Abstract
The gut microbiota plays a potential role in the pathophysiology of depression through the gut–brain axis. This cross-sectional study in 400 participants from the PREDIMED-Plus study investigates the interplay between gut microbiota and depression using a multi-omics approach. Depression was defined as antidepressant use or high Beck Depression Inventory-II scores. Gut microbiota was characterized by 16S rRNA sequencing, and faecal metabolites were analysed via liquid chromatography-tandem mass spectrometry. Participants with depression exhibited significant differences in gut microbial composition and metabolic profiles. Differentially abundant taxa included Acidaminococcus, Christensenellaceae R-7 group, and Megasphaera, among others. Metabolomic analysis revealed 15 significantly altered metabolites, primarily lipids, organic acids, and benzenoids, some of which correlated with gut microbial features. This study highlights the interplay between the gut microbiota and depression, paving the way for future research to determine whether gut microbiota influences depression pathophysiology or reflects changes associated with depression.
Similar content being viewed by others
Introduction
Depression is a common mental disorder characterized by a depressed mood or loss of pleasure or interest in activities for an extended period. A worldwide estimated 3.8% of the population experience depression, including 5% of adults (4% among men and 6% among women), and 5.7% of adults older than 60 years1. Depression results from a complex interplay of social, psychological, and biological factors and it is closely related to and affected by physical health. Many of the lifestyle factors that influence depression (such as physical inactivity and harmful alcohol use) are also known risk factors for chronic diseases such as diabetes, cancer, cardiovascular, and respiratory diseases, all of which impact the gut microbiota2,3,4,5.
Alterations or imbalances in the gut microbiome have been observed in patients with major depressive disorder6. This phenomenon can be explained by the gut–brain axis, a complex bidirectional communication network involving neural, endocrine, immune, and metabolic pathways that connect the gut microbiota and brain, influencing both physiological and psychological processes. At the core of this axis, the enteric nervous system and the central nervous system engage in reciprocal interactions that are essential for their respective physiological functions7. This interaction is characterized by a balanced, two-way communication that is essential for maintaining the normal operation of both neural systems8. The vagus nerve transmits signals from the brain to the gut, complemented by indirect methods like endocrine influences. The gut, particularly its microbiota, processes these signals and responds via several pathways9, including vagal activation (direct or via interactions with other nervous systems), modulation of the immune system, and the production of signalling molecules like cytokines and microbial metabolites. These metabolites can either travel through the bloodstream, affecting the immune system, or stimulate the vagus nerve and other tissues. Both cytokines and metabolites can cross the blood–brain barrier and influence brain function10. For example, pro-inflammatory cytokines such as interleukin-6 and tumour necrosis factor-alpha are known to modulate neural activity and have been implicated in the pathophysiology of depression11. Similarly, microbial metabolites such as short-chain fatty acids (SCFAs), including butyrate and acetate, can influence brain function by regulating neurotransmitter synthesis, modulating inflammation, and impacting the integrity of the blood–brain barrier12,13. Other metabolites, like tryptophan-derived compounds, are directly involved in serotonin synthesis and neural signalling14. These pathways underscore the complex interactions between the gut microbiota and the brain, highlighting potential mechanisms by which gut-derived signals contribute to mental health and disease. The metabolic signatures of the gut microbiome can be detected in stools and blood. Faecal metabolites provide a functional readout of the activities of the gut microbiota and are increasingly recognized as valuable indicators of both gut health and overall well-being15.
Clinical studies investigating the association between gut microbiome and depressive disorder have reported mixed results16. This is probably due to the differences in gut microbiota attributable to age, sex, geographical area, diet and other factors17. The objective of the present study is to characterize the gut microbial composition and the faecal metabolomic signature associated with depression in a senior Mediterranean population with metabolic syndrome.
Results
Characteristics of the study population
The characteristics of the study population by depression status are described in Table 1. Our analysis showed that among the 400 study participants, women represented 38% of the reference group (RG) (N = 331) and 71% of the Depression Group (DG) (N = 69). Participants in the RG were more likely to be current or former smokers, have a higher education, drink more alcohol, and engage in more leisure time physical activity. DG individuals were more likely to be unemployed or housekeepers. No statistically significant differences between groups were found between the groups regarding age, BMI, and adherence to the Mediterranean diet. The median BDI-II score was 6 and 17 for the RG and DG groups, respectively.
DG individuals have a less rich and diverse microbial signature than RG participants and the overall microbiota composition is different between groups
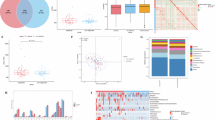
Alpha diversity indices showed significant variations in gut microbiota richness and diversity between the RG and DG groups. As depicted in Fig. 1, the Chao1 (richness) and Shannon (diversity) indices were significantly lower in the DG group (p = 0.004 and p = 0.01, respectively). In contrast, the Inverse Simpson index, which measures species evenness, showed no significant differences between groups (p = 0.097), suggesting that while overall richness and diversity differed, the relative distribution of taxa within samples remained comparable.
Wilcoxon Sum Rank was used to test the differences between groups and p ≤ 0.05 was considered as significant. DG depression group, RG reference group.
In the analysis of beta diversity, PCA performed on taxonomic counts at genus level revealed distinct variations in gut microbial composition between depression groups (Fig. 2). The first two principal components (PC1 and PC2) accounted for 21% and 18% of the total variation, respectively, illustrating group-level differences in microbial structure. Differences in gut microbial composition between groups were confirmed by PERMANOVA based on the Aitchison distance, adjusted for relevant covariates (p < 0.01).
DG depression group, PCA principal component analyses, RG reference group. The PERMANOVA model was adjusted by sex, age, BMI, smoking status, prevalence of diabetes and hypertension, education level, fibre consumption (g/day), alcohol consumption (g/day) and total physical activity (MET min/week).
Taking into account the DG group individuals, no differences in alpha and beta diversity were observed between participants under antidepressant treatment and those not taking antidepressants (Supplementary Figs. 1 and 2). Notably, the differences in alpha and beta diversity across depression groups remained consistent when stratifying by sex (Supplementary Figs. 3 and 4).
Differentially abundant genus between study groups
A differential abundance analysis was performed using ANCOM-BC, identifying 33 differentially abundant taxa between groups with an FDR ≤ 0.1 (Supplementary Table 1), among which 23 taxa remained significant at FDR ≤ 0.05. In contrast, MaAsLin2 revealed 9 differentially abundant taxa with FDR ≤ 0.1 (Supplementary Table 2), of which 6 were significant at FDR ≤ 0.05. As shown in Table 2, 8 taxa were identified as differentially abundant between groups across both methods, meeting an FDR threshold of FDR ≤ 0.05 in one method and FDR ≤ 0.1. Taxa with a negative effect size (LFC or coef.) were more abundant in the RG, whereas taxa with a positive effect size were more abundant in the DG. These differentially abundant taxa are visually represented in the taxonomic tree in Figure 3.
Letters with red background depict genera associated with reference group and letters with blue background depict genera associated with depression group. For significant taxa, the outermost layer depicts the phylum level followed by class, family and genus level.
