Abstract
Engineering functional CO2-concentrating mechanisms into C3 crops holds great potential for enhancing photosynthetic efficiency. Limited CO2-inducible A (LciA), a chloroplast envelope bicarbonate channel belonging to the formate/nitrite transporter (FNT) family, is a key algal CO2-concentrating mechanism component and has been considered as a prime candidate for introduction into C3 plants. However, its application has been hindered by an incomplete mechanistic understanding. Here we report the cryogenic electron microscopy structure of Chlamydomonas reinhardtii LciA. Combining structural analysis and growth assays, we determined key residues governing substrate access and permeation, and identified two substitutions (K136A/A114F) that enhance LciA activity. We found that bicarbonate selectivity is governed by electrostatic coordination mediated by Lys220 and steric constraint imposed by Ala117 and Val267 within the selectivity filter. Leveraging these insights, we successfully engineered the bacterial FNT family nitrite channel NirC through site-directed mutagenesis to gain bicarbonate transport activity, and we characterized the bicarbonate transport capacity of the Chlamydomonas nitrite channels NAR1.1/NAR1.5, which were amenable to further enhancement. Taken together, our study establishes LciA as a fundamental template for engineering and identifying FNT proteins with bicarbonate transport capability, thereby greatly expanding the molecular toolkit for synthetic biology approaches aimed at boosting photosynthetic efficiency in both algae and crops.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The cryo-EM density map and atomic coordinates of LciA have been deposited in the Electron Microscopy Data Bank and the Protein Data Bank under the accession numbers EMD-64722 and 9V2A, respectively. Source data are provided with this paper.
References
Field, C. B., Behrenfeld, M. J., Randerson, J. T. & Falkowski, P. Primary production of the biosphere: integrating terrestrial and oceanic components. Science 281, 237–240 (1998).
Falkowski, P. et al. The global carbon cycle: a test of our knowledge of Earth as a system. Science 290, 291–296 (2000).
Hennacy, J. H. & Jonikas, M. C. Prospects for engineering biophysical CO2 concentrating mechanisms into land plants to enhance yields. Annu. Rev. Plant Biol. 71, 461–485 (2020).
Rae, B. D., Long, B. M., Badger, M. R. & Price, G. D. Functions, compositions, and evolution of the two types of carboxysomes: polyhedral microcompartments that facilitate CO2 fixation in cyanobacteria and some proteobacteria. Microbiol. Mol. Biol. Rev. 77, 357–379 (2013).
He, S., Crans, V. L. & Jonikas, M. C. The pyrenoid: the eukaryotic CO2-concentrating organelle. Plant Cell 35, 3236–3259 (2023).
Im, C. S. & Grossman, A. R. Identification and regulation of high light-induced genes in Chlamydomonas reinhardtii. Plant J. 30, 301–313 (2002).
Duanmu, D., Miller, A. R., Horken, K. M., Weeks, D. P. & Spalding, M. H. Knockdown of limiting-CO2-induced gene HLA3 decreases HCO3− transport and photosynthetic Ci affinity in Chlamydomonas reinhardtii. Proc. Natl Acad. Sci. USA 106, 5990–5995 (2009).
Ohnishi, N. et al. Expression of a low CO2-inducible protein, LCI1, increases inorganic carbon uptake in the green alga Chlamydomonas reinhardtii. Plant Cell 22, 3105–3117 (2010).
Kono, A. et al. Structure and function of LCI1: a plasma membrane CO2 channel in the Chlamydomonas CO2 concentrating mechanism. Plant J. 102, 1107–1126 (2020).
Kono, A. & Spalding, M. H. LCI1, a Chlamydomonas reinhardtii plasma membrane protein, functions in active CO2 uptake under low CO2. Plant J. 102, 1127–1141 (2020).
Yamano, T., Sato, E., Iguchi, H., Fukuda, Y. & Fukuzawa, H. Characterization of cooperative bicarbonate uptake into chloroplast stroma in the green alga Chlamydomonas reinhardtii. Proc. Natl Acad. Sci. USA 112, 7315–7320 (2015).
Atkinson, N. et al. Introducing an algal carbon-concentrating mechanism into higher plants: location and incorporation of key components. Plant Biotechnol. J. 14, 1302–1315 (2015).
Miura, K. et al. Expression profiling-based identification of CO2-responsive genes regulated by CCM1 controlling a carbon-concentrating mechanism in Chlamydomonas reinhardtii. Plant Physiol. 135, 1595–1607 (2004).
Fang, W. et al. Transcriptome-wide changes in Chlamydomonas reinhardtii gene expression regulated by carbon dioxide and the CO2-concentrating mechanism regulator CIA5/CCM1. Plant Cell 24, 1876–1893 (2012).
