Abstract
Plant peptide hormones play essential roles in various physiological processes, including activating defence responses to biotic stress. Membrane-localized pattern recognition receptors (PRRs) typically detect peptide hormones to initiate signalling pathways. Through comprehensive peptide–receptor matching, we identified 63 peptide–receptor pairs in soybeans. Among these, the immunomodulatory phytocytokines GmPEP914 and GmPEP890 trigger robust immune responses that suppress a broad spectrum of pathogen infections, mediated by their identified receptors GmPEP914 and GmPEP890 RECEPTOR1 (GmP98R1) and GmP98R2. Biochemical assays combined with AI-based structural modelling demonstrated that both peptides directly bind GmP98Rs with nanomolar affinity, driven primarily by interactions between the receptors and the C-terminal residue of the peptides. The PEP914-P98R module is conserved across Fabales and Cucurbitales, underscoring its evolutionary importance. Collectively, our study establishes a scalable pipeline for systematic peptide–receptor discovery and elucidates the pivotal role of the PEP914-P98R module in soybean disease resistance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The raw data of RNA-seq experiments have been deposited in the Genome Sequence Archive (CNP0007338) in the CNCB-NGDC database. All sequences for phylogenetic analysis are available on Zenodo (https://doi.org/10.5281/zenodo.15746640)48. Any additional information required to reanalyse the data reported in this paper is available from the corresponding author. Source data are provided with this paper. Constructs and reagents in this study will be made available upon request, but a completed Materials Transfer Agreement may be required if there is potential for commercial application.
References
Olsson, V. et al. Look closely, the beautiful may be small: precursor-derived peptides in plants. Annu. Rev. Plant Biol. 70, 153–186 (2019).
Tavormina, P., De Coninck, B., Nikonorova, N., De Smet, I. & Cammue, B. P. A. The plant peptidome: an expanding repertoire of structural features and biological functions. Plant Cell 27, 2095–2118 (2015).
DeFalco, T. A. & Zipfel, C. Molecular mechanisms of early plant pattern-triggered immune signaling. Mol. Cell 81, 3449–3467 (2021).
Boutrot, F. & Zipfel, C. Function, discovery, and exploitation of plant pattern recognition receptors for broad-spectrum disease resistance. Annu. Rev. Phytopathol. 55, 257–286 (2017).
Furumizu, C. & Aalen, R. B. Peptide signaling through leucine-rich repeat receptor kinases: insight into land plant evolution. New Phytol. 238, 977–982 (2023).
Dievart, A., Gottin, C., Périn, C., Ranwez, V. & Chantret, N. Origin and diversity of plant receptor-like kinases. Annu. Rev. Plant Biol. 71, 131–156 (2020).
Tang, D., Wang, G. & Zhou, J.-M. Receptor kinases in plant–pathogen interactions: more than pattern recognition. Plant Cell 29, 618–637 (2017).
Liu, Z. et al. Phytocytokine signalling reopens stomata in plant immunity and water loss. Nature 605, 332–339 (2022).
Shiu, S.-H. et al. Comparative analysis of the receptor-like kinase family in Arabidopsis and rice. Plant Cell 16, 1220–1234 (2004).
Ngou, B. P. M., Heal, R., Wyler, M., Schmid, M. W. & Jones, J. D. G. Concerted expansion and contraction of immune receptor gene repertoires in plant genomes. Nat. Plants 8, 1146–1152 (2022).
Kileeg, Z., Haldar, A., Khan, H., Qamar, A. & Mott, G. A. Differential expansion and retention patterns of LRR-RLK genes across plant evolution. Plant Direct 7, e556 (2023).
Ngou, B. P. M., Wyler, M., Schmid, M. W., Kadota, Y. & Shirasu, K. Evolutionary trajectory of pattern recognition receptors in plants. Nat. Commun. 15, 308 (2024).
Rzemieniewski, J. & Stegmann, M. Regulation of pattern-triggered immunity and growth by phytocytokines. Curr. Opin. Plant Biol. 68, 102230 (2022).
Zhou, J.-M. & Zhang, Y. Plant immunity: danger perception and signaling. Cell 181, 978–989 (2020).
Snoeck, S., Johanndrees, O., Nürnberger, T. & Zipfel, C. Plant pattern recognition receptors: from evolutionary insight to engineering. Nat. Rev. Genet. 26, 268–278 (2025).
Li, T., Moreno-Pérez, A. & Coaker, G. Plant pattern recognition receptors: exploring their evolution, diversification, and spatiotemporal regulation. Curr. Opin. Plant Biol. 82, 102631 (2024).
