Abstract
The conserved target of rapamycin (TOR) kinase acts as a master regulator of growth by integrating nutrient and environmental signals in eukaryotes. However, how TOR influences chromatin remains poorly understood. Here we identified a multi-subunit complex in Arabidopsis thaliana, termed the chromatin-associated complex for growth (CACG). Our findings indicate that under nutrient-rich conditions, active TOR kinase enhances CACG mRNA translation, which is facilitated by pyrimidine-rich motifs in their 5′ untranslated regions. CACG components co-occupy stress-responsive genes marked by histone acetylation, repressing their transcription to promote growth. Conversely, under nutrient-deficient conditions, inactive TOR reduces CACG mRNA translation, relieving transcriptional repression of stress-responsive genes and leading to increased stress tolerance but impaired growth. These results indicate that the CACG complex acts as a critical nutrient-responsive transcriptional regulator that is required for coordinating plant growth and stress tolerance in a TOR-dependent manner. The molecular mechanism revealed here could aid in developing high-yield crops capable of thriving in adverse environments.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The raw RNA-seq and ChIP–seq data have been deposited in the Gene Expression Omnibus database under the accession code GSE276448. The gene symbols and locus codes of genes investigated in this study are as follows: WDR5A (AT3G49660), ZDP1 (AT1G09520), GTA1 (AT1G05860), GTA2 (AT3G53860), GTA3 (AT2G31600), TRB4 (AT1G17520), TRB5 (AT1G72740), GTE1 (AT2G34900), GTE6 (AT3G52280), CP2 (AT3G18210), GTB1 (AT4G37440), GTB2 (AT3G59670), GTB3 (AT3G50040), DP1 (AT1G09710), DRMY1 (AT1G58220), EMB1967 (AT3G54350), TOR (AT1G50030), DEAR5 (AT4G06746), DREB26 (AT1G21910), ERF1A (AT4G17500), ERF2 (AT5G47220), ERF018 (AT1G74930), ERF6 (AT4G17490), CBF2 (AT4G25470), ERF5 (AT5G47230), ERF104 (AT5G61600), DDF1 (AT1G12610), ERF13 (AT2G44840), ERF109 (AT4G34410), ERF022 (AT1G33760), ERF11 (AT1G28370) and CBF1 (AT4G25490). Source data are provided with this paper.
References
Bechtold, U. & Field, B. Molecular mechanisms controlling plant growth during abiotic stress. J. Exp. Bot. 69, 2753–2758 (2018).
Gong, Z. et al. Plant abiotic stress response and nutrient use efficiency. Sci. China Life Sci. 63, 635–674 (2020).
Zhang, H., Zhao, Y. & Zhu, J.-K. Thriving under stress: how plants balance growth and the stress response. Dev. Cell 55, 529–543 (2020).
Hermans, C., Hammond, J. P., White, P. J. & Verbruggen, N. How do plants respond to nutrient shortage by biomass allocation? Trends Plant Sci. 11, 610–617 (2006).
Wang, P. et al. Reciprocal regulation of the TOR kinase and ABA receptor balances plant growth and stress response. Mol. Cell 69, 100–112.e6 (2018).
Walley, J. W. & Dehesh, K. Molecular mechanisms regulating rapid stress signaling networks in Arabidopsis. J. Integr. Plant Biol. 52, 354–359 (2010).
Singh, K. B., Foley, R. C. & Oñate-Sánchez, L. Transcription factors in plant defense and stress responses. Curr. Opin. Plant Biol. 5, 430–436 (2002).
Rando, O. J. & Chang, H. Y. Genome-wide views of chromatin structure. Ann. Rev. Biochem. 78, 245–271 (2009).
Li, B., Carey, M. & Workman, J. L. The role of chromatin during transcription. Cell 128, 707–719 (2007).
Feng, S. & Jacobsen, S. E. Epigenetic modifications in plants: an evolutionary perspective. Curr. Opin. Plant Biol. 14, 179–186 (2011).
Guo, J. & He, X.-J. Composition and function of plant chromatin remodeling complexes. Curr. Opin. Plant Biol. 81, 102613 (2024).
