Abstract
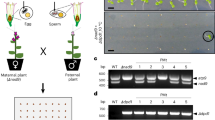
Mitochondria are inherited maternally in most plants as a classical paradigm of non-Mendelian inheritance, but the mechanism underlying paternal mitochondrial elimination (PME) remains almost unknown. We report here that angiosperms have evolved micromitophagy-mediated PME, in which vacuoles directly engulf paternal mitochondria via tonoplast invagination. We show that micromitophagy occurs specifically in male germline (MG) cells. To gain mechanistic insights, we used a vegetative-to-germline cell fate transition system to establish that micromitophagy is triggered by MG cell fate determination. We found evidence that ATG5 is translocated to vacuoles upon MG-cell-fate determination and interacts with mitochondrion-located HSP90.2 during mitochondrial engulfment by vacuoles, elucidating a cell-type-specific ATG neofunctionalization to mediate micromitophagy. This mechanism not only contributes to maternal inheritance of plant mitochondria but also supports the zygote-to-embryo transition. We further determined that micromitophagy is conserved in angiosperms but was continually optimized during evolution to support the best functioning of PME in MG cells with different properties. These findings bridge a long-standing gap in understanding plant PME with emerging mechanistic knowledge.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The sequence data from this article can be found in the TAIR database (www.arabidopsis.org) under the following accession numbers: At3g47440 (TIP5;1), At3g60460 (DUO1), At5g17290 (ATG5) and At5g56030 (HSP90.2). High-throughput sequencing data that support this study are available via the NCBI Gene Expression Omnibus (GSE162640). All unique/stable reagents and plasmids with transgene constructs generated in this study are available from the corresponding authors upon reasonable request with a completed materials transfer agreement. Source data are provided with this paper.
References
Birky, C. W. Jr. The inheritance of genes in mitochondria and chloroplasts: laws, mechanisms, and models. Annu. Rev. Genet. 35, 125–148 (2001).
Mittelsten Scheid, O. Mendelian and non-Mendelian genetics in model plants. Plant Cell 34, 2455–2461 (2022).
Correns, C. Vererbungsversuche mit blass (gelb) grünen und buntblättrigen Sippen bei Mirabilis jalapa, Urtica pilulifera und Lunaria annua. Z. Vererbungsl. 1, 291–329 (1909).
Birky, C. W. Jr. Uniparental inheritance of organelle genes. Curr. Biol. 18, R692–R695 (2008).
Nagata, N., Saito, C., Sakai, A., Kuroiwa, H. & Kuroiwa, T. The selective increase or decrease of organellar DNA in generative cells just after pollen mitosis one controls cytoplasmic inheritance. Planta 209, 53–65 (1999).
Wang, D. Y. et al. The levels of male gametic mitochondrial DNA are highly regulated in angiosperms with regard to mitochondrial inheritance. Plant Cell 22, 2402–2416 (2010).
Matsushima, R. et al. A conserved, Mg2+-dependent exonuclease degrades organelle DNA during Arabidopsis pollen development. Plant Cell 23, 1608–1624 (2011).
Ma, F. et al. The mitochondrial endonuclease M20 participates in the down-regulation of mitochondrial DNA in pollen cells. Plant Physiol. 178, 1537–1550 (2018).
Takami, T. et al. Organelle DNA degradation contributes to the efficient use of phosphate in seed plants. Nat. Plants 4, 1044–1055 (2018).
Tang, L. Y., Matsushima, R. & Sakamoto, W. Mutations defective in ribonucleotide reductase activity interfere with pollen plastid DNA degradation mediated by DPD1 exonuclease. Plant J. 70, 637–649 (2012).
Nagata, N. Mechanisms for independent cytoplasmic inheritance of mitochondria and plastids in angiosperms. J. Plant Res. 123, 193–199 (2010).
Mogensen, H. L. & Rusche, M. L. Quantitative ultrastructural analysis of barley sperm: I. Occurrence and mechanism of cytoplasm and organelle reduction and the question of sperm dimorphism. Protoplasma 128, 1–13 (1985).
Yu, H. S., Hu, S. Y. & Russell, S. D. Sperm cells in pollen tubes of Nicotians tabacum L.: three-dimensional reconstruction, cytoplasmic diminution, and quantitative cytology. Protoplasma 168, 172–183 (1992).