Yamaguchi, K. & Kawasaki, T. Pathogen- and plant-derived peptides trigger plant immunity. Peptides 144, 170611 (2021).
Yamaguchi, Y., Pearce, G. & Ryan, C. A. The cell surface leucine-rich repeat receptor for AtPep1, an endogenous peptide elicitor in Arabidopsis, is functional in transgenic tobacco cells. Proc. Natl Acad. Sci. USA 103, 10104–10109 (2006).
Huffaker, A., Pearce, G. & Ryan, C. A. An endogenous peptide signal in Arabidopsis activates components of the innate immune response. Proc. Natl Acad. Sci. USA 103, 10098–10103 (2006).
Yang, W. et al. Peptide REF1 is a local wound signal promoting plant regeneration. Cell 187, 3024–3038.e14 (2024).
Wang, L. et al. The systemin receptor SYR1 enhances resistance of tomato against herbivorous insects. Nat. Plants 4, 152–156 (2018).
Rhodes, J. et al. Perception of a divergent family of phytocytokines by the Arabidopsis receptor kinase MIK2. Nat. Commun. 12, 705 (2021).
Hou, S. et al. The Arabidopsis MIK2 receptor elicits immunity by sensing a conserved signature from phytocytokines and microbes. Nat. Commun. 12, 5494 (2021).
Yang, H. et al. Subtilase-mediated biogenesis of the expanded family of SERINE RICH ENDOGENOUS PEPTIDES. Nat. Plants 9, 2085–2094 (2023).
Rhodes, J. et al. Perception of a conserved family of plant signalling peptides by the receptor kinase HSL3. Elife 11, e74687 (2022).
Snoeck, S. et al. Evolutionary gain and loss of a plant pattern-recognition receptor for HAMP recognition. Elife 11, e81050 (2022).
Madeira, F. et al. The EMBL-EBI Job Dispatcher sequence analysis tools framework in 2024. Nucleic Acids Res. 52, W521–W525 (2024).
Boschiero, C. et al. MtSSPdb: the Medicago truncatula small secreted peptide database. Plant Physiol. 183, 399–413 (2020).
Butenko, M. A. et al. Tools and strategies to match peptide–ligand receptor pairs. Plant Cell 26, 1838–1847 (2014).
Hohmann, U. et al. Mechanistic basis for the activation of plant membrane receptor kinases by SERK-family coreceptors. Proc. Natl Acad. Sci. USA 115, 3488–3493 (2018).
Hou, S. et al. The secreted peptide PIP1 amplifies immunity through receptor-like kinase 7. PLoS Pathog. 10, e1004331 (2014).
Rzemieniewski, J. et al. CEP signaling coordinates plant immunity with nitrogen status. Nat. Commun. 15, 10686 (2024).
Yamaguchi, Y., Barona, G., Ryan, C. A. & Pearce, G. GmPep914, an eight-amino acid peptide isolated from soybean leaves, activates defense-related genes. Plant Physiol. 156, 932–942 (2011).
Ma, X., Xu, G., He, P. & Shan, L. SERKing coreceptors for receptors. Trends Plant Sci. 21, 1017–1033 (2016).
Snoeck, S. et al. Leveraging coevolutionary insights and AI-based structural modeling to unravel receptor–peptide ligand-binding mechanisms. Proc. Natl Acad. Sci. USA 121, e2400862121 (2024).
Helft, L. et al. LRR conservation mapping to predict functional sites within protein leucine-rich repeat domains. PLoS ONE 6, e21614 (2011).
Oldroyd, G. E. D., Murray, J. D., Poole, P. S. & Downie, J. A. The rules of engagement in the legume–rhizobial symbiosis. Annu. Rev. Genet. 45, 119–144 (2011).
Huang, L. & Schiefelbein, J. Conserved gene expression programs in developing roots from diverse plants. Plant Cell 27, 2119–2132 (2015).
Li, L., Habring, A., Wang, K. & Weigel, D. Atypical resistance protein RPW8/HR triggers oligomerization of the NLR immune receptor RPP7 and autoimmunity. Cell Host Microbe 27, 405–417.e6 (2020).
Schwab, R., Ossowski, S., Riester, M., Warthmann, N. & Weigel, D. Highly specific gene silencing by artificial microRNAs in Arabidopsis. Plant Cell 18, 1121–1133 (2006).