Liu, C., Lu, F., Cui, X. & Cao, X. Histone methylation in higher plants. Annu. Rev. Plant Biol. 61, 395–420 (2010).
Chen, Y., Guo, P. & Dong, Z. The role of histone acetylation in transcriptional regulation and seed development. Plant Physiol. 194, 1962–1979 (2024).
Scheid, R., Chen, J. & Zhong, X. Biological role and mechanism of chromatin readers in plants. Curr. Opin. Plant Biol. 61, 102008 (2021).
Ruthenburg, A. J., Li, H., Patel, D. J. & Allis, C. D. Multivalent engagement of chromatin modifications by linked binding modules. Nat. Rev. Mol. Cell Biol. 8, 983–994 (2007).
Du, J., Johnson, L. M., Jacobsen, S. E. & Patel, D. J. DNA methylation pathways and their crosstalk with histone methylation. Nat. Rev. Mol. Cell Biol. 16, 519–532 (2015).
Liu, Y. et al. Target of rapamycin (TOR): a master regulator in plant growth, development, and stress responses. Annu. Rev. Plant Biol. 76, 341–371 (2025).
Henriques, R., Bögre, L., Horváth, B. & Magyar, Z. Balancing act: matching growth with environment by the TOR signalling pathway. J. Exp. Bot. 65, 2691–2701 (2014).
Wullschleger, S., Loewith, R. & Hall, M. N. TOR signaling in growth and metabolism. Cell 124, 471–484 (2006).
Saxton, R. A. & Sabatini, D. M. mTOR signaling in growth, metabolism, and disease. Cell 168, 960–976 (2017).
Hagner, P. R. et al. Ribosomal protein S6 is highly expressed in non-Hodgkin lymphoma and associates with mRNA containing a 5′ terminal oligopyrimidine tract. Oncogene 30, 1531–1541 (2011).
Hopkins, K. C. et al. Virus-induced translational arrest through 4EBP1/2-dependent decay of 5′-TOP mRNAs restricts viral infection. Proc. Natl Acad. Sci. USA 112, E2920–E2929 (2015).
Philippe, L., van den Elzen, A. M. G., Watson, M. J. & Thoreen, C. C. Global analysis of LARP1 translation targets reveals tunable and dynamic features of 5′ TOP motifs. Proc. Natl Acad. Sci. USA 117, 5319–5328 (2020).
Hochstoeger, T. & Chao, J. A. Towards a molecular understanding of the 5′TOP motif in regulating translation of ribosomal mRNAs. Semin. Cell Dev. Biol. 154, 99–104 (2024).
Scarpin, M. R., Leiboff, S. & Brunkard, J. O. Parallel global profiling of plant TOR dynamics reveals a conserved role for LARP1 in translation. eLife 9, e58795 (2020).
Meng, Y. et al. TOR kinase, a GPS in the complex nutrient and hormonal signaling networks to guide plant growth and development. J. Exp. Bot. 73, 7041–7054 (2022).
Dong, Y. et al. Functional analogs of mammalian 4E-BPs reveal a role for TOR in global plant translation. Cell Rep. 42, 112892 (2023).
Tzeng, T.-Y., Kong, L.-R., Chen, C.-H., Shaw, C.-C. & Yang, C.-H. Overexpression of the lily p70s6k gene in Arabidopsis affects elongation of flower organs and indicates TOR-dependent regulation of AP3, PI and SUP translation. Plant Cell Physiol. 50, 1695–1709 (2009).
Xiong, Y. et al. Glucose–TOR signalling reprograms the transcriptome and activates meristems. Nature 496, 181–186 (2013).
Yuan, X., Xu, P., Yu, Y. & Xiong, Y. Glucose–TOR signaling regulates PIN2 stability to orchestrate auxin gradient and cell expansion in Arabidopsis root. Proc. Natl Acad. Sci. USA 117, 32223–32225 (2020).
Ye, R. et al. Glucose-driven TOR–FIE–PRC2 signalling controls plant development. Nature 609, 986–993 (2022).
Tan, L.-M. et al. The PEAT protein complexes are required for histone deacetylation and heterochromatin silencing. EMBO J. 37, e98770 (2018).
