Abstract
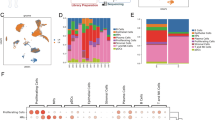
Cancer metastasis accounts for the majority of cancer-related deaths and lymph node metastasis is critical in cancer staging and prognosis. However, the mechanisms involved in lymph node metastatic non-small cell lung cancer (NSCLC) remain unclear. Here, we delineate the cellular and spatial landscape of metastatic lymph nodes and their matched primary tumors in NSCLC using multimodal omics. In results, tumor core and peripheral regions respectively exhibit high stemness and elevated expression of immunoglobulins. Increased expressions of CD274 (PD-L1) are observed in the immunoglobulin-high malignant cells and SPP1+IFI30+ macrophages. Notably, residual and tertiary lymphoid structures are identified. However, their anti-tumor immunity is compromised by CXCL13+ T cell exhaustion and immune exclusion. Furthermore, we identify the ferroptosis signaling, with its key factor FTH1 showing strong associations with cancer cell stemness and metastasis. This study elucidates the mechanisms of metastasis and immune evasion in lymph node metastatic NSCLC, providing insights for targeted therapies.
Similar content being viewed by others
Data availability
The raw scRNA-seq and ST data generated in this study have been deposited at the Genome Sequence Archive at the National Genomics Data Center (Beijing, China)136 under accession ID HRA009872. The raw sequencing data are available under restricted access for research purposes only, access can be obtained by the DAC (Data Access Committees) of the GSA-human database. According to the guidelines of GSA-human (https://ngdc.cncb.ac.cn/gsa-human/policy), all non-profit researchers can obtain access to the data, and the Principle Investigator of any research group is allowed to apply the data. The access authority can be obtained for Research Use Only. The user can also contact the corresponding author directly. Access requests are typically processed within 2 weeks. Once access has been approved, the data will be available to download for 2 months. The pre-processed scRNA-seq and ST data were deposited on Science Data Bank (https://doi.org/10.57760/sciencedb.19197; https://doi.org/10.57760/sciencedb.19202). The public single-cell RNA sequencing datasets used in this study can be found in Gene Expression Omnibus (GEO) with the accession number GSE127465, GSE123904, GSE131907 and GSE117529, in BioStudies with accession number E-MTAB-13526, from Zenodo (https://zenodo.org/records/8227624) or in the LungCancer website (https://lungcancer.chenlulab.com/#/download). The public ST data of NSCLC can be found in BioStudies with accession number E-MTAB-13530. The public ST data of mLNs in breast cancer can be found in GEO with accession number GSE190811. The ST data of nLN was from 10X Genomics website (https://www.10xgenomics.com/datasets/human-lymph-node-1-standard-1-1-0). Bulk RNA-seq data was from the cancer genome atlas (TCGA) and GEO (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE30219; https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE68465). Source data are provided with this paper.
Code availability
Code is available from GitHub (https://github.com/diChen310/mLN_NSCLC/, https://doi.org/10.5281/zenodo.18690124)137.
References
Tian, Y., He, Y., Li, X. & Liu, X. Novel nomograms to predict lymph node metastasis and distant metastasis in resected patients with early-stage non-small cell lung cancer. Ann. Palliat. Med. 10, 2548–2566 (2021).
Mani, K. et al. Causes of death among people living with metastatic cancer. Nat. Commun. 15, 1519 (2024).
Ji, H. et al. Lymph node metastasis in cancer progression: molecular mechanisms, clinical significance and therapeutic interventions. Signal Transduct. Target. Ther. 8, 367 (2023).
Nathanson, S. D., Kwon, D., Kapke, A., Alford, S. H. & Chitale, D. The role of lymph node metastasis in the systemic dissemination of breast cancer. Ann. Surg. Oncol. 16, 3396–3405 (2009).
Naxerova, K. et al. Origins of lymphatic and distant metastases in human colorectal cancer. Science 357, 55–60 (2017).
Steeg, P. S. Targeting metastasis. Nat. Rev. Cancer 16, 201–218 (2016).