Song, J., Tóth, K., Montes-Luz, B. & Stacey, G. Soybean hairy root transformation: a rapid and highly efficient method. Curr. Protoc. 1, e195 (2021).
Xie, J. et al. Tree Visualization By One Table (tvBOT): a web application for visualizing, modifying and annotating phylogenetic trees. Nucleic Acids Res. 51, W587–W592 (2023).
Kumar, S. et al. TimeTree 5: an expanded resource for species divergence times. Mol. Biol. Evol. 39, msac174 (2022).
Chen, C. et al. Extracellular plant subtilases dampen cold-shock peptide elicitor levels. Nat. Plants 10, 1749–1760 (2024).
Shen, W., Liu, J. & Li, J.-F. Type-II metacaspases mediate the processing of plant elicitor peptides in Arabidopsis. Mol. Plant 12, 1524–1533 (2019).
Yang, Z. et al. SoyMD: a platform combining multi-omics data with various tools for soybean research and breeding. Nucleic Acids Res. 52, D1639–D1650 (2024).
Tang, H. et al. JCVI: a versatile toolkit for comparative genomics analysis. Imeta 3, e211 (2024).
Li, L. Sequence for phylogeny [Data set]. Zenodo https://doi.org/10.5281/zenodo.15746640 (2025).
Acknowledgements
We thank D. Weigel (Max Planck Institute for Biology Tuebingen), J.-M. Zhou (Yazhouwan National Laboratory/Institute of Genetics and Developmental Biology, CAS), J.-W. Wang (CAS Center for Excellence in Molecular Plant Sciences) and F. Feng (Oklahoma State University) for critical comments on the manuscript; Y. Wang (Nanjing Agricultural University) for kindly sharing the P. sojae isolates; B. Ren and J. Feng (Institute of Genetics and Developmental Biology, CAS) for sharing soybean and Medicago materials; and J. Wang (Peking University) and W. Song (China Agricultural University) for technical assistance with protein purification and structure visualization. This study was funded by Biological Breeding-National Science and Technology Major Project 2023ZD04073 (L.L.), National Natural Science Foundation of China 32270298 (L.L.), and National Natural Science Foundation of China 32388201 (Z.T.).
Author information
Authors and Affiliations
Contributions
L.L. and L.Y. conceived and designed the research; L.L. and Z.T. supervised the project; L.L. and L.Y. participated in all experiments, design and data analysis; Y.G. generated the chimaeric constructs and initiated the receptor–peptide screening; Q.Y. performed protein purification from insect cells, ITC and MST assays; J.L. performed RNA-seq data analyses and synteny analyses; T.L., W.M. and H.L. contributed to the functional investigation of GmPEP914 and GmPEP890; L.L. wrote the original draft; all authors were involved in data analyses and reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Benjamin Petre and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Phylogenetic analysis of Arabidopsis and soybean LRR-RKs from the subfamily X, XI, and XII.
The neighbor-joining method with 1,000 bootstrap replicates was used for tree generation in MEGA11. Bootstrap values are indicated with only values higher than 50 shown. Full red cycles indicate cognate ligands of LRR-RKs in Arabidopsis. The tree was modified and displayed using TVBOT42.
Extended Data Fig. 2 Sequence alignment of identified unknown small secreted peptide family from soybean.
Small secreted peptides from soybean were predicted using an online pipeline developed based on protein length, signal peptide, homology with identified plant families, and transmembrane domain prediction28. Conserved amino acids were labeled with a black background. Fun, Family of unknown.
Extended Data Fig. 3 Protein expression of 140 chimeric GmRKs in N. benthamiana.
Immunoblots were performed using an anti-FLAG antibody to detect the expression of the 140 chimeric GmRKs. The expected protein sized were noted with asterisk. The experiments were repeated two times with similar results.
Extended Data Fig. 4 ROS production triggered by the synthesized peptide library in soybean.
2-week-old soybean leaf discs were treated with 1 μM of the indicated peptide. Cumulative ROS production was measured over 40 min. GmP1 and GmP2 (highlighted in red) correspond to GmPEP914 and GmPEP890, respectively. Mean cumulative relative luminescence unit (RLU) values (± SD) are shown (n = 8 discs). Statistical analysis versus mock was performed with one-way analysis of variance (ANOVA) followed by Tukey’s test. Significance values are indicated on plots. The experiments were repeated three times with similar results.
Extended Data Fig. 5 Calcium influx triggered by peptide-receptor pairs in N. benthamiana.