Zhou, Y. et al. Telobox motifs recruit CLF/SWN–PRC2 for H3K27me3 deposition via TRB factors in Arabidopsis. Nat. Genet. 50, 638–644 (2018).
Wang, M. et al. Arabidopsis TRB proteins function in H3K4me3 demethylation by recruiting JMJ14. Nat. Commun. 14, 1736 (2023).
Zheng, S.-Y. et al. Dual roles of the Arabidopsis PEAT complex in histone H2A deubiquitination and H4K5 acetylation. Mol. Plant 16, 1847–1865 (2023).
Shang, J.-Y. et al. COMPASS functions as a module of the INO80 chromatin remodeling complex to mediate histone H3K4 methylation in Arabidopsis. Plant Cell 33, 3250–3271 (2021).
Wang, Q. The role of forkhead-associated (FHA)-domain proteins in plant biology. Plant Mol. Biol. 111, 455–472 (2023).
Huo, H. H., Luo, M., Lee, Y.-R. J. & Liu, B. The Arabidopsis homolog of microspherule protein 1 is essential for embryogenesis and interacts with the Myb-like transcription factor DRMY1. Plant Cell Physiol. 66, 890–899 (2025).
Jin, J. et al. A mammalian chromatin remodeling complex with similarities to the yeast INO80 complex. J. Biol. Chem. 280, 41207–41212 (2005).
Wu, P. et al. DRMY1, a Myb-like protein, regulates cell expansion and seed production in Arabidopsis thaliana. Plant Cell Physiol. 60, 285–302 (2019).
Amiard, S. et al. The TELOMERE REPEAT BINDING proteins TRB4 and TRB5 function as transcriptional activators of PRC2-controlled genes to regulate plant development. Plant Commun. 5, 100890 (2024).
Jiang, D., Gu, X. & He, Y. Establishment of the winter-annual growth habit via FRIGIDA-mediated histone methylation at FLOWERING LOCUS C in Arabidopsis. Plant Cell 21, 1733–1746 (2009).
Raught, B., Gingras, A.-C. & Sonenberg, N. The target of rapamycin (TOR) proteins. Proc. Natl Acad. Sci. USA 98, 7037–7044 (2001).
Chresta, C. M. et al. AZD8055 is a potent, selective, and orally bioavailable ATP-competitive mammalian target of rapamycin kinase inhibitor with in vitro and in vivo antitumor activity. Cancer Res. 70, 288–298 (2010).
Hsieh, A. C. et al. The translational landscape of mTOR signalling steers cancer initiation and metastasis. Nature 485, 55–61 (2012).
Chua, Y. L., Channelière, S., Mott, E. & Gray, J. C. The bromodomain protein GTE6 controls leaf development in Arabidopsis by histone acetylation at ASYMMETRIC LEAVES1. Genes Dev. 19, 2245–2254 (2005).
Chen, H.-Y. et al. ORA47 (octadecanoid-responsive AP2/ERF-domain transcription factor 47) regulates jasmonic acid and abscisic acid biosynthesis and signaling through binding to a novel cis-element. N. Phytol. 211, 599–613 (2016).
Dubois, M. et al. ETHYLENE RESPONSE FACTOR6 acts as a central regulator of leaf growth under water-limiting conditions in Arabidopsis. Plant Physiol. 162, 319–332 (2013).
Park, S. et al. Regulation of the Arabidopsis CBF regulon by a complex low-temperature regulatory network. Plant J. 82, 193–207 (2015).
Licausi, F., Ohme-Takagi, M. & Perata, P. APETALA2/Ethylene Responsive Factor (AP2/ERF) transcription factors: mediators of stress responses and developmental programs. N. Phytol. 199, 639–649 (2013).
da Silva, E. C., Nogueira, R., da Silva, M. A. & de Albuquerque, M. B. Drought stress and plant nutrition. Plant Stress 5, 32–41 (2011).
Wang, Y. et al. The Arabidopsis DREAM complex antagonizes WDR5A to modulate histone H3K4me2/3 deposition for a subset of genome repression. Proc. Natl Acad. Sci. USA 119, e2206075119 (2022).
