Abstract
Aquatic environments absorb ~2.5 gigatonnes of atmospheric carbon each year1, more than the carbon stored in the atmosphere, soils, and all biomass combined. Primary producers transform this dissolved inorganic carbon into biomass that can subsequently flow into other trophic levels, or be released back into the environment through viral lysis. While there is substantial knowledge about the diversity and activity of viruses infecting photoautotrophic primary producers and the ecosystem impact, little is known about viruses infecting chemoautotrophs, representing a gap in our understanding of key processes driving microbial carbon cycling. Here, we combine metagenomics with quantitative 12/13C stable isotopic probing (qSIP) mesocosm experiments in a marine-derived meromictic pond to quantify population-specific isotopic enrichment, identify key chemoautotrophic primary producers, and virus-host dynamics. Isotopically enriched carbon is tracked from the genomes of chemoautotrophs to putative viruses, showing that active populations of hydrogen/sulfur-oxidizing chemoautotrophs (Thiomicrorhabdus, Hydrogenovibrio, Sulfurimonas, Sulfurovum) are targeted by viruses. This work provides the foundation for revealing the diversity and activity of viruses infecting globally-widespread chemoautotrophs. Our study sheds light on trophic interactions that impact microbial carbon cycling in aphotic environments and builds toward biogeochemical models that incorporate viral impacts on chemoautotrophic microbial communities.
Similar content being viewed by others
Data availability
Raw sequence data for this project is available on NCBI under Bioproject PRJNA1390687. Source data are provided with this paper. All other data is provided with the paper or appended as supplementary data. Source data are provided with this paper.
References
Friedlingstein, P. et al. Global Carbon Budget 2019. Earth Syst. Sci. Data 11, 1783–1838 (2019).
Ricci, F. & Greening, C. Chemosynthesis: a neglected foundation of marine ecology and biogeochemistry. Trends Microbiol 32, 631–639 (2024).
Hadas, O., Pinkas, R. & Erez, J. High chemoautotrophic primary production in Lake Kinneret, Israel: A neglected link in the carbon cycle of the lake. Limnol. Oceanogr. 46, 1968–1976 (2001).
Ulloa, O., Canfield, D. E., DeLong, E. F., Letelier, R. M. & Stewart, F. J. Microbial oceanography of anoxic oxygen minimum zones. Proc. Natl. Acad. Sci. 109, 15996–16003 (2012).
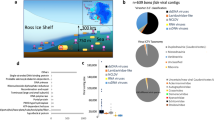
Domack, E. et al. A chemotrophic ecosystem found beneath Antarctic Ice Shelf. Eos Trans. Am. Geophys. Union 86, 269–272 (2005).
Overholt, W. A. et al. Carbon fixation rates in groundwater similar to those in oligotrophic marine systems. Nat. Geosci. 15, 561–567 (2022).
Farías, L., Fernández, C., Faúndez, J., Cornejo, M. & Alcaman, M. E. Chemolithoautotrophic production mediating the cycling of the greenhouse gases N2 O and CH4 in an upwelling ecosystem. Biogeosciences 6, 3053–3069 (2009).
Suttle, C. A. Cyanophages and Their Role in the Ecology of Cyanobacteria. In The Ecology of Cyanobacteria (eds Whitton, B. A. & Potts, P.) 563–589 (Kluwer Academic Publishers, 2002).
Fuhrman, J. A. Marine viruses and their biogeochemical and ecological effects. Nature 399, 541–548 (1999).
Wommack, K. E. & Colwell, R. R. Virioplankton: Viruses in Aquatic Ecosystems. Microbiol. Mol. Biol. Rev. 64, 69–114 (2000).
Wilhelm, S. & Suttle, C. A. Viruses and Nutrient Cycles in the Sea. BioScience 49, 781–788 (1999).
Hevroni, G., Vincent, F., Ku, C., Sheyn, U. & Vardi, A. Daily turnover of active giant virus infection during algal blooms revealed by single-cell transcriptomics. Sci. Adv. 9, eadf7971 (2023).
