Abstract
The assembly mechanisms of soil microbiome during plant disease progression remain incomplete. This study linked potato common scab (PCS) severity to destabilized soil microbiome dynamics in potato geocausphere soil (GS) through integrated metagenomic analysis and culture-based experiments. Across four Shandong fields, PCS-infected GS exhibited an 11.66% reduction in bacterial α-diversity (Shannon index) and elevated stochastic community assembly. Bacterial community structure explained 39.28% (GS) and 15.96% (bulk soil) of PCS variance, outperforming pathogen abundance contributions (14.39% GS, 7.33% BS). Two microbial interaction patterns emerged in GS: 1) synchronized shifts between pathogens and beneficial taxa as PCS intensified; 2) stochastic assembly governing entire communities but deterministic processes dominating specialized subgroups. These results propose a microbial stability framework connecting soil microbiome structural rules to plant disease progression, emphasizing community-level dynamics over pathogen-centric explanations.
Similar content being viewed by others
Introduction
Soil serves as the richest repository of microbial species for plants. While the health of plants is constantly influenced by the soil microbiota, they also depend on these microbes to sustain their own vitality1,2,3. The emergence of plant diseases is frequently preceded by alterations in the state of the microbial community, resulting in a disruption of its homeostasis4. Typically, this disruption manifests in a surge of microbial population, a decline in diversity, and shifts in the composition of microbial communities3,4. Nevertheless, the question remains unresolved whether there are discernible patterns in the variations of microbial community assembly within the intricate network of plant–microbe interactions, especially across heterogeneous soil environments. Furthermore, it remains unknown whether and how the microecological mechanisms, for instance, the stability of soil microbial communities contributes to the pathogen suppression.
How does the soil microbiome undergo alterations due to plant soil-borne diseases? In a healthy plant-soil ecosystem, plants regulate their rhizosphere microbiome via their “M” genes (Microbiome Genes, referring to key host genes that shape or regulate the composition and function of symbiotic or interacting microbial communities such as bacteria and fungi)2,3, by collaborating with root cells, immune proteins, antagonistic substances, and other factors to consolidate a defense barrier against invading pathogens5. This homeostatic system maintains a resilient network of cooperation and competition among soil microorganisms, preventing the unchecked proliferation of certain microorganisms, both exogenous and endogenous6. However, the emergence of soil-borne plant diseases frequently disrupts this rhizosphere microbial network, thus resulting in alterations in network size, connectivity, and complexity7. Additionally, these diseases can temporarily compromise the plant’s capacity to regulate its rhizosphere microbiome through the “amplification-selection” process4,8, leading to changes in microbial community assembly patterns7,9. In healthy hosts, the microbiota typically exhibit a relatively stable and consistent structure; however, when hosts are subjected to environmental stress or disease, the microbiota can become imbalanced to varying degrees, with changes among individuals displaying high variability and idiosyncrasy. Depending on the specific interaction mechanisms between plants and their microbiota, changes in the rhizosphere microbiome following disease occurrence can be described by Anna Karenina principle (AKP). This principle is used in microbial community studies to delineate a phenomenon whereby individuals under stressful conditions exhibit greater variability in microbial community composition compared to healthy individuals4. Correspondingly, Anti-AKP is used to denote that, under diseased conditions, the host’s microbiota tends to converge, typically driven by ubiquitous enrichment due to specific environmental selection pressures, resulting in a consistent dysbiotic pattern4. It is worth noting that the overall changes in the microbial community may not fully capture the intricate shifts occurring within all microbiota within the community. Even within the same microbial community, diseases can trigger diverse evolutionary trajectories among different microbial taxa. What is the role of alterations in soil microbiota assembly that accompany disease in disease development?
Within the microbial community, not all alterations in strain populations directly influence disease progression. In fact, most microorganisms may not contribute directly, and their alterations may stem more from changes in ecological niches. Notably, disruptions in microbial community homeostasis may lead to dysfunction of the microbiota, which is often harmful to the host10,11. Microbial communities are diverse assemblies of microorganisms, some of which maintain close mutualistic relationships with pathogens. These microorganisms, along with the pathogens, collectively constitute the pathobiome12. Some microorganisms can act as “helpers” to the pathogens, facilitating their growth and colonization13. During disease development, plants are not inactive. Numerous studies have observed that some specific beneficial taxa become enriched in diseased samples following disease onset7. With the involvement of plants, some beneficial strains may become key species in the microbial community, thereby creating more opportunities for restoring microbial community balance7,14,15. There are studies indicating that the development of certain plant diseases, such as potato common scab (PCS), can be influenced by soil disease suppression16. In disease-suppressive soils, despite exceeding pathogen thresholds, the presence of susceptible host plants, and favorable climatic conditions, disease incidence or severity typically remains low17. It has been reported that disease-suppressive soils are formed when plants accumulate beneficial microorganisms in the soil through a “cry for help” strategy after experiencing a severe disease outbreak18,19,20. These disease-suppressive microbial communities play a crucial role in the formation of disease-resistant soils. These microorganisms can serve as a disease-resistant soil legacy, which can be rapidly mobilized by the plant to ward off disease development after a future attack19,21. The MAMP-triggered immune pathways are activated to higher levels in disease-suppressive soils22. The pieces of evidence underscore the potential significance of the soil microbiome in influencing disease development. However, current research lacks the quantitative evidence to determine which factor has the greatest potential to influence disease development in soils - the status of the soil microbiome, pathogen populations, or soil properties.
In this study, we focused on PCS to investigate the interaction between soil microbiota assembly and disease development. The PCS is a worldwide serious soil-borne disease caused by several Streptomyces species, including S. scabies, S. acidiscabies, S. turgidiscabies, and others23. Importantly, PCS is not simply influenced by the pathogen abundance in the soil. Recent researches have reported links between soil microbial communities, soil physicochemical properties, and the development of PCS24,25. The PCS has relatively clear criteria for the delineation of disease severity levels, and there was a significant correlation between PCS severity and the microbial communities and pathogen abundance in potato geocausphere soils (GS)24. Our objectives were threefold: (i) to elucidate generic relationships between microbial community assembly and PCS severity across different soil conditions; (ii) to quantitatively assess the relative contributions of soil microbial community, pathogen abundance and physicochemical properties to PCS severity; and (iii) to delve into the microbiological mechanisms by which soil microbiota homeostasis suppresses PCS. For this study, experiments were conducted in four fields, representing two distinct climates and two soil types (Fig. 1A, B). All fields had experienced PCS for over three consecutive years, met the necessary bulk soil (BS) conditions for PCS occurrence. Plants exhibiting varying PCS severity were selected from each fields and categorized into two groups: H (PCS severity ≥4) and L (PCS severity 1–3). For each plant, two soil components, GS and BS, were analyzed. In addition, we measured the physicochemical properties of soils from each location (Supplementary Table 1) and used six different culture media to isolate bacterial strains (Supplementary Table 2).
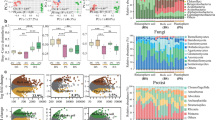
A Soil samples were collected from four fields, including TX and TZ in Tengzhou and JD and JF in Jiaozhou. B Two compartments of soil, such as GS and BS, were sampled for each selected plant. The GS represented the soil within 2 mm proximity of the tuber surfaces. The BS was collected from the plant growth area at a depth of 5–15 cm (from the cultivation hole after plant removal). Both PCoA and ADONIS showed that the soil physicochemical properties (C), bacterial communities (D). E PCoA revealed variations in GS bacterial, eukaryotic, and archaeal communities between fields or PCS severities.
Results
The response of GS bacterial community assembly to PCS exhibits characteristics of regional universality
PCoA and ADONIS analyses revealed significant differences among fields, which were observed in the physicochemical properties of BS (ADONIS, R² = 0.53, P = 0.001) (Fig. 1C), as well as in the community compositions of bacteria (Fig. 1D), eukaryotes (R² = 0.43, P = 0.001), and archaea (R² = 0.70, P = 0.001) (Supplementary Fig. 1). Soil conditions (Field types) significantly influenced the GS bacterial community compositions (R2 = 0.47, P = 0.001) (Fig. 1E). However, despite these soil condition variations, significant differences persisted in the GS bacterial communities between the high (H) and low (L) PCS severity groups (R2 = 0.15, P = 0.001) (Fig. 1E), indicating that disease-associated community shifts in GS maintained statistical independence from the variations in BS. This dual association is further supported by our functional annotation via PICRUSt2, which identified distinct KEGG pathways enriched in GS communities corresponding to different levels of PCS severity (Supplementary Table 3).
