Abstract
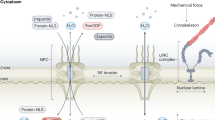
The nuclear pore complex (NPC) is a cornerstone of eukaryotic cell functionality, orchestrating the nucleocytoplasmic shuttling of macromolecules. Here we report that Plant Nuclear Envelope Transmembrane 1 (PNET1), a transmembrane nucleoporin, is an adaptable NPC component that is mainly expressed in actively dividing cells. PNET1’s selective incorporation into the NPC is required for rapid cell growth in highly proliferative meristem and callus tissues in Arabidopsis. We demonstrate that the cell cycle-dependent phosphorylation of PNET1 coordinates mitotic disassembly and post-mitotic reassembly of NPCs during the cell cycle. PNET1 hyperphosphorylation disrupts its interaction with the NPC scaffold, facilitating NPC dismantling and nuclear membrane breakdown to trigger mitosis. In contrast, nascent, unphosphorylated PNET1 is incorporated into the nuclear pore membrane in the daughter cells, where it restores interactions with scaffolding nucleoporins for NPC reassembly. The expression of the human PNET1 homologue is required for and markedly upregulated during cancer cell growth, suggesting that PNET1 plays a conserved role in facilitating rapid cell division during open mitosis in highly proliferative tissues.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The raw data files for the RNA-seq analysis have been deposited in the NCBI GEO under accession number GSE273725 and are publicly available. The raw data files for all mass spectrometry analyses have been deposited in the ProteomeXchange Consortium via the PRIDE partner repository (identifiers PXD054475 and PXD054530) and are publicly available. Source data are provided with this paper.
References
Mitchison, T. J. & Salmon, E. D. Mitosis: a history of division. Nat. Cell Biol. 3, E17–E21 (2001).
Inzé, D. & De Veylder, L. Cell cycle regulation in plant development. Annu. Rev. Genet. 40, 77–105 (2006).
Kastan, M. & Bartek, J. Cell-cycle checkpoints and cancer. Nature 432, 316–323 (2004).
Ankers et al. Dynamic NF-κB and E2F interactions control the priority and timing of inflammatory signalling and cell proliferation. eLife 5, e10473 (2016).
Chauhan, M., Modi, P. K. & Sharma, P. Aberrant activation of neuronal cell cycle caused by dysregulation of ubiquitin ligase Itch results in neurodegeneration. Cell Death Dis. 11, 441 (2020).
Sozzani, R. et al. Spatiotemporal regulation of cell-cycle genes by SHORTROOT links patterning and growth. Nature 466, 128–132 (2010).
Takahashi, N. et al. A regulatory module controlling stress-induced cell cycle arrest in Arabidopsis. eLife 8, e43944 (2019).
Ikeuchi, M., Sugimoto, K. & Iwase, A. Plant callus: mechanisms of induction and repression. Plant Cell 25, 3159–3173 (2013).
Chen, C. et al. Plant regeneration in the new era: from molecular mechanisms to biotechnology applications. Sci. China Life Sci. 67, 1338–1367 (2024).
Shen, J. et al. Translational repression by a miniature inverted-repeat transposable element in the 3′ untranslated region. Nat. Commun. 8, 14651 (2017).
Güttinger, S., Laurell, E. & Kutay, U. Orchestrating nuclear envelope disassembly and reassembly during mitosis. Nat. Rev. Mol. Cell Biol. 10, 178–191 (2009).
Kutay, U., Jühlen, R. & Antonin, W. Mitotic disassembly and reassembly of nuclear pore complexes. Trends Cell Biol. 31, 1019–1033 (2021).
Laurell, E. et al. Phosphorylation of Nup98 by multiple kinases is crucial for NPC disassembly during mitotic entry. Cell 144, 539–550 (2011).
Linder, M. I. et al. Mitotic disassembly of nuclear pore complexes involves CDK1- and PLK1-mediated phosphorylation of key interconnecting nucleoporins. Dev. Cell 43, 141–156.e7 (2017).
Ellenberg, J. et al. Nuclear membrane dynamics and reassembly in living cells: targeting of an inner nuclear membrane protein in interphase and mitosis. J. Cell Biol. 138, 1193–1206 (1997).