Aequorin-based calcium influx assays were performed in N. benthamiana leaves co-expressing the indicated GmRK chimeras and aequorin. GmP1 and GmP2 (highlighted in red) correspond to GmPEP914 and GmPEP890, respectively. Peak luminescence values elicited by the peptides are shown. Data are mean ± SEM (n = 8). Statistical analysis versus mock was performed by one-way analysis of variance (ANOVA) followed by Tukey’s test for comparisons involving more than two groups, or by two-sided Student’s t-test for pairwise comparisons. Significance values are indicated on plots.
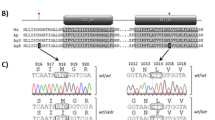
Extended Data Fig. 6 GmPEP914 and GmPEP890 trigger GmP98R1- and GmP98R2-dependent immunity.
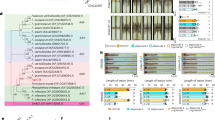
a, GmProPEP914 and GmProPEP890 are upregulated by flg22 treatment and pathogen infection. Soybean leaves that were treated with 1 μM flg22 or spray-inoculated with Pseudomonas syringae pv. glycinea and roots that were incubated with spores of Fusarium oxysporum were used for RT-qPCR analysis. Data are mean ± SEM (n = 3 biological replicates). Statistical analysis was performed with one-way analysis of variance (ANOVA) followed by Sidak’s test. b, Representative images of soybean etiolated hypocotyls inoculated with P. sojae P7076. Hypocotyls were pretreated with H2O (Mock), 1 μM GmPEP914, or 1 μM GmPEP914m for 24 hours. Images were taken 48 hours after inoculation with P. sojae P7076 (n = 12). Scale bars, 1 cm. c,d, Lesion length and relative biomass of P. sojae P7076-infected etiolated soybean hypocotyls as shown in (b). The experiments were repeated three times with similar results. Data are mean ± SD (n = 12 in c; n = 3 in d). Statistical analysis was performed with two-sided Student’s t-test. e, Symptom and quantification of Psg spray-inoculated onto soybean leaves. Leaves were pretreated with H2O (Mock), 1 μM flg22, or 1 μM GmPEP914 for 24 hours. Images were taken 4 days after inoculation with Psg. Bacterial growth was measured at 4 dpi. Data are mean ± SD (n = 8). Statistical analysis was performed with two-sided Student’s t-test. cfu, colony-forming units. f, The propep914/890 double mutants exhibited a slight increase in Psg growth compared to Wm82. Leaves of indicated genotypes were spray-inoculated with Psg. Bacterial growth was measured at 4 dpi (Left panel). Data are mean ± SD (n = 8). Statistical analysis was performed with two-sided Student’s t-test. cfu, colony-forming units. g,h, Sanger sequencing of two independent propep914/890 and p98r1r2 double mutant lines generated by CRISPR-Cas9. Deletions are indicated with red dashes. i, Protein expression in N. benthamiana of indicated GmRKs for ROS assays in (j). j, GmRKs-mediated ROS production triggered by GmPEP914 in N. benthamiana. Full-length of indicated GmRKs were transiently expressed in N. benthamiana. ROS production triggered by 1 μM GmPEP914 was measured after two days of expression. Mean RLU values (± SEM) are shown (n = 8). k, The p98r1r2 double mutants exhibited more susceptible to P. sojae P7076 compared to Wm82. Representative images of soybean etiolated hypocotyls inoculated with P. sojae P7076. Images were taken 48 hours after inoculation with P. sojae P7076 (n = 18). Scale bars, 1 cm. l,m, Lesion length and relative biomass of P. sojae P7076-infected etiolated soybean hypocotyls as shown in (k). The experiments were repeated three times with similar results. Data are mean ± SEM (n = 18 in l; n = 3 in m). Statistical analysis was performed with two-sided Student’s t-test.
Extended Data Fig. 7 GmPEP914 and GmPEP890 trigger defense genes expression in an GmP98R1- and GmP98R2-dependent manner.
a, Gene Ontology terms of GmPEP914-mediated upregulated and downregulated DEGs. b, Heatmap showing DEGs related to immunity in response to 1 μM GmPEP914 treatment for 2 hours in Wm82 (wild type) and p98r1r2 mutant (n = 3). c, RT-qPCR analysis of indicated gene expression in Wm82 and p98r1r2 mutant upon treatment without (Mock) or with 1 μM GmPEP914 for 2 hours. Data are mean ± SD (n = 3). Statistical analysis was performed with two-way analysis of variance (ANOVA) followed by Tukey’s test.