Chaffer, C. L. & Weinberg, R. A. A perspective on cancer cell metastasis. Science 331, 1559–1564 (2011).
Watanabe, S. & Asamura, H. Lymph node dissection for lung cancer: significance, strategy, and technique. J. Thorac. Oncol. 4, 652–657 (2009).
Trevaskis, N. L., Kaminskas, L. M. & Porter, C. J. From sewer to saviour-targeting the lymphatic system to promote drug exposure and activity. Nat. Rev. Drug Discov. 14, 781–803 (2015).
Liu, S., Chen, X. & Lin, T. Lymphatic metastasis of bladder cancer: molecular mechanisms, diagnosis and targeted therapy. Cancer Lett. 505, 13–23 (2021).
Cao, R. et al. PDGF-BB induces intratumoral lymphangiogenesis and promotes lymphatic metastasis. Cancer Cell 6, 333–345 (2004).
Follain, G. et al. Fluids and their mechanics in tumour transit: shaping metastasis. Nat. Rev. Cancer 20, 107–124 (2020).
Li, L. S. et al. Lymph node fibroblastic reticular cells preserve a tolerogenic niche in allograft transplantation through laminin α4. J. Clin. Invest. 132, e156994 (2022).
Reticker-Flynn, N. E. et al. Lymph node colonization induces tumor-immune tolerance to promote distant metastasis. Cell 185, 1924–1942.e1923 (2022).
Lee, C. K. et al. Tumor metastasis to lymph nodes requires YAP-dependent metabolic adaptation. Science 363, 644–649 (2019).
Pozniak, J. et al. A TCF4-dependent gene regulatory network confers resistance to immunotherapy in melanoma. Cell 187, 166–183.e125 (2024).
Chen, Y. et al. Spatiotemporal single-cell analysis decodes cellular dynamics underlying different responses to immunotherapy in colorectal cancer. Cancer Cell 42, 1268–1285.e1267 (2024).
Han, G. et al. An atlas of epithelial cell states and plasticity in lung adenocarcinoma. Nature 627, 656–663 (2024).
Qin, P. F. et al. Cancer-associated fibroblasts undergoing neoadjuvant chemotherapy suppress rectal cancer revealed by single-cell and spatial transcriptomics. Cell Rep. Med. 4, 101231 (2023).
Wang, Y. et al. Multidirectional characterization of cellular composition and spatial architecture in human multiple primary lung cancers. Cell Death Dis. 14, 462 (2023).
De Zuani, M. et al. Single-cell and spatial transcriptomics analysis of non-small cell lung cancer. Nat. Commun. 15, 4388 (2024).
He, D. et al. Single-cell RNA sequencing reveals heterogeneous tumor and immune cell populations in early-stage lung adenocarcinomas harboring EGFR mutations. Oncogene 40, 355–368 (2021).
Kim, N. et al. Single-cell RNA sequencing demonstrates the molecular and cellular reprogramming of metastatic lung adenocarcinoma. Nat. Commun. 11, 2285 (2020).
Larroquette, M. et al. Spatial transcriptomics of macrophage infiltration in non-small cell lung cancer reveals determinants of sensitivity and resistance to anti-PD1/PD-L1 antibodies. J. Immunother. Cancer 10, e003890 (2022).
Liu, B. L. et al. Temporal single-cell tracing reveals clonal revival and expansion of precursor exhausted T cells during anti-PD-1 therapy in lung cancer. Nat. Cancer 3, 108–121 (2022).
Salcher, S. et al. High-resolution single-cell atlas reveals diversity and plasticity of tissue-resident neutrophils in non-small cell lung cancer. Cancer Cell 40, 1503–1520.e1508 (2022).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Zhang, L. et al. Integrated single-cell RNA sequencing analysis reveals distinct cellular and transcriptional modules associated with survival in lung cancer. Signal Transduct. Target. Ther. 7, 9 (2022).
Leader, A. M. et al. Single-cell analysis of human non-small cell lung cancer lesions refines tumor classification and patient stratification. Cancer Cell 39, 1594–1609.e1512 (2021).
