Abstract
Probiotics have been widely tested for their effect on mental well-being, albeit with heterogeneous outcomes. Direct and indirect effects through the gut microbiome might lie at the basis of these observations. Here, in a post-hoc analysis, we assessed the effect of 4-week consumption of a probiotic candidate strain on the gut microbiome in students exposed to academic stress. Healthy students were randomized to consume a fermented milk product with Lacticaseibacillus rhamnosus CNCM I-3690 (N = 39) or an acidified non-fermented milk product (N = 40) twice daily for 4 weeks before academic exams. The gut microbiome was analysed by Quantitative Microbiome Profiling based on 16S rRNA gene amplicon and shotgun metagenomic sequencing. Stress and anxiety were assessed using both objective and self-reported markers. Changes of alpha-diversity markers and community shifts from baseline (beta diversity) were lower in L. rhamnosus treated individuals over controls, suggesting lower overall changes of gut microbiota during psychological stress in the Probiotic group. The intake of L. rhamnosus CNCM I-3690 induced differential abundance of some species, such as the maintenance of the quantitative abundance of Ruminococcus bicirculans, and co-varied with species, which differed according to visits (i.e., stress level), suggesting a potential beneficial effect of the strain before the highest increase of stress level. The higher quantitative abundance of F. prausnitzii induced by the probiotic intake was associated with lowered self-reported anxiety levels before the exam. Functional analysis revealed minor changes upon intake of the probiotic strain. Taken together, using a quantitative framework, we found that L. rhamnosus CNCM I-3690 has a potential effect on gut microbiome response to stress, although further studies are needed to better understand the precise interaction.
Similar content being viewed by others
Introduction
The prevalence of mental health disorders such as anxiety and depressive disorders is rising worldwide, affecting people at different stages of life1. The gut microbiome is a significant component in the two-way communication system between the gut and the nervous system, known as the microbiome-gut-brain axis, which has been extensively reviewed2,3,4. Pioneering studies in animal models have revealed various pathways connecting the gut microbiome to the central nervous system, including communication via nerves, the endocrine and immune system. Conversely, the central nervous system can influence the gut microbiome by regulating gut function and homeostasis. Animal studies reported a positive effect (behavior, physiology) of gut microbiota manipulation by probiotics; however, these translated poorly to humans5. Clinical studies have shown mixed and heterogeneous effects of probiotics on stress, cognition, and mood6,7,8,9,10,11. These effects may be mediated through various direct and indirect mechanisms that can vary between probiotic strains; these include production of neuroactive metabolites, protection of the gut epithelial barrier, regulation of host immunity, and modulation of the gut microbiome. Recent large cross-sectional studies have uncovered associations between the gut microbiome and mental health, particularly in anxiety and depression12,13,14,15,16. For example, several studies have reported a decrease in specific microbial taxa, including Coprococcus and Faecalibacterium in individuals with depression, however with heterogeneous adjustment of confounding factors between studies17. These findings have fueled interest to study the effect of gut microbiome modulators in various mental health disorders using (fermented) foods, probiotics and prebiotics separately or in combination18,19,20,21,22,23,24,25,26,27,28. When using a psychological stressor (academic exam) as a challenge, consumption of a fermented milk product containing L. paracasei Shirota during eight weeks mitigated the reduction in alpha-diversity of the gut microbiome concomitant to lower perceived stress markers26. In contrast, the intake of L. paracasei Lpc-37 during 10 weeks did not show an effect on stress, mood, anxiety nor gut microbiome20. These heterogenous effects may be due to various factors, such as trial duration, dose (109–1011 CFU/day), strain-specific features and inter-subject variability of the gut microbiome and/or levels of stress at baseline. So far, most studies have focused on the composition of the gut microbiome using non-quantitative approaches and lack functional insights both about the probiotic strain and gut microbiome. Genomic module-based analytical frameworks allow to assess the neuroactive metabolic potential of the gut microbiome, including neurotransmitter production (e.g., GABA, serotonin, dopamine, acetylcholine), short-chain fatty acid production, and tryptophan metabolism amongst others12,29. Using this framework, Berding et al. showed that a specific diet did not significantly alter the neuroactive potential of the gut microbiome in healthy subjects with moderate baseline stress level28. In contrast, a strain of Lactiplantibacillus (formerly Lactobacillus) plantarum increased the diversity of bacterial species predicted to synthesize neurotransmitters and the levels of some predicted microbial neuroactive metabolites in stressed adults27. However, to date, these studies have not been performed using quantitative microbiome assessment methods, nor were the dynamics gut microbiome changes during a specific psychological stress assessed, leaving doubt as to the physiological relevance of the observed (relative) shifts.
We previously reported that 4-week consumption of a fermented milk drink containing a probiotic candidate Lacticaseibacillus rhamnosus (L. rhamnosus) CNCM I-3690 (~2 x 10E11 cfu/ day) reduced anxiety and perceived stress levels in students exposed to a naturalistic psychological stressor (academic oral exam)30. Notably, this effect was independent of barrier stabilization, suggesting the contribution of other mechanisms. We hypothesize that the gut microbiome could contribute to this response. Therefore, in this study, we performed an exploratory investigation of the impact of L. rhamnosus CNCM I-3690 consumption on the gut microbiome before the academic exam. First, we explored the associations between baseline gut microbiome composition and function and markers of stress. We next analyzed quantitative changes in the compositional and functional aspects of the gut microbiome, considering the functional contribution of the probiotic strain. Overall, our study suggests that L. rhamnosus CNCM I-3690 may play a role in reducing perceived stress response, potentially by acting on the gut microbiome.
Results
Study cohort description
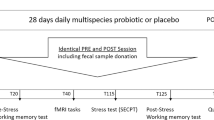
We previously performed a randomized, double-blind, controlled trial in students who were exposed to academic stress30. Subjects without psychiatric disorders (depression or general anxiety disorder, assessed by PHQ-9 and GAD-7 questionnaires) were enrolled. They were randomized to receive either a fermented milk product with L. rhamnosus CNCM I-3690 (N = 44, “Probiotic” group) or the control, an acidified but non-fermented milk product (N = 46, “Placebo” group) (Fig. 1). Samples and clinical metadata were collected at baseline (V1), 2 weeks (V2) and 4 weeks (V3, exam day) as previously described30. Self-reported markers of anxiety and perceived stress were measured by the State-Trait Anxiety Inventory-state (STAI) and Perceived Stress Scale (PSS) respectively. Objective markers of stress included cortisol, Salivary Alpha-Amylase (SAA), and secretory IgA (sIgA).
Created in BioRender. Raes, J. (2025) https://BioRender.com/t7iwlg5.