Mogensen, H. L. Exclusion of male mitochondria and plastids during syngamy in barley as a basis for maternal inheritance. Proc. Natl Acad. Sci. USA 85, 2594–2597 (1988).
Chung, K. P., Gonzalez-Duran, E., Ruf, S., Endries, P. & Bock, R. Control of plastid inheritance by environmental and genetic factors. Nat. Plants 9, 68–80 (2023).
Liu, P. et al. Mitopherogenesis, a form of mitochondria-specific ectocytosis, regulates sperm mitochondrial quantity and fertility. Nat. Cell Biol. 25, 1625–1636 (2023).
Mizushima, N. & Komatsu, M. Autophagy: renovation of cells and tissues. Cell 147, 728–741 (2011).
Huang, X. & Sun, M.-X. H3K27 methylation regulates the fate of two cell lineages in male gametophytes. Plant Cell 34, 2989–3005 (2022).
Yu, H. S. & Russell, S. D. Populations of plastids and mitochondria during male reproductive cell maturation in Nicotiana tabacum L.: a cytological basis for occasional biparental inheritance. Planta 193, 115–122 (1994).
Nakatogawa, H., Suzuki, K., Kamada, Y. & Ohsumi, Y. Dynamics and diversity in autophagy mechanisms: lessons from yeast. Nat. Rev. Mol. Cell Biol. 10, 458–467 (2009).
Gatica, D., Lahiri, V. & Klionsky, D. J. Cargo recognition and degradation by selective autophagy. Nat. Cell Biol. 20, 233–242 (2018).
Yoshimoto, K. & Ohsumi, Y. Unveiling the molecular mechanisms of plant autophagy—from autophagosomes to vacuoles in plants. Plant Cell Physiol. 59, 1337–1344 (2018).
Zhao, P., Zhou, X. M., Zhao, L. L., Cheung, A. Y. & Sun, M.-X. Autophagy-mediated compartmental cytoplasmic deletion is essential for tobacco pollen germination and male fertility. Autophagy 16, 2180–2192 (2020).
Schuck, S. Microautophagy-distinct molecular mechanisms handle cargoes of many sizes. J. Cell Sci. 133, jcs246322 (2020).
Zhou, J. et al. A non-canonical role of ATG8 in Golgi recovery from heat stress in plants. Nat. Plants 9, 749–765 (2023).
Zheng, X. et al. ATG8ylation-mediated tonoplast invagination mitigates vacuole damage. Nat. Commun. 16, 6621 (2025).
Kiššová, I. et al. Selective and non-selective autophagic degradation of mitochondria in yeast. Autophagy 3, 329–336 (2007).
Mijaljica, D., Prescott, M. & Devenish, R. J. Microautophagy in mammalian cells: revisiting a 40-year-old conundrum. Autophagy 7, 673–682 (2011).
Wang, L., Klionsky, D. J. & Shen, H. M. The emerging mechanisms and functions of microautophagy. Nat. Rev. Mol. Cell Biol. 24, 186–203 (2023).
Mizushima, N. et al. Dissection of autophagosome formation using Apg5-deficient mouse embryonic stem cells. J. Cell Biol. 152, 657–668 (2001).
Kang, B. H. et al. Regulation of tumor cell mitochondrial homeostasis by an organelle-specific Hsp90 chaperone network. Cell 131, 257–270 (2007).
Bao, F. et al. Arabidopsis HSP90 protein modulates RPP4-mediated temperature-dependent cell death and defense responses. New Phytol. 202, 1320–1334 (2014).
Borg, M. et al. Targeted reprogramming of H3K27me3 resets epigenetic memory in plant paternal chromatin. Nat. Cell Biol. 22, 621–629 (2020).
Williams, J. H., Taylor, M. L. & O’Meara, B. C. Repeated evolution of tricellular (and bicellular) pollen. Am. J. Bot. 101, 559–571 (2014).
Sanchez-Vera, V. et al. Autophagy is required for gamete differentiation in the moss Physcomitrella patens. Autophagy 13, 1939–1951 (2017).
Norizuki, T., Minamino, N., Sato, M., Tsukaya, H. & Ueda, T. Dynamic rearrangement and autophagic degradation of mitochondria during spermiogenesis in the liverwort Marchantia polymorpha. Cell Rep. 39, 110975 (2022).
Zhou, X., Zhao, P. & Sun, M.-X. Autophagy in sexual plant reproduction: new insights. J. Exp. Bot. 72, 7658–7667 (2021).
