abundant Bifidobacterium species largely do not function as putative keystone species in the classical ecological sense40. Instead, their loss is likely compensated by functional equivalent species, maintaining overall composition and functional stability despite an individual species’ absence at 2 months post delivery. However, after 6 months post delivery B. bifidum became “state-associated”, and so did B. adolescentis and B. breve 11 months post delivery (Fig. 1F).
Our analysis furthermore extends beyond presence/absence patterns to identify pairs of state-associated microbes that quantitatively influence each other’s abundance within the community. For each focal species (i) as defined above, we identified any species (j) whose abundance patterns significantly differed between matrices containing species (i) (σi+) and those without it (i) (σi-) using a Wilcoxon rank-sum test. At 2 months post delivery, most significant associations occurred among “state-dissociated” species, suggesting that these interactions were likely stochastic. In contrast, significant associations between “state-associated” species were rare but consistently negative across all time points (Fig. S2C), indicating that deterministic, competitive dynamics are limited to a small number of taxa.
Lastly, we filter our data to identify any non-stochastic instances where a “state-associated” focal species (σi) exerts a significant effect on any type of species (j) with involvements of dominant Bifidobacterium or E coli. Thereby, we found no significant interactions at 2 and 6 months post delivery. At 11 months post delivery, B. adolescentis was significantly more abundant in the presence of Gemella haemolysans. In maternal samples, B. breve was negatively affected by the presence of Lactobacillus fermentum and Enterococcus gallinarum. Importantly, across all timepoints we found no significant evidence of antagonism between highly-abundant Bifidobacterium and E. coli using our pipeline. However, given that they consistently co-occur in the infant gut, we conclude that their relationship must be ecologically neutral and that their co-occurrence is likely driven by shared environmental preferences and permissible niche overlap, rather than direct interactions. In contrast, E. coli exhibited significantly reduced abundance in the presence of Klebsiella pneumoniae at 11 months post delivery, pointing to a potential antagonistic relationship between these two Enterobacteriaceae (Fig. S2D).
Strain-resolved transmission routes in infant gut microbiota
To better understand the ecological strategies and transmission dynamics of key early-life gut microbes, we next investigated strain-level variation and gene-level diversity within highly-abundant Bifidobacterium species and E. coli. Therefore, we assembled metagenome-assembled genomes (MAGs) from shotgun sequencing data and overall acquired 458 unique dereplicated high-quality genomes (Supplementary Data 3). On average, these MAGs accounted for 75 ± 14% of metagenomic reads in infants at 2 months, 78 ± 5% at 6 months, and 75 ± 7% at 11 months post delivery, while covering 68 ± 7% of maternal reads. To verify that sequencing depth was sufficient for robust genome detection, we examined the relationship between genome coverage (average read depth) and breadth (fraction of the genome covered by reads)41. The resulting curve plateaued near ~0.85 breadth, indicating that additional sequencing would yield little new genomic information and that most genomes, including low-abundance taxa, were captured (Fig. S3A). Using recommended thresholds which minimize false positive detection from spurious read mappings (breadth > 0.5; coverage > 5)41, E. coli detection closely matched read-based profiles (Fig. S3B), confirming prevalence in infants at 2 and 6 months. Subsequently, we mapped all reads to our set of dereplicated genomes, and inStrain41 was used to profile the nucleotide diversity (π), assessing both within (intra-π) and across (inter-π) individual variation.
Intra-π for all MAGs did not correlate to the coverage of MAGs within samples (Spearman, p.adj > 0.05, respectively, Fig S3C), indicating that strain-level genetic variation in the infant gut is shaped by factors other than relative abundances of respective MAGs. Overall, intra-π remained stable between infants at 2 and 6 months post delivery, but increased significantly thereafter (t.test, p.adj < 0.05 for every other pairwise comparison), and E. coli MAGs most notably displayed elevated intra-π, suggesting the presence of multiple strains co-existing within individuals, whereas MAGs of highly-abundant Bifidobacterium (B. bifidum, B. longum subsp. longum, B. breve, and B. adolescentis) rarely showed such patterns, indicating reduced strain-level diversity (Fig. 2A).
A Dot-plot visualizing the distribution of overall nucleotide diversity (π) according to age-groups. Each dot represents a metagenome-assembled-genome (MAG) dereplicated at 95% average nucleotide identity (ANI), and size of dots represent mean coverage MAGs. MAGs of B. bifidum, B. breve, B. longum subsp. longum, B. adolescentis and E. coli are coloured in red, salmon, pink, orange, and black. Statistical significance was assessed using a two-sided Welch’s t-test with FDR correction. B Count of strain-persistence events where identical strains of B. bifidum, B. longum subsp. longum, B. breve, B. adolescentis, and E. coli re-occurred between adjacent timepoints within the same individual. C Violin / Dot-plot visualizing the distribution of overall gene nucleotide diversity (gene-π) in B. bifidum, B. breve, B. longum subsp. longum, B. adolescentis, and E. coli. Each dot represents a gene in respective MAGs. Statistical significance was assessed using a two-sided Welch’s t-test with FDR correction.
To investigate strain-level transmission patterns, we applied a threshold of pop-ANI 99.5% to distinguish between identical strains and strain-variants, and quantified strain-sharing events between individuals. Identical B. adolescentis strains were shared among 22% of maternal microbiota, while B. longum subsp. longum strains were shared among 31% of infants at 11 months post delivery (Fig. S3D). Furthermore, pop-ANI values were significantly higher between parenting mothers and their infants, as compared to random mother-infant pairs at two and 11 months post delivery (t-test, p.adj <0.0001 and 0.008, respectively; Fig. S3E). Notably, B. longum subsp. longum strains were frequently shared between infants and their mothers, whereas no such co-occurrence was observed for E. coli strains in mother-infant pairs (Fig. S3F). Next, we assessed strain persistence within infants across time for dominant Bifidobacterium and E. coli. Between 2 and 6 months post delivery, identical B. bifidum and B. longum subsp. longum strains persisted within individuals in 12.5% of cases, while no such persistence was observed for B. breve. From 6 to 11 months, identical E. coli strains persisted in 10% of cases, and from two to eleven, as well as from 6 to 11 months, B. longum subsp. longum strains persisted in 25% of infants (Fig. 2B). Overall, these findings reveal distinct transmission and persistence patterns among key infant gut microorganisms. B. adolescentis was frequently shared among maternal microbiota, while B. longum subsp. longum strains were commonly transmitted between infants and parenting mothers. In contrast, B. breve and B. bifidum were less frequently transmitted between mothers and their respective infants but were prevalent in the infant gut. E. coli displayed high microdiversity overall, and although not shared between mothers and infants, strains often persisted within infants across the first year post-delivery, implying other origins of colonization.