Extended Data Fig. 8 GmP98R1 and GmP98R2 are the receptors of GmPEP914 and GmPEP890.
a, AlphaFold-Multimer predicted structure of GmPEP914-GmP98R1-GmBAK1a complex. The side chains of C-terminal Y8 of GmPEP914 and R329/R331 of GmP98R1 are labeled. The predicted confidence scores are shown. b, Y8A mutant in GmPEP914 (GmPEP914m) affects its activity in triggering ROS production in soybean leaf. 1 μM peptides were used in the measurement. Mean RLU values (± SEM) are shown (n = 8). c, R329/R331 mutations in GmP98R1 affect GmPEP914-triggered ROS production in N. benthamiana. The indicated mutated GmP98R1 variants were transiently expressed in N. benthamiana. ROS production triggered by 1 μM GmPEP914 was measured after two days of expression. Mean RLU values (± SEM) are shown (n = 8). The inserted panel shows protein expression of indicated GmP98R1 variants in N. benthamiana. d, The ectodomains of GmP98Rs purified from insect cell. The experiment was repeated two times with similar results. e,f, Quantification of binding affinity by ITC analysis. The indicated proteins at 20 μM and peptides at 200 μM were used in the ITC analysis. Dissociation constants (Kd) and stoichiometries (N) are summarized in (c). All experiments were repeated three times with similar results. g, Quantification of binding affinity by MST analysis. The Kd determined by MST is indicated. Data are mean ± SD (n = 3 biological replicates). Please note that one same GmP98R1-GmPEP914 control from both panels was used to compare with the mutants. h, RT-qPCR analysis of GmBAK1a and GmBAK1b expression in transgenic hairy roots of soybean. Artificial microRNA (amiR-BAK1-1 and amiR-BAK1-2) targeting two different sites were expressed in transgenic hairy roots of soybean. Data are mean ± SD (n = 3 biological replicates). Statistical analysis was performed with one-way analysis of variance (ANOVA) followed by Tukey’s test.
Extended Data Fig. 9 PEP914-P98R module is present in Fabales and Cucurbitales.
a, ROS production triggered by PEP914 and PEP890 in the indicated plant species. ROS production in response to 1 μM peptides was measured. Mean RLU values (± SEM) are shown (n = 8). b, PEP914- and PEP890-induced ROS production in the indicated plant species. The species tree was generated using TimeTree 543. Species displaying positive and no ROS responses to the peptides are indicated by filled and open circle, respectively. NFNC, nitrogen fixing nodulation clade. c, ROS production mediated by CsP98Ra, MtP98Ra, and MtP98Rb in response to PEP914 or PEP890 in N. benthamiana. Full-length of indicated RKs were transiently expressed in N. benthamiana. ROS production in response to 1 μM peptides was measured after two days of expression. Cumulative RLU values (± SEM) are shown (n = 8). Statistical analysis was performed with two-sided Student’s t-test.
Extended Data Fig. 10 Sequence analysis of ProPEP914 from Fabales and Cucurbitales.
a, Amino acid sequence alignment of ProPEP914 from Fabales and Cucurbitales species. b, Phylogenetic analysis of ProPEP914 from Fabales and Cucurbitales species. The neighbor-joining method with 1,000 bootstrap replicates was used for tree generation in MEGA11.
Supplementary information
Supplementary Table 1 (download XLSX )
LRR-RK identified in soybean for ROS measurement.
Supplementary Table 2 (download XLSX )
Information on 55 synthesized peptides.
Supplementary Table 3 (download XLSX )
Candidate SSPs predicted by the MtSSPdb pipeline.
Supplementary Table 4 (download XLSX )
ROS production for matching peptide–receptor pairs by expressing chimaeric LRR-RKs with a fusion of the indicated ectodomain of LRR-RKs and the kinase domain of PEPR in N. benthamiana.
Supplementary Table 5 (download XLSX )
List of 2,738 DEGs identified in Wm82 treatment with GmPEP914 compared to Mock and TPM values in all samples.
Supplementary Table 6 (download XLSX )
Oligos used in this study.
Source data
Source Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Extended Data Fig. 6 (download PDF )
Unprocessed western blots and gels.
Source Extended Data Fig. 8 (download PDF )
Unprocessed western blots and gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yu, L., Gao, Y., Yang, Q. et al. Large-scale pairing identifies a soybean phytocytokine-receptor module conferring disease resistance. Nat. Plants 11, 1739–1747 (2025). https://doi.org/10.1038/s41477-025-02086-7
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-025-02086-7