Chanoca, A. et al. Anthocyanin vacuolar inclusions form by a microautophagy mechanism. Plant Cell 27, 2545–2559 (2015).
Ding, X. et al. Microautophagy mediates vacuolar delivery of storage proteins in maize aleurone cells. Front. Plant Sci. 13, 833612 (2022).
Izumi, M., Ishida, H., Nakamura, S. & Hidema, J. Entire photodamaged chloroplasts are transported to the central vacuole by autophagy. Plant Cell 29, 377–394 (2017).
Nakamura, S., Hidema, J., Sakamoto, W., Ishida, H. & Izumi, M. Selective elimination of membrane-damaged chloroplasts via microautophagy. Plant Physiol. 177, 1007–1026 (2018).
Samakovli, D., Margaritopoulou, T., Prassinos, C., Milioni, D. & Hatzopoulos, P. Brassinosteroid nuclear signaling recruits HSP90 activity. New Phytol. 203, 743–757 (2014).
Huang, S. et al. HSP90s are required for NLR immune receptor accumulation in Arabidopsis. Plant J. 79, 427–439 (2014).
Yamamoto, H., Zhang, S. & Mizushima, N. Autophagy genes in biology and disease. Nat. Rev. Genet. 12, 382–400 (2023).
Al Rawi, S. et al. Postfertilization autophagy of sperm organelles prevents paternal mitochondrial DNA transmission. Science 334, 1144–1147 (2011).
Sato, M. & Sato, K. Degradation of paternal mitochondria by fertilization-triggered autophagy in C. elegans embryos. Science 334, 1141–1144 (2011).
Politi, Y. et al. Paternal mitochondrial destruction after fertilization is mediated by a common endocytic and autophagic pathway in Drosophila. Dev. Cell 29, 305–320 (2014).
Zhou, Q. et al. Mitochondrial endonuclease G mediates breakdown of paternal mitochondria upon fertilization. Science 353, 394–399 (2016).
Ben-Hur, S. et al. Egg multivesicular bodies elicit an LC3-associated phagocytosis-like pathway to degrade paternal mitochondria after fertilization. Nat. Commun. 15, 5715 (2024).
Sato, M., Sato, K., Tomura, K., Kosako, H. & Sato, K. The autophagy receptor ALLO-1 and the IKKE-1 kinase control clearance of paternal mitochondria in Caenorhabditis elegans. Nat. Cell Biol. 20, 81–91 (2018).
Mori, T., Kuroiwa, H., Higashiyama, T. & Kuroiwa, T. GENERATIVE CELL SPECIFIC 1 is essential for angiosperm fertilization. Nat. Cell Biol. 8, 64–71 (2006).
Sprunck, S. et al. Egg cell-secreted EC1 triggers sperm cell activation during double fertilization. Science 338, 1093–1097 (2012).
Wang, W. et al. DMP8 and 9 regulate HAP2/GCS1 trafficking for the timely acquisition of sperm fusion competence. Proc. Natl Acad. Sci. USA 119, e2207608119 (2022).
Rotman, N. et al. A novel class of MYB factors controls sperm cell formation in plants. Curr. Biol. 15, 244–248 (2005).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Li, W. et al. Three STIGMA AND STYLE STYLISTs pattern the fine architectures of apical gynoecium and are critical for male gametophyte–pistil interaction. Curr. Biol. 30, 4780–4788 (2020).
Xin, H. P. et al. Expressed sequence-tag analysis of tobacco sperm cells reveals a unique transcriptional profile and selective persistence of paternal transcripts after fertilization. Sex. Plant Reprod. 24, 37–46 (2011).
Zhao, P. et al. A bipartite molecular module controls cell death activation in the basal cell lineage of plant embryos. PLoS Biol. 11, e1001655 (2013).
Shi, C. et al. Maternal control of suspensor programmed cell death via gibberellin signaling. Nat. Commun. 10, 3484 (2019).
Acknowledgements
We thank S. Xiao (Sun Yat-sen University) for providing the A. thaliana atg5-1 (SAIL_129_B07) mutant seed, S. Yang (China Agricultural University) for providing the A. thaliana hsp90.2-2 mutant seed, G. N. Drews (University of Utah) for offering the pDD45::GFP marker line and D. Li (the Core Facility, Wuhan University) for electron tomography analysis. This work was supported by the National Natural Science Foundation of China (grant number 32130031 to M.-X.S.); the Funding for Discipline Development of Guizhou Province (grant number XKBF(2025)030 to M.-X.S.); Fujian Provincial Natural Science Foundation of China (grant number 2025J01025 to X.H.); the Natural Science Foundation of Xiamen, China (grant number 3502Z202471012 to X.H.); and the Open Research Fund of the State Key Laboratory of Hybrid Rice (Wuhan University) (grant number KF202402 to X.H.).