In this post-hoc analysis, we analyzed the gut microbiome of 79 subjects (22. 9 ± 1.7 years, 55.6% female), with 39 subjects in Probiotic and 40 in Control groups. At baseline (V1), PSS and STAI (Mean + /− SD) were considered low for both groups (PSS Placebo 8.1 ± 5.1, PSS Probiotic 7.8 ± 4.8, STAI placebo 28.9 ± 6.6, STAI probiotic 29.9 ± 5.9). Our previous study (N = 90) showed that 4-week consumption of L. rhamnosus CNCM I-3690 lowered STAI and PSS induced by psychological stress30. In the current study cohort (N = 79), the effect was confirmed, but less significant due to reduced power (Visit 3 Wilcoxon rank-sum test, p = 0.08 for STAI, p = 0.04 for PSS).
Baseline gut microbiome and association with stress markers and clinical variables
We profiled the gut microbiome using quantitative 16S rRNA gene amplicon sequencing and shotgun metagenomics at three-time points (Fig. 1). To identify host factors which significantly contribute to microbiome variation in our study cohort, we performed a distance-based redundancy analysis (db-RDA) using both quantitative species abundance (mOTU) and function (Gut metabolic modules (GMM) and gut Brain modules (GBM)) at baseline (V1). Using Bray-Curtis dissimilarity, we observed that only moisture displayed a significant contribution to gut microbiome compositional variation (dbRDA R2 = 0.039, PERMANOVA, FDR = 0.012) (Fig. 2A and Supplementary Table 1), which was also observed for functional variation (Canberra dbRDA R2 = 0. 089, PERMANOVA, FDR = 0.013; Bray-Curtis dbRDA R2 = 0.131, PERMANOVA, FDR = 0.013). Overall gut microbiome composition (mOTU level) or functional potential (GMM-based) did not differ globally between groups based on beta-diversity (PERMANOVA, FDR = 0.2–0.4) (Fig. 2B, C and Supplementary Table 2) or alpha-diversity (richness, Pielou evenness, and the Inverse Simpson diversity index) (Supplementary Table 3). Differential analysis revealed one species (Clostridiales incertae sedis) that was higher in the Probiotic group at baseline (Wilcoxon rank-sum test, FDR = 0.09, Supplementary Table 4). At baseline, overall gut microbiome composition or functional potential did not differ between groups, ruling out pre-existing difference between groups.
A Effect sizes of moisture covariate on microbiome community variation (stepwise dbRDA on Bray-Curtis dissimilarity for the mOTU and Canberra distance for the GMM) B Principal Coordinate analysis of gut microbiome composition (mOTU level) and (C) function (GMM) D Significant association between richness and some microbial and host markers (Spearman’s ρ, FDR < 0.1) E Enterotyping of the gut microbiome in our baseline study cohort (N = 79), FGFP young adults (N = 199) and FGFP older adults (N = 2799).
Next, we assessed whether baseline gut microbiome descriptors were (cross-sectionally) associated with host and fecal parameters. We observed a positive correlation between species richness and total microbial load (Spearman’s ρ = 0.63, FDR = 1.10-8), species richness and the total STAI (Spearman’s ρ = 0.32, FDR = 0.06) and a negative correlation between species richness and moisture levels (Spearman’s ρ =−0.51, FDR = 3.10-5) (Fig. 2D, Supplementary Table 5). At species abundance level, there was no significant association with clinical parameters following FDR adjustment (Supplementary Table 6).
Enterotyping of the study cohort
We next assessed the variation of the gut microbiome by enterotyping using Dirichlet-Multinomial Mixtures (DMM). The gut microbiome exhibited optimal clustering into four distinct community types (enterotypes), determined by BIC (Supplementary Fig. 1), using a background dataset comprising the Belgian population-based cohort, Flemish Gut Flora Project (FGFP) subjects (N = 2998). In our cohort, as expected, most abundant species differed amongst enterotypes. For instance, Bact2 varied from others by higher quantitative abundance of Eggerthella lenta, [Ruminococcus] gnavus, Blautia obeum/wexlerae, and lower quantitative abundance of multiple taxa including Ruminococcus bromii, Faecalibacterium prausnitzii, B. adolescentis (Supplementary Table 7). The enterotype distribution at baseline differed from that observed in the whole FGFP (chi squared test, p = 0.003) (Fig. 2E), with more Bact2 compared to the FGFP. However, an analysis of age, BMI, and sex-matched subjects from FGFP and did not show this difference and indicated a higher prevalence of Bact2 in adults <30 years, compared to adults >30 years (Fig. 2E). Within the subgroup, there were no significant differences between the FGFP young adults and our study cohort (chi squared test p > 0.1). The distribution of enterotypes did not differ between the placebo group and the probiotic group at baseline (chi squared test, p = 0.6). Next, we assessed the enterotype distribution in relation to both clinical and microbial factors. Bact2 had a higher moisture than other enterotypes (Kruskal–Wallis, FDR = 0.01) (Supplementary Fig. 2 but there was no other difference in any other host parameters between enterotypes including calprotectin(Kruskal–Wallis, FDR = 0.92). As previously reported, Bact2 enterotype had a lower microbial load (Kruskal–Wallis, FDR < 0.001), species richness (Kruskal–Wallis test, FDR < 0.001) and Inverse Simpson diversity (Kruskal–Wallis test, FDR = 0.006), when compared to others (Supplementary Fig. 2, Supplementary Table 8).
Effect of L. rhamnosus CNCM I-3690 on gut host markers and microbiota composition during academic stress
We first assessed whether fecal moisture (a proxy for transit time) and calprotectin changed upon intervention. Using linear regression models, no change in time or between groups was observed during the intervention adjusted by baseline for both calprotectin and moisture (linear model, FDR > 0.5). At baseline, calprotectin was significantly higher in the probiotic group (Wilcoxon rank-sum test, False Discovery Rate (FDR) = 0.002) (Supplementary Table 9) but with a mean lower than 50 μg/g in both groups at all time points.