Suttle, C. A. Marine viruses — major players in the global ecosystem. Nat. Rev. Microbiol. 5, 801–812 (2007).
Lara, E. et al. Unveiling the role and life strategies of viruses from the surface to the dark ocean. Sci. Adv. 3, e1602565 (2017).
Han, Y. & Perner, M. The globally widespread genus Sulfurimonas: versatile energy metabolisms and adaptations to redox clines. Front. Microbiol. 6, 989 (2015).
Biderre-Petit, C. et al. A pan-genomic approach reveals novel Sulfurimonas clade in the ferruginous meromictic Lake Pavin. Mol. Ecol. Resour. 24, e13923 (2024).
Sass, K. & Perner, M. Characterization of Two Hydrogen-Oxidizing Hydrogenovibrio Strains From Kermadec Volcanic Island Arc Hydrothermal Vents. Front. Mar. Sci. 7, 295 (2020).
Scott, K. M. et al. Diversity in CO2 -Concentrating Mechanisms among Chemolithoautotrophs from the Genera. Hydrogenovibrio 85, e02096–18 (2019).
Klepac-Ceraj, V. et al. Microbial diversity under extreme euxinia: Mahoney Lake, Canada. Geobiology 10, 223–235 (2012).
Grote, J., Jost, G., Labrenz, M., Herndl, G. J. & Jürgens, K. Epsilonproteobacteria Represent the Major Portion of Chemoautotrophic Bacteria in Sulfidic Waters of Pelagic Redoxclines of the Baltic and Black Seas. Appl. Environ. Microbiol. 74, 7546–7551 (2008).
Glaubitz, S. et al. 13C-isotope analyses reveal that chemolithoautotrophic Gamma- and Epsilonproteobacteria feed a microbial food web in a pelagic redoxcline of the central Baltic Sea. Environ. Microbiol. 11, 326–337 (2009).
Labrenz, M. et al. Sulfurimonas gotlandica sp. nov., a chemoautotrophic and psychrotolerant epsilonproteobacterium isolated from a pelagic redoxcline, and an emended description of the genus Sulfurimonas. Int. J. Syst. Evol. Microbiol 63, 4141–4148 (2013).
Perner, M. et al. In situ chemistry and microbial community compositions in five deep-sea hydrothermal fluid samples from Irina II in the Logatchev field. Environ. Microbiol. 15, 1551–1560 (2013).
Akerman, N. H., Butterfield, D. A. & Huber, J. A. Phylogenetic diversity and functional gene patterns of sulfur-oxidizing subseafloor Epsilonproteobacteria in diffuse hydrothermal vent fluids. Front. Microbiol. 4, 185 (2013).
Perner, M. et al. Linking geology, fluid chemistry, and microbial activity of basalt- and ultramafic-hosted deep-sea hydrothermal vent environments. Geobiology 11, 340–355 (2013).
Laber, C. P. et al. Coccolithovirus facilitation of carbon export in the North Atlantic. Nat. Microbiol. 3, 537–547 (2018).
Tran, P. Q. & Anantharaman, K. Biogeochemistry Goes Viral: towards a Multifaceted Approach To Study Viruses and Biogeochemical Cycling. mSystems 6 (2021).
Locke, H. et al. Chapter Two - Marine viruses and climate change: Virioplankton, the carbon cycle, and our future ocean. In Advances in Virus Research, Viruses and Climate Change (ed. Roossinck, M. J.) 67–146 (Academic Press, 2022).
Luo, E. et al. Double-stranded DNA virioplankton dynamics and reproductive strategies in the oligotrophic open ocean water column. ISME J. 14, 1304–1315 (2020).
Labonté, J. M. et al. Single cell genomics-based analysis of gene content and expression of prophages in a diffuse-flow deep-sea hydrothermal system. Front. Microbiol. 10, 1262 (2019).