In the DG group, individuals under antidepressant treatment showed no significant differences compared to those not taking antidepressants in differentially abundant taxa (Supplementary Fig. 5). Furthermore, the direction of the results regarding differentially abundant genera remained consistent across sexes, with taxa enriched in the DG showing similar increases in both men and women, and those depleted in the DG exhibiting comparable reductions across sexes (Supplementary Fig. 6).
Predicted functional abundance differences
Supplementary Figure 7 shows the predicted pathways that significantly differ between depression groups at an FDR ≤ 0.05. These pathways were related to type II diabetes mellitus, bile secretion, biosynthesis of secondary metabolites, carbohydrate metabolism, and amino acid metabolism.
Faecal metabolites associated with depression
Figure 4 shows the metabolites robustly associated with depression, sorted by their elastic-net regression coefficient values. Out of the 518 faecal metabolites, 24 exhibited strong associations with depression. Specifically, eleven metabolites with negative coefficients were associated with the RG, while thirteen metabolites with positive coefficients were associated with the DG. According to the HMDB classification system, the elastic net selected metabolites were distributed across various superclasses as follows: 11 lipids and lipid-like molecules, 7 organic acids and derivatives, 3 organoheterocyclic compounds, 2 benzenoids, and 1 nucleosides, nucleotides, and analogues.
Values are means ± SDs for the sets of metabolites consistently selected 10 times after 10 iterations of the elastic net regression procedure with 10-fold cross-validation using the best set of alpha (0.6) and lambda (0.045) values that returned the best model accuracy across all cross-validated results employing the training data of the 400 participants. Metabolites with negative coefficients, associated with the reference group are plotted on the left, whereas those with positive coefficients, related to the depression group are plotted on the right. 20-Carboxy-LTB4 20-Carboxy-leukotriene B4, C16 Sphinganine 16-carbon Sphinganine, CE 20:4 20-carbon tetra-unsaturated cholesterol ester, TG 54:1 54-carbon monounsaturated triglyceride.
Among the 24 metabolites selected through the elastic net penalized regression, 15 remained significantly associated with depression after adjusting for confounding variables (FDR ≤ 0.05). Figure 5 shows the negative logarithm of the FDR value obtained from ANCOVA plotted against the β coefficient of the linear models of the elastic-net selected metabolites, where metabolites with stronger associations appear further from zero.
A positive β-coefficient indicates association with the DG while a negative β-coefficient indicates that the metabolite is associated with the reference group. The model was adjusted by sex, age, BMI, smoking status, the prevalence of diabetes and hypertension, education level, fibre consumption (g/day), alcohol consumption (g/day) and total physical activity (MET min/week). An FDR ≤ 0.05 was deemed statistically significant (up dotted line). RG reference group, DG depression group, BMI body mass index, 20-Carboxy-LTB4 20-carboxy-leukotriene B4, C16 Sphinganine 16-carbon Sphinganine, CE 20:4 20-carbon tetra-unsaturated cholesterol ester.
In the DG group, participants under antidepressant treatment showed no significant differences compared to those not taking antidepressants regarding significant faecal metabolites (Supplementary Fig. 8). The associations between these metabolites and depression remained consistent across sexes (Supplementary Fig. 9).
Correlation between gut microbial taxa and faecal metabolites related to depression
Figure 6 illustrates the correlation patterns between differentially abundant taxa by depression status group and the significant faecal metabolites identified in our study. Notably, we observed that the overall direction of these correlations was consistent across different groups. Specifically, the Eubacterium siraeum group, Oscillospiraceae NK4A214 group, Christensenellaceae R-7 group, Oscillospiraceae UCG-002 and Oscillospiraceae UCG-005 exhibited similar correlation patterns with the significant metabolites. The genera Streptococcus, Acidaminococcus and Megasphaera showed a weaker inverse correlation pattern compared to the previously mentioned genus. Regarding the metabolites, the bile acids derivatives Tyrosocholic Acid and Phenylalanocholic acid showed the strongest negative correlations, while 20-Carboxy-leukotriene B4 and N-Acteylglutamic acid exhibited the strongest positive correlations. Both gut microbial taxa and metabolites were clustered according to depression status.
Taxa abundance was stratified by depression groups. Left-side: Class and Family of each differentially abundant Genus. Right-side: CLR abundance of taxa per group represented with boxplots and significance between groups. Bottom-side: Group where the metabolite is more abundant. FDR false discovery rate, RG reference group, DG depression group, 20-Carboxy-LTB4 20-Carboxy-leukotriene B4, C16 Sphinganine 16-carbon Sphinganine, CE 20:4 20-carbon tetra-unsaturated cholesterol ester.
Discussion
This study represents the first human study investigation to simultaneously employ untargeted faecal metabolomics and 16S rRNA-based metataxonomic analysis to examine associations between depression and gut microbiota. This dual approach offers a comprehensive analysis of the microbial community and its metabolic functions, offering new insights into the potential role of the gut-brain axis in depression. As a cross-sectional study using baseline data from the PREDIMED-Plus trial, which enroled participants aged 55–75 years with at least three components of metabolic syndrome, this sub-study uniquely explores the relationship between gut microbiota and depression within a naturalistic, real-world context. Unlike many studies that compare controlled groups specifically selected for the presence or absence of depression18,19, our population includes individuals who naturally exhibit high depressive symptomatology and those who are treated or not for this condition. This approach enhances the applicability of our findings, reflecting genuine interactions and health challenges faced by a population of community-dwelling older adults with normal daily lives, capturing real-world understanding of the gut-brain axis in depression. Within this context, we identified 15 significant metabolites and observed 8 bacterial genera with differential abundance between those participants with and without depression. Genera such as Christensenellaceae R-7 group, Eubacterium siraeum group, Oscillospiraceae NK4A214 group, Oscillospiraceae UCG-002 and Oscillospiraceae UCG-005 were more abundant in the reference group while genera Streptococcus, Acidaminococcus and Megasphaera were enriched in the depression group. Notably, we detected associations between these differentially abundant taxa and depression-related metabolites, implying potential interactions between microbial composition and metabolic output. The predicted functional analysis further revealed 8 significant pathways, some of which align with the metabolomic signatures identified in the faecal samples. Our results also highlighted significant differences in alpha and beta diversity indices between study groups, supporting the hypothesis of an association between depression and gut microbial composition.
While greater bacterial diversity is often considered beneficial for human health, its enrichment in depression remains debated18. Some studies have found that depression and anxiety severity are negatively associated with bacterial richness and alpha diversity20, with lower alpha diversity observed in participants with high anxiety/depression symptoms compared to controls21. Conversely, other studies have reported no significant differences in microbiota diversity between individuals with Major Depressive Disorder and healthy controls22,23. In our cohort, we observed significant differences in the Chao1 and Shannon indices, with higher values in the reference group, suggesting greater species richness and diversity in these individuals. Additionally, the overall gut microbiota composition differed significantly between the studied groups, aligning with existing research that, even in the absence of statistically significant differences in richness and diversity, the gut microbiota of depressed participants often exhibits distinct clustering patterns when compared to controls16.