Author information
Authors and Affiliations
Contributions
X.H. designed and performed the research, analysed the data and wrote the draft. L.Z., Z.L., N.L., W.Z., F.G., T.C., C.S., X.Z., W.W. and H.C. performed the research. A.Y.C. analysed the data and finalized the paper. M.-X.S. designed the research, analysed the data and wrote and finalized the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Caiji Gao, Malgorzata Heidorn-Czarna and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The fission of paternal mitochondria during male gametogenesis.
a, The male gametogenesis process in the A. thaliana ProHTR10: HTR10-RFP line. The microspore (MSP) undergoes an asymmetrical division (pollen mitosis I) to form a larger VC and a smaller GC. VC exits the cell cycle, but GC completes pollen mitosis II (a symmetrical division) to produce two SCs. GC and SCs are collectively named as male germline cells. Thus, the male gametophyte (pollen) is a simplified plant body containing only two cell lineages, the VC wraps around the germline cells to form a “cell(s) within a cell” structure. The images are representative of at least three independent experiments with similar results. b-i, Paternal mitochondrial dynamic during male gametogenesis in A. thaliana. Mitochondrial morphology in microspore (b). Paternal mitochondrial morphology in the GC right after asymmetric microspore division (c-e). The paternal mitochondria in GC, which were inherited from microspore after asymmetric microspore division and then rapidly divided into the smaller mitochondria. With the formation of SC, paternal mitochondria in GC (f, g) and SC (h) were significantly smaller than their counterparts in VC. j-n, Paternal mitochondrial dynamic during male gametogenesis in O. sativa. Mitochondrial morphology in rice microspore(j). Paternal mitochondrial morphology in rice GC (k-m). The paternal mitochondria in GC, which were inherited from microspore after asymmetric microspore division and then also rapidly divided into the smaller mitochondria. The statistics of mitochondrial size during male gametogenesis of A. thaliana (i) and O. sativa (n). Results are from three independent experiments. The central lines represent the median value, the boxes represent the interquartile range and the whiskers extend to the minima and maxima. Statistical significance is determined by one-way ANOVA (P < 0.05, *; P < 0.001, ***). Scale bars, 5μm (a), 500 nm (b, c-h, j-m), 2μm (d, f). MSP, microspore; MN, microspore nucleus; GC, generative cell; GN, generative cell nucleus; SC, sperm cell; SN, sperm cell nucleus; VC, vegetative cell; VN, vegetative cell nucleus; EBP, early bicellular pollen; LBC, late bicellular pollen; ETP, early tricellular pollen; MP, mature pollen.
Extended Data Fig. 2 Gene expression profile analysis of ATG genes in wild-type vegetative cell (VC) and sperm cell (SC) of A. thaliana.
The data of expression profile come from the transcriptome database (GSE162640). The heatmap image was created by Origin software. The red box represents higher expression level, whereas the blue box represents lower expression level.
Extended Data Fig. 3 Inhibition of autophagosome formation by 3-MA did not block PME.
a, Schematic diagram of the 3-MA treatment experiments. Treatment ①: the pollen grains from a vacuole and mitochondrion double-labeled transgenic line were put into the PGM containing 3-MA, and the mitochondria dynamics in the SCs within germinated pollen tubes were observed after 5 h. Treatment ②: the pollen grains from the double-labeled transgenic line were put into the PGM containing 3-MA, and the mitochondria dynamics in the SCs within pollen tubes were observed after 8 h. Treatment ③: the pollen grains from the double-labeled transgenic line were placed on PGM without 3-MA and first cultured for 3 h, then continue cultured in PGM with 3-MA for another 5 h. b,d,f, The representative picture from treatment ① (b), treatment ② (d) and treatment ③ (f) showed that autophagy inhibitor 3-MA could not effectively interfere with PME to cause mitochondrial accumulation. c, The number statistics of mitochondria (Mt) in the SCs treated with 0 mM, 1 mM, 2.5 mM and 5 mM 3-MA after 5 h (treatment ①). e, The number statistics of mitochondria in the SCs treated with 0 mM and 2.5 mM 3-MA after 8 h (treatment ②). g, The number statistics of mitochondria in the SCs treated with 0 mM and 2.5 mM 3-MA after 5 h (treatment ③). Results are from three independent experiments. The central lines represent the median, the boxes represent the interquartile range and the whiskers extend to the minima and maxima. Statistical significance is determined by one-way ANOVA (P > 0.05, no significance). Scale bars, 5μm.