Rastelli, E. et al. High potential for temperate viruses to drive carbon cycling in chemoautotrophy-dominated shallow-water hydrothermal vents. Environ. Microbiol. 19, 4432–4446 (2017).
Jian, H. et al. Diversity and distribution of viruses inhabiting the deepest ocean on Earth. ISME J. 15, 3094–3110 (2021).
Kieft, K. et al. Ecology of inorganic sulfur auxiliary metabolism in widespread bacteriophages. Nat. Commun. 12, 3503 (2021).
Roux, S. et al. Ecology and evolution of viruses infecting uncultivated SUP05 bacteria as revealed by single-cell- and meta-genomics. eLife 3, e03125 (2014).
Castelán-Sánchez, H. G. et al. Extremophile deep-sea viral communities from hydrothermal vents: Structural and functional analysis. Mar. Genomics 46, 16–28 (2019).
Cheng, R. et al. Virus diversity and interactions with hosts in deep-sea hydrothermal vents. Microbiome 10, 235 (2022).
Li, X., Cheng, R., Zhang, C. & Shao, Z. Genomic diversity of phages infecting the globally widespread genus Sulfurimonas. Commun. Biol. 7, 1428 (2024).
Starr, E. P. et al. Stable-Isotope-Informed, Genome-Resolved Metagenomics Uncovers Potential Cross-Kingdom Interactions in Rhizosphere Soil. mSphere 6, e00085–21 (2021).
Trubl, G. et al. Active virus-host interactions at sub-freezing temperatures in Arctic peat soil. Microbiome 9, 208 (2021).
Greenlon, A. et al. Quantitative Stable-Isotope Probing (qSIP) with Metagenomics Links Microbial Physiology and Activity to Soil Moisture in Mediterranean-Climate Grassland Ecosystems. mSystems 7, e00417–e00422 (2022).
Lee, S. et al. Methane-derived carbon flows into host–virus networks at different trophic levels in soil. Proc. Natl. Acad. Sci. 118, e2105124118 (2021).
Wikner, J., Vallino, J. J., Steward, G. F., Smith, D. C. & Azam, F. Nucleic acids from the host bacterium as a major source of nucleotides for three marine bacteriophages. FEMS Microbiol. Ecol. 12, 237–248 (1993).
Ostrander, C. M. et al. Thallium isotope cycling between waters, particles, and sediments across a redox gradient. Geochim. Cosmochim. Acta 348, 397–409 (2023).
Vallino, J. J. & Huber, J. A. Using maximum entropy production to describe microbial biogeochemistry over time and space in a meromictic pond. Front. Environ. Sci. 6, 100 (2018).
Hungate, B. A. et al. Quantitative Microbial Ecology through Stable Isotope Probing. Appl. Environ. Microbiol. 81, 7570–7581 (2015).
Vyshenska, D. et al. A standardized quantitative analysis strategy for stable isotope probing metagenomics. mSystems e01280-22. https://doi.org/10.1128/msystems.01280-22. (2023).
Guo, J., Vik, D., Pratama, A. A., Roux, S. & Sullivan, M. Viral sequence identification SOP with VirSorter2 V.3. https://doi.org/10.17504/protocols.io.bwm5pc86.(2021).
Watanabe, T., Kubo, K., Kamei, Y., Kojima, H. & Fukui, M. Dissimilatory microbial sulfur and methane metabolism in the water column of a shallow meromictic lake. Syst. Appl. Microbiol. 45, 126320 (2022).
Updegraff, T. et al. Thiomicrorhabdus heinhorstiae sp. nov. and Thiomicrorhabdus cannonii sp. nov.: novel sulphur-oxidizing chemolithoautotrophs isolated from the chemocline of Hospital Hole, an anchialine sinkhole in Spring Hill, Florida, USA. Int. J. Syst. Evol. Microbiol. 72, 3 (2022).
Savvichev, A. S. et al. Microbial Processes and Microbial Communities in the Water Column of the Polar Meromictic Lake Bol’shie Khruslomeny at the White Sea Coast. Front. Microbiol. 11, 1945 (2020).