Yang, L., Guan, T. & Gerace, L. Integral membrane proteins of the nuclear envelope are dispersed throughout the endoplasmic reticulum during mitosis. J. Cell Biol. 137, 1199–1210 (1997).
Hashizume, C. et al. Nucleoporin Nup62 maintains centrosome homeostasis. Cell Cycle 12, 3804–3816 (2013).
Hartono et al. Nucleoporin Nup58 localizes to centrosomes and mid-bodies during mitosis. Cell Division 14, 7 (2019).
Goto, C., Hashizume, S., Fukao, Y., Hara-Nishimura, I. & Tamura, K. Comprehensive nuclear proteome of Arabidopsis obtained by sequential extraction. Nucleus 10, 81–92 (2019).
Tang, Y., Huang, A. & Gu, Y. Global profiling of plant nuclear membrane proteome in Arabidopsis. Nat. Plants 6, 838–847 (2020).
Fujitomo, T., Daigo, Y., Matsuda, K., Ueda, K. & Nakamura, Y. Critical function for nuclear envelope protein TMEM209 in human pulmonary carcinogenesis. Cancer Res. 72, 4110–4118 (2012).
Desvoyes, B., Arana-Echarri, A., Barea, M. D. & Gutierrez, C. A comprehensive fluorescent sensor for spatiotemporal cell cycle analysis in Arabidopsis. Nat. Plants 6, 1330–1334 (2020).
Simonini, S. et al. The Polycomb group protein MEDEA controls cell proliferation and embryonic patterning in Arabidopsis. Dev. Cell 56, 1945–1960.e7 (2021).
D’Ario, M. et al. Cell size controlled in plants using DNA content as an internal scale. Science 372, 1176–1181 (2021).
Cools, T., Iantcheva, A., Maes, S., Van den Daele, H. & De Veylder, L. A replication stress-induced synchronization method for Arabidopsis thaliana root meristems. Plant J. 64, 705–714 (2010).
Goldy, C. et al. The Arabidopsis GRAS-type SCL28 transcription factor controls the mitotic cell cycle and division plane orientation. Proc. Natl Acad. Sci. USA 118, e2005256118 (2021).
Ryu, K. H., Huang, L., Kang, H. M. & Schiefelbein, J. Single-cell RNA sequencing resolves molecular relationships among individual plant cells. Plant Physiol. 179, 1444–1456 (2019).
Wendrich, J. R. et al. Vascular transcription factors guide plant epidermal responses to limiting phosphate conditions. Science 370, eaay4970 (2020).
Zhang, T. Q., Xu, Z. G., Shang, G. D. & Wang, J. W. A single-cell RNA sequencing profiles the developmental landscape of Arabidopsis root. Mol. Plant 12, 648–660 (2019).
Denyer, T. et al. Spatiotemporal developmental trajectories in the Arabidopsis root revealed using high-throughput single-cell RNA sequencing. Dev. Cell 48, 840–852.e5 (2019).
Jean-Baptiste, K. et al. Dynamics of gene expression in single root cells of Arabidopsis thaliana. Plant Cell 31, 993–1011 (2019).
Shulse, C. N. et al. High-throughput single-cell transcriptome profiling of plant cell types. Cell Rep. 27, 2241–2247.e4 (2019).
Shahan, R. et al. A single-cell Arabidopsis root atlas reveals developmental trajectories in wild-type and cell identity mutants. Dev. Cell 57, 543–560.e9 (2022).
Xu, X. et al. Large-scale single-cell profiling of stem cells uncovers redundant regulators of shoot development and yield trait variation. Preprint at bioRxiv https://doi.org/10.1101/2024.03.04.583414 (2024).
Liu, W. et al. Transcriptional landscapes of de novo root regeneration from detached Arabidopsis leaves revealed by time-lapse and single-cell RNA sequencing analyses. Plant Commun. 3, 100306 (2022).
Ogura, N. et al. WUSCHEL-RELATED HOMEOBOX 13 suppresses de novo shoot regeneration via cell fate control of pluripotent callus. Sci. Adv. 9, eadg6983 (2023).