Our differential abundance analysis revealed that all significantly altered genera were part of the Firmicutes phylum, a dominant group in the gut microbiota. The genera more abundant in the reference group were classified within the Clostridia class, including members of the Oscillospiraceae (UCG-002, UCG-005, NK4A214 group) and the Ruminococcaceae (Eubacterium siraeum group) families. These families play a key role in fermenting dietary fibres, leading to the production of short-chain fatty acids (SCFAs), which are crucial for maintaining gut homoeostasis and are known for their anti-inflammatory properties by suppressing pro-inflammatory cytokines like IL-6 and TNF-α, and by enhancing regulatory T-cell activity, which is relevant given the growing body of evidence linking chronic inflammation to depression24,25. They also play a role in strengthening the gut barrier, reducing systemic inflammation, and modulating neurotransmitter production. The enrichment of these genera in the reference group suggests a potential protective effect against the chronic low-grade inflammation often observed in depression. The anti-inflammatory effects of SCFAs may thus represent a potential mechanism for mitigating depressive symptoms. The Christensenellaceae R-7 group, also enriched in the reference group, has been associated with a lean body phenotype and healthier metabolic profiles26, which could indirectly influence mental health by mitigating metabolic dysfunction, a common comorbidity in individuals with depression27. Furthermore, Christensenellaceae has been proposed to stabilize microbial communities28, potentially contributing to resilience against dysbiosis and its downstream effects on the gut-brain axis. In the depression group, genera such as Acidaminococcus, Streptococcus, and Megasphaera were more abundant. Acidaminococcus, a genus involved in amino acid fermentation, produces glutamate, a key excitatory neurotransmitter in the brain29. Dysregulation of glutamate levels is associated with excitotoxicity, neuroinflammation, and the pathophysiology of depression30, particularly through its influence on glutamatergic signalling and neural plasticity31. The overrepresentation of Acidaminococcus in the depression group may reflect disruptions in amino acid metabolism32, potentially exacerbating neurochemical imbalances and contributing to depressive symptomatology. Further research is warranted to understand whether this genus plays a causal role or is a consequence of gut dysbiosis in depression. Streptococcus, typically associated with the oral microbiome33 was also enriched in the depression group. Its presence in the gut might indicate disruptions with the oral-gut microbial axis. Streptococcus is known to metabolize tryptophan into kynurenine, a pathway that reduces serotonin availability while increasing neurotoxic metabolites14, which is essential for mood regulation34. This shift in tryptophan metabolism could disrupt serotonergic signalling, a mechanism central to mood regulation14. Furthermore, elevated Streptococcus levels have been associated with inflammatory cytokine production, such as IL-635, which can cross the blood-brain barrier and influence neuroinflammation36, further implicating this genus in depression. Megasphaera, another genus enriched in the depression group, is recognized for its role in propionate production through the fermentation of dietary carbohydrates37. While propionate has been associated with beneficial effects, such as improved gut barrier function and reduced inflammation in some contexts38, excessive production could disrupt gut-brain communication by altering signalling pathways and promoting a pro-inflammatory state12. Elevated levels of Megasphaera may therefore represent a maladaptive response to dietary or metabolic stressors in individuals with depression.
Our metabolomic findings revealed associations between several metabolites and depression. For example, 20-Carboxyleukotriene B4, an omega-oxidized metabolite of leukotriene B4, has been implicated in depressive states through its role in promoting inflammation and oxidative stress, processes known to affect neural function and mood regulation39. Elevated levels of phytanic acid are known to be neurotoxic in excessive concentrations and might contribute to neural dysfunctions associated with depression40. Among the metabolites with higher concentrations in the depression group, valeric acid, a short-chain fatty acid, was significantly enriched. In animal models of depression, elevated valeric acid levels have been linked to changes in gut microbiota composition and alterations in gut-brain communication41, suggesting a potential role in modulating depressive behaviours, aligning with our findings. Reduced proline metabolism in the depression group also points to amino acid pathway disruption. Proline is a key precursor for neurotransmitters such as glutamate, and its reduced availability may influence neural plasticity and mood regulation. Aligning with our findings, proline metabolism has been found to be reduced in depression42. Tyrosocholic and phenylalanocholic acids, which cannot be synthesized by humans, are products of the bacterial conversion of bile acids and evidence shows decreased bile acid concentrations in depressive mice models42, suggesting that alterations in bile acid metabolism may play a role in the gut-brain axis and the pathophysiology of depression. Our results are in line with the mentioned evidence, and therefore these results suggest that bacteria-derived metabolites may play a role in the relationship between gut microbiota and depression. These findings collectively suggest a multifaceted biochemical interplay in depression, involving several metabolic pathways, some of which are exclusive to bacteria.
The predicted functional abundance analysis supported our metabolomic findings, showing correlations with the actual metabolomic signature described in this study. For instance, glutathione metabolism is required for the biosynthesis of leukotrienes such as 20-Carboxy-leukotriene B443 and the bile secretion pathway is related to cholic acid, the precursor of tyroscholic acid and phenilalanocholic acid44. However, the biosynthesis of phenylalanine, tyrosine, and tryptophan pathway, despite its potential impact on depression45, did not correspond to the detection of actual metabolites measured from this pathway in our study.
The correlation matrix analysis showed that genera from the same class shared similar correlation profiles. Bile acids, such as phenylalanocholic and tyrosocholic acids, exhibited the most substantial negative correlations with members of the Clostridia class, while N-acetylglutamic acid and 20-Carboxyleukotriene B4 showed the strongest positive correlations. These patterns suggest that taxa in the Clostridia class may influence the synthesis or breakdown of these compounds, consistent with findings in other studies46,47,48. Notably, significant correlations were mostly found with taxa linked to the reference group, underscoring their potential role in the metabolic pathways relevant to depression.
Our findings showed no significant differences in gut microbiota composition or metabolomic profiles between participants taking antidepressants and those not. This raises important questions about the role of the gut microbiome in the treatment of depression. It is possible that traditional antidepressants do not substantially modify the gut microbiota or that depression-related gut microbiota changes persist despite pharmacological treatment. However, these results do not negate the potential of the gut microbiome as a therapeutic target. Recent randomized controlled trials have shown that probiotic supplementation can reduce depressive symptomatology, suggesting that microbiota-targeted interventions could serve as adjunctive therapies to conventional treatments49,50,51. Future longitudinal studies are needed to explore whether microbiota-targeted therapies might complement or enhance current treatments, particularly in individuals who do not respond to antidepressants.
The interpretation of the findings in this study has to be contextualized within some limitations. First, due to the cross-sectional design of the present study, we cannot infer a temporal or causal relationship between exposure and outcome. Second, this study is an observational analysis using baseline data from older Mediterranean adults with overweight/obesity and metabolic syndrome, limiting the generalizability of the findings to other populations. Although we have adjusted our models by different covariates that can explain the variability of our findings, we cannot discard residual confounding, especially in relation to dietary factors that can affect both the exposure and outcome. Third, the BDI-II, while reliable for screening depressive symptoms, is not a diagnostic tool, which may affect the interpretation of symptom severity. Fourth, the intrinsic nature of 16S sequencing limits the analysis to genus-level taxonomic resolution and lacks functional information, having to rely on predictive methods. Last, the use of faecal metabolomics also presents challenges, as faecal samples may not fully reflect systemic metabolic changes detectable in blood. Also, stool consistency, a known factor influencing gut microbiota composition, was not recorded in our study. Given that altered gastrointestinal transit times are associated with depression, future research should incorporate validated measures of stool consistency, such as the Bristol Stool Scale to control for this variable and better understand microbiome-depression associations.