We hypothesized that the observed differences in host-associated transmission patterns would be reflected in the genetic nucleotide diversity (gene-π) within Bifidobacterium and E. coli. Specifically, given that E. coli exhibited limited mother-to-infant transmission as well as higher pop-ANI values, we expected weaker host-specific selective pressures, leading to higher nucleotide diversity across a broader range of genes. Conversely, for Bifidobacterium, where frequent strain transmission and persistence between hosts occur, we anticipated stronger purifying selection on genes essential for host adaptation. To identify these genes, we applied EggNOG-mapper v6 for functional annotation42, and profiled gene-π in B. longum subsp. longum, B. breve, B. bifidum, B. adolescentis, and E. coli. As expected, E. coli displayed the highest gene-π on average. Additionally, gene-π did not significantly differ between B. breve, B. longum subsp. longum (t.test, p.adj > 0.05), suggesting that despite differences in transmission dynamics, their ecological roles and adaptive strategies are similar. However, B. bifidum exhibited a particularly narrow gene-π distribution that significantly differed from the remaining species (t.test, p.adj < 0.0001, Fig. 2C), indicating stronger genetic constraints on its adaption. Next, we categorized genes with gene-π values above the 99th percentile as “loose” and those below the 1st percentile as “restricted.” These labels are operational definitions intended to distinguish genes with unusually high or low within-species nucleotide diversity, rather than to imply predefined functional roles. Among the loose genes, we identified, for example, a starch-binding outer membrane protein from the SusD/RagB family in B. bifidum, which may contribute to HMO transport. Restricted genes, aside from several housekeeping functions, included a putative chitinase in B. bifidum and a multiple-sugar transporter in B. breve, both potentially involved in HMO acquisition (Supplementary Data 4).
In summary, our findings reveal distinct ecological strategies among key infant gut microbes. B. longum subsp. longum was frequently transmitted and persisted between infants and mothers, while B. bifidum exhibited strong genetic constraints, presumably reflecting host-specific adaptation. In contrast, E. coli showed high microdiversity, limited transmission from parenting mothers, but high persistence of singular strains in individual infants.
GH-driven co-evolution underpins microbial adaptation to HMOs
We next used our MAGs to investigate functional traits underpinning microbial adaptation to the infant gut environment. Lactose and HMOs, as the primary carbon sources for early colonizing microbes, represent key ecological drivers of primary succession in the infant gut43. To overcome the lack of traditional gene annotation and gain a more holistic understanding of microbial traits essential for the acquisition of carbohydrates in the infant gut, we employed a recently developed annotation-agnostic approach to infer microbial trophic strategies from genomic data44. We hypothesized that glycoside hydrolases (GHs) are necessary for the degradation of HMOs and lactose, that they are costly to produce, and that they benefit not only the GH producers but also all potential consumers of the resulting degradation products. As such, GHs function as model public goods, where their utility extends beyond enzyme synthesis to encompass a suite of associated traits, largely unknown but likely including chemotaxis, biofilm formation, and membrane transport, that enhance the efficient capture and utilization of GH-derived breakdown products. Public goods like GHs are subject to rapid evolutionary shifts between parasitism and cooperation due to trade-offs between production costs and competitive advantages. Furthermore, their evolutionary trajectories are shaped by high rates of gene deletion and horizontal transfer. By leveraging these dynamics, we can trace patterns of co-evolution between the GHs (public goods) and battery-traits across microbial species with distinct ecological strategies, providing deeper insights into how microbial communities adapt to HMO-rich environments44.
Because of their central role in infant gut microbiota, we sought to identify protein families that have co-evolved with GH2 family enzymes across our set of 458 unique dereplicated genomes. In total, we identified 2803 such protein families (εGH2), with an average of and 302 ± 284 per genome (εiGH2). To further investigate the degree of polymer degradation specialization in different microbial lineages and the associated strategies, we compared the number of GH2 enzymes encoded in a genome (OGH2) to two derived quantities based on the co-evolutionary patterns of GH2-related genes encoded within single genomes. The first, EGH2, was estimated using an elastic-net linear regression model fitted across all genomes. This value represents the expected number of GH2 enzymes a genome ‘should’ encode based on its genomic background of εiGH2. If true coevolution were occurring, EGH2 and OGH2 would be strongly correlated, as recently demonstrated for chitinases and their coevolved genes44. Indeed, we observed a strong correlation between EGH2 and OGH2, supporting the validity of our approach and the genuine co-evolution of εGH2 across diverse microbial phyla (Spearman, R = 0.83, p < 0.00001). However, our data also suggest that the specialized organisms for GH2 breakdown product utilization (high EGH2 and high OGH2) primarily include species from the genera Bacteroides and Parabacteroides, rather than typically dominant members of the infant gut microbiome (Supplementary Data 3).
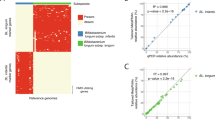
The final parameter we examined, fiCAZy, represents the fraction of genes coevolving with GH2 that are themselves CAZymes. If the selection of coevolved genes were random, the correlation between OGH2 and fCAZy would be expected to follow a diagonal line of identity. However, our analysis revealed a notable deviation, with Bifidobacterium forming a distinct cluster, alongside species from E. coli. We speculate that this clustering suggests a non-random coevolutionary dynamic, likely driven by shared selective pressures that require an expanded repertoire of CAZymes for efficient colonization of the infant gut. These findings highlight putative functional associations between these species. Notably, B. bifidum deviated from this pattern, as its GH2-associated genes were exclusively non-CAZymes, suggesting an alternative metabolic adaptation (Fig. 3A).
A Scattering of observed number of GH2 domain-containing enzymes against fraction of coevolved genes that are other carbohydrate active enzymes as well. Each dot represents a MAG, coloured by corresponding phylum. The most abundant bifidobacteria across infant and maternal samples (B. bifidum, B. breve, B. longum subsp. longum, B. adolescentis) as well as E. coli, are highlighted. B Bar-charts summarizing taxonomic origin of genes that co-evolved with GH-2 glycoside hydrolases. Bars represent the proportion (%) of GH-2 co-evolved genes assigned to different bacterial taxa, including B. adolescentis (orange), B. bifidum (red), B. breve (salmon), B. longum subsp. longum (pink), and E. coli (black). Only coevolved genes that were mapped to kegg-orthology (ko) groups via EggNOG mapper v6 are plotted (unknown were excluded).
To further explore the functional significance of these coevolved genes, we annotated them using EggNOG-mapper v642. Notably, while all genera displayed GH2 coevolving genes that could be attributed to amino acid metabolism, E. coli uniquely lacked such associations (Figs. 3B and S4A, B). This suggests that while E. coli mirrors the dynamics of Bifidobacterium, which are specialized in HMO-degradation, it adopts the ecological role of a versatile prototroph, and may additionally be less reliant on simultaneous amino acid degradation to access GH2 breakdown products, despite possessing the functional capacity to do so45.
B. bifidum and E. coli cooperate over the degradation of 2′FL
E. coli is unable to degrade HMOs26, but its ability to rapidly scavenge simple mono- and disaccharides, residual lactose in particular, likely explains its prevalence in breastfed infants. Another source of simple sugars is formed by extracellular degradation of HMOs, and notably, E. coli co-occurs with extracellularly degrading B. bifidum (Fig. S1D). However, it is unknown whether external degradation of HMOs by B. bifidum could lead to growth of E. coli.