Zhang, T. Q., Chen, Y. & Wang, J. W. A single-cell analysis of the Arabidopsis vegetative shoot apex. Dev. Cell 56, 1056–1074.e8 (2021).
Xu, J., Lee, Y. J. & Liu, B. Establishment of a mitotic model system by transient expression of the D-type cyclin in differentiated leaf cells of tobacco (Nicotiana benthamiana). New Phytol. 226, 1213–1220 (2020).
Tang, Y. et al. Proxiome assembly of the plant nuclear pore reveals an essential hub for gene expression regulation. Nat. Plants 10, 1005–1017 (2024).
Mosalaganti, S. et al. AI-based structure prediction empowers integrative structural analysis of human nuclear pores. Science 376, eabm9506 (2022).
Doucet, C. M., Talamas, J. A. & Hetzer, M. W. Cell cycle-dependent differences in nuclear pore complex assembly in metazoa. Cell 141, 1030–1041 (2010).
Mitchell, J. M., Mansfeld, J., Capitanio, J., Kutay, U. & Wozniak, R. W. Pom121 links two essential subcomplexes of the nuclear pore complex core to the membrane. J. Cell Biol. 191, 505–521 (2010).
Mansfeld, J. et al. The conserved transmembrane nucleoporin NDC1 is required for nuclear pore complex assembly in vertebrate cells. Mol. Cell 22, 93–103 (2006).
Gu, Y. et al. Nuclear pore permeabilization is a convergent signaling event in effector-triggered immunity. Cell 166, 1526–1538.e11 (2016).
Nkombo Nkoula, S. et al. Mechanisms of nuclear pore complex disassembly by the mitotic Polo-like kinase 1 (PLK-1) in C. elegans embryos. Sci. Adv. 9, eadf7826 (2023).
Nowack, M. K. et al. Genetic framework of cyclin-dependent kinase function in Arabidopsis. Dev. Cell 22, 1030–1040 (2012).
Braud, C., Zheng, W. & Xiao, W. LONO1 encoding a nucleoporin is required for embryogenesis and seed viability in Arabidopsis. Plant Physiol. 160, 823–836 (2012).
Park, G. T. et al. Nucleoporin MOS7/Nup88 is required for mitosis in gametogenesis and seed development in Arabidopsis. Proc. Natl Acad. Sci. USA 111, 18393–18398 (2014).
Ding, D., Muthuswamy, S. & Meier, I. Functional interaction between the Arabidopsis orthologs of spindle assembly checkpoint proteins MAD1 and MAD2 and the nucleoporin NUA. Plant Mol. Biol. 79, 203–216 (2012).
Iwase, A. et al. The AP2/ERF transcription factor WIND1 controls cell dedifferentiation in Arabidopsis. Curr. Biol. 21, 508–514 (2011).
Tang, Y., Ho, M. I., Kang, B. H. & Gu, Y. GBPL3 localizes to the nuclear pore complex and functionally connects the nuclear basket with the nucleoskeleton in plants. PLoS Biol. 20, e3001831 (2022).
Nomoto, M. & Tada, Y. Cloning-free template DNA preparation for cell-free protein synthesis via two-step PCR using versatile primer designs with short 3′-UTR. Genes Cells 23, 46–53 (2018).
Guan, S., Price, J. C., Prusiner, S. B., Ghaemmaghami, S. & Burlingame, A. L. A data processing pipeline for mammalian proteome dynamics studies using stable isotope metabolic labeling. Mol. Cell. Proteom. 10, M111.010728 (2011).