Caushi, J. X. et al. Transcriptional programs of neoantigen-specific TIL in anti-PD-1-treated lung cancers. Nature 596, 126–132 (2021).
Maynard, A. et al. Therapy-induced evolution of human lung cancer revealed by single-cell RNA sequencing. Cell 182, 1232–1251.e1222 (2020).
Liu, T. et al. Single cell profiling of primary and paired metastatic lymph node tumors in breast cancer patients. Nat. Commun. 13, 6823 (2022).
Quah, H. S. et al. Single cell analysis in head and neck cancer reveals potential immune evasion mechanisms during early metastasis. Nat. Commun. 14, 1680 (2023).
Jiang, S. et al. Generic Diagramming Platform (GDP): a comprehensive database of high-quality biomedical graphics. Nucleic Acids Res. 53, D1670–D1676 (2025).
Zhao, E. et al. Spatial transcriptomics at subspot resolution with BayesSpace. Nat. Biotechnol. 39, 1375–1384 (2021).
Fridman, W. H. et al. Tertiary lymphoid structures and B cells: an intratumoral immunity cycle. Immunity 56, 2254–2269 (2023).
Zhang, Y. et al. CCL19-producing fibroblasts promote tertiary lymphoid structure formation enhancing anti-tumor IgG response in colorectal cancer liver metastasis. Cancer Cell 42, 1370–1385 e1379 (2024).
Onder, L. & Ludewig, B. A fresh view on lymph node organogenesis. Trends Immunol. 39, 775–787 (2018).
Sautes-Fridman, C., Petitprez, F., Calderaro, J. & Fridman, W. H. Tertiary lymphoid structures in the era of cancer immunotherapy. Nat. Rev. Cancer 19, 307–325 (2019).
Travaglini, K. J. et al. A molecular cell atlas of the human lung from single-cell RNA sequencing. Nature 587, 619–625 (2020).
Li, F., Tiede, B., Massagué, J. & Kang, Y. B. Beyond tumorigenesis: cancer stem cells in metastasis. Cell Res. 17, 3–14 (2007).
Zakaria, N. et al. Human non-small cell lung cancer expresses putative cancer stem cell markers and exhibits the transcriptomic profile of multipotent cells. BMC Cancer 15, 84 (2015).
Zhang, W. C. et al. Glycine decarboxylase activity drives non-small cell lung cancer tumor-initiating cells and tumorigenesis. Cell 148, 259–272 (2012).
Bayir, H. et al. Achieving life through death: redox biology of lipid peroxidation in ferroptosis. Cell Chem. Biol. 27, 387–408 (2020).
Yu, R., Hang, Y. H., Tsai, H. I., Wang, D. Q. & Zhu, H. T. Iron metabolism: backfire of cancer cell stemness and therapeutic modalities. Cancer Cell Int. 24, 157 (2024).
Rodriguez, R., Schreiber, S. L. & Conrad, M. Persister cancer cells: iron addiction and vulnerability to ferroptosis. Mol. Cell 82, 728–740 (2022).
Yang, T. et al. Enolase 1 regulates stem cell-like properties in gastric cancer cells by stimulating glycolysis. Cell Death Dis. 11, 870 (2020).
Zhao, Z. et al. PKM2 promotes stemness of breast cancer cell by through Wnt/beta-catenin pathway. Tumour Biol. 37, 4223–4234 (2016).
Wallbillich, N. J. & Lu, H. Role of c-Myc in lung cancer: progress, challenges, and prospects. Chin. Med. J. Pulm. Crit. Care Med. 1, 129–138 (2023).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Sancho, P., Barneda, D. & Heeschen, C. Hallmarks of cancer stem cell metabolism. Br. J. Cancer 114, 1305–1312 (2016).
Peiris-Pagès, M., Martinez-Outschoorn, U. E., Pestell, R. G., Sotgia, F. & Lisanti, M. P. Cancer stem cell metabolism. Breast Cancer Res. 18, 55 (2016).
Bu, W. M. et al. Ferritin couples iron and fatty acid metabolism. Faseb J. 26, 2394–2400 (2012).