Zhao, S., Zhang, B., Yang, M., Zhu, J. & Li, H. Systematic profiling of histone readers in Arabidopsis thaliana. Cell Rep. 22, 1090–1102 (2018).
Guo, J. et al. The CBP/p300 histone acetyltransferases function as plant-specific MEDIATOR subunits in Arabidopsis. J. Integr. Plant Biol. 63, 755–771 (2021).
Wu, C.-J. et al. Arabidopsis histone acetyltransferase complex coordinates cytoplasmic histone acetylation and nuclear chromatin accessibility. Sci. Adv. 10, eadp1840 (2024).
Wu, C.-J. et al. Conserved and plant-specific histone acetyltransferase complexes cooperate to regulate gene transcription and plant development. Nat. Plants 9, 442–459 (2023).
Chang, Y. et al. Epigenetic regulation in plant abiotic stress responses. J. Integr. Plant Biol. 62, 563–580 (2020).
Feng, C. et al. Arabidopsis RPD3-like histone deacetylases form multiple complexes involved in stress response. J. Genet. Genom. 48, 369–383 (2021).
Chen, W. J. & Zhu, T. Networks of transcription factors with roles in environmental stress response. Trends Plant Sci. 9, 591–596 (2004).
Chen, X. et al. Protein kinases in plant responses to drought, salt, and cold stress. J. Integr. Plant Biol. 63, 53–78 (2021).
González, A. & Hall, M. N. Nutrient sensing and TOR signaling in yeast and mammals. EMBO J. 36, 397–408 (2017).
Dazert, E. & Hall, M. N. mTOR signaling in disease. Curr. Opin. Cell Biol. 23, 744–755 (2011).
Burkart, G. M. & Brandizzi, F. A tour of TOR complex signaling in plants. Trends Biochem. Sci. 46, 417–428 (2021).
Xiong, Y. & Sheen, J. Novel links in the plant TOR kinase signaling network. Curr. Opin. Plant Biol. 28, 83–91 (2015).
Hummel, M. et al. Proteomic LC–MS analysis of Arabidopsis cytosolic ribosomes: identification of ribosomal protein paralogs and re-annotation of the ribosomal protein genes. J. Proteom. 128, 436–449 (2015).
Wu, Q. & Bazzini, A. A. Translation and mRNA stability control. Ann. Rev. Biochem. 92, 227–245 (2023).
Kong, S. et al. DRMY1 promotes robust morphogenesis in Arabidopsis by sustaining the translation of cytokinin-signaling inhibitor proteins. Dev. Cell 59, 3141–3160.e7 (2024).
Nadi, R. et al. Overlapping roles of Arabidopsis INCURVATA11 and CUPULIFORMIS2 as Polycomb Repressive Complex 2 accessory proteins. Preprint at bioRxiv https://doi.org/10.1101/2024.03.15.585069 (2024).
Bloomer, R. H. et al. The Arabidopsis epigenetic regulator ICU11 as an accessory protein of Polycomb Repressive Complex 2. Proc. Natl Acad. Sci. USA 117, 16660–16666 (2020).
Mateo-Bonmatí, E. et al. INCURVATA11 and CUPULIFORMIS2 are redundant genes that encode epigenetic machinery components in Arabidopsis. Plant Cell 30, 1596–1616 (2018).
Lu, F. et al. Comparative analysis of JmjC domain-containing proteins reveals the potential histone demethylases in Arabidopsis and rice. J. Integr. Plant Biol. 50, 886–896 (2008).
Feng, K. et al. Advances in AP2/ERF super-family transcription factors in plant. Crit. Rev. Biotechnol. 40, 750–776 (2020).
Shoji, T. & Yuan, L. ERF gene clusters: working together to regulate metabolism. Trends Plant Sci. 26, 23–32 (2021).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Zang, C. et al. A clustering approach for identification of enriched domains from histone modification ChIP–seq data. Bioinformatics 25, 1952–1958 (2009).
Florens, L. et al. Analyzing chromatin remodeling complexes using shotgun proteomics and normalized spectral abundance factors. Methods 40, 303–311 (2006).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Warnes, M. G. R. et al. Package ‘gplots’. Various R programming tools for plotting data. https://cran.r-project.org/web/packages/gplots/ (2016).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis 2nd edn (Springer, 2016).