GS affected by high-PCS disease exhibit disruption in bacterial community homeostasis
Our findings indicate that following the occurrence of PCS, the GS bacterial community underwent a decline in α-diversity (Fig. 2A) and community similarity (Fig. 2B). Species whose niche width index exceeds the upper limit of the 95% confidence interval are defined as generalist species and species below the lower limit of the 95% confidence interval are defined as generalist species. Through the identification of specialized species, the shift in GS niche was characterized, demonstrating an relative abundance of specialized species in the GS bacterial community associated with PCS severity (Fig. 2C). MST results (MST < 0.5, deterministic processes dominate; When MST > 0.5, stochastic processes dominate) indicated an increased proportion of stochastic processes in the assembly of GS bacterial communities with high PCS severity compared to that with low PCS severity (Fig. 2D). Additionally, the GS bacterial community in high PCS severity exhibited a relatively weaker cooperative relationship, though the difference was not significant (Fig. 2E). Taken together, findings indicate that following the occurrence of PCS, the GS bacterial community underwent a decline in α-diversity and community similarity, an increase in stochastic processes during community assembly, and a rise in the relative abundance of specialized species (Fig. 2). These observations suggest that the changes in the GS bacterial community adhere to the AKP, indicating that PCS leads to a partial loss of plant control over the overall composition of the microbial community. In contrast, the eukaryotic and archaeal microbial communities did not display clear patterns in diversity, assembly processes, or the relative abundance of specialized species across varying PCS severity (Fig. 2).
A The variation patterns of Shannon diversity indices for bacterial, eukaryotic, and archaeal communities in GS and BS with respect to PCS severity. B Mantel correlations between the dissimilarity indices of bacterial, eukaryotic, and archaeal communities and PCS severity. C Variation patterns of specialized species relative abundance in bacterial, eukaryotic, and archaeal communities with PCS severity. D Distribution patterns of the MST for bacterial, eukaryotic, and archaeal communities. E Cohesion analysis present the patterns of changes in microbial (bacterial, eukaryotic, and archaeal) network complexity (total cohesion), cooperation (positive cohesion/total cohesion), and competition (negative cohesion/total cohesion) between GSH and GSL.
Soil bacterial community structure is a more accurate indicator of PCS severity than pathogen population
A total of 63 amplicon sequence variant (ASVs) were identified as Streptomyces spp. in the microbial profiles. A comparison of the representative sequences of these ASVs with PCS Streptomyces species revealed a perfect match between ASV9 and S. acidiscabies, with 100% sequence identity (Supplementary Table 4). When analyzing the relative abundance of the Streptomyces genus and ASV9 in the GS, it can be found that the difference in ASV9 relative abundance between GSH and GSL was more significant than in Streptomyces genus, except in the JF region (Fig. 3A, B). TxtAB genes were able to accurately indicate the population of pathogenic Streptomyces spp.26. We examined the copy number of the txtAB gene in GS and found a similar distribution pattern (Fig. 3C) as well as a significant correlation between the txtAB gene copy number and the ASV9 relative abundance (R2 = 0.57, P < 0.001). Therefore, ASV9 was considered to represent the pathogenic Streptomyces in this study. Additionally, to characterize the main features of soil physicochemical properties, we utilized the values from each BS sample on the PCo1 axis of Fig. 1 (labeled as BS-Soil-PCo1). The main characteristics of the microbial communities (bacterial, eukaryotic, and archaeal) in the GS and BS were delineated using the values from each sample on the PCo1 axis of Fig. 1 and Supplementary Fig. 1 (Supplementary Table 5).
A Relative abundance distribution of PCS-associated Streptomyces ASV (Bacterial ASV9, BASV9) in GS and BS. B Relative abundance of the Streptomyces genus in GS and BS. C Quantification of txtAB gene copies per gram of soil in GS and BS. D Hierarchical partitioning analysis revealed the explanatory power of microbial community characteristics and pathogen population indices in GS for PCS severity. E Hierarchical partitioning analysis revealed the explanatory power of microbial community characteristics, pathogen population indices, and soil physicochemical properties in BS for PCS severity.
For GS, PCS severity showed the strongest correlation with the bacterial community (Spearman, |ρ | = 0.77, P < 0.001), surpassing the correlation with pathogen relative abundance (Spearman, |ρ | = 0.73, P < 0.001) and txtAB gene copy number (Spearman, |ρ | = 0.62, P < 0.001) (Supplementary Table 6). The GS microbial communities (bacteria and eukaryotes), pathogen relative abundance and txtAB gene copy number collectively accounted for 68% of the variation in PCS severity (Fig. 2D, Supplementary Table 7). Interestingly, the GS bacterial community composition dominated the PCS severity (individual interpretation rate: 39.28%), outweighing the effect of the pathogen population (individual interpretation rate for ASV9: 14.39%, txtAB: 7.01%).
For BS, pathogen relative abundance and txtAB gene copy number demonstrated that a higher pathogen population did not necessarily result in higher levels of PCS severity. For instance, the copy number of the txtAB gene was higher in the Tengzhou sample site with low-PCS severity compared to the Jiaozhou site with high-PCS severity (Fig. 3C). BS bacterial community, pathogen relative abundance and soil physicochemical properties all showed significant associations with PCS severity, but BS bacterial community exhibited stronger correlations with PCS severity than did pathogen population (Fig. 3E, Supplementary Table 6, 7). Collectively, these observations suggest that the bacterial community structure in both GS and BS serves as a more effective indicator of PCS severity compared to the pathogen population.
AKP and anti-AKP may coexist in the disrupted GS microbial communities
The core bacterial community of GS was classified into nine distinct ecological clusters using weighted gene co-expression network analysis (WGCNA). Remarkably, the Green cluster emerged as the key microbial consortium distinguishing the high and low PCS severity groups (Fig. 4B). Across all four fields, a highly significant correlation was observed universally between the community composition of the Green cluster and the PCS severity (JD: R2 = 0.87, P < 0.001; JF: R2 = 0.78, P < 0.001; TX: R2 = 0.81, P < 0.001; JF: R2 = 0.79, P < 0.001) (Fig. 3C). Subsequently, the analysis of the Green cluster’s community assembly revealed that those with higher PCS severity exhibited a greater tendency towards deterministic processes (R2 = 0.59, P < 0.001) (Fig. 4D), indicating that the changes in the Green cluster followed anti-AKP. This phenomenon contrasts with the previous finding where the overall bacterial community changes adhered to AKP (Fig. 2). These evidences suggest that following the infection of potato plants by PCS Streptomyces spp., both AKP and anti-AKP paradigm coexist within the microbial community.
The variable importance projection (VIP) scores (A) and random forest variable importance metrics (B) indicated that the Green cluster served as a key microbial consortium indicative of PCS severity. C Significant correlations between the Green cluster and PCS severity. D The MST of the Green cluster decreased with the increased of PCS severity. E The taxonomic composition of the Green cluster. F The top 15 bacterial genera with the highest number of ASVs in the Green cluster.
The Green cluster comprised 157 ASVs, accounting for a total relative abundance of 17.18% (Supplementary Table 8). Of these, 127 ASVs (15.59%) were annotated to the genus level and were distributed among 36 genera (Supplementary Table 7). ASV3, identified as Flavobacterium sp., held the highest relative abundance within the Green cluster, accounting for 2.33%. Following were ASV9 (1.56%, Streptomyces), ASV8 (1.22%, Dyadobacter) and ASV6 (1.13%, Agrobacterium).