Koskenkorva-Frank, T. S., Weiss, G., Koppenol, W. H. & Burckhardt, S. The complex interplay of iron metabolism, reactive oxygen species, and reactive nitrogen species: insights into the potential of various iron therapies to induce oxidative and nitrosative stress. Free Radic. Bio Med. 65, 1174–1194 (2013).
Zhao, J. et al. Current insights into the expression and functions of tumor-derived immunoglobulins. Cell Death Discov. 7, 148 (2021).
Cords, L. et al. Cancer-associated fibroblast classification in single-cell and spatial proteomics data. Nat. Commun. 14, 4294 (2023).
Schupp, J. C. et al. Integrated single-cell atlas of endothelial cells of the human lung. Circulation 144, 286–302 (2021).
Dai, D. N. et al. Elevated expression of CST1 promotes breast cancer progression and predicts a poor prognosis. J. Mol. Med. 95, 873–886 (2017).
Zhang, L., Wang, D., Zhang, L. & Zhu, L. CST1 promoted gastric cancer development by activating the AKT pathway. Clinics 80, 100561 (2025).
Dinh, H. Q. et al. Integrated single-cell transcriptome analysis reveals heterogeneity of esophageal squamous cell carcinoma microenvironment. Nat. Commun. 12, 7335 (2021).
Gharbaran, R. Advances in the molecular functions of syndecan-1 (SDC1/CD138) in the pathogenesis of malignancies. Crit. Rev. Oncol. Hematol. 94, 1–17 (2015).
Hamidi, H. & Ivaska, J. Every step of the way: integrins in cancer progression and metastasis. Nat. Rev. Cancer 18, 533–548 (2018).
Qin, Y. Q., Liu, S. Y., Lv, M. L. & Sun, W. L. Ambra1 in cancer: implications for clinical oncology. Apoptosis 27, 720–729 (2022).
Jeung, H. C. et al. PLEKHA7 signaling is necessary for the growth of mutant KRAS driven colorectal cancer. Exp. Cell Res. 409, 112930 (2021).
Mariathasan, S. et al. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554, 544–548 (2018).
Dong, L. L., Wang, H., Gao, Y., Wang, S. & Wang, W. B. Long non-coding RNA PVT1 promotes the proliferation, migration and EMT process of ovarian cancer cells by regulating CTGF. Oncol. Lett. 25, 71 (2023).
Chi, C. H. et al. ISLR affects colon cancer progression by regulating the epithelial-mesenchymal transition signaling pathway. Anti Cancer Drug 33, E670–E679 (2022).
Feng, D. et al. SMOC2 promotes an epithelial-mesenchymal transition and a pro-metastatic phenotype in epithelial cells of renal cell carcinoma origin. Cell Death Dis. 13, 639 (2022).
Nieto, P. et al. A single-cell tumor immune atlas for precision oncology. Genome Res. 31, 1913–1926 (2021).
Matsubara, E. et al. The Significance of SPP1 in lung cancers and its impact as a marker for protumor tumor-associated macrophages. Cancers 15, 2250 (2023).
Saade, M., de Souza, G. A., Scavone, C. & Kinoshita, P. F. The role of GPNMB in inflammation. Front. Immunol. 12, 674739 (2021).
Perea, L. et al. Reduced airway levels of fatty-acid binding protein 4 in COPD: relationship with airway infection and disease severity. Resp. Res. 21, 21 (2020).
Zhang, S. et al. Interferon Gamma Inducible Protein 30: from biological functions to potential therapeutic target in cancers. Cell Oncol. 47, 1593–1605 (2024).
Li, L. et al. IFI30 as a key regulator of PDL1 immunotherapy prognosis in breast cancer. Int. Immunopharmacol. 133, 112093 (2024).
Amin, A. et al. Dissection of paracrine/autocrine interplay in lung tumor microenvironment mimicking cancer cell-monocyte co-culture models reveals proteins that promote inflammation and metastasis. BMC Cancer 23, 926 (2023).