To test this hypothesis, we focused on the metabolism of 2′-O-fucosyl-lactose (2’FL), the most abundant breastmilk HMO46. First, we isolated B. bifidum from infant stool and assessed its growth on 0.1% 2’FL (w/v) as the primary carbon source in modified M9-medium (m-M9)47. B. bifidum was able to grow at 37 °C under anoxic conditions (Fig. 4A), but not in m-M9 lacking 2’FL (Fig. S5A). This confirmed that the additives to m-M9 were insufficient to support growth without an external carbon source. B. bifidum also failed to grow in m-M9 + 0.1% 2’FL without casamino acids or cysteine, consistent with the well-established cysteine requirement for Bifidobacterium48, and indicating additional metabolic dependencies in this isolate (Fig. S5B). Additionally, we isolated highly-abundant HMO-importers from infant stool (B. longum subsp. longum and B. breve), but neither grew on m-M9 + 2’FL (Fig. 4A). We then analyzed whether E. coli K-12 could directly metabolize 2’FL or its individual monosaccharides - fucose, galactose, and glucose as carbon sources. While E. coli K-12 could not metabolize 2’FL, it exhibited slight growth in m-M9 only, in the absence of 2’FL, suggesting either a preference for amino acid metabolism in the absence of polysaccharides or possibly inhibitory effects of 2’FL on growth (Fig. S5C). However, E. coli K-12 efficiently grew when provided with either one of the three monomers, including fucose (Fig. S5D). Next, we tested whether E. coli K-12 could grow in a spent medium from B. bifidum cultures, observing growth only when the pH was adjusted to its initial value of 6.3 (Fig. 4B). Repeating the experiments with higher 2’FL concentrations (0.5% w/v) confirmed that E. coli could utilize degradation products released by B. bifidum (Fig. S5E). Using thin-layer chromatography (TLC) and high-performance anion exchange chromatography (HPAEC) we determined that B. bifidum efficiently degraded 2’FL, but left galactose and fucose unconsumed, which were subsequently utilized by E. coli K12 (Figs. 4C and S6A–D).
A Growth of B. bifidum (red), B. breve (salmon), B. longum subsp. longum (pink) in modified M9-medium (m-M9) with 2’-O-fucosyl-lactose (2’FL) as the only source of carbon, measured by optical density (OD-600). Data represent the mean ± s.e.m. of 3 incubation vials per species. B Growth of E. coli in supernatant of B. bifidum previously grown in m-M9 with 2’FL as the only carbon-source. Data represent the mean ± s.e.m. of 6 incubation vials. C Concentration of fucose and galactose in m-M9 + 2’FL B. bifidum supernatants spent by E. coli at pH 6,3. Data represent the mean ± s.e.m. of 10 incubation vials. D Relative abundances of B. bifidum and E. coli in co-culture in m-M9 + 2’FL without any additions of cysteine or other amino acids. Relative abundances were estimated by fluorescent gram-stain and manual counting, averaging 9 fields of view per timepoint. Data represent the mean ± s.e.m. of 3 co-cultures. E Growth of B. bifidum and E. coli co-cultures with respective controls, including a cysteine mutant (E. coli ∆-cysE) which is incapable of cysteine production. Growth was assessed by optical density measurements (OD-600) after 24 h. Blanks included sterile m-M9 + 2’FL medium without cysteine or other amino acids. Boxplots represent the mean ± s.e.m. of 3 to 6 biological replicates. Boxplots show the median (center line), interquartile range (box; 25th–75th percentile), and whiskers extending to the most extreme values within 1.5×IQR. Points represent individual measurements. Differences in OD-600 were assessed using two-sided pairwise Welch’s t-tests with FDR correction for multiple comparisons (p.adj = 0.01). F Visual conclusion of the article. Asterisks represent adjusted p-values: ∗p < 0.05. Dots in line-plots show group median, error bars show standard deviation. Box plots show group median and interquartile range.
We next hypothesized that under nutrient-limiting conditions, prototrophic E. coli K12 could provide essential amino acids to an auxotrophic partner, thus facilitating coexistence. To test this, we co-cultured E. coli K12 and our cysteine auxotroph B. bifidum isolate in m-M9 + 0.5% 2’FL(v/v) without added cysteine and casamino acids. After 24 hours, we observed turbidity in this co-culture, suggesting growth (Fig. S5F). Fluorescent Gram-staining confirmed co-existence of both species throughout the experiment, until B. bifidum became dominant after 12 h (Fig. 4D, and Supplementary Data 5). Furthermore, no significant growth was observed when one partner was ethanol-killed before inoculation, confirming the necessity of active metabolic interaction. Additionally, we repeated the co-culture using an E. coli K12 mutant (∆-cysE) lacking cysteine biosynthesis, observing a significant impairment of growth (t.test, p.adj = 0.01; Fig. 4E). We therefore conclude that prototrophic E. coli K12 can facilitate the growth and metabolic activity of auxotrophic B. bifidum under nutrient-limiting conditions by alleviating its cysteine dependency. In turn, B. bifidum enables the degradation of 2’FL, providing E. coli with essential carbon sources, demonstrating a reciprocal cross-feeding interaction.
Discussion
Bifidobacterium and E. coli are both core members of neonatal gut microbiota18,49,50. Here, we examined their co-existence in exclusively breastfed, term-born infants during the initial year post-delivery using strain-resolved metagenomics, a novel computational pipeline (MAJIC), and an annotation-agnostic approach to infer shared ecological constraints during early-life microbiome assembly. We found that E. coli frequently occurs alongside one or more highly-abundant Bifidobacterium species (B. longum subsp. longum, B. breve, or B. bifidum) as long as breastmilk constitutes the primary source of nutrition (Figs. 1 and S1). Because this co-occurrence is not unique to extracellular HMO-degraders, we argue that residual lactose is likely the primary determinant of E. coli persistence. However, both genera share ecological constraints related to carbohydrate acquisition in the infant gut, evidenced by their non-random enrichment of fiCAZy (Fig. 3A). Lastly, we demonstrate in vitro that E. coli supplies cysteine to auxotrophic B. bifidum, enabling joint degradation of 2’FL (Fig. 4F), suggesting that HMO-derived cross-feeding may represent a secondary source of simple sugars for E. coli.
Traditionally, interactions between Bifidobacterium and E. coli in the healthy infant gut had been interpreted as antagonistic, in part due to acidification during HMO fermentation51,52. The metabolic bifid-shunt yields acetate and lactate, which can lower environmental pH and are thought to restrict E. coli growth53. Also, in our batch-culture experiments, E. coli was unable to grow in B. bifidum spent medium (pH = 4.3) without prior pH adjustment. However, fecal pH in breastfed infants does not fall below 4.3 and has remained above 5.5 for the past decades54, well above the threshold for lethal intracellular accumulation of undissociated organic acids in E. coli. In vivo, luminal pH is further buffered by host bicarbonate secretion55 and microbial cross-feeding of fermentation end-products56. Furthermore, E. coli possesses multiple acid tolerance mechanisms, including proton pumps, amino acid decarboxylation systems, and the production of alkaline metabolites, which enable survival and colonization under moderately acidic conditions57. Here, we highlight that E. coli is present in most breastfed infants within our cohort and co-occurs consistently with Bifidobacterium, which argues against strict exclusion by acidification in term-born infants and instead points to stable ecological conditions that support E. coli at low relative abundance. This raises the question of which resources sustain E. coli populations in the healthy infant gut.
Freter’s nutrient niche hypothesis offers a plausible explanation: to co-colonize the infant gut, each species must use at least one limiting nutrient better than all other species58,59. Breastmilk introduces two sources of carbohydrates to colonizing microbiota, HMOs and lactose. E. coli is particularly competitive for the acquisition of lactose due to its high-affinity transport system and rapid growth kinetics27,28,29. Since most lactose is absorbed by the host in the small intestine, only limited amounts reach the distal gut30, likely having a key role in constraining E. coli to low relative abundances. In contrast, Bifidobacterium are specialized in degradation of HMOs, which are not absorbed by the host and therefore constitute a large and exclusive carbohydrate pool supporting their high relative abundance60,61. Moreover, the extensive structural diversity of HMOs62 enables niche partitioning among Bifidobacterium species, facilitating their stable co-occurrence. Formula-fed infants lack HMOs, diminishing the selective advantage of HMO specialists and favoring metabolism of simple sugars, consistent with higher E. coli abundances in these infants63. However, our experimental data show that cross-feeding of HMO-degradation products can sustain E. coli growth in vitro, suggesting that cross-feeding could act as an additional secondary mechanism influencing E. coli abundances in breastfed infants, given the presence of extracellular HMO-degraders such as B. bifidum.