Indirect contributions of BS to the PCS severity via influencing the key consortium in GS
The results showed that the BS’s comprehensive characteristics, encompassing microbial communities, pathogen population, and physicochemical properties, explain only 33.5% of the PCS severity (Fig. 3E). Random forest (RF) analysis also showed that the BS bacterial community predicted PCS severity with about 70%, with a significant portion unexplained (Supplementary Fig. 3). Therefore, we hypothesize that there is a significant correlation between the dissimilarity of microbial community structures in BS and GS (Fig. 5A). The bacterial community in the BS was the most significant contributor to the GS bacterial community, explaining 42.0% of its variation (Fig. 5B, Supplementary Table 9), while accounting for only 5.6% of the variation in GS pathogen population (Fig. 5C, Supplementary Table 9). Notably, high PCS severity significantly reduced the stochastic processes of bacterial community assembly between the two soil types (Fig. 5D). A bidirectional mediation analysis revealed that the model only sustained a pathway where the BS bacterial community directed the GS key consortium and subsequently influenced PCS severity (Fig. 5E). Conversely, the model does not support a reversed directionality. In essence, the abundant microbial resources in the BS may promote the assembly of key consortiums in the GS, thus potentially mitigating the PCS severity.
A A mantel test demonstrated a significant correlation between the BS bacterial community and the GS bacterial community. B A hierarchical partitioning analysis revealed the explanatory power of microbial community characteristics, pathogen population indices, and soil physicochemical properties in BS in shaping the GS bacterial community. C A hierarchical partitioning analysis revealed the explanatory power of microbial community characteristics, pathogen population indices, and soil physicochemical properties in BS for the GS pathogen population. D The pattern of MST values between the BS and GS bacterial communities varied with changes in PCS. Different colors represent different levels of disease severity. E The bidirectional mediation model solely supports the GS key consortium as a mediating factor in the process of BS bacterial community influencing PCS severity, but not the reverse direction.
The microbes of key consortium suppress the PCS severity
We isolated 420 strains belonging to 81 species, 45 genera, and 4 phyla from potato epidermis and its surface soil (Supplementary Fig. 4). Among the 36 genera of Green cluster, 12 were obtained. From these 12 genera, 60 strains were chosen for evaluation of their ability to inhibit S. scabies (Fig. 6A). Our results showed that one-third of these strains (20 strains), belonging to the genera Chryseobacterium, Stenotrophomonas, Sphingobacterium, Sphingobium, and Bacillus, exhibited varying degrees of inhibitory or antagonistic activity against S. scabies on agar plates. Furthermore, these strains alleviated the inhibitory effect of the PCS pathogen on plant seedling growth (Supplementary Figs. 5, 6).
A Sixty strains originating from the Green cluster were examined for their ability to inhibit PCS S. scabies. The diameter of the red dot represents the size of the inhibition zone achieved by each strain. B Growth curves of four beneficial strains, including Flavobacterium nitrogenifigens HT2-2, Microbacterium phyllosphaerae HR3-14, Stenotrophomonas maltophilia TL3-11, and Stenotrophomonas rhizophila DL3-15. C The suppressive effects of the four strains and their various combinations against PCS S. scabies at 12 h, 24 h, and 36 h, showcasing the dynamic effectiveness of these synthetic communities. D, E The synthetic community FMSS demonstrates its efficacy in suppressing PCS severity (D) and promoting potato yield (E). F The FMSS down-regulated the relative expression of txtAB gene in PCS S. scabies. G The FMSS also exhibits influences on various physiological indices of potatoes, including activities of catalase (CAT), peroxidase (POD), superoxide dismutase (SOD), aseorbateperoxidase (APX), polyphenol oxidase (PPO), phenylalanine ammonia lyase (PAL), O2- accumulation and soluble protein content.
We performed sequence alignment between the isolated strains and the ASVs from the green ecological cluster identified by WGCNA. Based on this, we selected strains with inhibitory effects on PCS pathogens and constructed a synthetic community (coded as FMSS, 1/4 + 1/4 + 1/4 + 1/4) consisting of four strains: Flavobacterium nitrogenifigens HT2-2, Microbacterium phyllosphaerae HR3-14, Stenotrophomonas maltophilia TL3-11, and Stenotrophomonas rhizophila DL3-15 (Supplementary Table 10). Growth curves analysis showed that all strains were in the logarithmic phase at 24 h, based on measurements of their seed suspension at 12, 24, 36, and 48 h (Fig. 6B). Further antagonistic experiments revealed that the strains exhibited the highest antagonistic ability against the PCS pathogen after 24 h of cultivation (Fig. 6C). Eight co-cultivation protocols were designed, considering the cultivation time of the seed suspension (Supplementary Fig. 7). Among these, the optimal protocol, resulting in the largest inhibitory zone diameter of 21.87 mm, entailed sequential inoculation of TL3-11 and DL3-15 for 24 h, followed by HT2-2 for another 24 h, and finally HR3-14 for an additional 24 h of fermentation cultivation.
Inoculation with beneficial strains significantly alleviated the inhibitory effect of pathogen on the growth of radish seedlings (Supplementary Fig. 8A–C). Specifically, after a 7-day growth period, the fresh weight of plant seedlings increased by 3.0-fold, 2.0-fold, 2.9-fold, 3.2-fold, and 4.4-fold following inoculation with HT2-2, HR3-14, TL3-11, DL3-15, and the synthetic consortium FMSS, respectively, in contrast to single inoculation with the PCS pathogen (Supplementary Fig. 8C). Meanwhile, inoculation with FMSS enhanced the enzyme activity of polyphenol oxidase (PPO) by 2.2-fold and phenylalanine ammonia-lyase (PAL) by 1.1-fold compared to the pathogen treatment, and exceeded the average of all individual strains (Supplementary Fig. 8D, E).
The pot experiment showed significant reductions (P < 0.05) in both the incidence and disease index of PCS at harvest, attributable to the application of the FMSS (Fig. 6D). Specifically, the incidence decreased from 45.45 to 22.22%, while the disease index decreased from 23.94 to 19.05%, yielding a control effect of 29.53%. The FMSS also enhanced potato yield by 38.10% (Fig. 6E). Interestingly, inoculation of FMSS fermentation broth significantly down-regulated the relative expression of pathogen toxin gene txtAB gene (Fig. 6F). The FMSS significantly elevated the activities of CAT, POD, APX, PPO, and PAL, reduced O2- accumulation, and increased soluble protein content in potato tubers (P < 0.05) (Fig. 6G). This suggests that the FMSS mitigated the disease stress imposed by PCS on plants by enhancing their antioxidant capacity and disease resistance.
Discussion
Plants can recruit beneficial microbes from distant soil (such as BS) to the proximal soil by secreting root metabolites, etc., to assist in their own stress resistance. In other words, our study believes that the structural differences of BS, this natural “microbial repository” for plants to obtain beneficial microbes, may be able to affect the process of plants’ selective acquisition of beneficial microbes. Despite the establishment of research frameworks to examine interactions between individual microbes and plants, our comprehension of the relationships between microbial communities and plant diseases, particularly at a broader level, remains incomplete and confronts several challenges. For instance, the extent to which microbial communities can influence disease severity and the underlying microecological mechanisms are still quite confusing. In this study, we focused on PCS and systematically investigated the dynamic interplay between soil microbial community states and PCS severity under varying soil conditions. This study, based on hierarchical partitioning analysis, considers different types of soil microbial communities as explanatory factors and the disease index of PCS as the response factor to analyze the contribution of various microbial communities to the severity of PCS. The results of hierarchical segmentation show that the composition of soil microbial communities, especially bacterial communities, offered a more effective indicator of PCS severity than the mere pathogen population (Fig. 3D, E). This may be due to the fact that bacteria typically have faster growth rates and higher metabolic activity, allowing them to quickly respond to environmental changes and rapidly alter community structure and function. Such dynamic changes help influence the colonization of pathogens and the occurrence of plant diseases. In contrast, fungal growth is relatively slower, and changes in their community structure are often less sensitive than those of bacteria. Notably, within bacterial communities linked to high PCS severity in GS, we observed a loss of homeostasis in line with the AKP paradigm, yet an exceptional consortium of beneficial microbes emerged among them, exhibiting an anti-AKP paradigm. These findings are likely to be prevalent in microbiota-plant disease interactions and hold positive implications for controlling plant diseases. Plants can recruit beneficial microbes from distant soil (such as BS) to the proximal soil by secreting root metabolites, etc., to assist in their own stress resistance. In other words, our study believes that the structural differences of BS, this natural “microbial repository” for plants to obtain beneficial microbes, may be able to affect the process of plants’ selective acquisition of beneficial microbes.