Salah, F. S. et al. Tumor suppression in mice lacking GABARAP, an Atg8/LC3 family member implicated in autophagy, is associated with alterations in cytokine secretion and cell death. Cell Death Dis. 7, e2205 (2016).
Rizzo, M. B. D. et al. Oral squamous carcinoma cells promote macrophage polarization in an MIF-dependent manner. QJM Int. J. Med. 111, 769–778 (2018).
Chen, R. et al. TNFSF13 is a novel onco-inflammatory marker and correlates with immune infiltration in gliomas. Front Immunol. 12, 713757 (2021).
Cotzomi-Ortega, I. et al. Autophagy inhibition induces the secretion of macrophage migration inhibitory factor (MIF) with autocrine and paracrine effects on the promotion of malignancy in breast cancer. Biology 9, 20 (2020).
Li, H., Ding, J. Y., Zhang, M. J., Yu, H. J. & Sun, Z. J. Tertiary lymphoid structures and cytokines interconnections: the implication in cancer immunotherapy. Cancer Lett. 568, 216293 (2023).
Su, S. C. et al. Blocking the recruitment of naive CD4 T cells reverses immunosuppression in breast cancer. Cell Res. 27, 461–482 (2017).
Schraufstatter, I. U., Zhao, M., Khaldoyanidi, S. K. & DiScipio, R. G. The chemokine CCL18 causes maturation of cultured monocytes to macrophages in the M2 spectrum. Immunology 135, 287–298 (2012).
Gu-Trantien, C. et al. CXCL13-producing T cells link immune suppression and adaptive memory in human breast cancer. Jci Insight 2, e91487 (2017).
Oliveira, G. et al. Landscape of helper and regulatory antitumour CD4+ T cells in melanoma. Nature 605, 532–538 (2022).
Lüthje, K. et al. The development and fate of follicular helper T cells defined by an IL-21 reporter mouse. Nat. Immunol. 13, 491–498 (2012).
Jiang, Y., Li, Y. & Zhu, B. T-cell exhaustion in the tumor microenvironment. Cell Death Dis. 6, e1792 (2015).
Andrews, L. P. et al. LAG-3 and PD-1 synergize on CD8+T cells to drive T cell exhaustion and hinder autocrine IFN-y-dependent anti-tumor immunity. Cell 187, 4355–4372.e4322 (2024).
Chang, Y. C., Yang, Y. C., Tien, C. P., Yang, C. J. & Hsiao, M. Roles of aldolase family genes in human cancers and diseases. Trends Endocrinol. Metab. 29, 549–559 (2018).
Guo, M. et al. Molecular, metabolic, and functional CD4 T cell paralysis in the lymph node impedes tumor control. Cell Rep. 42, 113047 (2023).
Costa, C. et al. The RacGAP ArhGAP15 is a master negative regulator of neutrophil functions. Blood 118, 1099–1108 (2011).
Faroudi, M. et al. Critical roles for Rac GTPases in T-cell migration to and within lymph nodes. Blood 116, 5536–5547 (2010).
Lin, W. P., Li, H. & Sun, Z. J. T cell exhaustion initiates tertiary lymphoid structures and turbocharges cancer-immunity cycle. Ebiomedicine 104, 105154 (2024).
Roe, K. NK-cell exhaustion, B-cell exhaustion and T-cell exhaustion-the differences and similarities. Immunology 166, 155–168 (2022).
Ma, C. et al. Immunoglobulin G expression and its potential role in primary and metastatic breast cancers. Curr. Mol. Med. 13, 429–437 (2013).
Meylan, M., Sautès-Fridman, C. & Fridman, W. H. Tertiary lymphoid structures generate and propagate anti-tumor antibody-producing plasma cells in renal cell cancer. Med. Sci. 38, 536–538 (2022).
Xiao, Z. et al. Desmoplastic stroma restricts T cell extravasation and mediates immune exclusion and immunosuppression in solid tumors. Nat. Commun. 14, 5110 (2023).
Zilionis, R. et al. Single-cell transcriptomics of human and mouse lung cancers reveals conserved myeloid populations across individuals and species. Immunity 50, 1317–1334.e1310 (2019).