On the one hand, HMO cross-feeding interactions may increase substrate use efficiency64, resulting in enhanced SCFA production65, a hallmark of a healthy infant gut ecosystem. On the other hand, cross-feeding interactions involve extracellular release of public goods that enlarge niche space but become susceptible to exploitation by pathogens8,66,67. These opposing effects raise a critical question: what is the role of cross-feeding for the stability of microbial communities during early-life assembly? Ecological theory predicts that obligate syntrophic interactions are destabilizing, since the mutual dependence between cooperating species leads to disruptive positive feedback loops whereby coupled species either bloom or collapse together, resulting in highly dynamic and unstable microbial communities67. This means that, although cooperation can increase the overall metabolic efficiency of microbial communities, it comes at the cost of ecosystem stability. Generally, there are several mechanisms by which cooperation can be maintained in the gut microbiota in order to optimize these trade-offs. Competitive interactions among other microbial species can dampen ‘boom-and-bust’ dynamics by limiting resource availability to cooperating partners68. Immunological constraints regulate microbial composition69. Spatial heterogeneity of the intestinal ecosystem can lead to the separation of interacting species70,71. Finally, functional redundancy replaces single strong interactions with multiple weaker ones, while simultaneously buffering communities against loss of metabolic functions67.
Our in vitro observation of cross-feeding between E. coli and B. bifidum is an extreme case of obligate mutual dependence, imposed by low-nutrient conditions lacking both easily accessible lactose and cysteine. Conceptually, this interaction is consistent with a snowdrift game, where the success of E. coli is maximized under conditions of high enzymatic costs and low capture efficiency by B. bifidum, whereas increased nutrient availability would alleviate dependence and shift the mutualistic interaction to either commensalistic behavior or exclusion of one partner72. How does this theory relate to early-life microbiota? In the healthy breastfed infant gut, the coupling between E. coli and B. bifidum is attenuated by multiple stabilizing forces, including host- and breastmilk-mediated immunological constraints, spatial separation of taxa along the developing mucosal landscape, functional redundancy among both genera, and competition arising from residual lactose, a byproduct of incomplete host digestion that may promote stability by increasing competitive overlap among early colonizers. However, in diseased or premature infants, the nutrient and redox landscape is altered73, lactase function and transit are perturbed74, and cysteine availability can be limited75, potentially shifting interactions towards stronger metabolic interdependence among gut microbes. Consistent with this framework, extracellularly feeding B. bifidum and E. coli are often detected in gut microbiotas of extremely premature infants76, although this pattern likely reflects multiple concurrent drivers.
Lastly, the “restaurant hypothesis” extends Freter’s nutrient niche framework by proposing that cross-feeding interactions expand the available niche space: extracellular degraders act as “restaurants” that release metabolic byproducts, creating new nutritional opportunities for species that cannot access the primary substrate77,78. As discussed above, such public goods are inherently susceptible to exploitation, but resident commensal E. coli may contribute to colonization resistance by limiting access to these nutritional niches. This can occur either through metabolically flexible strains that concurrently utilize residual lactose and HMO-derived monosaccharides79, or through the coexistence of multiple strains with distinct monosaccharide preferences, both of which reduce carbohydrate availability to other potentially pathogenic Enterobacteriaceae. However, while shotgun metagenomics sensitively profiles microbial genomes, without targeted qPCR we cannot reliably quantify Enterobacteriaceae below ~0.1% relative abundance or evaluate the ecological relevance of ultra-low-abundance strains that fall below our detection limit.
In summary, our data indicate that early-life gut microbiotas are structured primarily by competition for host-derived sugars, while cross-feeding of HMO-degradation products may additionally modulate ecosystem stability and accessibility, given the presence of extracellular HMO-degraders. However, it remains unclear whether cross-feeding interactions originate from evolved traits or passive processes, such as cell lysis or overflow metabolism. Future research should prioritize understanding how microbes interact and compete for key nutrients within their metabolic networks. Stable isotope–based tracing of carbon fluxes across taxa offers a powerful approach to uncover these dynamics, helping guide the rational design of probiotics and/or dietary interventions to support child health.
Methods
Experimental model, clinical definitions, stool sample collection
Stool samples were collected within the context of the LucKi Gut study, an ongoing longitudinal study31 among newborns and their families. Written informed consent was obtained from both parents/legal caregivers prior to enrolment in the study. This research confirmed to the principles of the Helsinki Declaration. Ethical approval was obtained by the Medical Ethical Committee of Maastricht University Medical Center (study number: METC-15-4-237). Pregnant women residing in the South Limburg region of the Netherlands were recruited through obstetrics and gynecology clinics, lactation information sessions, and advertisements at pregnancy yoga classes, baby clothing stores, and on social media. Infants born prematurely (gestational age ≥32 weeks) were excluded. One maternal sample and 9 infant fecal samples were collected throughout the first 14 months of life for the overarching LucKi study. Participants received fecal sampling starter kits consisting of stool collection tubes (Sarstedt, REF 80.623.022), cold transport containers (Sarstedt, REF 95.1123), safety bags, gloves, faeces collection devices (Fe-Col, REF FC2010), questionnaires, instructions and consent forms. The samples were collected at home and immediately stored at −20 °C in their home freezers. Samples were thereafter transported to the family’s well-baby clinic using frozen transport container to preserve the cold chain. From there, samples were transported to the laboratory, where frozen fecal matter was aliquoted and stored at −80 °C until further analyzes.
At each fecal sampling time-point, parents also completed a questionnaire gathering information on the infant’s lifestyle, health, development, medication use, and feeding practices, as well as maternal health (during pregnancy), diet, and medication use. Samples collected at 2 weeks (maternal sample), and 2, 6, and 11 months (infant samples) post partum were selected for the purpose of the present study. All selected infants were healthy and born after 38 weeks of gestation and were exclusively breastfed initially.
Isolation of DNA, metagenomic sequencing, and read-based processing
DNA was extracted using the Power Soil Pro Kit (Qiagen) following the manufacturer’s protocol for stool samples, with the inclusion of one negative control per extraction batch, consisting of nuclease-free water to allow for assessment of contamination. DNA was eluted in 40 μl nuclease free water and stored at −20 °C until further analysis. DNA was quantified using Qubit 2.0 dsDNA BR assay kit (ThermoFisher Scientific, Q32850). Library preparation was done following the manufacturer’s instructions using the DNA flex kit with 8-mer UDI. Shotgun metagenome sequencing (short-read) on DNA samples was performed at Azenta Life Sciences on Illumina NovaSeq platform at paired-end reads 2× 150 bp. Short-read metagenomic data were processed using HUMAnN 380 with default setting. Briefly, HUMAnN 3 first estimates community composition with MetaphlAn 481, second it maps reads to a community pangenome with bowtie282, and third it aligns unmapped reads to a protein database using DIAMOND83.