The influence of soil microbiota on plant disease severity is widely acknowledged7, yet quantitative evidence detailing its extent remains limited. In this study, soil bacterial communities were identified as the primary driver of PCS severity, outweighing even the impact of pathogen populations (Fig. 3D, E). This finding could significantly contribute to future innovations in PCS control strategies focused on manipulating soil microbial communities. We recognize that these results may vary across diverse disease systems. Multiple factors could account for our findings. Firstly, the severity of PCS is susceptible to regulation by soil microbial communities, as evidenced by reports of PCS-suppressive soils decades ago27. Secondly, the severity of PCS is easily quantifiable, and there exists a strong correlation between PCS severity, pathogen population, and soil bacterial communities24. Thirdly, the impact of PCS on GS bacterial communities was profound enough that variations in soil conditions at a regional scale failed to hide the differences in GS bacterial communities among samples with different PCS severities (Fig. 1). Lastly, while PCS disrupted the homeostasis of GS bacterial communities, it concomitantly triggered the accumulation of certain beneficial microorganisms (Fig. 4E, F). Regarding why the state of soil microbiota affected PCS severity, we speculated that a significant factor lies in the mutualistic relationships established among beneficial microorganisms in healthy soil communities, which more effectively suppress pathogen virulence and bolster plant immunity. Our observations revealed that certain strain combinations within the key consortium linked to PCS severity exhibit more pronounced mutualistic phenomena, inhibiting pathogen growth and augmenting the activities of potato enzymes, such as CAT, POD, and PAL (Fig. 6E). The pathogenicity of Streptomyces spp. primarily stems from its attack on plants through the production of the toxin thaxtomin A26. Notably, we found that the expression of the txtAB gene, responsible for thaxtomin A synthesis, could be repressed by certain strains (Fig. 6F). This finding aligns with another study that demonstrates a strain of Pseudomonas inhibiting the expression of thaxtomin A toxin genes in PCS Streptomyces spp.28. These discoveries indicate that multiple soil microorganisms may possess the ability to suppress the thaxtomin A-producing capacity of pathogens. In summary, these beneficial interactions may explain why soil microbiota exerted a greater influence on PCS severity compared to pathogen populations in the PCS system.
The findings of our study revealed two contrasting coexistence patterns in the GS bacterial community. First, we observed a positive correlation between the abundance of pathogen and beneficial microorganisms that serve as its inhibitors (Fig. 4C). Second, we noted the coexistence of AKP and anti-AKP paradigms within the system (Fig. 4D). The emergence of PCS disrupts the potato’s ability to ‘select and amplify’ favorable bacterial communities in GS, resulting in the proliferation of pathogens and other microorganisms. Intriguingly, as PCS severity escalated, we observed an enrichment of beneficial microorganisms that possess the capability to restrain the PCS-causing pathogens (Fig. 6A, C). Therefore, these two events occurring on the same timescale exhibited a similar trend in the population dynamics of both pathogen and beneficial microorganisms. This coexistence of contrasting individuals is not an anomaly; similar patterns have been documented in other disease systems. For instance, in tomato bacterial wilt, rhizobacteria producing inhibitory siderophores coexist robustly with the pathogen Ralstonia solanacearum, exhibiting a positive correlation in their respective populations29. Furthermore, within the GS bacterial community, we identified a consortium that exhibited a heightened deterministic assembly process (anti-AKP) against the backdrop of an overall stochastic process (AKP) (Fig. 4D). Regarding the underlying mechanisms, we posit that while these opposing individuals/consortia coexist temporally, they may occupy distinct spatial scales. Microbial colonization on plant surfaces is inherently uneven, potentially leading to island effects. Studies on Arabidopsis thaliana roots have shown that rhizosphere microbial communities and root metabolites exhibited a spatial pattern along the root axis, indicating a dependency on root segments30,31. Therefore, further research on the spatiotemporal regulation of microbiome assemblages may be essential to elucidate these phenomena. With regard to the significance of this opposing coexistence, we hypothesize that it serves as a cornerstone for restoring the homeostasis of soil microbial communities. As evidenced in the bacterial wilt system of tomatoes, rhizosphere competitive bacterial communities hinder soil pathogen invasion by fiercely competing for nutrients, secreting antibacterial compounds, and reducing environmental niches13. Introducing beneficial bacterial communities into disrupted microbial ecosystems can enhance the community’s competitive balance, ultimately strengthening its disease-suppression capabilities32. Therefore, in our study, the coexistence of beneficial and harmful microbiomes may constrain their respective niches, paving the way for the restoration of soil microbial community homeostasis.
The BS conditions of this study guaranteed the occurrence of PCS. The question arises: why do we observe varying disease severities within the same field, and how does BS contribute to this? Across the four studied fields, we noticed slight variations in the bacterial communities within BS of different potato plants (Supplementary Fig. 1D). More importantly, the BS bacterial community had a significant effect on PCS severity, but only to a limited extent of 15.96% (Fig. 3E). RF analysis also indicated that the accuracy of BS bacterial community in predicting PCS severity stands at 70%, leaving a considerable proportion unexplained (Supplementary Fig. 3A). Notably, the GS bacterial community exhibited an extremely high explanatory rate for PCS severity (Fig. 3D); and the predictive accuracy in RF analysis reached 100% (Supplementary Fig. 3B, C). Moreover, a significant correlation existed between BS and GS bacterial communities (Fig. 5A); and the mediation model where the BS bacterial community impacted PCS severity by influencing the key consortium of the GS bacterial community was supported (Fig. 5E). Therefore, we deduced that BS primarily contributed to PCS severity by shaping the GS bacterial community, particularly its key consortium. Undoubtedly, BS is the primary source of GS microbial communities, as soil serves as the richest seed bank in ecosystems33. The assembly of plant-associated microbial communities follows the “priority effect,” wherein the BS microbial communities played a pivotal role in the initial assembly of GS microbial communities34. However, as the intermediate process of BS bacterial communities influencing PCS severity elongates, more room is created for other factors to come into play. The formation of GS microbial communities was also influenced by various factors, such as the colonization process of the microorganisms35. Studies have reported that beneficial microorganisms in the BS microbiota must undergo chemotaxis, rhizosphere adhesion, and colonization into biofilms to establish themselves in the rhizosphere35. When considering the potential of artificially manipulating soil microbiota, there are promising avenues, including introducing highly competitive beneficial bacteria into disrupted communities to bolster their overall competitiveness32. Our prior research showed an approach, as inoculating a plant-derived strain, Brevibacillus laterosporu BL12, significantly inhibited the PCS severity14. This strain and other potentially beneficial microorganisms, like Pseudomonas, became keystone species within the community14,24. In summary, BS bacterial communities likely contributed to the PCS severity primarily by shaping key consortium in GS. While multiple factors still influence the shaping of GS communities by BS bacterial communities, the significant role of BS in the formation of GS bacterial communities is undeniable. Exploring intermediary methods to facilitate the successful establishment and stabilization of BS-beneficial bacterial communities in GS could serve as a potential strategy to bolster soil immunity.
Methods
Field experiments
Four field trials were conducted in the Xiaofole (JF, 36.2202° N, 119.8412° E) and Dingguan Villages (JD, 36.2371° N, 119.8541° E) of Jiaozhou City, and the Lizihang (TZ, 35.1479° N, 117.1143° E) and Xiaolonghe Villages (TX, 35.1805° N, 117.0704° E) of Tengzhou City, in Shandong Province, China in late August 2018 (Fig. 1A). Both JF and JD fields experience a warm temperate monsoon climate, while having a brown-type soil. However, TZ and TX fields have a continental monsoon climate, while having a cinnamon-type soil. All experimental fields exceeded 600 m2 in area, with a documented history of PCS incidence persisting for over three consecutive years. The physicochemical properties of all soils are given (Supplementary Table 1). The investigation was conducted during the autumn cultivation cycle from August to November 2018, employing standardized planting configurations with interplant spacing of 20–25 cm and row spacing of 70–80 cm. All field management practices adhered to region-specific integrated pest management protocols.