Next, we assessed the effect of the probiotic strain on the gut microbiome at two weeks and at one month after intake. We assessed the detection of L. rhamnosus CNCM I-3690 using StrainPhlAn. The probiotic strain was the only L. rhamnosus species in the gut microbiome within our study cohort (Supplementary Fig. 3A). Quantitative analysis of mOTUs assigned to L. rhamnosus confirmed the exclusive increase in the probiotic group at both visits (Wilcoxon rank-sum test, FDR < 0.001) with variable abundance between subjects (Supplementary Fig. 3B, C), and no difference between enterotypes (Kruskal–Wallis test, FDR > 0.1). Then, we assessed the change of 16 S rRNA genus level alpha-diversity (Richness, Shannon, Inverse Simpson, Pielou), as 16S rRNA gene amplicon sequencing has been tested for most probiotics studies21,22,23,25,26,27. The probiotic intervention prevented a decrease of genus-level alpha-diversity (Inverse Simpson (Wilcoxon signed-rank test, W = 2129, p-val = 0.003, FDR = 0.005) and Shannon diversity (Wilcoxon signed-rank test, W = 2129, p-val = 0.003, FDR = 0.005) and the Pielou’s evenness (Wilcoxon signed-rank test, W = 2248, p-val = 0.010, FDR = 0.013) when comparing the delta between baseline (V1) and other visits (not significant for richness; Wilcoxon signed-rank test, W = 2700.5, p-val = 0.341, FDR = 0.341) (Fig. 3A).This was also observed using mixed linear models over time (Fig. 3B), but not at the species level (Shotgun-based; Supplementary Table 10). Within-subject Bray-Curtis dissimilarity to baseline was lower in Probiotic group versus Placebo group both using genus-based 16S rRNA gene (Wilcoxon rank-sum test, p-value = 0.009) (Fig. 3C) and species-level shotgun metagenomics (Wilcoxon rank-sum test, FDR = 0.03). Excluding the genus Lactobacillus did not change the results (Supplementary Fig. 4), which suggests lower overall changes of gut microbiota during psychological stress in Probiotic group were not driven by artificial technical effects of adding an additional strain to the community structure. Since some subjects had relatively high calprotectin levels (>100 µg/g), we repeated the analysis excluding these individuals. The results remained consistent (Supplementary Fig. 5).
A Change of alpha-diversity indices for each subject versus baseline for the placebo and the probiotic group (Wilcoxon test). B Linear mixed-effect models of the effect of the probiotic in the diversity; the model takes as the baseline the “Placebo” and the “Visit 1” as the reference levels for Group and Visit variables. The coefficients and interaction terms are the differences from the baselines. C Within-subject Bray-Curtis dissimilarity to baseline in both groups with lower distance indicating less changes (Wilcoxon test).
We further assessed the effect of L. rhamnosus CNCM I-3690 intake by comparing the changes in beta-diversity over time, taking covariates into account. Bray-Curtis dissimilarity at the species level showed significant differences between groups at the different timepoints (PERMANOVA, FDR = 0.003) (Fig. 4A), with a larger effect size than stool moisture, while taking subject into account, (non-redundant dbRDA, adjusted Visit 2 R2 = 0.0199, Visit 3 R2 = 0. 0195) (Fig. 4B). Self-reported markers of stress (PSS) contributed significantly to gut microbiome variation at V2 (non-redundant dbRDA, adjusted R2 = 0.0078). Using a Negative binomial mixed effect model adjusted by stool moisture, group and visit interaction revealed significant increase in Lactobacillus rhamnosus [ref_mOTU_v3_00710] g_Pseudoflavonifractor and members from the genus Ruminococcus, namely Ruminococcus species incertae sedis [ext_mOTU_v3_26371] and Ruminococcus bicirculans [ref_mOTU_v3_02792] in the Probiotic group (Fig. 4C). No change in enterotype distribution was observed over time in any group (McNemar’s chi-squared FDR > 0.1).
A Bray-Curtis dissimilarity changed over time in both groups. B Confounding variables indicated as significant and non-redundant according to a distance-based redundancy analysis. R2 (purple) indicates the variance explained by each variable, while Cum_R2 (green) represents the cumulative variance explained. C Negative binomial mixed-effect model of the taxa that showed a change in the abundance after the probiotic intake, the model was corrected by the effect of moisture. The model takes as the baseline the “Placebo” and the “Visit 1” as the reference levels for Group and Visit variables. The coefficients and interaction terms are the differences from the baselines.
We further explored the co-variation of L. rhamnosus mOTU with resident species using an approach tailored to analysis of quantitative microbiome data (Fig. 5). Following 2-week consumption (i.e. at the V2 timepoint), we observed that Semi-Parametric Rank-based INference in Graphical models (SPRING)31 showed that L. rhamnosus positively associated with several butyrate producers such as Coprococcus and Faecalibacterium sp, and lactate/ acetate producers Bifidobacterium longum and B. bifidum. It negatively correlated with species including Flavonifractor, amongst others (Fig. 5A). Before the exam (V3), the L. rhamnosus mOTU covaried with different species, positively with Clostridiales, and B. longum (Fig. 5B). Overall, our analysis suggests that intake of L. rhamnosus CNCM I-3690 during psychological stress induced differential abundance of some species, and with different covariation over time. By analyzing various network centrality metrics (reviewed in ref. 32) and comparing them between the placebo and probiotic groups, we observed significant differences in degree centrality (the number of direct connections a node has), closeness centrality (how efficiently a node can reach all others in the network), and eigenvector centrality (which reflects a node’s influence based on the connectivity of its neighbors) (Supplementary Fig. 6A). Notably, at Visit 3, the probiotic group exhibited a lower degree centrality but higher closeness and eigenvector centrality, suggesting a shift towards a more structured and hierarchical microbial network. This pattern implies that L. rhamnosus administration may restructure the community by either becoming a key hub itself or promoting the prominence of other influential taxa. Interestingly, L. rhamnosus showed marked increases in degree, betweenness, closeness, and eigenvector centrality from Visit 2 to Visit 3. While initially below the median for all metrics, by Visit 3 it had increased above the median, indicating its emergence as a central and influential member of the microbial community (Supplementary Fig. 6B)
A 2 weeks after consumption of the probiotic strain (V2) and (B) 4 weeks after consumption of the probiotic strain, and during exposure to stress (V3). Red color indicates negative associations, and blue color positive associations.
Effect of L. rhamnosus CNCM I-3690 on gut microbiota functional potential during academic stress
Given the effect of L. rhamnosus CNCM I-3690 on gut microbial ecology, we further explored the functional patterns emerging from the metagenomic profiling. We first profiled the neuroactive potential of the probiotic strain by genomic analysis. A total of 15 GBM modules were identified (completeness >50%) including prevalent and less prevalent ones in the gut microbiome (Supplementary Fig. 7). Amongst the gut modules encoded by L. rhamnosus, MGB040 (Inositol degradation pathway) was present in less than 15% of 533 representative gut genomes12. We also examined whether the neuroactive potential of L. rhamnosus was strain-specific by pangenomic analysis on 124 L. rhamnosus genomes. All GBM genes were detected in the L. rhamnosus core genome, suggesting the detected neuroactive potential is shared among current known members of this species. Next, we studied the effect of the intake of L. rhamnosus CNCM I-3690 on the total gut microbiome functional potential. Functional beta diversity analysis (Canberra distance and Bray-Curtis dissimilarity) revealed a significant group-time interaction effect, even after correcting for moisture (Canberra: R2 = 0.039/p-val = 0.022) (Supplementary Fig. 8A). Moisture explained most functional variation for each of the individual time points V1 (Canberra: R2 = 0.089/p-val = 0.001), V2 (Canberra: R2 = 0.052/p-val = 0.001), and V3 (Canberra R2 = 0.055/p-val = 0.002) (Supplementary Fig. 8B). Beyond this, the effect of L. rhamnosus CNCM I-3690 on overall function was observed only at V2 (Canberra: R2 = 0.081/p-val = 0.004) (Supplementary Fig. 8B), in contrast to results for the taxonomic composition (see above). L. rhamnosus consumption was reflected in the increase of pathways related to the bacterial strain itself, such as the case of MF0007 (lactose and galactose degradation), which remained significant even after correcting for moisture (Supplementary Fig. 8C). Collectively, these results indicate that L. rhamnosus CNCM I-3690 had a minor effect on function with mostly contributions from its own genome rather than altering the community functional potential through ecological interactions.