De novo assembly and processing of metagenome-assembled genomes (MAGs)
Raw sequence reads were quality filtered (-q 20) with fastp v.0.20.084 before de novo genome assembly using SPAdes v.3.14.185 at default parameters. CheckM v.1.1.386 was used for the assessment of metagenome assembled genomes (MAGs). Low-quality genomes (contamination > 20% and completion < 70%) excluded from further analysis. These were taxonomically classified using GTDBtk (v. 2.1.0)87. The MAG dataset was dereplicated with standard settings via dRep88, resulting in a final set of 458 unique quality MAGs (Supplementary Data 3). Dereplicated MAGs were furthermore concatenated into a single fasta file, and bowtie282 was used to create a mapping index from it, as well as to calculate the percentage of reads per sample that were mapping to our set of dereplicated MAGs. A “scaffold-to-bin” was created using “parse_stb.py” from dRep88, and Prodigal89 was used to profile all genes from the concatenated MAG file, thereby creating a “gene” file. Combining these files, “inStrain profile” was used to screen within-sample nucleotide diversity (π) and gene-nucleotide diversity (gene-π), and “inStrain compare” was used to profile across-sample population average nucleotide identity (popANI)41. The ‘gene’ file was furthermore used for annotation of all genes using EggNOG mapper v. 6.042.
Detection of glycoside-hydrolase-linked genes
We expanded a previous analysis that detected genes that coevolve with chitinases44 by analyzing 21,559 high-quality species-representative genomes from GTDB r214 (>99% completeness, <1% contamination). Genes were called with Pyrodigal90, and full-length proteins were clustered at 70% similarity using diamond deepclust followed by diamond recluster (both with --approx-id 70)83. Partial reading frames were then added to clusters using diamond blastp (--query-cover 100 --id 80 -b 20 -c 1 -k 10). Glycosyl hydrolases were called with dbcan491. All genes were annotated using eggnog-mapper v592. To look for genes that co-evolve with GH2, we only considered protein clusters that appear in >9 genomes, which reduces computational load and false positive rates due to random fluctuations44. Ancestral states were reconstructed using maximum parsimony, as implemented in the mpr function of the PhyloTools Julia package93. We detected co-evolution as correlated gain-loss events using canonical correlation analysis (CCA). We used CCA because there are many GH2 protein clusters, and CCA uncovers how the optimal linear combination of their gain/loss vectors correlates with a target vector. These correlations were compared to null distributions generated from randomized vectors, where events from different genes were randomized, therefore controlling for the mean tendency of nodes to display gain/loss events44. To analyze MAGs that were assembled de novo from our own data, we first assign MAG proteins to the protein homology clusters defined above using diamond blastp (--query-cover 80 --subject-cover 80 -k 3 -c 1). Each MAG protein was assigned to the best-matching protein cluster based on e-value. The number predicted number of GH2 enzymes in each MAG was calculated using the random forest regression model from above, and compared to the observed number of GH2 proteins encoded by the MAG. To identify the fraction of GH2 co-evolved genes that themselves are carbohydrate-active enzymes, we compared annotated coevolving genes against the PFAM database using PFAM.db v3.8.294.
Growth assays with Bifidobacterium and E. coli
B. bifidum, B. longum subsp. longum, and B. breve were isolated from infant stool samples by spreading 10-7 dilutions of fecal slurries onto MRS agar with cysteine (0.1% w/v; Sigma-Aldrich) and mupirocin (50 mg/l; Sigma-Aldrich) (m-MRS). Plates were incubated at 37 °C under anaerobic conditions for 48 h. Per plate, several colonies were picked at random and grown in liquid m-MRS, repeating the 48 h-long incubation. From incubations with visible growth, DNA was extracted using the Power Soil Pro Kit (Qiagen) following the manufacturer’s protocol and finally eluted in 40 µl nuclease-free water. DNA isolated from pure bacterial cultures was subjected to whole-genome sequencing via Illumina NextSeq 2000. Obtained sequencing reads were pre-processed with fastp v.0.23.284. Unicycler v.0.4.995 with the “–mode conservative” option was used to produce assemblies, after which contigs below 1000 bp were filtered out. Genome completeness and contamination were estimated using CheckM v.1.2.086, and we retained sequences with completeness >99% and contamination < 1%. GTDB-Tk v.2.1.096 was used to classify assemblies to the strain level. Thereby, we identified several B. bifidum, B. longum subsp. longum, and B. breve, and selected one representative respectively for our growth assays.
Bifidobacterium isolates were pre-grown in MRS-medium + cysteine (0.1% w/v; Sigma-Aldrich) for 48 h. Thereof, 2% (v/v) was transferred to modified M9 medium47 with 0.1% or 0.5% 2’-O-fucosyl-lactose (2’FL; Glycome, IRE), following a 24 h incubation at 37 °C under anaerobic conditions (Coy Laboratory Products, USA) to probe their capabilities of metabolizing 2’FL. Modified M9 had the following additions: biotin (10 mg/l), para-aminobenzoic acid (10 mg/l), thiamine (400 mg/l), nicotinic acid (400 mg/l), pyridoxine (400 mg/l), pantothenate calcium (200 mg/l), riboflavin (200 mg/l), and a diluted casamino-acid extract (0.01% w/v; Sigma-Aldrich). pH was adjusted to 6.3, and media were filter-sterilized (Sartorius 0.22 µm filter; Sigma-Aldrich) before inoculation. E. coli K12 was pre-grown in brain-heart infusion (BHI; Sigma Aldrich) for 24 h before being inoculated (1% v/v) in m-M9 or m-M9 spent by B. bifidum. To generate supernatants of spent media, cultivates were centrifuged at 13,000 rpm for 10 min, supernatants were collected, pH was adjusted to 6.3, and lastly filter-sterilized (Sartorius 0.22 µm filter; Sigma-Aldrich). Growth of all cultures was monitored under anaerobic conditions in a plate-reader (Thermo Multiskan, Thermo Fisher, USA). For end-point assessments, optical density (OD-600) of cultures was measured by spectrophotometry (Camspec, Spectronic, UK). E. coli BW25113 ∆-cysE was obtained from the Keio collection97. For co-culture experiments, B. bifidum and E. coli K12 were pre-grown in m-MRS or BHI for 48 or 24 h under anaerobic conditions at 37 °C, respectively. Cell density of inoculates was assessed by Luna FX7 automated cell counter (Thermo Fisher USA) to ensure close to 1:1 ratios of all inoculates. E. coli ∆-cysE was pre-grown in BHI + cysteine. Relative abundances of B. bifidum and E. coli K12 in co-culture were assessed by fluorescent Gram stain (LIVE BacLight, Thermo Fisher, USA) and manual counting. Nine representative pictures were taken per timepoint (Zeiss, GER) to estimate the average relative abundance of either microbe in co-culture (Supplementary File 1).
Quantification of 2’-O-fucosyl-lactose and its monomers
For TLC, Culture supernatant samples (3 μL) were spotted onto silica plates (Sigma Z740230) and resolved in running buffer containing 2:1:1 butanol, acetic acid and dH2O, respectively. TLCs were subsequently stained using diphenylamine–aniline–phosphoric acid stain98 and developed by heating.