Sampling
Each experimental field was systematically divided into six sampling subplots (each ~100 m2). Within each subplot, 2–3 potato plants were randomly selected for sampling. Sampling occurred between November 10 to 13, 2018 (~90 days after planting), and two compartments of the soil, GS (within 2 mm of tuber surface) and BS (collected from the plant growth area at a depth of 5–15 cm), were selected for sampling from each plant (Fig. 1B). A total of 58 potato plants were selected, including 14 plants in JD, 13 in JF, 16 in TX, and 15 in TZ. The sample with abnormal size, pests, mechanical damage, and marginal growth were excluded. The potato tubers were carefully collected with an alcohol-sterilized shovel, and one tuber was chosen from each plant. All samples were stored in ice packs ~4 °C) and then transported to the laboratory within 8 h. GS was gently collected with a sterile brush, and then stored at –80 °C. After homogenization, one part of each BS sample was stored at –80 °C until DNA extraction, and the rest was immediately subjected to the physicochemical analysis. The PCS severity of the tubers was assessed on a scale of 1–9. This scale was based on the percentage of the surface covered by the PCS lesions: 1: no scab; 2: 0.1–0.8%; 3: 0.9–2.8%; 4: 2.9–7.9%; 5: 8.0–18.0%; 6: 18.1–34.0%; 7: 34.1–55.0%; 8: 55.1–77.0%; and 9: 77.1–100%24. According to the severity, the tubers were grouped and named as H (PCS severity level ≥4) or L (PCS severity level 1–3) (Supplementary Fig. 2).
Physicochemical analysis
To further investigate the explanatory power of soil physicochemical properties on disease index and explore the potential relationship between soil physicochemical properties and microbial communities, we measured multiple soil physicochemical indicators from four regions. The physicochemical properties of BS were analyzed in three technical replicates from each sample to evaluate soil conditions in the study fields (Fig. 1). The soil pH and electrical conductivity (EC) were measured using a mixture of soil and deionized water free of CO2 at a ratio of 1:2.5 (w/v). The soil moisture content, organic matter, total potassium, total phosphorus, total nitrogen, ammonium (NH4+-N), nitrate (NO3−-N), total sulfur, sulfate (SO42−) and iron ions (Fe3+) were determined following the previously established methods36,37.
Isolation of strains from potato epidermis and testing their ability to suppress Streptomyces
To identify potential beneficial microorganisms in the soil that could control PCS, we isolated bacterial strains from potato epidermis using enrichment and dilution techniques. The potato epidermis was peeled with a sterile scalpel. Considering the location of the potato epidermis, the isolated strains comprised epidermal endophytes, and those residing in the epidermal surface and proximity of the geocaulosphere. We used six different culture media to isolate bacterial strains (Supplementary Table 2). Briefly, the soil samples were homogenized and then 1 g of the soil was mixed with 9 mL of sterile water. The soil slurry was incubated at room temperature for 30 min and then diluted to 100, 1000, and 10,000 times with sterile water. These dilutions were plated on solid plates at 37 °C for 3–7 days. Single colony-forming units were selected and subcultured three times on solid plates. The purified single colonies and 1 mL of sterile liquid media were incubated at 37 °C and 180 rpm for 24 h in a shaker. Then, the purified strains were stored on slants of the corresponding media at 4 °C for standby use.
Though we isolated thousands of strains, 420 strains were selected according to their morphology. The 16S rRNA gene was amplified by the primers Eub 27 F (5’-AGAGTTTGATCMTGGCTCAG-3’) and Eub 1492 R (5’-ACGGYTACCTTGTTACGACTT-3’). The final volume of the amplification reaction solution was 25 mL containing 18.35 mL of ddH2O, 2.5 mL of 10 buffer, 2 mL of dNTP, 0.5 mL of each primer, 0.15 mL of Taq polymerase, and 1 mL of a template solution. The conditions of the PCR were as follows: initial denaturation at 95 °C for 10 min, then 35 cycles containing thermal denaturation at 95 °C for 40 s, annealing at 53 °C for 40 s, extension at 72 °C for 90 s, and the final extension at 72°C for 10 min. The sequence alignment was performed at NCBI to determine the approximate phylogenetic affiliation of the strains. Taxonomy was confirmed if the maximum identity of the sequence reported by NCBI BLAST was >97% and was the greatest of all listed matches. A phylogenetic tree was constructed to visualize the phylogenetic relationships among microbes using the neighbour-joining method with MEGA7 software (https://www.kent.ac.uk/software/mega-7)24.
The antagonistic ability of the strains against S. scabies was assessed using the drilling method according to the size of the inhibition zone. The S. scabies, stored on Actinomycetes culture medium plates, was combined with 1 mL of sterile liquid Actinomycetes culture medium and incubated on a shaker at 28 °C and 180 rpm for 3–5 days. From this culture, a 100 μL aliquot of the S. scabies strain suspension containing mycelium was evenly coated onto each Actinomycetes culture medium plate, and then four small holes were evenly punched into the plates. The strain suspensions were added to the small holes. The plates were sealed and incubated at 28 °C for 7–10 days. The inhibition zone size was observed and recorded. All experiments were performed in a laminar flow cabinet.
Soil DNA extraction
We extracted soil DNA using the E.Z.N.A.™ Soil DNA Kit (Omega, USA) according to the manufacturer’s instructions, while a NanoDrop 2000 spectrophotometer (Thermo Scientific, USA) was used to determine the quantity and quality of DNA. The DNA was stored at −80 °C until further analysis.
Quantitative PCR (qPCR)
The txtAB is a key gene involved in the biosynthesis of PCS phytotoxin thaxtomin A, and it also enables the accurate quantification of pathogenic Streptomyces strains26. The qPCR method was used to determine the copy numbers of the txtAB gene in three technical replicates to determine the absolute abundance of Streptomyces PCS pathogens. The primers StrepF (5’-GCAGGACGCTCACCAGGTAGT-3’) and StrepR (5’-ACTTCGACACCGTTGTCCTCAA-3’) were used to amplify a 72 bp fragment of the txtAB gene from Streptomyces PCS pathogens38. The reaction was performed on a CFX96™ real-time system (Bio-Rad, USA) using 96-well plates. The amplification reaction mixture had a final volume of 20 μL, and it contained 10 μL of SYBR® Premix Ex Taq™ (Tli RNaseH Plus; Takara, China), 5 μL of DNA, 0.2 μL of each 10 μM primer, and 4.6 μL of ultrapure water. The plasmids containing the txtAB gene fragment were constructed to prepare the standard curve, and the plasmid copy numbers were automatically calculated using an online calculator (cels.uri.edu/gsc/cndna.html) based on the concentration, size, and average weight of the base pairs. A standard curve equation that showed the relationship between gene copy numbers and Ct values was generated with serial dilutions of the above plasmids with known copy numbers after qPCR. The copy numbers of the samples were calculated based on the standard curve equation and their own Ct values. All PCRs included no-template controls and positive controls with known Ct values. The thermocycling conditions for amplification of the txtAB genes were as follows: 95 °C for 5 min; 35 cycles of 95 °C for 5 s, 53 °C for 10 s, and 72 °C for 10 s; and a final melting cycle to produce a melting curve for the quality control. The standard efficiency curves for all runs were r2 > 0.997, and the efficiencies were 90–110%.
Illumina MiSeq sequencing and analysis
The soil bacterial, eukaryotic microorganism, and archaeal communities were detected using the Illumina MiSeq PE250 sequencer (Illumina, USA). To pinpoint microbial communities closely associated with the occurrence of scab disease in the soil, we performed high-throughput amplicon sequencing on the collected soil samples. The bacterial 16S rRNA gene’s V3-V4 hypervariable region was PCR-amplified and a library was constructed using primers 338 F (5’-ACTCCTACGGGAGGCAGCA-3’) and 806 R (5’-GGACTACHVGGGTWTCTAAT-3’)37. Similarly, the V4 hypervariable region of the eukaryotic microbial 18S rRNA gene was PCR-amplified and a library was built with primers 528 F (5’-GCGGTAATTCCAGCTCCAA-3’) and new 706 R (5’-AATCCRAGAATTTCACCTCT-3’). The V4 hypervariable region of the archaeal 16S rRNA gene underwent the same procedure, using nested primers: Arch340F (5’-CCCTAYGGGGYGCASCAG-3’) and Arch1000R (5’-GGCCATGCACYWCYTCTC-3’) in the first round, and Uni519F (5’-CAGYMGCCRCGGKAAHACC-3’) and Arch806R (5’-GGACTACNSGGGTMTCTAAT-3’) in the second round. The NEB Next® UltraTM DNA Library Prep Kit (NEB, USA) was used for the construction of amplicon libraries (https://www.neb.cn/zh-cn/products/e7645-nebnext-ultra-ii-dna-library-prep-kit-for-illumina). During the PCR amplification process, unique Barcode sequences were appended to both ends of the amplified gene fragments in each sample, serving as sample identifiers. The bacterial amplicon library generated 18.3 GB of raw sequencing data, with an average of 161.5 MB per sample; the eukaryotic microbial amplicon library produced 9.8 GB of raw sequencing data, with an average of 86.5 MB per sample; and the archaeal amplicon library generated 9.8 GB of raw sequencing data, with an average of 87.0 MB per sample.