Association between gut microbiome, L. rhamnosus and stress markers
Last, we examined the connection between L. rhamnosus mOTU, the gut microbiome, and self-reported stress markers PSS and STAI, both in the Placebo and/or Probiotic groups specifically between V3 and other visits, where the increase of stress markers was observed together with significant differences between groups. Correlations were evaluated by comparing changes in the abundance of taxa and GMM over time against stress markers (Fig. 6). In the Probiotic group, we found more significant correlations (Spearman’s ρ, FDR < 0.1) compared to the Placebo group. For instance, lower levels of Faecalibacterium sp were linked to higher STAI level, while Clostridium and Colinsella species showed a positive correlation with increased PSS (Supplementary Table 11). A linear regression analysis to showed that these observed significant correlations were not confounded by moisture (ANOVA FDR < 0.1) (Supplementary Table 12). We complemented the analysis with cortisol, which increased significantly upon stress, but with no difference between Probiotic and Placebo30. Two additional taxa negatively (within Clostridiales) correlated with change of cortisol in the Probiotic group compared to baseline (Fig. 6). Overall, our analysis suggests that 1) certain gut microbiome taxa correlated with the probiotic in the network analysis (see above), are negatively associated with stress responses and 2) most microbiome – stress associations are driven by poorly characterized microbes.
Association between changes of self-reported stress markers and cortisol is associated with some resident taxa in both Placebo and Probiotic groups (Spearman’s ρ, FDR < 0.1).
Discussion
Probiotics have gained considerable attention for their purported benefits on mental well-being, yet their effects remain inconsistent across studies. Here, we studied microbiome effects in a trial in which 4-week consumption of L. rhamnosus CNCM I-3690 reduced self-reported stress markers in students exposed to an academic exam. Our first objective was to investigate the association between baseline gut microbiome composition and clinical markers including those related to stress and anxiety. Following that, we explored how the consumption of L. rhamnosus CNCM I-3690 for four weeks altered the composition and function of the gut microbiome, and how changes were associated with lower self-reported stress response.
First, we explored the baseline gut microbiome and the association with clinical and stress markers. Given the large inter-subject variability in the gut microbiome, partitioning into community types is an approach that facilitates identifying associations of gut microbiome with environmental and host factors. We observed a higher prevalence of the Bacteroides enterotypes in young adults compared to middle aged participants of the Belgian Flemish Gut Flora Project (FGFP) population cohort. Consistent with prior studies, Bact2-enterotype harbored higher moisture (a proxy for decreased transit time33) and lower gut microbiome alpha-diversity34,35. We did not find any specific associations between baseline gut microbiome member abundance and perceived stress and anxiety; however, a positive association of STAI with richness was observed. Given the low sample size and the low-moderate anxiety level at baseline, this needs to be further substantiated in larger and more focused cohorts.
Next, we assessed the effect of 4-week consumption of a fermented milk with L. rhamnosus CNCM I-3690 on the gut microbiome during the course of an academic exam; for the first time, a quantitative microbiome profiling approach was used. We found that, in the probiotic group, the microbiome changed less within individuals upon stress intervention using both alpha and beta-diversity suggesting maintenance of gut microbiome structure upon academic stress. Lower changes in alpha - diversity was observed in previous studies using a single strain in healthy subjects exposed to an academic stressor26. In another study, a lower change in alpha-diversity, depression and anxiety (STAI) symptoms was observed in depressed subjects following the intake of a probiotic mixture containing eight different strains of lactic acid bacteria and bifidobacteria as add-on to therapies24. In this study the effect of the probiotic mixture on gut microbiota was limited to an increase in Lactobacillus, indicating that the effect was likely due to specific probiotic strains rather than a broader modulation of the gut microbiome. We further found a higher abundance of some bacterial species following intake of L. rhamnosus CNCM I-3690 such as R. bicirculans, which has been recently negatively associated with major depressive disorder36. Next, using quantitative co-occurrence network analysis, we found that L. rhamnosus positively covaried with a.o. butyrate-producers (Faecalibacterium, Coprococcus) and some Bifidobacterium species (B. longum, B. bifidum), and negatively covaried with Flavonifractor. Before the exam, in the period with the highest stress level, co-variation with B. longum was maintained. Recently, a strain of Bifidobacterium longum was shown to reduce PSS in healthy adults with mild-to-moderate stress-levels37.
We compared network metrics, and found that L. rhamnosus administration may impact the community by either becoming a key hub itself or promoting the prominence of other taxa. We observed significant changes in degree, closeness, and eigenvector centrality. After 1-month consumption, the probiotic group showed fewer direct connections (lower degree centrality) but increased connectivity to influential nodes (higher closeness and eigenvector centrality). This suggests that L. rhamnosus may reorganize the microbial network into a more structured and hierarchical community, either by becoming an influential node itself or by enhancing the role of key taxa. Mechanistic work of the effect of Lactobacillus rhamnosus strain using synthetic communities, or spent media experiments would be needed to study its precise direct or indirect metabolic interactions with specific gut members38,39
When we specifically studied the direct correlation between quantitative taxon abundance change and stress markers that were different between groups (STAI and PSS), we found that more species significantly associated with STAI following intake of L. rhamnosus CNCM I-3690 over placebo. Examples include a lower abundance of Faecalibacterium with higher STAI levels, and inversely for Collinsella. Altogether, results from the above mentioned complementary analyses suggest that the previously described link between L. rhamnosus CNCM I-3690 and lowered perceived stress and anxiety could be explained by modulation or interaction with resident microbiota members. Interestingly, most association between microbiome and stress markers (self reported and cortisol) were driven by poorly characterized microbes, suggesting that further isolation and description of strains in subjects exposed to stress are needed.