To analyze and quantify monosaccharide release in culture supernatants, we used high-performance anion exchange chromatography with pulsed amperometric detection (HPAEC-PAD). Sugars were separated using a CarboPac PA-1 anion exchange column with a PA-1 guard using a Dionex ICS-6000 (Thermo Fisher) and detected using PAD. Flow was 0.75 mL min−1 and elution conditions were 0–25 min 5 mM NaOH and then 25–40 min 5–100 mM NaOH. Software used was the Chromeleon Chromatography Data System. Monosaccharide/disaccharide standards were included at the following concentrations: Maltose = 0.1 mg/ml, Glucose = 0.1 mM, Fucose = 0.1, 0.075, 0.05, 0.025, and 0.01 mM, N-acetylglucosamine = 0.2 mM, Galactose = 0.2 mM, N-acetylgalactosamin = 0.1 mM, and Lactose 0.2 mM. All data were obtained by diluting supernatant samples 1/10 prior to injection.
Statistics, reproducibility, and data visualization
Statistical analysis (Student’s T-test, ANOVA, repeated measures ANOVA, Wilcoxon Test, Chi-square test) was performed in “R version 4.0” and the R package “rstatix version 0.7.0”99. All p-values were adjusted using Bonferroni’s method. Data was visualized via “R version 4.0” and R package “ggplot2 version 3.3.340”100. “Ampvis2”101 and “phyloseq”102 were used for handling of metagenomic counts, metadata, and taxonomy files, including species filtering, analysis of alpha and beta diversity, as well as PCA. “co_occurrence()” in phylosmith103 was used for co-occurrence analysis of microbial species. For randomization of matrices, we used the “randomizeMatrix()” function in “Picante”104 with “null.model = “frequency” over 10,000 iterations. Kolmogorow–Smirnov-Test – “ks.test()” in base R – was used to test for whether differences in randomized and observed matrices were statistically different. “c.score()” function in “bipartite”105 was used for calculation of checkerboard indices. For MAJIC, randomized and observed matrices were transformed into presence/absence matrices using “phyloseq_standardize_otu_abundance()” with “method = pa” in “metagMisc”106, and average Jaccard dissimilarities as well as cross-sample Shannon diversity (CSI) index per species were calculated using the functions “vegdist()” and “diversity()” in “vegan”107, respectively. MAJIC was coded “R version 4.0”, looping through focal species σi to obtain σi+ (µi+), and in σi- (µi-) - as described in Fig. S2A.
Data availability
Source data are provided with this paper. Further information for resources and reagents should be directed to and will be fulfilled by lead contacts David Seki (david.seki@univie.ac.at) or Lindsay J. Hall (l.hall.3@bham.ac.uk). Infant fecal sample metagenome sequencing raw reads are publicly available in the NCBI Sequence Read Archive (SRA) under accession no. PRJNA1230889. All 458 MAGs recovered from gut metagenome samples are available via Zenodo108 (https://doi.org/10.5281/zenodo.18848062), and all high-quality MAGs with completeness > 95% (n = 279) are also available via PRJNA1230889. Source data are provided with this paper.
Code availability
The code for analysis of the data109 can be accessed via https://github.com/sekid666/MAJIC and https://doi.org/10.5281/zenodo.18833415.
References
Olm, M. R. et al. Robust variation in infant gut microbiome assembly across a spectrum of lifestyles. Science 376, 1220–1223 (2022).
Reyman, M. et al. Effects of early-life antibiotics on the developing infant gut microbiome and resistome: a randomized trial. Nat. Commun. 13, 893 (2022).
Vatanen, T. et al. Mobile genetic elements from the maternal microbiome shape infant gut microbial assembly and metabolism. Cell 185, 4921–4936.e15 (2022).
Valles-Colomer, M. et al. The person-to-person transmission landscape of the gut and oral microbiomes. Nature 614, 125–135 (2023).
Baumann-Dudenhoeffer, A. M., D’Souza, A. W., Tarr, P. I., Warner, B. B. & Dantas, G. Infant diet and maternal gestational weight gain predict early metabolic maturation of gut microbiomes. Nat. Med. 24, 1822–1829 (2018).
Cheng, Y. et al. Influence of geographical location on maternal-infant microbiota: study in two populations from Asia and Europe. Front. Cell. Infect. Microbiol. 11, 663513 (2022).
A Talbert, J. & D. Townsend, S. Human milk as a complex natural product. Nat. Prod. Rep. 42, 406–420 (2025).
Charbonneau, M. R. et al. A microbial perspective of human developmental biology. Nature 535, 48–55 (2016).
Lawson, M. A. E. et al. Breast milk-derived human milk oligosaccharides promote Bifidobacterium interactions within a single ecosystem. ISME J. 14, 635–648 (2020).
Henrick, B. M. et al. Bifidobacteria-mediated immune system imprinting early in life. Cell 184, 3884–3898.e11 (2021).
Fukuda, S. et al. Bifidobacteria can protect from enteropathogenic infection through production of acetate. Nature 469, 543–547 (2011).
Huda, M. N. et al. Stool microbiota and vaccine responses of infants. Pediatrics 134, e362–e372 (2014).
Masi, A. C. & Stewart, C. J. Untangling human milk oligosaccharides and infant gut microbiome. iScience 25, 103542 (2022).
Flores Ventura, E. et al. Mother-to-infant vertical transmission in early life: a systematic review and proportional meta-analysis of Bifidobacterium strain transmissibility. npj Biofilms Microbiomes 11, 121 (2025).
Stewart, C. J. et al. Temporal development of the gut microbiome in early childhood from the TEDDY study. Nature 562, 583–588 (2018).
Ferretti, P. et al. Assembly of the infant gut microbiome and resistome are linked to bacterial strains in mother’s milk. Nat. Commun. 16, 11536 (2025).
Charbonneau, M. R. et al. Sialylated milk oligosaccharides promote microbiota-dependent growth in models of infant undernutrition. Cell 164, 859–871 (2016).
Ioannou, A. et al. Resource sharing of an infant gut microbiota synthetic community in combinations of human milk oligosaccharides. ISME J. 18, wrae209 (2024).
Dalby, M. J. et al. Faecal microbiota and cytokine profiles of rural Cambodian infants linked to diet and diarrhoeal episodes. npj Biofilms Microbiomes 10, 85 (2024).
Chu, D. M. et al. Maturation of the infant microbiome community structure and function across multiple body sites and in relation to mode of delivery. Nat. Med. 23, 314–326 (2017).
Bäckhed, F. et al. Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe 17, 690–703 (2015).
Samarra, A., Cabrera-Rubio, R., Martínez-Costa, C. & Collado, M. C. The role of Bifidobacterium genus in modulating the neonate microbiota: implications for antibiotic resistance acquisition in early life. Gut Microbes 16, 2357176 (2024).
Jochum, L. & Stecher, B. Label or concept – what is a pathobiont? Trends Microbiol. 28, 789–792 (2020).
Fagundes-Neto, U. & Andrade, J. A. B. d Acute diarrhea and malnutrition: lethality risk in hospitalized infants. J. Am. Coll. Nutr. 18, 303–308 (1999).
Nji, E. et al. High prevalence of antibiotic resistance in commensal Escherichia coli from healthy human sources in community settings. Sci. Rep. 11, 3372 (2021).
Salli, K. et al. Selective utilization of the human milk oligosaccharides 2′-fucosyllactose, 3-fucosyllactose, and difucosyllactose by various probiotic and pathogenic bacteria. J. Agric. Food Chem. 69, 170–182 (2021).
Conway, T. & Cohen, P. S. Commensal and pathogenic Escherichia coli metabolism in the gut. Microbiol. Spectr. 3 10.1128/microbiolspec.mbp-0006–2014 (2015).