The quality of the raw sequence was evaluated and low-quality cut-offs for forward and reverse readings were determined. The software suite Quantitative Insights into Microbial Ecology 2 (QIIME2) facilitated quality control measures and the creation of an ASV feature table39. The code for processing the raw data has been provided in Supplementary Note 1. The DADA2 plugin within QIIME2 was employed for the removal of noise, the identification and exclusion of chimeric sequences40. Additionally, we eliminated singleton ASVs and those with a total count below five across all samples. Bacterial, eukaryotic and archaeal sequences were aligned and taxonomically annotated using the SILVA 138 database (https://www.arb-silva.de/). For a consistent comparison of microbial community diversity, sample sequencing depths were equalized to the smallest sequencing count obtained among them. For bacterial samples, sequence counts were standardized to 28,666, while for eukaryotic microbial samples, the counts were standardized to 21,477. For archaeal samples, the counts were standardized to 66,469. All analyses in the study proceeded using these normalized datasets.
Construction of PCS biocontrol synthetic communities
Based on the guiding insights from the results of microbiome analysis and preliminary antagonistic experiments, we selected microorganisms significantly associated with the PCS severity from the GS bacterial community. Artificial synthetic communities for inhibiting PCS were then constructed through a random combination strategy. To assess the compatibility and interactions between the two strains, the cross-hatching method was employed on LB plates in a 28 °C constant temperature incubator for an inverted culture over 48 h. Prior to inoculation, it was ensured that all strains possessed a consistent concentration, specifically during the logarithmic growth phase (OD600 = 0.5). Subsequently, through the successive removal of one strain at a time, the simplest synthetic microbial community was gradually constructed. By comparing the antagonistic abilities of each synthetic microbial community and individual strains against PCS S. scabies, we screened out the simplest synthetic microbial community with strong antagonistic ability. Finally, we constructed a synthetic microbial community named FMSS, which consists of four strains, Flavobacterium nitrogenifigens HT2-2, Microbacterium phyllosphaerae HR3-14, Stenotrophomonas maltophilia TL3-11, and Stenotrophomonas rhizophila DL3-15, demonstrating significantly enhanced inhibitory capabilities compared with individual strains.
To optimize the co-cultivation sequence of FMSS, we tested eight cultivation protocols. (1) F. nitrogenifigens HT2-2 was cultured individually for 24 h. (2) S. maltophilia TL3-11 was cultured individually for 24 h. (3) S. rhizophila DL3-15 was cultured individually for 24 h. (4) M. phyllosphaerae HR3-14 was cultured individually for 24 h. (5) F. nitrogenifigens HT2-2 was inoculated first and cultured for 24 h, followed by S. maltophilia TL3-11 and S. rhizophila DL3-15 inoculated simultaneously and cultured for 24 h, and finally M. phyllosphaerae HR3-14 inoculated and cultured for 24 h. (6) M. phyllosphaerae HR3-14 was inoculated first and cultured for 24 h, followed by S. maltophilia TL3-11 and S. rhizophila DL3-15 inoculated simultaneously and cultured for 24 h, and finally F. nitrogenifigens HT2-2 inoculated and cultured for 24 h. (7) S. maltophilia TL3-11 and S. rhizophila DL3-15 were inoculated simultaneously and cultured for 24 h, followed by F. nitrogenifigens HT2-2 inoculated and cultured for 24 h, and finally M. phyllosphaerae HR3-14 inoculated and cultured for 24 h. (8) F. nitrogenifigens HT2-2 was inoculated first and cultured for 24 h, followed by M. phyllosphaerae HR3-14 inoculated and cultured for 24 h, and finally S. maltophilia TL3-11 and S. rhizophila DL3-15 inoculated simultaneously and cultured for 24 h.
Radish seedling growth experiment
We assessed the alleviating or exacerbating effect of the strains on the pathogens phytotoxicity using radish seedling growth tests. Radish seeds (Korean Long White Radish) were surface sterilized by soaking in 0.26% NaClO solution for 10 min, then rinsed three times with sterile water and air-dried. The treated seeds were evenly placed on a three-layer sterile filter paper in a petri dish for germination, kept moist for 3–5 days (ensuring the petri dish remained moist throughout the germination process). Germinating seeds with consistent sprouting and growth were selected and gently transferred into 50 mL test tubes containing 10 mL of 1% water agar medium using sterile tweezers, with 3 seeds placed per tube. The pathogens and each strain were co-inoculated in the test tubes, with a control group inoculated with an equal amount of culture medium. The test tubes were then incubated in a culture chamber with intermittent light exposure, set for 16 h of light, and cultured for 7 days. The pathogens were co-inoculated with each strain in test tubes, and a control group was treated with the same amount of culture medium only. The tubes were incubated in a growth chamber, and the radish seedlings were gently removed from the tubes for subsequent phenotypic and physiological measurements.
Determination of the relative expression of the txtAB gene in S. scabies
The impact of the synthetic community on the expression of the key toxin synthesis gene txtAB in PCS pathogens was assessed through gene quantification experiments. To prepare sterile FMSS fermentation broth, the fermentation broth was first transferred into centrifuge tubes and centrifuged at 6000 rpm for 5 min at 4 °C. The supernatant was carefully collected, while the cell pellet was discarded. The collected supernatant was then filtered twice using a sterile filter with a pore size of 0.45 μm to obtain a sterile fermentation broth for subsequent experiments. For the preparation of the S. scabies pathogen seed solution, spores of the PCS strain were gently scraped from Gauze’s I agar plates and inoculated into TDMc liquid medium. The cultures were incubated at 28°C with shaking at 170 rpm for 4–7 days until spore germination was observed. Next, 1% (v/v) of the pathogen seed solution was transferred to 20 mL of fresh TDMc medium and cultured under the same conditions for an additional 4 days to enhance pathogen growth. Once visible colonies were established, the sterile FMSS fermentation broth was inoculated with the pathogen culture at a 3% (v/v) ratio and further incubated for 3 days to ensure successful co-cultivation. Collect the pathogen colonies, wash them 1–2 times with pH 7.4 PBS, and use sterile filter paper to absorb excess moisture, ensuring the colonies remain as dry as possible. RNA of the pathogen was extracted using the MolPure® Bacterial RNA Kit. cDNA was synthesized from the extracted RNA using the HiScript II Q RT SuperMix for qPCR Reverse Transcription Kit. Using the cDNA as a template for amplification of the txtAB gene of the pathogen using primers Strep F: (5’-GCAGGACGCTCACCAGGTAGT-3’) and Strep R: (5’-ACTTCGACACCGTTGTCCTCAA-3’). Finally, calculate the relative expression level of the txtAB gene using the 2-ΔΔCT method.
Pot experiment treatment
To evaluate the inhibitory effect of the synthetic community on PCS and demonstrate that potatoes have enriched strains with the ability to suppress the disease, this potting experiment was conducted on August 13, 2021 at Shandong Agricultural University, Tai’an, China. The experiment comprised two treatments, with 24 plants planted in each. The treatments were: one with PCS S. scabies alone (P) and the other with PCS S. scabies combined with a synthetic community (P + FMSS). Prior to planting, composite fertilizer was applied to the soil, which was then tilled and thoroughly mixed. Germinated seed potatoes were then sown, with a depth of approximately 10 cm in both treatments. Following planting, each plant was watered with 100 mL of a 3 × 108 CFU/mL pathogen fermentation solution. During the seedling stage of potato growth, a second irrigation with the pathogen solution was performed. In the early stage of tuber expansion, the P + FMSS treatment was watered with 100 mL of FMSS fermentation solution, while the P treatment received an equal amount of medium. On November 3, 2021, the mature potatoes were harvested. The tubers were collected, counted, assessed for disease severity, and weighed accordingly.