Extended Data Fig. 4 Vacuolar dynamic during male gametogenesis in A. thaliana.
a-c, Vacuolar dynamic in microspore. Vacuoles are monolayer membrane structures with light osmic acid-staining, and are various in shape and size. Vacuoles are indicated by black asterisk (*). As the microspore nucleus moved peripherally to a position near the cell wall (b and c), a large central vacuole appeared, which was formed by the fusion of pre-existing small vacuoles. d-i, Vacuolar dynamic in bicellular pollen. After asymmetric microspore division, the large vacuole was again divided into small vacuoles (indicated by black filled arrows) which were assigned to both GC (indicated by black hollow arrows) and VC (d). Then, the vacuoles in VC became gradually undetectable, while those in GC were clearly visible (black filled arrows in e-i). j-l, Vacuolar dynamic in mature pollen. The spherical vacuoles (indicated by black filled arrows) existed uniquely in SC (indicated by black hollow arrows), while no longer clearly visible in VC at the mature pollen stage. The black dotted line indicated the SC membrane, which is distinguished between SC and VC. By identifying the plasma membrane (indicated by hollow arrows in d, e, f, h, k) and cell size, we can simply distinguish between germline cells (GC and SC) and the VC. In addition, VC contains specifically lipid bodies (tiny white dots in (j)), which are a monolayer membrane structure surrounded by the endoplasmic reticulum and are different from the vacuole; also see Extended Data Fig. 4b. Lipid bodies accumulate significantly in the mature pollen, so that GC and SC can be distinguished easily according to existence of lipid bodies. (i and l) The higher magnification view of the regions enclosed in the black boxes in (h and k). The conceivable degradation of paternal mitochondria displaying fragmentary bilayer membrane structure and cristae was observed in vacuoles within GC (i). The images are representative of at least three independent experiments with similar results. Scale bars, 2μm (a-e, g, h, j, k, n), 0.5μm (f, i, l, o). MN, microspore nucleus; GC, generative cell; VC, vegetative cell; SC, sperm cell; SN, sperm cell nucleus.
Extended Data Fig. 5 Organelles degraded in vegetative cell (VC) via macroautophagy pathway.
The growing phagophore wrapped VC-specific lipid body (left) and then formed an autophagosome structure (right). The arrows indicate phagophore (left) and autophagosome (right). Scale bars, 200 nm. Lb, lipid body; Er, endoplasmic reticulum.
Extended Data Fig. 6 The transcript abundance of ATG5 and HSP90.2.
ATG5 and HSP90.2 were predominantly expressed in sperm cell (SC) and increased as the developmental fate of vegetative cell (VC) shifted to male germline cells. FPKM (Fragments Per Kilobase of transcript per Million mapped reads) was used to complete the normalization calculation. The data of expression profile come from the transcriptome database (GSE162640), and results are from three independent biological replicates and the error bars show standard deviation (SD).
Extended Data Fig. 7 The dysfunction of ATG5 or HSP90.2 disrupts the paternal mitochondria entering into vacuoles.
a,b, Paternal mitochondrial morphology in atg5 (left) and hsp90.2 (right). Similar to wild type (WT), the paternal mitochondria in the GCs of atg5 and hsp90.2 divided into the smaller mitochondria (a). The paternal mitochondria in the SCs of atg5 and hsp90.2 were also significantly smaller than their counterparts in the VCs (b). The magenta dotted lines indicate the SC membrane. c, The statistics of mitochondrial size during male gametogenesis of WT control, atg5 and hsp90.2. Results are from three independent experiments. The central lines represent the median, the boxes represent the interquartile range and the whiskers extend to the minima and maxima. Statistical significance is determined by two-way ANOVA (P > 0.05, no significance; P < 0.001, ***). d, The representative images from the SC pairs of WT control, atg5 and hsp90.2 with the background of the vacuole and mitochondrion double-labeled line show that the absence of ATG5 or HSP90.2 disturbed the paternal mitochondria entering into vacuoles (manifested as Mito-GFP and TIP5;1-RFP co-localization and indicated by white arrows). e, The frequency statistics of the mitochondrion-vacuole co-localization observed in every 30 SC pairs. The 30 SC pairs were randomly photographed and calculated for the frequency of mitochondrion-vacuole co-localization event. The same method was applied in WT and two mutants for data acquisition and difference comparison. Results are from nine independent experiments. Statistical significance is determined by one-way ANOVA (P < 0.001, ***) and the error bars show standard deviation (SD). Scale bars, 500 nm (a, b), 5μm (d). GC, generative cell; SC, sperm cell; VC, vegetative cell; EBP, early bicellular pollen; MP, mature pollen.