Wang, Y. & Spalding, M. H. Acclimation to very low CO2: contribution of limiting CO2 inducible proteins, LCIB and LCIA, to inorganic carbon uptake in Chlamydomonas reinhardtii. Plant Physiol. 166, 2040–2050 (2014).
Mariscal, V. et al. Differential regulation of the Chlamydomonas Nar1 gene family by carbon and nitrogen. Protist 157, 421–433 (2006).
Mukherjee, A. et al. Thylakoid localized bestrophin-like proteins are essential for the CO2 concentrating mechanism of Chlamydomonas reinhardtii. Proc. Natl Acad. Sci. USA 116, 16915–16920 (2019).
Mackinder, L. C. M. et al. A spatial interactome reveals the protein organization of the algal CO2-concentrating mechanism. Cell 171, 133–147.e114 (2017).
Vikramathithan, J. et al. Overexpression of Chlamydomonas reinhardtii LCIA (CrLCIA) gene increases growth of Nannochloropsis salina CCMP1776. Algal Res. https://doi.org/10.1016/j.algal.2020.101807 (2020).
McGrath, J. M. & Long, S. P. Can the cyanobacterial carbon-concentrating mechanism increase photosynthesis in crop species? A theoretical analysis. Plant Physiol. 164, 2247–2261 (2014).
Fei, C., Wilson, A. T., Mangan, N. M., Wingreen, N. S. & Jonikas, M. C. Modelling the pyrenoid-based CO2-concentrating mechanism provides insights into its operating principles and a roadmap for its engineering into crops. Nat. Plants 8, 583–595 (2022).
Long, B. M. et al. Carboxysome encapsulation of the CO2-fixing enzyme Rubisco in tobacco chloroplasts. Nat. Commun. 9, 3570 (2018).
Chen, T. et al. Engineering α-carboxysomes into plant chloroplasts to support autotrophic photosynthesis. Nat. Commun. https://doi.org/10.1038/s41467-023-37490-0 (2023).
Atkinson, N., Mao, Y., Chan, K. X. & McCormick, A. J. Condensation of Rubisco into a proto-pyrenoid in higher plant chloroplasts. Nat. Commun. 11, 6303 (2020).
Atkinson, N., Stringer, R., Mitchell, S. R., Seung, D. & McCormick, A. J. SAGA1 and SAGA2 promote starch formation around proto-pyrenoids in Arabidopsis chloroplasts. Proc. Natl Acad. Sci. USA 121, e2311013121 (2024).
Price, G. D. & Howitt, S. M. Plant science: towards turbocharged photosynthesis. Nature 513, 497–498 (2014).
Nölke, G. et al. The integration of algal carbon concentration mechanism components into tobacco chloroplasts increases photosynthetic efficiency and biomass. Biotechnol. J. 14, e1800170 (2019).
Förster, B. et al. The Chlamydomonas reinhardtii chloroplast envelope protein LCIA transports bicarbonate in planta. J. Exp. Bot. 74, 3651–3666 (2023).
Pengelly, J. J. et al. Transplastomic integration of a cyanobacterial bicarbonate transporter into tobacco chloroplasts. J. Exp. Bot. 65, 3071–3080 (2014).
Rolland, V., Badger, M. R. & Price, G. D. Redirecting the cyanobacterial bicarbonate transporters BicA and SbtA to the chloroplast envelope: soluble and membrane cargos need different chloroplast targeting signals in plants. Front. Plant Sci. 7, 185 (2016).
Uehara, S., Adachi, F., Ito-Inaba, Y. & Inaba, T. Specific and efficient targeting of cyanobacterial bicarbonate transporters to the inner envelope membrane of chloroplasts in Arabidopsis. Front. Plant Sci. 7, 16 (2016).
Uehara, S., Sei, A., Sada, M., Ito-Inaba, Y. & Inaba, T. Installation of authentic BicA and SbtA proteins to the chloroplast envelope membrane is achieved by the proteolytic cleavage of chimeric proteins in Arabidopsis. Sci. Rep. 10, 2353 (2020).
Rottet, S. et al. Engineering the cyanobacterial ATP-driven BCT1 bicarbonate transporter for functional targeting to C3 plant chloroplasts. J. Exp. Bot. 75, 4926–4943 (2024).
Fernandez, E. & Galvan, A. Inorganic nitrogen assimilation in Chlamydomonas. J. Exp. Bot. 58, 2279–2287 (2007).