Enzyme activity assay
In biological control, one of the key mechanisms by which biocontrol bacteria exert their protective effects is by inducing plant resistance. Reactive oxygen species (ROS) have been recognized as toxic byproducts of plant metabolic processes, capable of causing damage to macromolecules such as lipids, proteins, and DNA. Accordingly, to protect cells from ROS-induced damage, plants have developed a defense system, which primarily consists of enzymatic antioxidant mechanisms (such as SOD, CAT, and POD) and non-enzymatic antioxidant systems. The enzyme activities of radish seedling and potato epidermis was determined according to Wang et al. 37. Grind the collected radish seedlings and potato skin samples on ice using a mortar and pestle. Add 10 mL of 50 mM sodium phosphate buffer (pH 7.8) to form a homogenate. Centrifuge the homogenate at 10,000 g for 10 min at 4 °C. Collect the supernatant as the enzyme extract for analyzing the ROS generation rate and antioxidant enzyme activity. ROS levels were quantified using a colorimetric assay. A reaction mixture comprising 0.5 mL of Solution A, 0.5 mL of phosphate-buffered saline (PBS, pH 7.8), and 1 mL of 1 mM hydroxylamine hydrochloride was incubated at 25 °C for 1 h. Afterward, 1 mL of 17 mM p-aminobenzenesulfonic acid and 1 mL of 7 mM α-naphthylamine were added to the mixture and thoroughly mixed. Following a 20 min incubation at 25 °C, the absorbance was measured at 530 nm. The steps for measuring the SOD activity in radish seedling and potato skin samples are as follows:The assay reaction mixture (3.3 mL total) consisted of 1.5 mL of phosphate buffer (pH 7.8), 0.3 mL of 65 mM methionine, 0.3 mL of 500 µM nitroblue tetrazolium, 0.3 mL of 100 µM EDTA-Na₂, 0.3 mL of 200 µM riboflavin, 0.1 mL of enzyme solution, and 0.5 mL of distilled water. Control tubes were kept in the dark, while test tubes were exposed to 4000 lx light for 20–30 min. The dark control was used as a blank reference, and absorbance was recorded at 560 nm. To measure the catalase (CAT) activity in radish seedlings and potato skin, prepare a reaction mixture containing 0.1 mL enzyme solution and 2.9 mL CAT reaction solution, and immediately measure the absorbance at 240 nm. To measure the peroxidase (POD) activity in radish seedlings and potato skin, add 20 µL of enzyme solution to 3 mL POD reaction solution to initiate the reaction. Record the absorbance at 470 nm for 0–30 s. To measure the ascorbate peroxidase (APX) activity in radish seedlings and potato skin, prepare a reaction mixture containing 0.1 mL enzyme solution and 2.9 mL APX reaction solution, and measure the absorbance at 290 nm, with a time interval of 0–40 s. To measure the PPO activity in radish seedlings and potato skin, mix 6 mL phosphate-citrate buffer (pH 6.0) with 1 mL 0.5% hydroquinone solution, and preheat in a water bath at 30 °C. Then add 1 mL solution A and continue incubation for 5 min. The reaction is terminated by heating to inactivate the enzymes. Subsequently, centrifuge at 5000 rpm for 10 min, collect the supernatant, and measure the absorbance at 410 nm. To measure the PAL activity in radish seedlings and potato skin, take 0.5 g of fresh radish seedlings and potato skin tissues and grind them with 5 mL of 5 mM mercaptoethanol-borate buffer, 0.1 g polyvinylpyrrolidone, and a small amount of quartz sand in a precooled mortar. Centrifuge the homogenate at 8000 rpm for 15 min at 4 °C to obtain the crude enzyme extract. Mix 0.1 mL of the supernatant with 3 mL of 0.02 M phenylalanine-borate buffer (pH 8.8) and 2.9 mL distilled water. Incubate the reaction mixture at 30 °C for 30 min and measure the absorbance at 290 nm. The enzyme activity is defined as the amount of enzyme required to cause a 0.01 absorbance change per minute at 290 nm.
Statistical analysis
Statistical analysis was conducted to decipher the microbial assembly processes under different disease severities, identifying potential key microbial groups that inhibit PCS occurrence. We performed a one-way analysis of variance (ANOVA) using IBM SPSS statistical software, supplemented by a Duncan’s multiple range test at a significance threshold of P < 0.05. The “vegan” R package was utilized to calculate the microbial community’s α and β-diversity indices. Furthermore, the “vegan” package was also employed to conduct PERMANOVA analysis to assess the differences in microbial community structure between different groupings. The “vegan” and “linkET” packages were used for the Mantel test. Subsequently, we carried out hierarchical partitioning analysis using the “rdacca.hp” package to further investigate the explanatory of microbial community structure, pathogens, txtAB gene, and soil physicochemical properties on the PCS severity41. The KEGG pathway (ko) functional profiles of GS and BS bacterial communities were predicted using the PICRUSt2 based on the community composition42. A total of 306 pathways were annotated, which belonged to six categories: “environmental information processing”, “metabolism”, “genetic information processing”, “cellular processes”, “organismal systems” and “human diseases.” Pathways belonging to “environmental information processing” and “metabolism” were selected, and pathways not belonging to bacteria were removed to calculate the differences using the STAMP software43. The calculation of microbial community cohesion indices was performed using the R packages “SpiecEasi”, “phyloseq”, and “Matrix”. WGCNA was conducted using the online “BIC” platform (https://www.bic.ac.cn/BIC/#/)44. During the network construction process, ASVs with a total abundance less than the number of samples (58) were excluded to ensure the robustness of the network. Visualization of network graphs was performed using the Gephi software. RF analysis and PLS-DA analysis were both conducted on the Biozeron cloud platform (http://www.cloud.biomicroclass.com/CloudPlatform) to assess the importance of different microbial taxonomic features. Mediation effect analysis was performed using the “mediation” package in R to explore the potential influence of soil community structure on PCS severity. Simple correlation calculations were done using the “Hmisc” R package. Origin2021 software (https://www.originlab.com/2021) was utilized for the visualization of statistical data as well as for conducting two-sided Wilcoxon rank-sum tests and post hoc Dunn’s multiple comparison tests.
Data availability
The raw sequencing data are publicly available in the NCBI Sequence Read Archive (SRA) under the Bioproject number PRJNA703762. The code for processing the raw data has been provided in Supplementary Note 1. Other codes may be made available to qualified researchers upon reasonable request from the corresponding author.
Code availability
The code for processing the raw data has been provided in Supplementary Note 1. Other codes may be made available to qualified researchers upon reasonable request from the corresponding author.
Abbreviations
- PCS:
-
potato common scab
- GS:
-
geocaulosphere soil
- BS:
-
bulk soil
- H:
-
(the group with) high common scab severity
- L:
-
(the group with) low common scab severity
- JF:
-
Xiaofole Village of Jiaozhou City, Shandong Province, China
- JD:
-
Dingguan Village of Jiaozhou City, Shandong Province, China
- TZ:
-
Lizihang Village of Tengzhou City, Shandong Province, China
- TX:
-
Xiaolonghe Village of Tengzhou City, Shandong Province, China
- AKP:
-
Anna Karenina principle.
References
He, J., Dai, H., Zhang, X. & Wang, E. Mycorrhizal signals promote root development dependent on LysM-receptor like kinases in rice. N. Crops 1, 100009 (2024).
Zhan, C. & Wang, M. Disease resistance through M genes. Nat. Plants 10, 352–353 (2024).
Lv M. H., et al. Ms gene and Mr gene: microbial-mediated spatiotemporal communication between plants. iMeta 3, e210 (2024).
Arnault, G., Mony, C. & Vandenkoornhuyse, P. Plant microbiota dysbiosis and the anna karenina principle. Trends Plant Sci. 28, 18–30 (2023).
Jiang, G. et al. Effects of plant tissue permeability on invasion and population bottlenecks of a phytopathogen. Nat. Commun. 15, 62 (2024).