Doranga, S., Krogfelt, K. A., Cohen, P. S. & Conway, T. Nutrition of Escherichia coli within the intestinal microbiome. EcoSal Plus 12, eesp-0006-2023 (2024).
Jang, J. et al. Environmental Escherichia coli: ecology and public health implications-a review. J. Appl. Microbiol. 123, 570–581 (2017).
Lifschitz, C. H., Smith, E. O. & Garza, C. Delayed complete functional lactase sufficiency in breast-fed infants. J. Pediatr. Gastroenterol. Nutr. 2, 478–482 (1983).
de Korte-de Boer, D. et al. LucKi birth cohort study: rationale and design. BMC Public Health 15, 934 (2015).
Shao, Y. et al. Stunted microbiota and opportunistic pathogen colonization in caesarean-section birth. Nature 574, 117–121 (2019).
Vatanen, T. et al. A distinct clade of Bifidobacterium longum in the gut of Bangladeshi children thrives during weaning. Cell 185, 4280–4297.e12 (2022).
A descriptive study investigating the use and nature of baby-led weaning in a UK sample of mothers. https://onlinelibrary.wiley.com/doi/10.1111/j.1740-8709.2010.00243.x.
Chapman, J. A. et al. Clostridia from preterm infants metabolize human milk oligosaccharides to suppress pathobionts and modulate intestinal function in organoids. Nat Microbiol 11, 940–959 (2026)
Lordan, C. et al. Linking human milk oligosaccharide metabolism and early life gut microbiota: bifidobacteria and beyond. Microbiol. Mol. Biol. Rev. 88, e00094–23 (2024).
Buzun, E. et al. A bacterial sialidase mediates early-life colonization by a pioneering gut commensal. Cell Host Microbe 32, 181–190.e9 (2024).
Coppa, G. V. et al. Human milk oligosaccharides inhibit the adhesion to Caco-2 cells of diarrheal pathogens: Escherichia coli, Vibrio cholerae, and Salmonella fyris. Pediatr. Res. 59, 377–382 (2006).
Ojima, M. N. et al. Priority effects shape the structure of infant-type Bifidobacterium communities on human milk oligosaccharides. ISME J. 16, 2265–2279 (2022).
Power, M. E. et al. Challenges in the quest for keystones. BioScience 46, 609–620 (1996).
Olm, M. R. et al. inStrain profiles population microdiversity from metagenomic data and sensitively detects shared microbial strains. Nat. Biotechnol. 39, 727–736 (2021).
Hernández-Plaza, A. et al. eggNOG 6.0: enabling comparative genomics across 12 535 organisms. Nucleic Acids Res. 51, D389–D394 (2023).
Laursen, M. F. & Roager, H. M. Human milk oligosaccharides modify the strength of priority effects in the Bifidobacterium community assembly during infancy. ISME J. 17, 2452–2457 (2023).
Pollak, S. et al. Public good exploitation in natural bacterioplankton communities. Sci. Adv. 7, eabi4717 (2021).
Zampieri, M., Hörl, M., Hotz, F., Müller, N. F. & Sauer, U. Regulatory mechanisms underlying coordination of amino acid and glucose catabolism in Escherichia coli. Nat. Commun. 10, 3354 (2019).
Castanys-Muñoz, E., Martin, M. J. & Prieto, P. A. 2′-fucosyllactose: an abundant, genetically determined soluble glycan present in human milk. Nutr. Rev. 71, 773–789 (2013).
Miller, J. H. Experiments in Molecular Genetics (Cold Spring Harbor Laboratory, [Cold Spring Harbor, N.Y.], 1972).
Ferrario, C. et al. Exploring amino acid auxotrophy in Bifidobacterium bifidum PRL2010. Front. Microbiol. 6, 1331 (2015).
Li, X. et al. The infant gut resistome associates with E. coli, environmental exposures, gut microbiome maturity, and asthma-associated bacterial composition. Cell Host Microbe 29, 975–987.e4 (2021).
Yin, Q. et al. Ecological dynamics of Enterobacteriaceae in the human gut microbiome across global populations. Nat. Microbiol. 10, 541–553 (2025).
Presser, K. A., Ratkowsky, D. A. & Ross, T. Modelling the growth rate of Escherichia coli as a function of pH and lactic acid concentration. Appl. Environ. Microbiol. 63, 2355–2360 (1997).
Pinhal, S., Ropers, D., Geiselmann, J. & de Jong, H. Acetate Metabolism and the Inhibition of Bacterial Growth by Acetate. J. Bacteriol. 201, https://doi.org/10.1128/jb.00147-19 (2019).
FUSHINOBU, S. Unique sugar metabolic pathways of bifidobacteria. Biosci. Biotechnol. Biochem. 74, 2374–2384 (2010).
Henrick, B. M. et al. Elevated fecal pH indicates a profound change in the breastfed infant gut microbiome due to reduction of bifidobacterium over the past century. mSphere 3, https://doi.org/10.1128/msphere.00041-18 (2018).
Becker, H. M. & Seidler, U. E. Bicarbonate secretion and acid/base sensing by the intestine. Pflüg. Arch. Eur. J. Physiol. 476, 593–610 (2024).
Detman, A. et al. Cell factories converting lactate and acetate to butyrate: Clostridium butyricum and microbial communities from dark fermentation bioreactors. Microb. Cell Factories 18, 36 (2019).
Xu, Y. et al. An acid-tolerance response system protecting exponentially growing Escherichia coli. Nat. Commun. 11, 1496 (2020).
Freter, R., Brickner, H., Botney, M., Cleven, D. & Aranki, A. Mechanisms that control bacterial populations in continuous-flow culture models of mouse large intestinal flora. Infect. Immun. 39, 676–685 (1983).
Freter, R., Brickner, H., Fekete, J., Vickerman, M. M. & Carey, K. E. Survival and implantation of Escherichia coli in the intestinal tract. Infect. Immun. 39, 686–703 (1983).
Brand-Miller, J. C., McVeagh, P., McNeil, Y. & Messer, M. Digestion of human milk oligosaccharides by healthy infants evaluated by the lactulose hydrogen breath test. J. Pediatr. 133, 95–98 (1998).
Kujawska, M. et al. Succession of Bifidobacterium longum strains in response to a changing early life nutritional environment reveals dietary substrate adaptations. iScience 23, 101368 (2020).
Ninonuevo, M. R. et al. A strategy for annotating the human milk glycome. J. Agric. Food Chem. 54, 7471–7480 (2006).
Penders, J. et al. Quantification of Bifidobacterium spp., Escherichia coli and Clostridium difficile in faecal samples of breast-fed and formula-fed infants by real-time PCR. FEMS Microbiol. Lett. 243, 141–147 (2005).
Bäckhed, F., Ley, R. E., Sonnenburg, J. L., Peterson, D. A. & Gordon, J. I. Host-bacterial mutualism in the human intestine. Science 307, 1915–1920 (2005).
Fischbach, M. A. & Sonnenburg, J. L. Eating for two: how metabolism establishes interspecies interactions in the gut. Cell Host Microbe 10, 336–347 (2011).
Culp, E. J. & Goodman, A. L. Cross-feeding in the gut microbiome: ecology and mechanisms. Cell Host Microbe 31, 485–499 (2023).
Coyte, K. Z., Schluter, J. & Foster, K. R. The ecology of the microbiome: Networks, competition, and stability. Science 350, 663–666 (2015).