Mukherjee, M., Vajpai, M. & Sankararamakrishnan, R. Anion-selective formate/nitrite transporters: taxonomic distribution, phylogenetic analysis and subfamily-specific conservation pattern in prokaryotes. BMC Genom. 18, 560 (2017).
Lü, W. et al. The formate/nitrite transporter family of anion channels. Biol. Chem. 394, 715–727 (2013).
Waight, A. B., Czyzewski, B. K. & Wang, D. N. Ion selectivity and gating mechanisms of FNT channels. Curr. Opin. Struct. Biol. 23, 499–506 (2013).
Suppmann, B. & Sawers, G. Isolation and characterization of hypophosphite-resistant mutants of Escherichia coli: identification of the FocA protein, encoded by the pfl operon, as a putative formate transporter. Mol. Microbiol. 11, 965–982 (1994).
Waight, A. B., Love, J. & Wang, D.-N. Structure and mechanism of a pentameric formate channel. Nat. Struct. Mol. Biol. 17, 31–37 (2009).
Wang, Y. et al. Structure of the formate transporter FocA reveals a pentameric aquaporin-like channel. Nature 462, 467–472 (2009).
Lü, W. et al. pH-dependent gating in a FocA formate channel. Science 332, 352–354 (2011).
Lü, W. et al. The formate channel FocA exports the products of mixed-acid fermentation. Proc. Natl Acad. Sci. USA 109, 13254–13259 (2012).
Beyer, L. et al. Coordination of FocA and pyruvate formate-lyase synthesis in Escherichia coli demonstrates preferential translocation of formate over other mixed-acid fermentation products. J. Bacteriol. 195, 1428–1435 (2013).
Clegg, S., Yu, F., Griffiths, L. & Cole, J. A. The roles of the polytopic membrane proteins NarK, NarU and NirC in Escherichia coli K-12: two nitrate and three nitrite transporters. Mol. Microbiol. 44, 143–155 (2002).
Jia, W., Tovell, N., Clegg, S., Trimmer, M. & Cole, J. A single channel for nitrate uptake, nitrite export and nitrite uptake by Escherichia coli NarU and a role for NirC in nitrite export and uptake. Biochem. J. 417, 297–307 (2008).
Lü, W. et al. Structural and functional characterization of the nitrite channel NirC from Salmonella typhimurium. Proc. Natl Acad. Sci. USA 109, 18395–18400 (2012).
Czyzewski, B. K. & Wang, D.-N. Identification and characterization of a bacterial hydrosulphide ion channel. Nature 483, 494–497 (2012).
Marchetti, R. V. et al. A lactate and formate transporter in the intraerythrocytic malaria parasite, Plasmodium falciparum. Nat. Commun. https://doi.org/10.1038/ncomms7721 (2015).
Wu, B. et al. Identity of a Plasmodium lactate/H+ symporter structurally unrelated to human transporters. Nat. Commun. https://doi.org/10.1038/ncomms7284 (2015).
Wiechert, M., Erler, H., Golldack, A. & Beitz, E. A widened substrate selectivity filter of eukaryotic formate-nitrite transporters enables high-level lactate conductance. FEBS J. 284, 2663–2673 (2017).
Peng, X. et al. Structural characterization of the Plasmodium falciparum lactate transporter PfFNT alone and in complex with antimalarial compound MMV007839 reveals its inhibition mechanism. PLoS Biol. https://doi.org/10.1371/journal.pbio.3001386 (2021).
Lyu, M., Su, C. C., Kazura, J. W. & Yu, E. W. Structural basis of transport and inhibition of the Plasmodium falciparum transporter PfFNT. EMBO Rep. https://doi.org/10.15252/embr.202051628 (2021).
Fang, S. et al. Molecular mechanism underlying transport and allosteric inhibition of bicarbonate transporter SbtA. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.2101632118 (2021).
Merlin, C., Masters, M., McAteer, S. & Coulson, A. Why is carbonic anhydrase essential to Escherichia coli? J. Bacteriol. 185, 6415–6424 (2003).
Du, J., Förster, B., Rourke, L., Howitt, S. M. & Price, G. D. Characterisation of cyanobacterial bicarbonate transporters in E. coli shows that SbtA homologs are functional in this heterologous expression system. PLoS ONE https://doi.org/10.1371/journal.pone.0115905 (2014).
Rexach, J., Fernández, E. & Galván, A. The Chlamydomonas reinhardtii Nar1 gene encodes a chloroplast membrane protein involved in nitrite transport. Plant Cell 12, 1441–1453 (2000).
Margulis, L. Symbiotic theory of the origin of eukaryotic organelles; criteria for proof. Symp. Soc. Exp. Biol. 29, 21–38 (1975).