Li, M. et al. Facilitation promotes invasions in plant-associated microbial communities. Ecol. Lett. 22, 149–158 (2019).
Gao, M. et al. Disease-induced changes in plant microbiome assembly and functional adaptation. Microbiome 9, 187 (2021).
Wang, X. et al. An amplification-selection model for quantified rhizosphere microbiota assembly. Sci. Bull. 65, 983–986 (2020).
Wen, T. et al. Specific metabolites drive the deterministic assembly of diseased rhizosphere microbiome through weakening microbial degradation of autotoxin. Microbiome 10, 177 (2022).
Kou, C. et al. A necessary considering factor for crop resistance: Precise regulation and effective utilization of beneficial microorganisms. N. Crops 1, 100023 (2024).
Shi, X. et al. Insights into plant–microbe interactions in the rhizosphere to promote sustainable agriculture in the new crops era. N. Crops 1, 100004 (2024).
Bass, D., Stentiford, G. D., Wang, H. C., Koskella, B. & Tyler, C. R. The pathobiome in animal and plant diseases. Trends Ecol. Evol. 34, 996–1008 (2019).
Li, M. et al. Indirect reduction of Ralstonia solanacearum via pathogen helper inhibition. ISME J. 16, 868–875 (2022).
Li, C. et al. Biocontrol of potato common scab by Brevibacillus laterosporus BL12 is related to the reduction of pathogen and changes in soil bacterial community. Biol. Control153, 104496 (2021).
Qiao, Y. et al. Core species impact plant health by enhancing soil microbial cooperation and network complexity during community coalescence. Soil Biol. Biochem. 188, 109231 (2024).
Rosenzweig, N., Tiedje, J. M., Quensen, J. F., Meng, Q. & Hao, J. J. Microbial communities associated with potato common scab-suppressive soil determined by pyrosequencing analyses. Plant Dis. 96, 718–725 (2012).
Janvier, C. et al. Soil health through soil disease suppression: which strategy from descriptors to indicators? Soil Biol. Biochem. 39, 1–23 (2007).
Raaijmakers, J. M. & Mazzola, M. Soil immune responses. Science 352, 1392–1393 (2016).
Yuan, J. et al. Root exudates drive the soil-borne legacy of aboveground pathogen infection. Microbiome 6, 156 (2018).
Rolfe, S. A., Griffiths, J. & Ton, J. Crying out for help with root exudates: adaptive mechanisms by which stressed plants assemble health-promoting soil microbiomes. Curr. Opin. Microbiol. 49, 73–82 (2019).
Bakker, P. A. H. M., Pieterse, C. M. J., de Jonge, R. & Berendsen, R. L. The soil-borne legacy. Cell 172, 1178–1180 (2018).
Trivedi, P., Leach, J. E., Tringe, S. G., Sa, T. & Singh, B. K. Plant-microbiome interactions: from community assembly to plant health. Nat. Rev. Microbiol. 18, 607–621 (2020).
Bignell, D. R. D., Fyans, J. K. & Cheng, Z. Phytotoxins produced by plant pathogenic Streptomyces species. J. Appl. Microbiol. 116, 223–235 (2014).
Shi, W. et al. The occurrence of potato common scab correlates with the community composition and function of the geocaulosphere soil microbiome. Microbiome 7, 14 (2019).
Kopecky, J. et al. Bacterial, archaeal and micro-eukaryotic communities characterize a disease-suppressive or conducive soil and a cultivar resistant or susceptible to common scab. Sci. Rep. 9, 14883 (2019).
Qu, X., Wanner, L. A. & Christ, B. J. Using the TxtAB operon to quantify pathogenic Streptomyces in potato tubers and soil. Phytopathology 98, 405–412 (2008).
Hiltunen, L. H. et al. Soil bacterial community in potato tuberosphere following repeated applications of a common scab suppressive antagonist. Appl. Soil Ecol. 167, 104096 (2021).
Arseneault, T., Goyer, C. & Filion, M. Pseudomonas fluorescens LBUM223 increases potato yield and reduces common scab symptoms in the field. Phytopathology 105, 1311–1317 (2015).
Gu, S. et al. Competition for iron drives phytopathogen control by natural rhizosphere microbiomes. Nat. Microbiol. 5, 1002–1010 (2020).
Loo, E. P.-I. et al. Sugar transporters spatially organize microbiota colonization along the longitudinal root axis of Arabidopsis. Cell Host Microbe 32, 543–556.e6 (2024).
Fan, X., Ge, A.-H. & Wang, E. Spatially distributed metabolites SWEETen the root for microbes. Cell Host Microbe 32, 445–447 (2024).
Li, M. et al. Synthetic microbial communities: sandbox and blueprint for soil health enhancement. iMeta 3, e172 (2024).
Ling, N., Wang, T. & Kuzyakov, Y. Rhizosphere bacteriome structure and functions. Nat. Commun. 13, 836 (2022).
Debray, R. et al. Priority effects in microbiome assembly. Nat. Rev. Microbiol. 20, 109–121 (2022).
Xu, Z. et al. Chemical communication in plant–microbe beneficial interactions: a toolbox for precise management of beneficial microbes. Curr. Opin. Microbiol. 72, 102269 (2023).
Lv, M. et al. The moderate substitution of Astragalus sinicus returning for chemical fertilizer improves the N cycle function of key ecological bacterial clusters in soil. Front. Microbiol. 13, 1067939 (2022).
Wang, Y. et al. Suaeda salsa root-associated microorganisms could effectively improve maize growth and resistance under salt stress. Microbiol. Spectr. 10, e0134922 (2022).
Sagova-Mareckova, M. et al. Determination of factors associated with natural soil suppressivity to potato common scab. PLoS One 10, e0116291 (2015).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Lai, J., Zou, Y., Zhang, J. & Peres-Neto, P. R. Generalizing hierarchical and variation partitioning in multiple regression and canonical analyses using the rdacca.hp R package. Methods Ecol. Evol. 13, 782–788 (2022).
Douglas, G. M. et al. PICRUSt2 for prediction of metagenome functions. Nat. Biotechnol. 38, 685–688 (2020).
Parks, D. H., Tyson, G. W., Hugenholtz, P. & Beiko, R. G. STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30, 3123–3124 (2014).
Chen, T., Liu, Y. X. & Huang, L. ImageGP: an easy-to-use data visualization web server for scientific researchers. iMeta 1, e5 (2022).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (42407427, 42377309, and 42077027), the Shandong Provincial Natural Science Foundation, China (ZR2023QD075), the Key R&D Program of Ningxia Hui Autonomous Region of China (2023BCF01015), the 'First Class Discipline' Construction Project of Shandong Agricultural University (SKL81103), the Shandong Provincial “811” Project of First-class Discipline Construction, the National Key R&D Program of China (2023YFD20014003), Shandong Province Key R&D Program (2021CXGC010804), the China Postdoctoral Science Foundation (2024M751875), the Potato Innovation Program for the Chief Expert of Shandong Province (SDAIT-16), and the State Key Laboratory of Microbial Technology Open Projects Fund (M2023-07). We sincerely thank all those who have assisted me with any part of this manuscript and the WeChat subscription ID “meta-genome” and “NAU_LorMe” for the analysis methods.
Author information
Authors and Affiliations
Contributions
Z.G., B.Z., and T.N. conceived and supervised the project. W.S., M.h.L., R.W., Z.G., B.Z., and T.N. designed the experiment. W.S., M.h.L., R.W., L.W., and M.c.L. collected the samples. M.h.L., R.W., L.W., B.W., and R.L. performed parts of the experiment. W.S., M.h.L., R.W., M.c.L. and M.S. analyzed the data. W.S., M.h.L., M.S., B.X., C.Y., T.N., B.Z., and Z.G. edited the manuscript. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shi, W., Lv, M., Wang, R. et al. Geocaulosphere soil bacterial community drives potato common scab outcomes beyond pathogen abundance. npj Biofilms Microbiomes 11, 134 (2025). https://doi.org/10.1038/s41522-025-00774-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41522-025-00774-y
This article is cited by
-
Evaluation of Bacillus velezensis strain N23 for the control of potato common scab
BMC Plant Biology (2026)
-
Rare Microbial Taxa Dominate the Microecological Landscape of Cadmium Exposure in Rice Rhizosphere
Microbial Ecology (2026)