We next studied the relation between functional shifts (induced or contributed by) the strain and stress response. Based on genome analysis, several gut brain modules were detected in L. rhamnosus, amongst which inositol degradation, which is rare in gut microbes12 and was found to be the most expressed pathway by L. rhamnosus CNCM I-3690 in small intestinal samples following consumption of the strain40. Interestingly, inositol degradation was recently found as a module that correlated negatively with anxiety in a cross-sectional cohort36. Here, we did not observe a differential abundance of inositol degradation during the highest increase of stress (before exam), which may be related to low sample size or the use of quantitative approaches. The lower effect of the probiotic strain on function of gut microbiome is consistent with the concept of functional redundancy41. Further stratification of subjects according to stress levels could reveal differential functional enrichment. In a previous study in healthy subjects, the functional contribution of multi-strain probiotics was shown to differ according to strains and subjects’ gut microbiome42. Previous findings have shown that L. rhamnosus CNCM I-3690 persists longer than two other probiotic strains in fecal samples of certain individuals43,44, possibly through adherence to host cells, metabolic interaction with other resident bacteria42,45,46, or functional enrichment42, which may indicate fitness in lower GI tract.
Altogether, our results show that L. rhamnosus CNCM I-3690 administration (1) is linked to prevention of microbiota perturbation upon academic stress, (2) causes abundance changes in some gut taxa, and (3) possibly complements the functioning of the gut microbiome for probiotic-specific pathways. Our study has some limitations. First, it does not capture dietary habits and has a modest sample size. While diet is a significant factor of microbiome variation47,48, the contribution is lower than other covariates, especially moisture/transit time49,50,51. Second, our functional analysis relies solely on shotgun metagenomics and should be coupled with metabolomics, especially to investigate production of neuroactive compounds, which will allow to associate taxa-metabolites with lower stress markers. Third, to preserve blinding, it is critical that the test and control products closely resemble each other in appearance, taste, and texture. Carboxymethylcellulose (CMC) was used as a texturizing agent. A previous study in healthy individuals suggested a detrimental impact of CMC on the intestinal microbiota52 raising the possibility that it may have influenced the reduced microbial diversity observed in the placebo control group. However, the daily intake used in the present study (1.2 g) was substantially lower than the dose tested by Chassaing et al. (15 g), suggesting that any such effect at this dose is likely negligible. Nevertheless, our study complements previous mental health cross-sectional studies and interventional studies with an analysis at quantitative and functional level during psychological stress. Further mechanistic studies are needed to elucidate how L. rhamnosus CNCM I-3690 interacts with specific members of gut microbiome using for instance synthetic communities or ex-vivo fecal fermentation combined with production of neuroactive compounds. In this study, L. rhamnosus CNCM I-3690 was tested in a preventive approach in subjects with low baseline level of anxiety or stress. It is unknown whether similar findings on gut microbiome would be observed in subjects with variable baseline stress levels. The higher prevalence of Bact2 in young adults (students) is intriguing and needs to be confirmed in larger studies and especially in association with diet, as subjects were excluded for anxiety disorders and depression. Follow-up studies will make it possible to better assess the effect of the strain on mental health taking account of variation of gut microbiome, stress markers and dietary habits. Such studies will guide more rationale microbiome-based approaches for mental health.
Methods
Study design and participants
This randomized, controlled study (registered on Clinicaltrials.gov on January 17, 2018, NCT03408691) was described by Wauters et al.30. Participants were healthy male or female students, aged 20 to 30, recruited from the faculties of (bio)medical and pharmaceutical sciences or industrial engineering at the bachelor’s or master’s level, with a scheduled oral thesis defense. Students with scores of 10 or higher on the General Anxiety Disorder 7-item (GAD-7) or Patient Health Questionnaire 9-item (PHQ-9) scales were excluded from the study. After an initial screening visit for eligibility, a run-in period of at least 15 days occurred before randomization. A baseline visit was scheduled more than one month before the defense (D-35 to D-27), and a second visit took place two weeks prior (D-14 ± 1 day), with sample and questionnaire collection. On the day of the thesis defense (D0), further samples and questionnaires were obtained to assess the impact of the stressor. The trial adhered to the Declaration of Helsinki and Good Clinical Practice guidelines, receiving approval from the Ethics Committee of University Hospitals Leuven (S60969). Written informed consent was obtained from all participants before their inclusion. Data collection took place at KU Leuven and University Hospitals Leuven (Leuven, Belgium).
Product intervention
Test product was a fermented milk containing L. rhamnosus CNCM I-3690 strain with a count of 1011 CFU/100 g). Control product was an acidified milk, depleted in lactose, containing phosphoric acid, and Carboxy Methyl Cellulose. Both products were manufactured by Danone Research, France, and were similar in sweetness, flavor (multi-fruit), texture, color, packaging, and nutritional content (isocaloric). Subjects were randomized to L. rhamnosus-containing (probiotic) or acidified (placebo) milk consumed twice daily for 4 weeks. Subjects ingested two bottles (100 g/bottle) of Test or Control product per day (one at breakfast, one at dinner), for 28 days.
Clinical outcomes
The clinical endpoints were described in Wauters et al.30. The primary endpoint was the effect of the test product containing L. rhamnosus CNCM I-3690 compared to the control product (placebo) on the stress-induced change on small intestinal permeability from baseline. Objective and self-reported stress/anxiety were measured on each visit using salivary markers (cortisol, Salivary Alpha-Amylase (SAA), and secretory IgA (sIgA) and psychological questionnaires: Momentary anxiety levels were measured with the state version of the validated State-Trait Anxiety Inventory (STAI) questionnaire. Perceived stress in the preceding week was assessed with the 10-item Perceived Stress Scale (PSS). Both questionnaires were collected before the urine collection on each visit with an additional STAI immediately after the thesis. Other questionnaires included the Generalized Anxiety Disorder Assessment (GAD-7) (all visits), and Patient Health Questionnaire 9 (PHQ9), at screening as exclusion criterium.
Stool collection and fecal parameters measurement
Stool samples were collected at V1 (baseline), V2 (14 days of consumption of L. rhamnosus CNCM I-3690 / control product), and V3 (28 days of consumption of L. rhamnosus CNCM I-3690 / control product thesis defense) from 79 subjects. Two subjects had 2 samples (drop-out after V2) and 1 sample (drop-out after V1), resulting in a total of 233 samples. Fecal samples were analyzed for microbial load, calprotectin, moisture and gut microbiome53
The microbial load of the study cohort was measured by flow cytometry as described previously9. Based on the exact weight of the aliquots analyzed, cell counts were converted to microbial loads per gram of fecal material. The fecal moisture content was determined as the percentage of mass loss after lyophilization from around 100 mg frozen aliquots of non-homogenized fecal material as previously done7. Fecal calprotectin concentrations were determined using the fCAL ELISA Kit (Bühlmann, Amherst, USA). The measurements were done on frozen fecal material (−80 °C).