As an epidemiological study, our focus was on identifying associations that could be the base of future studies without investigating mechanisms. Replication in other cohorts would be essential to confirm the robustness and generalizability of the results. Future studies should specifically include clinical cohorts comprising individuals formally diagnosed with depression. Such studies would allow for validation of the identified key bacterial genera and metabolites in a clinical context and help establish whether these associations are consistent across different populations and diagnostic criteria. In addition, studies using model organisms are warranted in the future to validate findings reported in this study and explore causal relationships. This type of studies would provide controlled environments to dissect the interactions between gut microbiota, metabolites, and depressive behaviours, enhancing the applicability of these findings.
This study also has notable strengths. Although our sample is not representative of the general population, individuals with overweight or obesity at high risk of cardiometabolic diseases make up a significant portion of the global population. Thus, our findings may be relevant to similar community-dwelling older adults who may benefit from interventions supporting mental health. The well-defined population and extensive collection of high-quality standardized data allowed for rigorous statistical adjustments. The BDI-II is a well-standardized measure with strong psychometric properties, including high reliability and validity. It has been extensively tested across different populations and settings, which helps ensure that its results are consistent and accurate. Finally, the use of faecal metabolomics in this specific context provides direct insights into the gut microbiota’s metabolic activities, offering a comprehensive view of the gut environment and its interactions with the host.
Methods
Study design and population
This research is a cross-sectional analysis utilizing baseline data from a cohort of 400 individuals enrolled in the PREDIMED-Plus study52. Eligible participants included men and women aged 55-75 years who had no documented history of cardiovascular disease at enrolment. All participants were classified as overweight or obese (body mass index (BMI) between 25 and 40 kg/m²) and met at least three criteria of the metabolic syndrome at baseline53. The trial was registered at the International Standard Randomized Controlled Trial (Number: ISRCTN89898870, registration date: 2014) and received approval from the ethical review boards of all participating institutions. A detailed protocol has been published previously54 and can be accessed on the study website http://www.predimedplus.com/.
The current study focuses on a subset of participants from the PREDIMED-Plus centres in Reus, Barcelona-IMIM, Alicante, and Valencia who provided stool samples. At the time of faecal sample collection, participants were asked whether they had been undergoing antibiotic treatment or taking probiotics within the last two weeks. Participants who reported taking antibiotics during this period were excluded, and no participants reported probiotic use. Additionally, data on antibiotic and probiotic prescriptions were obtained from the questionnaire used to determine participant eligibility for the study. These data were cross-referenced with the participants’ responses provided before faecal sample collection to ensure accuracy. A flowchart outlining the study population is provided in Supplementary Fig. 10.
General evaluations, anthropometric data and blood biochemistry measurements
Participants completed a comprehensive questionnaire to gather data on socio-demographics and medical history. Physical activity during leisure time was estimated using the validated Regicor Short Physical Activity Questionnaire55. Waist circumference was measured twice at the midpoint between the lowest rib and the iliac crest with an anthropometric tape. Body weight and height were recorded twice using calibrated electronic scales and a wall-mounted stadiometer, respectively. Dietary fibre intake (g/day) was estimated using a validated semi-quantitative food frequency questionnaire56.
Depression status
Participants were considered to have depression if they were under antidepressant treatment, or they had a Beck Depression Inventory-II (BDI-II) score equal to or higher than 20 (Depression Group, DG). Participants with a score lower than 20 who were not taking antidepressants were categorized as reference groups (Reference Group, RG). The BDI-II is a standardized self-report questionnaire designed to assess the severity of depressive symptoms in adolescents and adults57. Comprising 21 items, participants indicate how they have felt over the past two weeks on a 4-point scale, resulting in a total score ranging from 0 to 63. The BDI-II encompasses a wide spectrum of depressive symptomatology, including cognitive, affective, somatic, and motivational domains. Scores can be categorized as reflecting minimal (0–13), mild (14–19), moderate (20–28), or severe (29–63) depression57. The instrument has demonstrated high internal consistency, with Cronbach’s alpha values often exceeding 0.90, and good test-retest reliability57. Moreover, its construct validity has been established through correlations with other established measures of depression and clinical diagnoses. Our categorization method based on the DBI-II score, while different from the more common cut-off of 14 used for screening mild depression, better distinguishes participants with clinically impactful depressive symptomatology, supporting the study’s focus on understanding relationships involving more severe depressive states. The cut-off of 20 has also been used in other studies to identify depressive states58,59. Including participants with BDI-II scores over 20 and those taking antidepressants allows for a clear distinction based on both self-reported depressive symptoms as indicated by the BDI-II score and clinical treatment status.
Stool samples collection, DNA extraction, 16S rRNA gene amplicon sequencing and faecal metabolomics analysis
Participants collected stool samples at home before the intervention began and stored them frozen until delivery to the laboratory. The samples were then divided into 250 mg aliquots and stored at −80 °C until analysis. Microbial DNA extraction was performed using the QIAamp PowerFecal DNA kit (Qiagen, Hilden, Germany) following the manufacturer’s instructions, with an initial 5-min lysis step using the FastPrep-24™ 5G Homogenizer (MP Biomedicals, Santa Ana, CA, USA). Extracted DNA was stored at −20 °C until further processing and its concentration and purity were measured using the Qubit 2.0 Fluorometer with the dsDNA BR Assay Kit (Thermo Fisher Scientific, Waltham, MA, USA). Triplicate PCR reactions were performed to amplify the V4 region (515F-806R) of the prokaryotic 16S ribosomal RNA (rRNA) gene.
Each 35 μl reaction contained 0.7 μl of unique barcoded primers (515F-n and 806R-n), 0.7 μl dNTPs, 0.35 μl Phusion Green Hot Start II High-Fidelity DNA Polymerase (Thermo Scientific, Landsmeer, The Netherlands), 7 μl 5× Phusion Green HF Buffer, and 25.5 μl DNAse- and RNAse-free water (Promega, Madison, WI, USA). The PCR cycle included an initial denaturation at 98 °C for 30 s, followed by 25 cycles of denaturation at 98 °C for 10 s, annealing at 50 °C for 10 s, elongation at 72 °C for 10 s, and a final extension at 72 °C for 7 min. PCR products were visualized on 1% agarose gel (~290 bp), purified using the CleanPCR kit (CleanNA, Alphen Aan den Rijn, The Netherlands), and quantified with the Qubit dsDNA BR Assay Kit. Sequencing libraries were constructed using 200 ng of purified PCR product and sequenced on an Illumina Novaseq platform. To ensure data quality, artificial mock communities, and negative control samples were included in each library, with seven stool samples sequenced in duplicates. Negative control samples were included in each library to control for potential contaminant sequences using nuclease-free water or material from DNA extraction blanks.