Gregersen, L. H. et al. Dominance of a clonal green sulfur bacterial population in a stratified lake: Clonal dominance in microbial lake community. FEMS Microbiol. Ecol. 70, 30–41 (2009).
Lauro, F. M. et al. An integrative study of a meromictic lake ecosystem in Antarctica. ISME J. 5, 879–895 (2011).
Tran, P. Q. et al. Depth-discrete metagenomics reveals the roles of microbes in biogeochemical cycling in the tropical freshwater Lake Tanganyika. ISME J. 15, 1971–1986 (2021).
Block, K. R., O’Brien, J. M., Edwards, W. J. & Marnocha, C. L. Vertical structure of the bacterial diversity in meromictic Fayetteville Green Lake. Microbiol. Open 10, e1228 (2021).
Phillips, A. A. et al. Microbial succession and dynamics in meromictic Mono Lake, California. Geobiology 19, 376–393 (2021).
Saini, J. S. et al. Bacterial, phytoplankton, and viral distributions and their biogeochemical contexts in meromictic lake cadagno offer insights into the proterozoic ocean microbial loop. mBio 13, e00052–22 (2022).
Cabello-Yeves, P. J., Picazo, A., Roda-Garcia, J. J., Rodriguez-Valera, F. & Camacho, A. Vertical niche occupation and potential metabolic interplay of microbial consortia in a deeply stratified meromictic model lake. Limnol. Oceanogr. 68, 2492–2511 (2023).
Overmann, J., Cypionka, H. & Pfenning, N. An extremely low-light adapted phototrophic sulfur bacterium from the Black Sea. Limnol. Oceanogr. 37, 150–155 (1992).
Manske, A. K., Glaeser, J., Kuypers, M. M. M. & Overmann, J. Physiology and phylogeny of green sulfur bacteria forming a monospecific phototrophic assemblage at a depth of 100 meters in the black sea. Appl. Environ. Microbiol. 71, 8049–8060 (2005).
Bao, N. & Gao, K. Interactive Effects of Elevated CO2 Concentration and Light on the Picophytoplankton Synechococcus. Front. Mar. Sci. 8, 634189 (2021).
Puxty, R. J., Millard, A. D., Evans, D. J. & Scanlan, D. J. Viruses inhibit CO2 fixation in the most abundant phototrophs on earth. Curr. Biol. 26, 1585–1589 (2016).
Zhong, K. X., Wirth, J. F., Chan, A. M. & Suttle, C. A. Mortality by ribosomal sequencing (MoRS) provides a window into taxon-specific cell lysis. ISME J. 17, 105–116 (2023).
Eren, A. M. et al. Anvi’o: an advanced analysis and visualization platform for ‘omics data. PeerJ 3, e1319 (2015).
Weinbauer, M. G. Ecology of prokaryotic viruses. FEMS Microbiol. Rev. 28, 127–181 (2004).
Suttle, C. A. & Chen, F. Mechanisms and rates of decay of marine viruses in seawater. Appl. Environ. Microbiol. 58, 3721–3729 (1992).
Martínez Martínez, J., Talmy, D., Kimbrel, J. A., Weber, P. K. & Mayali, X. Coastal bacteria and protists assimilate viral carbon and nitrogen. ISME J. 18, wrae231 (2024).
Amend, A. From dandruff to deep-sea vents: Malassezia-like fungi are ecologically hyper-diverse. PLoS Pathog. 10, e1004277 (2014).
Kleindienst, S. et al. Diverse sulfate-reducing bacteria of the Desulfosarcina/Desulfococcus clade are the key alkane degraders at marine seeps. ISME J. 8, 2029–2044 (2014).
Dunford, E. A. & Neufeld, J. D. DNA Stable-Isotope Probing (DNA-SIP). J. Vis. Exp. 2027. https://doi.org/10.3791/2027.(2010).