Bacterial DNA extraction and sequencing
Fecal DNA was extracted according to the protocol outlined by Falony et al.49. In brief, DNA was isolated from around 100 mg of frozen samples using the MagAttract PowerMicrobiome DNA/RNA KF kit (QIAGEN, Hilden, Germany) following the manufacturer’s guidelines. The V4 region of 16S rRNA genes was amplified using the 515 F/806 R primer pair and purified with the QIAquick PCR Purification Kit.
16S rRNA gene amplicon sequencing and processing
Nucleic acid concentration was measured using DropQuant, followed by amplification of the V4 region of the 16S rRNA gene using 515 F/806 R primers (GTGYCAGCMGCCGCGGTAA and GGACTACNVGGGTWTCTAAT, respectively), which were modified to include Illumina adaptors and dual-index barcodes54. PCR amplicons were quantified with a Fragment Analyzer (Advanced Analytical Technologies). Sequencing was performed on the Illumina MiSeq platform (MiSeq Reagent Kit v2, Illumina, San Diego, USA) at the VIB Nucleomics core laboratory (Leuven, Belgium). The 16S rRNA gene amplicon data was analyzed using the DADA2 pipeline8. Specifically, the first 30 base pairs were trimmed, and sequence lengths were set to 130 bp for the forward strand and 200 bp for the reverse. DADA2’s default parameters were used for sequence error rate estimation, dereplication, inferred sample composition, and chimera removal. Taxonomic assignment was performed using the DADA2 RDP implementation (R package “dada2,” function “assignTaxonomy”) with the GTDB_bac-arc_ssu_r86 database. The identification of L. rhamnosus ASVs was conducted by comparing all ASVs from the Lactobacillus genus against the NCBI Nucleotide collection (nt) database (updated on 2024/06/03) using Megablast with default parameters. ASVs were identified as L. rhamnosus if both the alignment and identity percentages were 100%.
Shotgun metagenomic data processing
Sequencing libraries were prepared with the Illumina sample preparation kit in accordance with the manufacturer’s instructions. The sequencing was performed using NovaSeq 6000 paired-end Illumina sequencing. Quality control to remove low-quality reads, adapters, and human reads was performed with BBDuk (http://jgi.doe.gov/data-and-tools/bb-tools/). A mean of 9.349/2.24 ( ± 2.24) Gb were kept per sample. For each sample, remaining high quality reads were assembled into contigs with metaSPAdes55. Genes were predicted from assembled contigs with Prodigal56. A gut microbiome gene catalog was constructed by clustering ORFs from all samples with > 95% identity with CD-HIT-EST57. The gene catalog was functionally annotated by eggNOG-mapper v258. KEGG Orthology (KO) annotations from genes were used to quantify the abundance of Gut Metabolic Modules (GMMs)41 and Gut-Brain Modules (GBMs)12 for functional pathway analysis.
Quantitative microbiome profiling
The quantitative microbiome profiling (QMP) matrix was constructed following a previously described method9. In summary, samples were adjusted to an even sampling depth, defined as the ratio between the sampling size (16S rRNA gene copy number-corrected sequencing depth) and microbial load (the average total cell count per gram of frozen fecal material). 16S rRNA gene copy numbers were obtained from the rRNA operon copy number database rrnDB. The resulting matrices were represented as the “QMP” matrix and the “even sample depth rarefied matrix,” which reflects the number of reads per sample rarefied according to the sampling depths derived from cell counts.
For shotgun metagenomic analysis, the QMP matrix was estimated as described previously59. using decontaminated sequencing data and cell counts. Briefly, the shotgun sampling size was calculated based on the average abundance of ten universal single-copy marker genes from the MOCAT60 pipeline (COG0012, COG0016, COG0018, COG0172, COG0215, COG0495, COG0525, COG0533, COG0541, COG0552). Paired-end reads were normalized to achieve an even sampling depth, defined as the ratio between the sampling size and microbial load (i.e., the average total cell count per gram of frozen fecal material). Normalization was performed by randomly selecting reads to match the minimum observed sampling depth in the dataset. The QMP rarefied samples were then taxonomically classified and quantified using mOTUs v2.40 Additionally, the QMP rarefied samples were mapped against the gene catalog using BWA61.
Ecological network
Three ecological networks were constructed using the species annotations provided by the mOTU classification tool. The networks were built for each independent time point using a semi-parametric rank-based correlation method31 (R package “SPRING”, function “SPRING” parameters “nlambda = 100, rep.num = 50, subsample.ratio = developer recommendation”). The resulting networks were analyzed using the R package Igraph. For each graph, the taxa directly linked with L. rhamnosus mOTU were subset from the network and represented into an individual plot. Network statistics including degree, betweenness, closeness, and eigenvector centrality were calculated for all timepoints. We estimated the degree centrality (R package “igraph”, function “degree”), which represents the average number of edges per node (species). The betweenness centrality (R package “igraph”, function “betweenness”) quantifies the number of shortest paths passing through a node. The closeness centrality (R package “igraph”, function “closeness”) is defined as the inverse of the sum of the shortest path lengths from a node to all other nodes in the network. Finally, the eigenvector centrality (R package “igraph”, function “evcent”) measures node importance based on the principle that connections to highly connected nodes contribute more to a node’s score.
Neuroactive potential of Lacticaseibacillus rhamnosus CNCM I-3690 and strain-level detection in metagenomes
A total of 199 available L. rhamnosus genomes were downloaded from NCBI RefSeq database (20th December 2021) using PanACoTA prepare module62. After PanACoTA quality control and deduplication a total of 123 genomes were kept for pangenomics analysis. The Panaroo pipeline63 was used to identify the pangenome of L. rhamnosus using the 123 selected genomes and the L. rhamnosus CNCMI-3690 strain. Pangenome genes were functionally annotated with eggNOG-mapper58. KEGG Orthology (KO) annotations from these genes were used to identify the presence of Gut-Brain Modules (GBMs)12 and Gut Metabolic Modules (GMMs)41. 533 other genomes from representative species in the gut microbiome were also annotated for the presence of GBMs and GMMs. >= 50% coverage of a pathway was used as threshold to be considered as present in the genome. Strainphlan64 a computational tool for performing metagenomic strain-level population genomics on large metagenomic datasets, was used to identify L. rhamnosus CNCM I-3690 strain retrieved from metagenomic reads. RAxML65 was used to generate the L. rhamnosus phylogenetic tree.