Yang, Z. et al. Molecular mechanism underlying regulation of Arabidopsis CLCa transporter by nucleotides and phospholipids. Nat. Commun. 14, 4879 (2023).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D Struct. Biol. 74, 531–544 (2018).
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Pravda, L. et al. MOLEonline: a web-based tool for analyzing channels, tunnels and pores (2018 update). Nucleic Acids Res. 46, W368–w373 (2018).
Acknowledgements
We thank L. Liu from the University of Liverpool for critical reading and discussion of this manuscript. We thank H. Zhao and A. Dong at the cryo-EM centre of Fudan University and M. Zhang at the Cryo-EM Facility at the CAS Center for Excellence in Molecular Plant Sciences for their technical assistance on cryo-EM data collection. This work was supported by grants from the National Natural Science Foundation of China (nos 32530054 and 32025020 to P.Z. and no. 32401000 to Z.Y.), the Chinese Academy of Sciences (nos 317GJHZ2022023GC and XDB0630100 to P.Z.), the Shanghai Science and Technology Commission (no. 23310710100 to P.Z.), the National Key R&D Program of China (no. 2023YFA0914600 to J.H.), the Postdoctoral Fellowship Program of the China Postdoctoral Science Foundation (no. GZC20232665 to Z.Y.) and the Shanghai ‘Super Postdoctoral’ Incentive Program (no. 2023647 to Z.Y.).
Author information
Authors and Affiliations
Contributions
Z.Y. and J.G. designed and performed the bulk of the experiments. J.G. and Z.Y. carried out the protein expression and purification and sample preparation. Z.Y. and X.Z. carried out the cryo-EM data collection and structure determination. J.G., Z.Y. and F.L. carried out the growth assay. M.M. and F.Y. contributed to the protein purification. Z.Y., P.Z. and J.H. wrote the manuscript with input from the other authors. P.Z. and Z.Y. conceived the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Alistair McCormick, Takashi Yamano and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Sequence alignment of LciA and FNT homologs from bacteria and parasite.
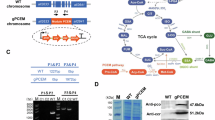
The structural architecture of LciA is shown on the top. Truncation variants LciAN39 (Δ1-38) and LciAN73 (Δ1-72) were designed based on sequence alignment with bacterial FNT homologs to delete the putative chloroplast-targeting peptide. Scissor symbols indicate the sites of N-terminal truncations corresponding to LciAN39 and LciAN73, with critical residues highlighted with boxes. Chlamydomonas reinhardtii LciA (UniProt: Q75NZ3), Salmonella typhimurium NirC (UniProt: E8XEH9), Escherichia coli FocA (UniProt: P0AC23), Clostridium difficile HSC (UniProt: Q186B7), Plasmodium falciparum PfFNT (UniProt: O77389).
Extended Data Fig. 2 Expression detected in Escherichia coli.
a–f, Western blots of LciA truncations and mutants. All mutations were performed in the LciAN39 truncation). (a) LciA full-length (FL) and truncations, related to Fig. 1b. (b) LciA K136 and R231 mutants, related to Fig. 2c,d. (c, d) LciA mutants in the permeation pathway, related to Fig. 3e. (e, f) LciA mutants in the selectivity filter, related to Fig. 4b,c. g–i, Western blots of bacterial and parasite FNT homologs. (g) NirC mutants, related to Fig. 5b. (h, i) FocA, HSC and PfFNT mutants, related to Extended Data Fig. 8d. j–m, Western blots of Chlamydomonas NAR1.1 and NAR1.5, related to Fig. 6c,f. (j, l) The NAR1.1 and NAR1.5 chimeras and mutants exhibit low expression levels, and strong signals are only detectable when they are immunoblotted individually. (k, m) When probed on the same membrane alongside LciAN39, only weak bands for NAR1.1 and NAR1.5 variants were visible, even though LciAN39 signal has been overexposed. Ponceau staining controls are included to verify equal loading. Each experiment was independently repeated three times with similar results.
Extended Data Fig. 3 Protein purification and structure determination.
a, Gel filtration profile on Superose 6 column and Coomassie-blue-stained SDS-PAGE analysis of LciA purified in buffer containing 0.03% DDM and 0.003% CHS. b, Gel filtration profile on Superose 200 column and Coomassie-blue-stained SDS-PAGE analysis of LciA-nanodiscs. For (a) and (b), Each experiment was independently repeated three times with similar results. c, Flowchart for Cryo-EM data processing. d, Representative cryo-EM micrograph (left) and 2D class averages (right). e, Local resolution estimation. The labels on the right are in unit Å. f, The gold-standard Fourier shell correlation curve. g, Cryo-EM densities of LciA protomer at a contour level of 6 σ.