Ratzke, C., Barrere, J. & Gore, J. Strength of species interactions determines biodiversity and stability in microbial communities. Nat. Ecol. Evol. 4, 376–383 (2020).
Zheng, D., Liwinski, T. & Elinav, E. Interaction between microbiota and immunity in health and disease. Cell Res. 30, 492–506 (2020).
Zhang, Z. et al. Spatial heterogeneity and co-occurrence patterns of human mucosal-associated intestinal microbiota. ISME J. 8, 881–893 (2014).
Kim, H. J., Boedicker, J. Q., Choi, J. W. & Ismagilov, R. F. Defined spatial structure stabilizes a synthetic multispecies bacterial community. Proc. Natl. Acad. Sci. USA 105, 18188–18193 (2008).
Gore, J., Youk, H. & van Oudenaarden, A. Snowdrift game dynamics and facultative cheating in yeast. Nature 459, 253–256 (2009).
Letourneau, J., Walker, L., Han, S. H., David, L. A. & Younge, N. A pilot study of fecal pH and redox as functional markers in the premature infant gut microbiome. PLoS ONE 19, e0290598 (2024).
Montgomery, R. K., Mulberg, A. E. & Grand, R. J. Development of the human gastrointestinal tract: twenty years of progress. Gastroenterology 116, 702–731 (1999).
Miller, R. G., Jahoor, F. & Jaksic, T. Decreased cysteine and proline synthesis in parenterally fed, premature infants. J. Pediatr. Surg. 30, 953–958 (1995).
Seki, D. et al. Gut microbiota genome features associated with brain injury in extremely premature infants. Gut Microbes 16, 2410479 (2024).
Adediran, J. et al. An Escherichia coli Nissle 1917 Missense mutant colonizes the streptomycin-treated mouse intestine better than the wild type but is not a better probiotic. Infect. Immun. 82, 670–682 (2014).
Leatham-Jensen, M. P. et al. The streptomycin-treated mouse intestine selects Escherichia coli envZ missense mutants that interact with dense and diverse intestinal microbiota. Infect. Immun. 80, 1716–1727 (2012).
Fabich, A. J. et al. Comparison of carbon nutrition for pathogenic and commensal Escherichia coli strains in the mouse intestine. Infect. Immun. 76, 1143–1152 (2008).
Beghini, F. et al. Integrating taxonomic, functional, and strain-level profiling of diverse microbial communities with bioBakery 3. eLife 10, e65088 (2021).
Blanco-Miguez, A. et al. Extending and improving metagenomic taxonomic profiling with uncharacterized species with MetaPhlAn 4. Nat Biotechnol. 41, 1633–1644 (2023).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Buchfink, B., Reuter, K. & Drost, H.-G. Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat. Methods 18, 366–368 (2021).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
SPAdes: A New Genome Assembly Algorithm and Its Applications to Single-Cell Sequencing - PMC. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3342519/.
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Chaumeil, P.-A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk: a toolkit to classify genomes with the Genome Taxonomy Database. Bioinformatics 36, 1925–1927 (2020).
Olm, M. R., Brown, C. T., Brooks, B. & Banfield, J. F. dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J. 11, 2864–2868 (2017).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 11, 119 (2010).
Larralde, M. Pyrodigal: Python bindings and interface to Prodigal, an efficient method for gene prediction in prokaryotes. J. Open Source Softw. 7, 4296 (2022).
Zheng, J. et al. dbCAN3: automated carbohydrate-active enzyme and substrate annotation. Nucleic Acids Res. 51, W115–W121 (2023).
Cantalapiedra, C. P., Hernández-Plaza, A., Letunic, I., Bork, P. & Huerta-Cepas, J. eggNOG-mapper v2: functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol. Biol. Evol. 38, 5825–5829 (2021).
Narushima, H. & Hanazawa, M. A more efficient algorithm for MPR problems in phylogeny. Discrete Appl. Math. 80, 231–238 (1997).
PFAM.db. Bioconductor http://bioconductor.org/packages/PFAM.db/.
Wick, R. R., Judd, L. M., Gorrie, C. L. & Holt, K. E. Unicycler: resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput. Biol. 13, e1005595 (2017).
Chaumeil, P.-A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk v2: memory friendly classification with the genome taxonomy database. Bioinformatics 38, 5315–5316 (2022).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006.0008 (2006).
Zhang, Z., Xie, J., Zhang, F. & Linhardt, R. J. Thin layer chromatography for the analysis of glycosaminoglycan oligosaccharides. Anal. Biochem. 371, 118–120 (2007).
Pipe-Friendly Framework for Basic Statistical Tests • rstatix. https://rpkgs.datanovia.com/rstatix/.
Wickham, H. Ggplot2: Elegant Graphics for Data Analysis (Springer, New York, NY, 2009). https://doi.org/10.1007/978-0-387-98141-3.
Andersen, K. S., Kirkegaard, R. H., Karst, S. M. & Albertsen, M. ampvis2: an R package to analyse and visualise 16S rRNA amplicon data. Preprint at https://doi.org/10.1101/299537 (2018).
McMurdie, P. J. & Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217 (2013).
Smith, S. D. phylosmith: an R-package for reproducible and efficient microbiome analysis with phyloseq-objects. J. Open Source Softw. 4, 1442 (2019).
Kembel, S. W. et al. Picante: R tools for integrating phylogenies and ecology. Bioinformatics 26, 1463–1464 (2010).
Dormann, C. How to be a specialist? Quantifying specialisation in pollination networks. Netw. Biol. 1, 1–20 (2011).
Mikryukov, V. vmikk/metagMisc: v.0.0.4. Zenodo https://doi.org/10.5281/zenodo.1172500 (2018).
Oksanen, J. et al. Vegan Community Ecology Package Version 2.6-2 April 2022. (2022).
Seki, D. Wellcome Dereplicated MAGs (with metadata). Zenodo https://doi.org/10.5281/zenodo.18848062 (2026).
sekid666. sekid666/MAJIC: MAJIC. Zenodo https://doi.org/10.5281/zenodo.18833415 (2026).
Acknowledgements
We thank all families for their participation during this study. We would like to thank the QIB sequencing team for technical assistance with shotgun metagenomics library preparation. This project was funded by the Wellcome Trust Investigator Award no. 220876/Z/20/Z to L.J.H., and a Biotechnology and Biological Sciences Research Council (BBSRC) Institute Strategic Programme, Gut Microbes and Health BB/R012490/1, and its constituent projects BBS/E/F/000PR10353 and BBS/E/F/ 000PR10356, and by the BBSRC Institute Strategic Programme Food Microbiome and Health BB/X011054/1 and its constituent project BBS/E/F/000PR13631 to L.J.H. D.S. and S.P. acknowledges funding from the Austrian Science Fund (FWF; doi.org/10.55776/COE7).
Author information
Authors and Affiliations
Contributions
Conceptualization, D.S. and L.J.H; methodology, D.S., M.K., R.K., S.P., L.I.C., C.R.B., A.AG.; software, D.S., S.P; investigation, D.S.; writing – original draft, D.S. and L.J.H.; writing – review & editing, D.S., L.J.H., S.P.; resources, L.J.H., L.I.C., N.v.B., J.P., P.T.C., M.M.; and funding acquisition, L.J.H.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Seki, D., Pollak, S., Kujawska, M. et al. Human milk oligosaccharide mediates mutualism between Escherichia coli and Bifidobacterium bifidum. Nat Commun 17, 3489 (2026). https://doi.org/10.1038/s41467-026-71764-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-71764-7