Buckley, D. H., Huangyutitham, V., Hsu, S.-F. & Nelson, T. A. Stable Isotope Probing with15 N Achieved by Disentangling the Effects of Genome G+C Content and Isotope Enrichment on DNA Density. Appl. Environ. Microbiol. 73, 3189–3195 (2007).
Bushnell, B. BBMap: A. Fast, accurate, splice-aware aligner in Report Number: LBNL-7065E, (17 AD).
Li, D., Liu, C.-M., Luo, R., Sadakane, K. & Lam, T.-W. MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31, 1674–1676 (2015).
Guo, J. et al. VirSorter2: a multi-classifier, expert-guided approach to detect diverse DNA and RNA viruses. Microbiome 9, 37 (2021).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075 (2013).
Nayfach, S. et al. CheckV assesses the quality and completeness of metagenome-assembled viral genomes. Nat. Biotechnol. 39, 578–585 (2021).
Luo, E., Leu, A. O., Eppley, J. M., Karl, D. M. & DeLong, E. F. Diversity and origins of bacterial and archaeal viruses on sinking particles reaching the abyssal ocean. ISME J. 16, 1627–1635 (2022).
Camargo, A. P. et al. Identification of mobile genetic elements with geNomad. Nat. Biotechnol. 42, 1303–1312 (2024).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinforma. 11, 119 (2010).
Sayers, E. W. et al. GenBank 2024 Update. Nucleic Acids Res. 52, D134–D137 (2024).
Camargo, A. P. et al. IMG/VR v4: an expanded database of uncultivated virus genomes within a framework of extensive functional, taxonomic, and ecological metadata. Nucleic Acids Res. 51, D733–D743 (2023).
Buchfink, B., Reuter, K. & Drost, H.-G. Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat. Methods 18, 366–368 (2021).
Figueroa, J. L. III, Dhungel, E., Bellanger, M., Brouwer, C. R. & White, R. A. III MetaCerberus: distributed highly parallelized HMM-based processing for robust functional annotation across the tree of life. Bioinformatics 40, btae119 (2024).
Kanehisa, M. & Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27–30 (2000).
Galperin, M. Y. et al. COG database update 2024. Nucleic Acids Res. 53, D356–D363 (2025).
Grazziotin, A. L., Koonin, E. V. & Kristensen, D. M. Prokaryotic Virus Orthologous Groups (pVOGs): a resource for comparative genomics and protein family annotation. Nucleic Acids Res. 45, D491–D498 (2017).
Terzian, P. et al. PHROG: families of prokaryotic virus proteins clustered using remote homology. NAR Genomics Bioinforma. 3, lqab067 (2021).
Mistry, J. et al. Pfam: The protein families database in 2021. Nucleic Acids Res. 49, D412–D419 (2021).
Luo, E., Aylward, F. O., Mende, D. R. & DeLong, E. F. Bacteriophage Distributions and Temporal Variability in the Ocean’s Interior. mBio 8, e01903–e01917 (2017).
Couvin, D. et al. CRISPRCasFinder, an update of CRISRFinder, includes a portable version, enhanced performance and integrates search for Cas proteins. Nucleic Acids Res. 46, W246–W251 (2018).
Skennerton, C. T., Imelfort, M. & Tyson, G. W. Crass: identification and reconstruction of CRISPR from unassembled metagenomic data. Nucleic Acids Res. 41, e105–e105 (2013).
Menzel, P., Ng, K. L. & Krogh, A. Fast and sensitive taxonomic classification for metagenomics with Kaiju. Nat. Commun. 7, 11257 (2016).
Uritskiy, G. V., DiRuggiero, J. & Taylor, J. MetaWRAP—a flexible pipeline for genome-resolved metagenomic data analysis. Microbiome 6, 158 (2018).
Wu, Y.-W., Simmons, B. A. & Singer, S. W. MaxBin 2.0: an automated binning algorithm to recover genomes from multiple metagenomic datasets. Bioinformatics 32, 605–607 (2016).