Laughney, A. M. et al. Regenerative lineages and immune-mediated pruning in lung cancer metastasis. Nat. Med. 26, 259–269 (2020).
Wu, L. H., Li, M. & Hung, H. Y. NRF2 upregulates GPX2 to inhibit ferroptosis and enhance immune escape of melanoma. J. Biol. Regul. Homeost. Agents 38, 4375–4385 (2024).
Tang, D. L., Chen, X., Kang, R. & Kroemer, G. Ferroptosis: molecular mechanisms and health implications. Cell Res. 31, 107–125 (2021).
Yan, Y. et al. Multi-omic profiling highlights factors associated with resistance to immuno-chemotherapy in non-small-cell lung cancer. Nat. Genet. 57, 126–139 (2025).
Chu, X. et al. Cancer stem cells: advances in knowledge and implications for cancer therapy. Signal Transduct. Target Ther. 9, 170 (2024).
Ubellacker, J. M. et al. Lymph protects metastasizing melanoma cells from ferroptosis. Nature 585, 113–118 (2020).
Li, Y. X. et al. 7-Dehydrocholesterol dictates ferroptosis sensitivity. Nature 626, 411–418 (2024).
Sun, C., Mezzadra, R. & Schumacher, T. N. Regulation and Function of the PD-L1 Checkpoint. Immunity 48, 434–452 (2018).
Miao, S. et al. Cancer cell-derived immunoglobulin G activates platelets by binding to platelet FcgammaRIIa. Cell Death Dis. 10, 87 (2019).
Liu, Y. et al. Single-cell and spatial transcriptome analyses reveal tertiary lymphoid structures linked to tumour progression and immunotherapy response in nasopharyngeal carcinoma. Nat. Commun. 15, 7713 (2024).
Xiaoxu, D., Min, X. & Chengcheng, C. Immature central tumor tertiary lymphoid structures are associated with better prognosis in non-small cell lung cancer. BMC Pulm. Med. 24, 155 (2024).
Sun, X. et al. Maturation and abundance of tertiary lymphoid structures are associated with the efficacy of neoadjuvant chemoimmunotherapy in resectable non-small cell lung cancer. J. Immunother. Cancer 10, e005531 (2022).
Wei, C. Y. et al. Delineating the early dissemination mechanisms of acral melanoma by integrating single-cell and spatial transcriptomic analyses. Nat. Commun. 14, 8119 (2023).
Cha, S. M. et al. SPP1+ macrophages in HR+ breast cancer are associated with tumor-infiltrating lymphocytes. NPJ Breast Cancer 10, 83 (2024).
Wu, Y. C. et al. Spatiotemporal immune landscape of colorectal cancer liver metastasis at single-cell level. Cancer Discov. 12, 134–153 (2022).
Zhang, Y. et al. Single-cell analyses reveal key immune cell subsets associated with response to PD-L1 blockade in triple-negative breast cancer. Cancer Cell 39, 1578–1593.e1578 (2021).
Chen, J. et al. Single-cell profiling of tumor immune microenvironment reveals immune irresponsiveness in gastric signet-ring cell carcinoma. Gastroenterology 165, 88–103 (2023).
Zhang, Q. et al. Iron promotes ovarian cancer malignancy and advances platinum resistance by enhancing DNA repair via FTH1/FTL/POLQ/RAD51 axis. Cell Death Dis. 15, 329 (2024).
Zhang, X. Y. et al. CircPIAS1 promotes hepatocellular carcinoma progression by inhibiting ferroptosis via the miR-455-3p/NUPR1/FTH1 axis. Mol. Cancer 23, 113 (2024).
Dura, B. et al. scFTD-seq: freeze-thaw lysis based, portable approach toward highly distributed single-cell 3 mRNA profiling. Nucleic Acids Res. 47, e16 (2019).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Germain, P. L., Lun, A., Garcia, M. eixideC., Macnair, W. & Robinson, M. D. Doublet identification in single-cell sequencing data using scDblFinder. F1000Res. 10, 979 (2021).