Du, P. et al. WRKY transcription factors and OBERON histone‐binding proteins form complexes to balance plant growth and stress tolerance. EMBO J. 42, e113639 (2023).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Picard. Broad Institute http://broadinstitute.github.io/picard/ (accessed 10 June 2020).
Zhang, Y. et al. Model-based Analysis of ChIP–Seq (MACS). Genome Biol. 9, R137 (2008).
Ramírez, F., Dündar, F., Diehl, S., Grüning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (grant numbers 32025003 and 32470628 to X.-J.H.).
Author information
Authors and Affiliations
Contributions
X.-J.H. and X.W. conceived the study and designed the experiments. X.W., Z.-Z.L., Y.-J.L., L.L. and S.C. performed the experiments. D.-Y.Y. performed the bioinformatics analyses. X.W. and X.-J.H. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Identification of the CACG components by AP-MS.
The heatmap indicates the identified components in CACG. The color gradient represents the NSAF values.
Extended Data Fig. 2 Determination of the interactions among CACG components by Y2H assays.
Yeast strains expressing the indicated GAL4-AD and GAL4-BD fusion proteins were cultivated on SD medium lacking Trp and Leu (SD-W-L) and on SD medium lacking Trp, Leu, and His (SD-W-L-H) supplemented with 3 mM 3-AT. Yeast strains containing empty GAL4-AD or GAL4-BD vectors (VEC) serve as negative controls. Positive strains indicating protein-protein interactions are highlighted with dashed boxes. The experiments were repeated independently twice and showed similar results.
Extended Data Fig. 3 Morphological phenotypes of single and multiple CACG mutants.
a - c, Morphological phenotypes of 20-day-old plants (a), 24-day-old plants (b), and 30-day-old plants (c). Scale bar, 1 cm. d, Statistical analysis of the days to bolting (n = 24, left) and the number of rosette leaves (n = 24, right). Values are mean ± SD. P values were determined by two-tailed Student’s t-test.
Extended Data Fig. 4 Analysis of lethal CACG mutants.
a, Phenotypes of mature siliques in the indicated mutants. b, The ratio of aborted seeds (n = 10). Values are mean ± SD. c, Genotypes of the progeny of self-bred gte1−/−gte6−/−trb4+/−trb5−/−, dp1−/−drmy1+/−, and emb1967+/−. P values were determined by chi-square test.
Extended Data Fig. 5 Growth defects observed in the CACG mutants.
a, Morphological phenotypes of 40-day-old plants. Scale bar, 1 cm. b, Statistical analyses of plant height of 40-day-old plants (n = 20, left), fresh weight of 20-day-old plants (n = 20, middle), and primary root length of 12-day-old plants (n = 20, right). Values are mean ± SD. P values were determined by two-tailed Student’s t-test. c, Phenotypic observation of siliques from 40-day-old plants (left) and rosette leaves from 28-day-old plants (right) in CACG mutants. Scale bar, 1 cm. d, Statistical analysis of the silique length in 40-day-old plants. Values are mean ± SD. P values were determined by two-tailed Student’s t-test. e, Statistical analysis of CACG mutant growth rates. Seedlings were initially cultivated in solid MS for 12 days before transferring to soil. The daily count of rosette leaves was recorded after transplanting (n = 20). Values are mean ± SD.
Extended Data Fig. 6 Morphological phenotypes of gtb1 gtb2 gtb3, trb4 trb5-c, and their complementation lines.
a, The morphological defects in gtb1 gtb2 gtb3 and trb4 trb5-c were complemented by the GTB1-Flag and TRB5-Myc transgenes, respectively. Scale bar, 1 cm. b, Immunoblot analysis shows the protein levels of GTB1-Flag and TRB5-Myc in the transgenic lines. The actin protein level is shown as a loading control. The experiments were independently repeated twice and showed similar results.
Extended Data Fig. 7 Effect of different nutrient components on CACG protein levels.