Gao, M., Nakajima An, D., Parks, J. M. & Skolnick, J. AF2Complex predicts direct physical interactions in multimeric proteins with deep learning. Nat. Commun. 13, 1744 (2022).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Acknowledgements
We thank C. Gutierrez at Centro de Biologia Molecular Severo Ochoa and D. Bergmann at Stanford University for sharing PlaCCI seeds and B. Liu at UC Davis for sharing the CYCD3;1–Myc construct and GFP–TUA6 N. benthamiana transgenic seeds. We also thank J. Shen from UC Berkeley for assistance with the callus induction assay, M. Nomoto and Y. Tada from Nagoya University for providing reagents for cell-free protein synthesis, L. Xu from Chinese Academy of Sciences for sharing the detached leaf regeneration scRNA-seq data35 and D. Schichnes from the Rausser College of Natural Resources Biological Imaging Facility at UC Berkeley for assistance with fluorescence imaging. This work was supported by the National Institute of General Medical Sciences of the National Institutes of Health (grant no. R35GM154623-01 to Y.G., grant nos R01GM135706 and S10OD030441 to S.X. and a diversity supplement to A.V.R.), the National Institute of Food and Agriculture (HATCH project nos CA-B-PLB-0243-H to Y.G. and CA-D-PLB-2850-H to X.X.), the Carnegie Endowment Fund to the Carnegie Mass Spectrometry Facility, and the US National Science Foundation (grant no. 2049931 to Y.G.).
Author information
Authors and Affiliations
Contributions
Y.F., Y.T. and Y.G. conceived the project. Y.F. and P.X. imaged and quantified the root phenotypes. Y.F. and K.H. performed the callus induction assay. Y.F. and H.N. performed the GUS staining assay. M.J. contributed to RNA extraction and qPCR. A.V.R. and S.X. performed the phosphorylation site identification through mass spectrometry. Y.F., Y.L. and X.X. performed scRNA-seq reanalysis on shoot inflorescence and regeneration tissue from the detached leaf data. Y.F. conducted all the rest of the experiments and data analyses. Y.F. and Y.G. wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Katja Graumann, Mónica Pradillo and Kentaro Tamura for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 PNET1 is a conserved cell cycle regulator.
a, AlphaFold predictions of human TMEM209 and Arabidopsis PNET1 structures (https://alphafold.ebi.ac.uk/). pLDDT score shows the confidence of the prediction. b, The pnet1 T-DNA and CRISPR mutant alleles. c, Root length measurement of 6-day-old seedlings (n = 30 for each background, two-tailed t-tests). The lower and upper whiskers indicate the minimum and maximum values, respectively. The upper boundary, middle line, and the lower boundary of the box indicate the 75th percentile, medium, and 25th percentile, respectively. Scale bar, 5 mm. d, Fluorescence imaging of the RAM in Dex::PNET1–GFP / pnet1-1 seedling with dex treatment. Scale bar, 20 μm. e, Quantification of the percentage of cells at different cell cycle stages within the epidermal layer of the RAM under homeostatic conditions (n = 10 seedlings for each background, two-tailed t-tests). Data are presented as mean values ± SD.
Extended Data Fig. 2 PNET1 is specifically and highly expressed in actively dividing cells.
a, GUS staining of 14-day-old proPNET1::GUS and WT non-transgenic seedlings. No specific and consistent staining pattern was observed. Scale bars, 5 mm. b, PNET1 expression pattern in de novo root tissues regenerated from detached Arabidopsis leave. Data was obtained by reanalyzing scRNA-seq data published previously35. H3.1 and CDKB2;1 are cell cycle markers for the S and G2/M phase, respectively. c, GUS staining of WT and proPNET1::GUS plants at 0 and 4 days past shoot removal (dpsr). Scale bars, 0.1 mm. Panels a and c represent results from two independent experiments conducted with separate reporter lines, yielding similar outcomes.
Extended Data Fig. 3 PNET1 regulates cell proliferation rate.
a, Relative expression levels of PNET1 in WT and two independent PNET1 overexpression lines. RT-qPCR was performed using 6-day-old seedlings (n = 3 biological replicates, two-sided t-tests). Data are presented as mean values ± SD. b, Representative images of calli derived from hypocotyl explants after growing on CIM2 for 4 days and then SIM for 12 days. Arrowheads indicate regenerated shoots. Scale bars, 1 mm. c, The number of regenerated shoots per callus was measured 12 days after growing on SIM (n = 40 for each genetic background). Statistical analysis was performed using one-way ANOVA followed by Tukey’s multiple comparison tests (P < 0.05). Exact P values are provided in the Source Data. The lower and upper whiskers indicate the minimum and maximum values, respectively. The upper boundary, middle line, and the lower boundary of the box indicate the 75th percentile, medium, and 25th percentile, respectively. d, Leaves of GFP–TUA6 transgenic N. benthamiana were co-infiltrated with Agrobacteria carrying 35S::PNET1–CFP and Dex::CYCD3;1–mCherry constructs. Representative images of epidermal cells undergoing cell division at 48 hours after dex treatment. Arrows label dividing cells with observable mitotic spindle structures. The experiment was repeated three times with similar results. Scale bar, 50 µm.