Sequencing libraries were demultiplexed and only read pairs with perfectly matching barcodes were used for downstream steps. Amplicon sequence variants (ASVs) were determined with the default settings of the DADA2 pipeline60 and taxonomy was assigned to each ASV using the Silva database (v138.1)61. A total of 1.080.337 paired-end reads per sample on average (median: 1.120.481, IQR: [379.416–1.645.145]) were generated across all study samples. After filtering, denoising, and chimera removal, an average of 948.831 non-chimeric reads per sample (median: 969.828, IQR: [334.276–1.415.867]) were retained, with 11% of sequences identified as chimeric and removed. Importantly, no samples were excluded due to low sequencing depth, as all exceeded a 10.000-read threshold that was used in this study.
Metabolomic profiling of the stool samples was conducted at the Broad Institute of Harvard University and MIT, employing four liquid chromatography-tandem mass spectrometry (LC-MS) techniques to profile endogenous metabolites in 400 baseline faecal water samples. The techniques used were: 1) hydrophilic interaction liquid chromatography for positive ion polar metabolites (HILIC-pos), 2) hydrophilic interaction liquid chromatography for negative ion metabolites (HILIC-neg), 3) reversed-phase chromatography for positive ion lipid profiling (C8-pos), and 4) reversed-phase chromatography with a T3 ultra-performance liquid chromatography column for negative ion detection of free fatty acids, bile acids, and metabolites of intermediate polarity (C18-neg). The methodologies for these four techniques have been detailed in previous publications62. For each technique, samples from each subject were randomized and analysed as a pool. To ensure quality control and standardization, paired pooled reference samples were inserted at intervals for analysis.
Microbiome data analysis
For the assessment of within-sample diversity (alpha diversity), Chao1, Shannon, and inverse-Simpson indices63,64,65 were computed using absolute counts of microbial taxa. Significance in alpha diversity between groups was assessed with the Wilcoxon signed-rank test66.
For between-sample diversity (beta diversity), a filtering threshold of 10% prevalence and a detection level of 0.1% at the genus level on relative abundance data was implemented. Beta diversity was evaluated with Euclidean distances using centered log-ratio (CLR) transformed counts, providing Aitchison distances67. This approach provides an appropriate measure of compositional dissimilarity, reducing the impact of unequal sampling depths and enhancing interpretability. The significance of the observed beta diversity patterns across depression groups was determined using permutational multivariate analysis of variance (PERMANOVA)68 adjusting for covariates. The results were visualized through principal component analysis (PCA) to provide an interpretable summary of the data structure based on compositional dissimilarities.
Differential abundance analysis (DAA) was performed using the same filtering criteria as applied in the beta diversity analysis and covariates were used to adjust the models. Two methods were employed for DAA: MaAsLin269 and ANCOM-BC70. We chose these methods because they are widely used in microbiome analysis, including studies where both are applied together to enhance robustness in differential abundance testing71,72. For MaAsLin2, a linear model was fitted to CLR-transformed absolute counts, while ANCOM-BC was executed also on CLR-transformed counts and default parameters without additional filtering. The Benjamini-Hochberg false discovery rate (FDR) procedure was utilized to adjust for multiple comparisons in both methods. Taxa achieving an FDR threshold of ≤0.05 in one method and ≤0.1 in the other were identified as differentially abundant between the groups under investigation. (i.e., a genus was called differentially abundant if it achieved FDR ≤ 0.05 by both MaAsLin2 and ANCOM-BC, or FDR ≤ 0.05 in MaAsLin2 and FDR ≤ 0.1 in ANCOM-BC, or FDR ≤ 0.05 in ANCOM-BC and FDR ≤ 0.1 in MaAsLin2). The application in parallel of both methods ensures the robustness of the results to methodological variations, as they offer different approaches to handling compositional data.
DADA2-generated ASVs were imported to PICRUSt 2.073, and the Kyoto Encyclopaedia of Genes and Genomes (KEGG)74 database was used to predict the functional gene content of the microbial communities present in the study samples. DAA of predicted pathways was performed using MaAsLin2 adjusting for covariates. A Linear Model was fitted to CLR-transformed pathway abundance counts, and an FDR procedure was used to adjust for multiple comparisons. FDR ≤ 0.05 was used to assess significance.
Faecal metabolomics data analysis
Prior to the analytical modelling, our metabolomic data underwent a pre-processing phase for ensuring the robustness of subsequent analyses. Acknowledging the typically skewed nature of metabolite distributions, we employed a rank-based inverse normal transformation (INT)75 for each metabolite. The INT procedure helps normalize data distributions, reducing the impact of skewness and outliers, which can otherwise compromise the assumptions of parametric statistical models. The INT method has been previously used in metabolomic studies76 and performs well with other nonnormal data77. The INT can be broken down into a two-step procedure. In the first, the observations are transformed onto the probability scale using the empirical cumulative distribution function (ECDF). In the second, the observations are transformed onto the real line, as Z-scores.
Metabolomics analysis began with an initial set of 537 identified faecal metabolites. Drug and drug-related metabolites were excluded resulting in 518 metabolites for downstream analysis.
Due to the high dimensionality and collinear nature of the metabolomics data, we employed a penalized regression model using the Elastic Net regularization technique78 to discern the relationship between depression status and faecal metabolite concentrations. Penalized regression models have been widely applied in metabolomics research, particularly in untargeted metabolomics studies requiring robust feature selection in high-dimensional datasets79,80,81. The Elastic Net approach was chosen for its capacity to combine the benefits of LASSO and Ridge penalties, which enables effective feature selection while managing multicollinearity among predictor variables.
Training and validation procedures were performed. The model training and hyperparameters alpha and lambda tuning were executed using the caret package in the R programming environment. A resampling strategy of 10-fold cross-validation (10-fold CV) was implemented, with the process being repeated 10 times to enhance the reliability of the model evaluation. This resampling was conducted on 80% of the collected dataset designated as the training set, with the remaining 20% held out for final model validation. The model with the best accuracy was accomplished with alpha and lambda values of 0.6 and 0.045, respectively. In a subsequent robustness check, 10-fold cross-validation elastic net models were run using the defined alpha and lambda values with different random seeds. Only metabolites with coefficients consistently different from zero across all 10 runs were retained, ensuring the selection of stable predictors for the outcome of interest.
The significance of the selected metabolites in relation to depression was further investigated using Analysis of Covariance (ANCOVA). This analysis was carried out controlling for covariates and to establish the independent associations of the metabolite concentrations on the outcome variable (study groups). Significance was determined based on an FDR-adjusted p-value threshold of ≤0.05. Pearson correlation analyses were conducted to assess the relationship between the relative abundance of selected significant gut microbial features identified and the rank-based inverse normal transformed concentrations of metabolites with FDR ≤ 0.05. This correlation analysis was stratified by depression status. The assessment of correlation significance was performed using the ‘cor.test’ function of the R ‘stats’ package and the resulting p-values were FDR adjusted. A heatmap depicting significant correlations was generated using the “ComplexHeatmap” package, providing a visual representation of these relationships.