Kang, D. D. et al. MetaBAT 2: an adaptive binning algorithm for robust and efficient genome reconstruction from metagenome assemblies. PeerJ 7, e7359 (2019).
Alneberg, J. et al. Binning metagenomic contigs by coverage and composition. Nat. Methods 11, 1144–1146 (2014).
Olm, M. R., Brown, C. T., Brooks, B. & Banfield, J. F. dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J. 11, 2864–2868 (2017).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Katoh, K. & Standley, D. M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 30, 772–780 (2013).
Minh, B. Q. et al. IQ-TREE 2: New Models and Efficient Methods for Phylogenetic Inference in the Genomic Era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Parks, D. H. et al. GTDB: an ongoing census of bacterial and archaeal diversity through a phylogenetically consistent, rank normalized and complete genome-based taxonomy. Nucleic Acids Res. 50, D785–D794 (2022).
Shaffer, M. et al. DRAM for distilling microbial metabolism to automate the curation of microbiome function. Nucleic Acids Res. 48, 8883–8900 (2020).
Tenenbaum, D. & Maintainer, B. KEGGREST: Client-side REST access to the Kyoto Encyclopedia of Genes and Genomes (KEGG). R Package Version 1480 (2025).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
R: A language and environment for statistical computing. R Found. Stat. Comput. Vienna Austria (2023).
Okasen, J. et al. vegan: Community Ecology Package. R Package Version 26-10 (2024).
Shen, W. et al. SeqKit: A Cross-Platform and Ultrafast Toolkit for FASTA/Q File Manipulation. PLOS ONE 11, e0163962 (2016).
Muggeo, V. M. R. Interval estimation for the breakpoint in segmented regression: a smoothed score-based approach. Aust. N. Z. J. Stat. 59, 311–322 (2017).
Lind, A. L. & Pollard, K. S. Accurate and sensitive detection of microbial eukaryotes from whole metagenome shotgun sequencing. Microbiome 9, 58 (2021).
Xie, C., Goi, C. L. W., Huson, D. H., Little, P. F. R. & Williams, R. B. H. RiboTagger: fast and unbiased 16S/18S profiling using whole community shotgun metagenomic or metatranscriptome surveys. BMC Bioinforma. 17, 508 (2016).
Acknowledgements
We would like to acknowledge Gretta Serres, Alex Worden, Matthew Johnson, Sabrina Elkassas, Cynthia Becker, and Amy Apprill for discussions and/or methodology relevant to this manuscript. The data generation, analyses, and manuscript were supported by the University of North Carolina at Charlotte (startup funds to E.L.), the Hypothesis Fund (to E.L.), and the National Science Foundation (OCE-2513189 to E.L.). Wet lab analyses were supported by the Woods Hole Oceanographic Institution (Weston Howland Jr. Postdoctoral Fellowship to E.L.), the National Oceanic and Atmospheric Administration (NA19OAR4320072 subaward 0007525/102212019 to J.A.H.), and the National Science Foundation (OCE-1947776 to J.A.H.). B.E.B. was supported by the National Science Foundation (PRFB2010963). JJV was supported by the Simons Foundation (549941FY22). G.T. was supported by the US Department of Energy (SCW1632) and was conducted under Contract DE-AC52-07NA27344.
Author information
Authors and Affiliations
Contributions
E.L. conceptualized, supervised, and acquired funding for the study with input from J.A.H.; E.L. and B.E.B. collected the samples with help from JV and JAH. EL conducted experiments and wet lab analyses with input from G.T. and J.A.H.; N.P., T.J.R., M.M.S., J.J.V., and E.L. conducted the data analyses. E.L. wrote the manuscript with input from all co-authors. Correspondence and requests for materials should be addressed to elaine.luo@charlotte.edu.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Niculina Musat and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Luo, E., Pham, N.D., Rogers, T.J. et al. Quantitative stable isotope probing (qSIP)-informed metagenomics identifies viruses infecting chemoautotrophs. Nat Commun (2026). https://doi.org/10.1038/s41467-026-71833-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-71833-x