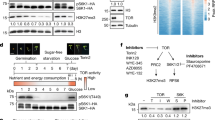
a, The protein levels of CACG in response to treatment with different nutrient components. Twelve-day-old plants grown on solid MS medium were transferred into liquid MS medium lacking macroelements, microelements, and/or vitamins. The protein levels of indicated CACG components were detected by immunoblot analysis after the plants were incubated for 4 days. Ma, macroelements; Mi, microelements; V, vitamins. The actin protein level is shown as a loading control. TOR activity was monitored by the phosphorylation state of S6K. The experiments were independently repeated twice and showed similar results. b, Statistical analysis of pS6K protein levels. Actin was used as a loading control. Each black dot represents an independent biological replicate.
Extended Data Fig. 8 The protein levels of CACG components were not influenced by inhibitors of proteasome or autophagy.
a, Immunoblot analysis shows the effect of the proteasome inhibitor MG132 on the protein levels of CACG components. Twelve-day-old plants grown in solid MS medium were transferred to H2O, liquid MS, or H2O containing 50 µM MG132 for 4 days. The actin protein level is shown as a loading control. b, Immunoblot analysis shows the effect of the autophagy inhibitor 3-MA on the protein levels of CACG components. Twelve-day-old plants grown in solid MS medium were incubated in H2O, liquid MS, or H2O containing 5 mM 3-MA for 4 days. The actin protein level is shown as a loading control. The experiments in (a) and (b) were independently repeated twice and showed similar results.
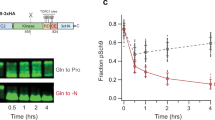
Extended Data Fig. 9 Pyrimidine-rich motifs in the 5’ UTRs of GTE1 and DP1 mRNAs facilitate target selection by TOR.
a, The expression of LUC driven by wild-type GTE1 and DP1 promoters along with wild-type or modified 5’ UTRs (5’ UTR-GTE1-R and 5’ UTR-DP1-R) were determined by luminescence imaging in tobacco leaves. Agrobacterium strains carrying the indicated plasmids were injected into tobacco leaves, either with (right) or without 6 µM AZD8055 (left). b, c, Statistical analysis of the expression levels of LUC driven by the wild-type GTE1 and DP1 promoters, along with their wild-type or modified 5’-UTRs. Protein (b) and transcript (c) levels were determined by luminescence imaging and quantitative RT-PCR, respectively. Each black dot represents an independent biological replicate. Values are mean ± SD from three independent biological replicates. P values were determined by two-tailed Student’s t-test.
Extended Data Fig. 10 The bromodomain-containing protein GTE1 exhibits a higher affinity for acetylated histone peptides than non-acetylated peptides.
a, The interaction of GTE1 and its bromodomain mutant with the indicated histone peptides was determined by pull-down assays. The experiments were independently repeated three times and showed similar results. b, Statistical analyses of the affinity of GTE1 with the indicated histone peptides. Values are means ± SD of three independent biological replicates. P values were determined by two-tailed Student’s t-test.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–7.
Supplementary Data 1 (download XLSX )
Full list of co-purified proteins as determined by AP–MS.
Supplementary Data 2 (download XLSX )
DEGs in gte1 gte6, gta1 gta2 gta3, gtb1 gtb2 gtb3, drmy1 and wdr5a-r mutants relative to the wild type.
Supplementary Data 3 (download XLSX )
DEGs in Col-H2O, Col-AZD, drmy1-MS and gtb1 gtb2 gtb3-MS relative to Col-MS.
Supplementary Data 4 (download XLSX )
DEGs in Col-H2O, Col-AZD, drmy1-MS and gtb1 gtb2 gtb3-MS relative to Col-MS.
Supplementary Data 5 (download XLSX )
Primers used in this study.
Source data
Source Data Fig. 1 (download PDF )
Unprocessed western blots.
Source Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Data Fig. 4 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 6 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 7 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 8 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 10 (download PDF )
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, X., Liu, ZZ., Yuan, DY. et al. Nutrient-driven TOR signalling controls a chromatin-associated complex for orchestrating plant growth and stress tolerance. Nat. Plants 11, 2115–2129 (2025). https://doi.org/10.1038/s41477-025-02107-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-025-02107-5
This article is cited by
-
Functional characterisation of Target of Rapamycin (TOR) signalling in Physcomitrella
Plant Cell Reports (2026)