A flowchart detailing the experimental methodology is provided in Supplementary Fig. 11.
Mediterranean diet adherence assessment
Adherence to the Mediterranean diet was assessed using the erMEDAS questionnaire82, a validated tool specifically designed to quantify Mediterranean dietary patterns. The erMEDAS score ranges from 0 to 17, with higher scores indicating greater adherence. This tool evaluates the consumption of key Mediterranean food groups, including olive oil, fruits, vegetables, legumes, nuts, fish, whole grains, and fermented dairy products, while also considering moderate wine consumption and limiting the intake of red and processed meats, sugary beverages, and ultra-processed foods. The minimal clinically important difference in the score is one point.
Confounder adjustments
The following statistical analyses were adjusted for covariates to control for potential confounding factors: PERMANOVA, differential abundance analyses (MaAsLin2 and ANCOM-BC), functional pathway predictions (PICRUSt2), and ANCOVA models for metabolomics analysis. All mentioned analyses were adjusted for sex, age (in years), BMI (in kg/m2), recruitment centre (Reus, Barcelona, Alicante, Valencia), adherence to the Mediterranean diet (range, 0–17; higher scores indicate greater adherence; minimal clinically important difference, 1 point), smoking status (current, former and never smoked), diabetes (yes/no), hypertension (yes/no), educational level (Higher Education, Secondary School and Primary School or less), dietary fibre intake (g/day), alcohol consumption (g/day) and total physical activity (MET·min/week) for both microbiome data and faecal metabolomics data. The individual contribution of each covariate to gut microbiota composition was assessed using PERMANOVA, and the results are presented in Supplementary Table 3.
Data availability
The datasets generated and analysed during the current study are not publicly available due to data regulations and for ethical reasons, considering that this information might compromise research participants’ consent because our participants only gave their consent for the use of their data by the original team of investigators. However, collaboration for data analyses can be requested by sending a letter to the PREDIMED-Plus steering Committee (predimed_plus_scommittee@googlegroups.com). The request will then be passed to all the members of the PREDIMED-Plus Steering Committee for deliberation.
References
Depressive Disorder (Depression) https://www.who.int/news-room/fact-sheets/detail/depression.
Kunasegaran, T., Balasubramaniam, V. R. M. T., Arasoo, V. J. T., Palanisamy, U. D. & Ramadas, A. The modulation of gut microbiota composition in the pathophysiology of gestational diabetes mellitus: a systematic review. Biology (Basel) 10, (2021).
Li, J. et al. Microbial and metabolic profiles unveil mutualistic microbe-microbe interaction in obesity-related colorectal cancer. Cell Rep. Med. 101429 https://doi.org/10.1016/J.XCRM.2024.101429 (2024).
Garcia-Fernandez, H. et al. Sex-specific differences in intestinal microbiota associated with cardiovascular diseases. Biol. Sex Differ. 15, (2024).
Shimizu, K. et al. Dysbiosis of gut microbiota in patients with severe COVID-19. Acute Med. Surg. 11, (2024).
Nikolova, V. L. et al. Perturbations in gut microbiota composition in psychiatric disorders: a review and meta-analysis. JAMA Psychiatry 78, 1 (2021) .
Montagnani, M. et al. The crosstalk between gut microbiota and nervous system: a bidirectional interaction between microorganisms and metabolome. Int. J. Mol. Sci. 24, (2023).
Cryan, J. F. et al. The microbiota–gut–brain axis. Physiol. Rev. 99, 1877–2013 (2019).
Wang, F. B. & Powley, T. L. Vagal innervation of intestines: afferent pathways mapped with new en bloc horseradish peroxidase adaptation. Cell Tissue Res. 329, 221–230 (2007) .
Zhu, F., Tu, H. & Chen, T. The microbiota–gut–brain axis in depression: the potential pathophysiological mechanisms and microbiota combined antidepression effect. Nutrients 14, (2022).
Miller, A. H. & Raison, C. L. The role of inflammation in depression: from evolutionary imperative to modern treatment target. Nat. Rev. Immunol.16, 22–34 (2015).
Dalile, B., Van Oudenhove, L., Vervliet, B. & Verbeke, K. The role of short-chain fatty acids in microbiota–gut–brain communication. Nat. Rev. Gastroenterol. Hepatol. 16, 461–478 (2019).
Braniste, V. et al. The gut microbiota influences blood-brain barrier permeability in mice. Sci. Transl. Med. 6, (2014).
O’Mahony, S. M., Clarke, G., Borre, Y. E., Dinan, T. G. & Cryan, J. F. Serotonin, tryptophan metabolism and the brain–gut–microbiome axis. Behav. Brain Res. 277, 32–48 (2015).
Zierer, J. et al. The fecal metabolome as a functional readout of the gut microbiome. Nat. Genet. 50, 790–795 (2018).
Gao, M. et al. Gut microbiota composition in depressive disorder: a systematic review, meta-analysis, and meta-regression. Transl. Psychiatry 13, 1–18 (2023).
Liu, X. et al. Comparative analysis of gut microbiota and fecal metabolome features among multiple depressive animal models. J. Affect. Disord. 314, 103–111 (2022).
Chen, Y. et al. Gut microbiota dysbiosis in depressed women: the association of symptom severity and microbiota function. J. Affect. Disord. 282, 391–400 (2021).
Lai, W. T. et al. Shotgun metagenomics reveals both taxonomic and tryptophan pathway differences of gut microbiota in major depressive disorder patients. Psychol. Med. 51, 90–101 (2021).
Madan, A. et al. The gut microbiota is associated with psychiatric symptom severity and treatment outcome among individuals with serious mental illness. J. Affect. Disord. 264, 98–106 (2020).
Simpson, C. A., Mu, A., Haslam, N., Schwartz, O. S. & Simmons, J. G. Feeling down? A systematic review of the gut microbiota in anxiety/depression and irritable bowel syndrome. J. Affect. Disord. 266, 429–446 (2020).
Naseribafrouei, A. et al. Correlation between the human fecal microbiota and depression. Neurogastroenterol. Motil. 26, 1155–1162 (2014).
Zheng, P. et al. Gut microbiome remodeling induces depressive-like behaviors through a pathway mediated by the host’s metabolism. Mol. Psychiatry 21, 786–796 (2016).
Mansuy-Aubert, V. & Ravussin, Y. Short chain fatty acids: the messengers from down below. Front. Neurosci. 17 (2023).
Faugere, M. et al. Quality of life is associated with chronic inflammation in depression: a cross-sectional study. J. Affect. Disord. 227, 494–497 (2018).
Oki, K. et al. Comprehensive analysis of the fecal microbiota of healthy Japanese adults reveals a new bacterial lineage associated with a phenotype characterized by a high frequency of bowel movements and a lean body type. BMC Microbiol. 16, 1–13 (2016).
Luo, H., Jiang, Z. L. & Ren, Y. Therapy management of metabolic disorder comorbidity with depression. Front. Psychol. 12, 683320 (2021).