Enterotyping
The 16S rRNA gene bacterial profiles were collapsed at the genus level and integrated along with the Belgian Flemish Gut Flora Project (FGFP) cohort49. Identification of the enterotypes was accomplished with the Dirichlet-multinomial Model approach66. Briefly, the genus-level count matrix was rarefied to 10000 reads and merged alongside the 2998 samples of the FGFP cohort, adding the estimated fraction of unobserved genera (n = 265) according to the asymptotic maximum number of species inferred from the Lomolino model67 (R package vegan, function = “fitspecaccum”, model = “lomolino”). Dirichlet-multinomial Mixtures Model (DMM) building was done using the R library “DirichletMultinomial” function “dmn”. The optimal number of enterotypes was chosen by minimizing the BIC score. To compare enterotype distributions between our cohort and younger subjects from FGFP, a group of 199 healthy subjects matched by age, BMI and sex with the study sample was taken from the FGFP cohort.
Statistical analysis
Diversity analysis was performed using the R statistical software (v4.3.1). Beta diversity analyses from the mOTU species annotations and the GMM functional annotation were done using the “vegan” R library. The Bray-Curtis index (library “vegan”, function “vegdist”) was used to estimate the dissimilarities between samples in the QMP mOTU species and the GMM abundance table, respectively. For the GMM, the Canberra distance was also run to mitigate horse-shoe effects in the PCoA representation. Low frequent taxa and GMM (80% of zero data) were removed before the dissimilarity estimation. A distance-based redundancy analysis (dbRDA) (library “vegan” function “capscale”) was performed to reduce dimensionality in the taxonomic and functional distance matrix. The Permutational Multivariate Analysis of Variance Using Distance Matrices (ADONIS test) (library “vegan” function “adonis”) was used to check for significant associations between the microbial composition and metadata variables. For those variables that were significant in the ADONIS test, we performed a stepwise dbRDA model building to identify the non-redundant variables that best predicted the microbiome variation (Library “vegan, function “ordistep”). Clinical measures were correlated into the ordination using the function “envfit” (library “vegan”). The adonis and envfit p-values were adjusted using the Benjamini-Hochberg method (library “stats” function “p.adjust”). Observed richness, Shannon and Inverse Simpson index (library “microbiome” function “diversity”) and Pielou’s evenness (library “microbiome” function “evenness”) indices were estimated at the species level for each sample.
The effect of the interaction between the, Observed richness, Shannon, Inverse Simpson, STAI, and PSS was determined by a mixed effect model. The model considered the visit*group interaction as the fixed effect; subject ID was modelled using a random intercept (R library “ lmerTest” function “lmer”).
Delta analyses were performed by subtracting the abundance of consecutive time points for the different diversity, species, genera, clinical variables or GMM abundance. Identification of species, genera and GMM that significantly increased after probiotic intervention was done using a Wilcoxon test, within the different time points between placebo and probiotic intervention (R library “stats” function “wilcox.test”) and between the time points using Wilcoxon test (R library “stats” function “wilcox.test”, “paired = TRUE”). Associations between bacterial features (species, genus and GMM) and group/time were confirmed using a zero-inflated mixed effect negative binomial model68. The model considered the visit*group interaction as the fixed effect; subject ID was modelled using a random intercept (R library “glmmTMB” function “glmmTMB”). An ANOVA test (library “car” function “Anova”) was used to determine the significance of the visit*group interaction; a Wald test was used to determine the significance of the different factors of the model (R library “base” function “summary”). All feature*time interactions were deconfounded by fecal moisture using the step function (library “stats” function “step”). All correlations were performed using Spearman correlation (R library “stats”, function “cor.test”, “method = spearman”) and represented using a corplot (R library “corplot”, function “corplot”). Results were visualized using the R package ggplot2. All the p-values were corrected using the Benjamini-Hodge method (R library “stats” function “p.adjust”). FDR) was applied and significance was defined at FDR < 0.1.
Data availability
Raw amplicon sequencing and shotgun data have been deposited in European Genome-phenome Archive EGAC00001003263.
Code availability
Code to replicate key analyses and figures from this manuscript is available on GitHub (https://github.com/raeslab/).
References
McGrath, J. J. et al. Age of onset and cumulative risk of mental disorders: a cross-national analysis of population surveys from 29 countries. Lancet Psychiatry 10, 668–681 (2023).
Martin, C. R., Osadchiy, V., Kalani, A. & Mayer, E. A. The Brain-Gut-Microbiome Axis. Cell. Mol. Gastroenterol. Hepatol. 6, 133–148 (2018).
Shoubridge, A. P. et al. The gut microbiome and mental health: advances in research and emerging priorities. Mol. Psychiatry 27, 1908–1919 (2022).
Cryan, J. F. et al. The Microbiota-Gut-Brain Axis. Physiological Rev. 99, 1877–2013 (2019).
Kelly, J. R. et al. Lost in translation? The potential psychobiotic Lactobacillus rhamnosus (JB-1) fails to modulate stress or cognitive performance in healthy male subjects. Brain, Behav., Immun. 61, 50–59 (2017).
Chao, L. et al. Effects of Probiotics on Depressive or Anxiety Variables in Healthy Participants Under Stress Conditions or With a Depressive or Anxiety Diagnosis: A Meta-Analysis of Randomized Controlled Trials. Front. Neurol. 11, 421 (2020).
Huang, R. et al. Efficacy of Probiotics on Anxiety: A Meta-analysis of Randomized Controlled Trials. Neuropsychiatry 7, 862–871 (2017).
Le Morvan De Sequeira, C., Hengstberger, C., Enck, P. & Mack, I. Effect of Probiotics on Psychiatric Symptoms and Central Nervous System Functions in Human Health and Disease: A Systematic Review and Meta-Analysis. Nutrients 14, 621 (2022).
Liu, R. T., Walsh, R. F. L. & Sheehan, A. E. Prebiotics and probiotics for depression and anxiety: A systematic review and meta-analysis of controlled clinical trials. Neurosci. Biobehav. Rev. 102, 13–23 (2019).
Zhang, N. et al. Efficacy of probiotics on stress in healthy volunteers: A systematic review and meta-analysis based on randomized controlled trials. Brain Behav. 10, e01699 (2020).
Reis, D. J., Ilardi, S. S. & Punt, S. E. W. The anxiolytic effect of probiotics: A systematic review and meta-analysis of the clinical and preclinical literature. PLoS ONE 13, e0199041 (2018).
Valles-Colomer, M. et al. The neuroactive potential of the human gut microbiota in quality of life and depression. Nat. Microbiol 4, 623–632 (2019).
Bosch, J. A. et al. The gut microbiota and depressive symptoms across ethnic groups. Nat. Commun. 13, 7129 (2022).
Radjabzadeh, D. et al. Gut microbiome-wide association study of depressive symptoms. Nat. Commun. 13, 7128 (2022).