Extended Data Fig. 4 Structural comparison.
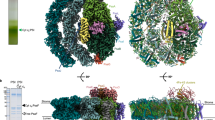
a, b, Density map (a) and electrostatic potential surface (b) of LciA pentamer. The intermembrane-space view (left) and cut-open view from the membrane plane (right) are shown. Each protomer is colored distinctly. The elongated lipid-like densities in the central tunnel of pentamer are colored yellow. c, Electrostatic potential surface of FNT homologs, shown in periplasmic or extracellular view. Structural models of Escherichia coli FocA (PDB: 3KCU), Salmonella typhimurium NirC (PDB: 4FC4), Clostridium difficile HSC (PDB: 3TDO) and Plasmodium falciparum PfFNT (PDB: 7E26) are shown. d, Structural comparison of a protomer between LciA and FNT homologs.
Extended Data Fig. 5 Functional characterization of functional enhanced LciA and engineered NirC mutants in the E. coli Δcan strain at pH 7.0.
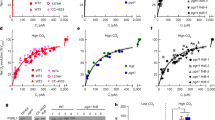
Phenotypes were recorded from the same plate at 24 h (a) and 36 h (b), with growth strength evaluated by dilution gradient and colony intensity. LciAN39 displayed a time-dependent improvement from weak to moderate growth, whereas the LciA mutants (K136A, K136A/A114F) sustained moderate growth, and NirC (I45A/I191V) consistently demonstrated the strongest growth.
Extended Data Fig. 6 Permeation pathway of LciA.
a, Cut-open electrostatic potential surface viewed from the membrane plane shows the channel architecture, including the external vestibule, central chamber and internal vestibule divided by two constrictions. The zoomed-in view shows the residues constituting the constrictions. b, Functional analysis of the LciA mutations in the E. coli Δcan strain at pH 7.0. All mutations were performed in the LciAN39 truncation.
Extended Data Fig. 7 Functional characterization of LciA mutations and chimeric NAR1.1/NAR1.5 in the E. coli Δcan strain.
a, Functional characterization of the A117S/V267I mutant in the E. coli Δcan strain at pH 9.0. b, Functional characterization of chimeric NAR1.1/NAR1.5 in the E. coli Δcan strain at pH 7.0.
Extended Data Fig. 8 Engineering of bacterial and parasite FNT homologs.
a–c, Structural alignments of LciA and EcFocA (a), CdHSC (b) and PfFNT (c) show the difference of the selectivity filter. Residues constituting the selectivity filter are shown as sticks. d, Functional characterization of engineered FNT-homolog variants in the E. coli Δcan strain at pH 9.0. e, Functional characterization of StNirC-I42F mutation (equivalent to LciA-A114F) and the NirC-I42F/I45A/I191V triple mutations in the E. coli Δcan strain at pH 7.0.
Extended Data Fig. 9 Sequence alignment of Chlamydomonas NAR1 homologs.
The structural architecture of LciA is shown on the top. Scissor symbols indicate the sites of N-terminal truncations corresponding to LciAN39 and LciAN73, with critical residues highlighted with boxes. Cr: Chlamydomonas reinhardtii. LciA/NAR1.2 (UniProt: Q75NZ3), NAR1.1 (UniProt: Q9LE25), NAR1.3 (UniProt: Q6IYG1), NAR1.4 (UniProt: Q6IYG4), NAR1.5 (UniProt: Q6IYG3), NAR1.6 (UniProt: Q6IYG2).
Extended Data Fig. 10 Engineering of Chlamydomonas NAR1 homologs.
a, c, e, Structural alignments of LciA with AlphaFold2-predicted NAR1.3 (a), NAR1.4 (c) and NAR1.6 (e) show the difference of the selectivity filter. Residues constituting the selectivity filter are shown as sticks. b, d, f, Functional characterization of engineered NAR1.3 (b), NAR1.4 (d) and NAR1.6 (f) in the E. coli Δcan strain at pH 9.0.
Supplementary information
Supplementary Information (download PDF )
Supplementary Table 1.
Source data
Source Data Fig. 1 (download PDF )
Unprocessed growth assays.
Source Data Extended Data Fig. 1 (download PDF )
Unprocessed gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Guo, J., Yang, Z., Zhang, X. et al. Structure of Chlamydomonas reinhardtii LciA guided the engineering of FNT family proteins to gain bicarbonate transport activity. Nat. Plants 12, 231–240 (2026). https://doi.org/10.1038/s41477-025-02200-9
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-025-02200-9