Ang, W. S. et al. A keystone gut bacterium christensenella minuta—a potential biotherapeutic agent for obesity and associated metabolic diseases. Foods 12, 2485 (2023).
Boiangiu, C. D. et al. Sodium ion pumps and hydrogen production in glutamate fermenting anaerobic bacteria. J. Mol. Microbiol. Biotechnol. 10, 105–119 (2005).
Serafini, G., Rihmer, Z. & Amore, M. The role of glutamate excitotoxicity and neuroinflammation in depression and suicidal behavior: focus on microglia cells. Neuroscience 2, 127–130 (2015).
Sanacora, G. & Banasr, M. From pathophysiology to novel antidepressant drugs: glial contributions to the pathology and treatment of mood disorders. Biol. Psychiatry 73, 1172–1179 (2013).
Neis, E. P. J. G., Dejong, C. H. C. & Rensen, S. S. The role of microbial amino acid metabolism in host metabolism. Nutrients 7, 2930–2946 (2015).
Escobar-Arregocés, F. et al. Characterization of the oral microbiota and the relationship of the oral microbiota with the dental and periodontal status in children and adolescents with nonsyndromic cleft lip and palate. Systematic literature review and meta-analysis. Clin. Oral Investig. 28, (2024).
Liu, Z. et al. Regulation of serotonin production by specific microbes from piglet gut. J. Anim. Sci. Biotechnol. 14, (2023).
Moreira, L. B. et al. Presence of Streptococcus mutans and interleukin-6 and -10 in amniotic fluid. J. Matern. Fetal Neonatal Med. 35, 9463–9469 (2022).
Shan, C., Zhang, C. & Zhang, C. The role of IL-6 in neurodegenerative disorders. Neurochem. Res. 49, 834–846 (2024).
Louis, P. & Flint, H. J. Formation of propionate and butyrate by the human colonic microbiota. Environ. Microbiol. 19, 29–41 (2017).
Tong, L. C. et al. Propionate ameliorates dextran sodium sulfate-induced colitis by improving intestinal barrier function and reducing inflammation and oxidative stress. Front. Pharmacol. 7, 202262 (2016).
Regulska, M., Szuster-Głuszczak, M., Trojan, E., Leśkiewicz, M. & Basta-Kaim, A. The emerging role of the double-edged impact of arachidonic acid- derived eicosanoids in the neuroinflammatory background of depression. Curr. Neuropharmacol. 19, 278–293 (2020).
Chaudhary, S., Sahu, U., Kar, S. & Parvez, S. Phytanic acid-induced neurotoxicological manifestations and apoptosis ameliorated by mitochondria-mediated actions of melatonin. Mol. Neurobiol. 54, 6960–6969 (2017).
Chen, M., Wang, L., Lou, Y. & Huang, Z. Effects of chronic unpredictable mild stress on gut microbiota and fecal amino acid and short-chain fatty acid pathways in mice. Behav. Brain Res. 464, 114930 (2024).
Xie, Z. et al. Integrated multi-omics analysis reveals gut microbiota dysbiosis and systemic disturbance in major depressive disorder. Psychiatry Res. 334, 115804 (2024).
Wheelan, P. & Murphy, R. C. Metabolism of 6-trans-Isomers of leukotriene B4 in cultured hepatoma cells and in human polymorphonuclear leukocytes. J. Biol. Chem. 270, 19845–19852 (1995).
Guzior, D. V. & Quinn, R. A. Review: microbial transformations of human bile acids. Microbiome 9, 1–13 (2021).
Strasser, B., Sperner-Unterweger, B., Fuchs, D. & Gostner, J. M. Mechanisms of inflammation-associated depression: immune influences on tryptophan and phenylalanine metabolisms. Curr. Top. Behav. Neurosci. 31, (2017).
Bremm, K. D. et al. Generation of leukotrienes from human granulocytes by alveolysin from Bacillus alvei. Infect. Immun. 44, 188–193 (1984).
Shetty, N., Wren, M. W. D. & Coen, P. G. The role of glutamate dehydrogenase for the detection of Clostridium difficile in faecal samples: a meta-analysis. J. Hosp. Infect. 77, 1–6 (2011).
Coleman, J. P. & Hudson, L. L. Cloning and characterization of a conjugated bile acid hydrolase gene from Clostridium perfringens. Appl. Environ. Microbiol. 61, 2514 (1995).
Nikolova, V. L., Cleare, A. J., Young, A. H. & Stone, J. M. Acceptability, tolerability, and estimates of putative treatment effects of probiotics as adjunctive treatment in patients with depression: a randomized clinical trial. JAMA Psychiatry 80, 842–847 (2023).
Tian, P. et al. Bifidobacterium breve CCFM1025 attenuates major depression disorder via regulating gut microbiome and tryptophan metabolism: a randomized clinical trial. Brain Behav. Immun. 100, 233–241 (2022).
Kim, C. S. et al. Probiotic supplementation improves cognitive function and mood with changes in gut microbiota in community-dwelling older adults: a randomized, double-blind, placebo-controlled, multicenter trial. J. Gerontol. A Biol. Sci. Med. Sci. 76, 32–40 (2021).
Salas-Salvadó, J. et al. Effect of a lifestyle intervention program with energy-restricted mediterranean diet and exercise on weight loss and cardiovascular risk factors: one-year results of the PREDIMED-Plus Trial. Diabetes Care 42, 777–788 (2019).
Alberti, K. G. M. M. et al. Harmonizing the metabolic syndrome: a joint interim statement of the International Diabetes Federation Task Force on Epidemiology and Prevention; National Heart, Lung, and Blood Institute; American Heart Association; World Heart Federation; International Atherosclerosis Society; and International Association for the Study of Obesity. Circulation 120, 1640–1645 (2009).
Martínez-González, M. A. et al. Cohort Profile: design and methods of the PREDIMED-Plus randomized trial. Int. J. Epidemiol. 48, 387–388o (2019).
Molina, L. et al. Validation of the regicor short physical activity questionnaire for the adult population. PLoS ONE 12, e0168148 (2017).
Fernández-Ballart, J. D. et al. Relative validity of a semi-quantitative food-frequency questionnaire in an elderly Mediterranean population of Spain. Br. J. Nutr. 103, 1808–1816 (2010).
Beck, A., Steer, R. & Brown, G. W. Beck Depression Inventory–II (BDI-II). APA PsycTests https://doi.org/10.1037/T00742-000 (1996).
Maksimovskiy, A. L., Okine, C., Cataldo, A. M. & Dillon, D. G. Sluggish retrieval of positive memories in depressed adults. Cogn. Affect. Behav. Neurosci. 22, 1172–1182 (2022).
Gehrman, P. R., Bartky, E. J., Travers, C. & Lapidus, K. A Fully remote randomized trial of transcranial alternating current stimulation for the acute treatment of major depressive disorder. J. Clin. Psychiatry 85, (2024).
Callahan, B. J. et al. DADA2: high resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581 (2016).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590 (2013).
Kostic, A. D. et al. The dynamics of the human infant gut microbiome in development and in progression towards type 1 diabetes. Cell Host Microbe 17, 260 (2015).