Extended Data Fig. 8 The micromitophagy was activated in the A. thaliana sperm-like cell.
a, A vegetative to sperm cell fate switch system (Huang and Sun, 2022) allowed the monitoring of micromitophagy upon MG cell fate determination. Immuno-gold labeling with ATG5 antibody showed that in these sperm-like cells ATG5 localized to vacuole during the micromitophagy process. The magenta arrows, anti-ATG5 (10 nm gold particles). b, Tomographic slice images of micromitophagy in the sperm-like cell. The corresponding 3D models are shown in Fig. 3n. Scale bars, 200 nm. Va, vacuole; Mt, mitochondrion.
Extended Data Fig. 9 Paternal micromitophagy during male gametogenesis in O. sativa and L. longiflorum.
a-i, The micromitophagy occurred in O. sativa GC (a-f) and SC (g-i). Mitochondria are labeled as Mt. There were many small dispersed vacuoles (indicated by the magenta asterisks (*) or labeled as Va) in GC and SC, but only one large central vacuole (CV) in VC. Micromitophagy occurred only in the small vacuoles (c-f, h, i), but not in the central vacuole (a, g). The trapped paternal mitochondria (PM) in a vacuole and formation of the micromitophagic vesicle (c). The blue arrows indicate the membrane of micromitophagic vesicles. Several mitochondria could be trapped in the same vacuoles (d, h, i). The micromitophagic vesicle with PM was degrading in the vacuole (e, f). (f and i) The higher magnification view of the regions enclosed in the dashed boxes in (e and h). j-l, The micromitophagy occurred in L. longiflorum GC. Type ➀: the vacuole membrane invagination-mediated PM microautophagy was exhibited in L. longiflorum GC (j, k). (k) shows the degrading mitochondrion in the vacuole. Type ➁: the micromitophagy mediated direct fusion of vacuole and PM in L. longiflorum GC (l). Scale bars, 5μm (a), 2μm (b, g, h), 500 nm (c-f, i-l). The images are representative of at least three independent experiments with similar results. GC, generative cell; SC, sperm cell; CV, central vacuole; Va, vacuole; Mt, Mitochondria.
Extended Data Fig. 10 The colocalization of paternal mitochondrion and vacuole in the fertilized egg cell.
Paternal mitochondrion (PMt) and vacuole (PVa) were able to colocalize in the fertilized egg cell (FEC). The ProTIP5;1: TIP5;1-RFP ProDUO1:Mito-GFP line (Two-marker line) were used as the paternal parent (♂), and WT was used as the maternal parent (♀). The images are representative of at least three independent experiments with similar results. Scale bars, 10μm. DSY, degraded synergid cell; FCC, fertilized central cell.
Supplementary information
Supplementary Information (download PDF )
Supplementary Table 1.
Supplementary Video 1 (download AVI )
Live-cell imaging showing that the mitochondria can enter the vacuole in sperm cells.
Supplementary Video 2 (download AVI )
3D models from electron tomography analysis of micromitophagy.
Supplementary Video 3 (download AVI )
3D models from electron tomography analysis of micromitophagy.
Supplementary Video 4 (download AVI )
3D models from electron tomography analysis of micromitophagy.
Supplementary Video 5 (download AVI )
3D models from electron tomography analysis of micromitophagy.
Source data
Source Data Fig. 2 (download PDF )
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Huang, X., Zhao, L., Liu, Z. et al. ATG5–HSP90.2-mediated micromitophagy as a cytological basis for maternal inheritance of plant mitochondria. Nat. Plants 12, 417–431 (2026). https://doi.org/10.1038/s41477-025-02216-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-025-02216-1