Extended Data Fig. 4 Identification of PNET1 interactors using proximity labeling and Alphafold prediction.
a, Total protein extracts from 35S::PNET1–3xHA–TurboID seedlings treated with biotin were immunoblotted with streptavidin and anti-HA antibody. Samples from WT seedlings were used as control. b, Predicted protein-protein interaction between PNET1 and Nup43 using Alphafold2. The interaction surface details are shown in the lower panel.
Extended Data Fig. 5 Phosphorylation of PNET1 affects its interaction with Nup43 and the cell cycle rate.
a, Y2H analysis with Nup43 as prey and PNET1D and PNET1A as bait. Diploid yeasts were grown on DDO and TDO supplemented with 0.5 mM 3-AT media. EV, empty vector. The experiment was repeated twice with similar results. b, Localization of PNET1A–CFP, GFP–TUA6, and CYCD3;1–mCherry in N. benthamiana epidermal cells. Scale bar, 10 µm. c, The dex-inducible CYCD3;1–mCherry was coexpressed with PNET–CFP, PNET1A–CFP, or free CFP in N. benthamiana epidermal cells. Percentage of dividing cells after 24, 36, or 48 hours of dex treatment was quantified. Two hundred cells were examined per replicate (n = 10 replicates, two-tailed t-tests). Data are presented as mean values ± SD. d, Measurements of PNET1D–mCherry or PNET1A–mCherry signal intensity on the NE using the same microscope settings (n = 20 nuclei). Statistical analysis was performed using one-way ANOVA followed by Tukey’s multiple comparison tests. Exact P values are provided in the Source Data. e, Multiple sequence alignment of PNET1 and its homologs in indicated eukaryotic species using ClustalW. Positions of detected phosphorylation sites on AtPNET1 are outlined by red boxes, and conserved phosphorylation sites are colored with an orange background. The design of partial phospho-mimic PNET1 mutant variants (PNET1Da–e) is labeled below the sequence alignment. f, Transient coexpression of PNET1 and PNET1 phospho-mimic/deficient variants fused to mCherry with Nup160–GFP in N. benthamiana. The experiment was repeated twice with similar results. Scale bars, 10 μm. g and h, Representative images of 6-day-old WT and 35S::PNET1D–YFP transgenic seedlings in WT (g) and pnet1 mutant (h) background, along with measurements of their root length (n = 40 for each line, two-tailed t-tests). Scale bars, 1 cm. For all box plots, the lower and upper whiskers indicate the minimum and maximum values, respectively. The upper boundary, middle line, and the lower boundary of the box indicate the 75th percentile, medium, and 25th percentile, respectively.
Supplementary information
Supplementary Tables 1–4 (download XLSX )
Supplementary Tables 1–4.
Source data
Source Data Figs. 3 and 5 and Extended Data Figs. 3 and 5 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download PDF )
Unprocessed western blots for Fig. 4d.
Source Data Fig. 5 (download PDF )
Unprocessed western blots for Fig. 5a,b,d.
Source Data Extended Data Fig. 4 (download PDF )
Unprocessed western blots for Extended Data Fig. 4a.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fang, Y., Tang, Y., Xie, P. et al. Nucleoporin PNET1 coordinates mitotic nuclear pore complex dynamics for rapid cell division. Nat. Plants 11, 295–308 (2025). https://doi.org/10.1038/s41477-025-01908-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-025-01908-y
This article is cited by
-
PNET1 is a key regulator of NPC dynamics and cell division
Nature Plants (2025)