Chao, A. Estimating the population size for capture-recapture data with unequal catchability. Biometrics 43, 783 (1987).
Shannon, C. E. A mathematical theory of communication. Bell Syst. Tech. J. 27, 623–656.
Simpson, E. H. Measurement of diversity [16]. Nature 163, 688 (1949).
Wilcoxon, F. Individual comparisons by ranking methods. Biometr. Bull. 1, 80 (1945).
Aitchison, J., Barceló-Vidal, C., Martín-Fernández, J. A. & Pawlowsky-Glahn, V. Logratio analysis and compositional distance. Math. Geol. 32, 271–275 (2000).
Anderson, M. J. A new method for non-parametric multivariate analysis of variance. Austral Ecol. 26, 32–46 (2001).
Mallick, H. et al. Multivariable association discovery in population-scale meta-omics studies. PLoS Comput. Biol. 17, e1009442 (2021).
Lin, H. & Peddada, S.D. Analysis of compositions of microbiomes with bias correction. Nat. Commun. 11, 1–11 (2020).
Wallen, Z. D. et al. Metagenomics of Parkinson’s disease implicates the gut microbiome in multiple disease mechanisms. Nat. Commun. 13, (2022).
Faggiani, L. D. et al. Effect of ultra-processed food consumption on the gut microbiota in the first year of life: findings from the MINA-Brazil birth cohort study. Clin. Nutr. 46, 181–190 (2025).
Douglas, G. M. et al. PICRUSt2 for prediction of metagenome functions. Nat. Biotechnol. 38, 685 (2020).
Kanehisa, M. & Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27 (2000).
McCaw, Z. R., Lane, J. M., Saxena, R., Redline, S. & Lin, X. Operating characteristics of the rank-based inverse normal transformation for quantitative trait analysis in genome-wide association studies. Biometrics 76, 1262–1272 (2020).
García-Gavilán, J. F. et al. Olive oil consumption, plasma metabolites, and risk of type 2 diabetes and cardiovascular disease. Cardiovasc. Diabetol. 22, 340 (2023).
Bishara, A. J. & Hittner, J. B. Testing the significance of a correlation with nonnormal data: comparison of Pearson, Spearman, transformation, and resampling approaches. Psychol. Methods 17, 399–417 (2012).
Zou, H. & Hastie, T. Regularization and variable selection via the elastic net. J. R. Stat. Soc. Ser. B Stat. Methodol. 67, 301–320 (2005).
Zhang, Y. et al. Metabolomic profiles of sleep-disordered breathing are associated with hypertension and diabetes mellitus development. Nat. Commun. 15, (2024).
Li, J. et al. The Mediterranean diet, plasma metabolome, and cardiovascular disease risk. Eur. Heart J. 41, 2645–2656 (2020).
Bever, A. M. et al. Metabolomic signatures of inflammation and metabolic dysregulation in relation to colorectal cancer risk. J. Natl Cancer Inst. 116, 1126–1136 (2024).
Schröder, H. et al. Validity of the energy-restricted Mediterranean Diet Adherence Screener. Clin. Nutr. 40, 4971–4979 (2021).
Acknowledgements
The authors thank all PREDIMED-Plus participants and investigators. CIBEROBN, CIBERESP, and CIBERDEM are initiative of the Instituto de Salud Carlos III (ISCIII), Madrid, Spain. The authors also thank the PREDIMED-Plus Biobank Network as a part of the National Biobank Platform of the ISCIII for storing and managing the PREDIMED-Plus biological samples. This work was supported by the official Spanish Institutions for funding scientific biomedical research, CIBER Fisiopatología de la Obesidad y Nutrición (CIBEROBN) and Instituto de Salud Carlos III (ISCIII), through the Fondo de Investigación para la Salud (FIS), which is co-funded by the European Regional Development Fund (six coordinated FIS projects lead by J.S.-S. and J.Vi., including the following projects: PI13/00233, PI13/00728, PI13/00462, PI14/01206, PI14/ 00696, PI16/00533, PI16/00366, PI16/00501, PI17/01441, PI17/00855, PI19/00017, PI19/00781, PI19/00576, PI20/00557, PI21/0046; the Especial Action Project entitled: Implementación y evaluación de una intervención intensiva sobre la actividad física Cohorte PREDIMED-Plus grant to J.S.-S.; the Recercaixa (number 2013ACUP00194) grant to J.S.-S.; grants from the Consejería de Salud de la Junta de Andalucía (PI0458/2013, PS0358/2016, PI0137/2018); the PROMETEO/ 2017/017 and PROMETEO/2021/21 grants from the Conselleria de Innovación, Universidades, Ciencia y Sociedad Digital from the Generalitat Valenciana; and by NIH grant R01DK127601. This research was also partially funded by the Eat2beNICE/H2020-SFS-2016-2 EU- H2020 European grant, and the Horizon 2020 PRIME study (Prevention and Remediation of Insulin Multimorbidity in Europe; grant agreement #847879). J.S.-S., the senior author of this paper, was partially supported by ICREA under the ICREA Academia programme. A.H.-C. is supported by a predoctoral grant from Martí Franquès— INVESTIGO research fellowship funded and supported by NextGenerationEU, Servicio Público de Empleo Estatal and Universitat Rovira i Virgili (2022PMF-INV-01). I.M.-I. was supported by a Miguel Servet type II grant from Instituto de Salud Carlos III, Madrid, Spain (CPII21/00013). J.N. is supported by a predoctoral grant from Ministerio de Ciencia, Innovación y Universidades (FPU 20/00385). None of the funding sources took part in the design, collection, analysis, interpretation of the data, writing the report, or in the decision to submit the manuscript for publication.
Author information
Authors and Affiliations
Contributions
All the principal PREDIMED-Plus investigators contributed to the study concept and design and to data extraction from the PREDIMED-Plus participants. A.H.-C., J.F.G.-G. and J.S.-S. contributed to the concept and design of the present study. A.H.-C. wrote the first draft and performed the statistical analyses under the supervision of J.F.G.-G. and A.A. A.H.-C., J.F.G.-G. and J.S.-S. are the guarantors of this work and, as such, had full access to all the data in the study and take responsibility for the integrity of the data and the accuracy of the data analysis. All authors reviewed the manuscript for important intellectual content and approved the final version to be published.
Corresponding authors
Ethics declarations
Competing interests
J.S.-S. reports serving on the board of and receiving grant support through his institution from the International Nut and Dried Fruit Council, serving on the board of the Instituto Danone Spain and the International Danone institute. None of the other authors declare competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hernández-Cacho, A., García-Gavilán, J.F., Atzeni, A. et al. Multi-omics approach identifies gut microbiota variations associated with depression. npj Biofilms Microbiomes 11, 68 (2025). https://doi.org/10.1038/s41522-025-00707-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41522-025-00707-9
This article is cited by
-
Fecal metaproteomics reveals alterations in gut microbiota and intestinal proteins in adolescents with bipolar depression
Translational Psychiatry (2026)
-
Beyond the Gut: Integrating Oral Microbiota into the Microbiota–Brain Axis in Depression
Molecular Neurobiology (2026)