Qiu, X. et al. Single-cell mRNA quantification and differential analysis with Census. Nat. Methods 14, 309–315 (2017).
Yu, L., Cao, Y., Yang, J. Y. H. & Yang, P. Benchmarking clustering algorithms on estimating the number of cell types from single-cell RNA-sequencing data. Genome Biol. 23, 49 (2022).
Haghverdi, L., Lun, A. T. L., Morgan, M. D. & Marioni, J. C. Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors. Nat. Biotechnol. 36, 421–427 (2018).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
Gulati, G. S. et al. Single-cell transcriptional diversity is a hallmark of developmental potential. Science 367, 405–411 (2020).
Vogl, T. et al. S100A12 is expressed exclusively by granulocytes and acts independently from MRP8 and MRP14. J. Biol. Chem. 274, 25291–25296 (1999).
Cooper, M. A. et al. Human natural killer cells: a unique innate immunoregulatory role for the CD56(bright) subset. Blood 97, 3146–3151 (2001).
Fitzsimons, E. et al. A pan-cancer single-cell RNA-seq atlas of intratumoral B cells. Cancer cell 42, 1784–1797.e1784 (2024).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e3529 (2021).
Ma, Y. & Zhou, X. Spatially informed cell-type deconvolution for spatial transcriptomics. Nat. Biotechnol. 40, 1349–1359 (2022).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Wilkerson, M. D. & Hayes, D. N. ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics 26, 1572–1573 (2010).
Tay, J. K., Narasimhan, B. & Hastie, T. Elastic net regularization paths for all generalized linear models. J. Stat. Softw. 106, 1–31 (2023).
Silva, T. C. et al. TCGA Workflow: analyze cancer genomics and epigenomics data using Bioconductor packages. F1000Res. 5, 1542 (2016).
Sean, D. & Meltzer, P. S. GEOquery: a bridge between the gene expression omnibus (GEO) and BioConductor. Bioinformatics 23, 1846–1847 (2007).
Members & Partners, C.-N. Database resources of the national genomics data center, china national center for bioinformation in 2024. Nucleic Acids Res. 52, D18–D32 (2024).
Chen, D. Dissecting origin factors of lymph node metastasis in non-small cell lung cancer via multimodal omics. Github. https://doi.org/10.5281/zenodo.18690124 (2025).
Acknowledgements
This study is supported by National Natural Science Foundation of China grants (No. 32470832 to H.-l.P., No. 82073286 to H.-X.L.), Liaoning Revitalization Talents Program (XLYC2002035 to H.-l.P.); Liaoning Science and Technology Innovation Funding (20230101-JH2/1013 to H.-l.P.); The Construction of Liaoning Cancer Research Center (Lung Cancer) (2019JH6/10200011 to H.-X.L.); Technological Special Project of Liaoning Province of China (2019020176-JH1/103 to H.-X.L.); Liaoning Provincial Joint Science and Technology Program (Technical R&D Project) (2024JH2/102600187 to S.L.); Innovation program of science and research from the DICP, CAS (DICP I202209 to H-l.P., DICP I202414 to D.C.).
Author information
Authors and Affiliations
Contributions
H.-l.P., D.C. and H.-X.L. conceived the project, H.-l.P. and H.-X.L. supervised the project. D.C., Yawei W., H-.X.L. and H.-l.P. the designed project and Y.L., D.S., S.L. and D.C. performed most of the experiments, D.C. performed computational data analysis, D.C., H.-X.L. and H.-l.P. analyzed data. Q.W., T.F., S.P., Yuhan W., Y.Z., X.S., Z.H., J.L., Y.Z., N.S., W.W., L.Z. and Y.M. provided significant intellectual input. D.C., H.-X.L. and H.-l.P. wrote the manuscript with input from all other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks anonymous reviewers for their contribution to the peer review of this work. [A peer review file is available.]
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, D., Liu, Y., Shi, D. et al. Dissecting origin factors of lymph node metastasis in non-small cell lung cancer via multimodal omics. Nat Commun (2026). https://doi.org/10.1038/s41467-026-71819-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-71819-9