Qin, Y. et al. Combined effects of host genetics and diet on human gut microbiota and incident disease in a single population cohort. Nat. Genet 54, 134–142 (2022).
Yang, J. et al. Landscapes of bacterial and metabolic signatures and their interaction in major depressive disorders. Sci. Adv. 6, eaba8555 (2020).
Simpson, C. A. et al. The gut microbiota in anxiety and depression – A systematic review. Clin. Psychol. Rev. 83, 101943 (2021).
Mysonhimer, A. R., Cannavale, C. N., Bailey, M. A., Khan, N. A. & Holscher, H. D. Prebiotic Consumption Alters Microbiota but Not Biological Markers of Stress and Inflammation or Mental Health Symptoms in Healthy Adults: A Randomized, Controlled, Crossover Trial. J. Nutr. 153, 1283–1296 (2023).
Jackson, P. P. et al. Inulin-type fructans and 2’fucosyllactose alter both microbial composition and appear to alleviate stress-induced mood state in a working population compared to placebo (maltodextrin): the EFFICAD Trial, a randomized, controlled trial. Am. J. Clin. Nutr. S0002916523661143 https://doi.org/10.1016/j.ajcnut.2023.08.016 (2023).
Mäkelä, S. M. et al. Efficacy and safety of Lacticaseibacillus paracasei Lpc-37® in students facing examination stress: A randomized, triple-blind, placebo-controlled clinical trial (the ChillEx study). Brain, Behav., Immun. - Health 32, 100673 (2023).
Moloney, G. M. et al. Improvements in sleep indices during exam stress due to consumption of a Bifidobacterium longum. Brain, Behav., Immun. - Health 10, 100174 (2021).
Ng, Q. X. et al. Effect of Probiotic Supplementation on Gut Microbiota in Patients with Major Depressive Disorders: A Systematic Review. Nutrients 15, 1351 (2023).
Bloemendaal, M. et al. Probiotics-induced changes in gut microbial composition and its effects on cognitive performance after stress: exploratory analyses. Transl. Psychiatry 11, 300 (2021).
Schaub, A.-C. et al. Clinical, gut microbial and neural effects of a probiotic add-on therapy in depressed patients: a randomized controlled trial. Transl. Psychiatry 12, 227 (2022).
Liu, G., Chong, H.-X., Chung, F. Y.-L., Li, Y. & Liong, M.-T. Lactobacillus plantarum DR7 Modulated Bowel Movement and Gut Microbiota Associated with Dopamine and Serotonin Pathways in Stressed Adults. IJMS 21, 4608 (2020).
Kato-Kataoka, A. et al. Fermented Milk Containing Lactobacillus casei Strain Shirota Preserves the Diversity of the Gut Microbiota and Relieves Abdominal Dysfunction in Healthy Medical Students Exposed to Academic Stress. Appl Environ. Microbiol 82, 3649–3658 (2016).
Ma, T. et al. Probiotic consumption relieved human stress and anxiety symptoms possibly via modulating the neuroactive potential of the gut microbiota. Neurobiol. Stress 14, 100294 (2021).
Berding, K. et al. Feed your microbes to deal with stress: a psychobiotic diet impacts microbial stability and perceived stress in a healthy adult population. Mol. Psychiatry 28, 601–610 (2023).
Kaur, H., Bose, C. & Mande, S. S. Tryptophan Metabolism by Gut Microbiome and Gut-Brain-Axis: An in silico Analysis. Front. Neurosci. 13, 1365 (2019).
Wauters, L. et al. Lactobacillus rhamnosus CNCM I-3690 decreases subjective academic stress in healthy adults: a randomized placebo-controlled trial. Gut Microbes 14, 2031695 (2022).
Yoon, G., Gaynanova, I. & Müller, C. L. Microbial Networks in SPRING - Semi-parametric Rank-Based Correlation and Partial Correlation Estimation for Quantitative Microbiome Data. Front. Genet. 10, 516 (2019).
Oña, L., Shreekar, S. K. & Kost, C. Disentangling microbial interaction networks. Trends Microbiol. S0966842X25000277 https://doi.org/10.1016/j.tim.2025.01.013 (2025).
Vandeputte, D. et al. Stool consistency is strongly associated with gut microbiota richness and composition, enterotypes and bacterial growth rates. Gut 65, 57–62 (2016).
Vieira-Silva, S. et al. Quantitative microbiome profiling disentangles inflammation- and bile duct obstruction-associated microbiota alterations across PSC/IBD diagnoses. Nat. Microbiol 4, 1826–1831 (2019).
MetaCardis Consortium et al. Statin therapy is associated with lower prevalence of gut microbiota dysbiosis. Nature 581, 310–315 (2020).
Brushett, S. et al. Gut feelings: the relations between depression, anxiety, psychotropic drugs and the gut microbiome. Gut Microbes 15, 2281360 (2023).
Boehme, M. et al. Bifidobacterium longum subsp. longum Reduces Perceived Psychological Stress in Healthy Adults: An Exploratory Clinical Trial. Nutrients 15, 3122 (2023).
Lebas, M., Garault, P., Carrillo, D., Codoñer, F. M. & Derrien, M. Metabolic Response of Faecalibacterium prausnitzii to Cell-Free Supernatants from Lactic Acid Bacteria. Microorganisms 8, 1528 (2020).
Weiss, A. S. et al. In vitro interaction network of a synthetic gut bacterial community. ISME J. 16, 1095–1109 (2022).
Zaccaria, E. et al. L. rhamnosus CNCM I-3690 survival, adaptation, and small bowel microbiome impact in human. Gut Microbes 15, 2244720 (2023).
Vieira-Silva, S. et al. Species–function relationships shape ecological properties of the human gut microbiome. Nat. Microbiol 1, 16088 (2016).
Alvarez, A.-S. et al. Safety and functional enrichment of gut microbiome in healthy subjects consuming a multi-strain fermented milk product: a randomised controlled trial. Sci. Rep. 10, 15974 (2020).
FitzGerald, J. et al. Improved gut microbiome recovery following drug therapy is linked to abundance and replication of probiotic strains. Gut Microbes 14, 2094664 (2022).
Guillemard, E. et al. A Randomised, Controlled Trial: Effect of a Multi-Strain Fermented Milk on the Gut Microbiota Recovery after Helicobacter pylori Therapy. Nutrients 13, 3171 (2021).
Martín, R. et al. Over-production of exopolysaccharide by Lacticaseibacillus rhamnosus CNCM I-3690 strain cutbacks its beneficial effect on the host. Sci. Rep. 13, 6114 (2023).
Martín, R. et al. The potential probiotic Lactobacillus rhamnosus CNCM I-3690 strain protects the intestinal barrier by stimulating both mucus production and cytoprotective response. Sci. Rep. 9, 5398 (2019).
Cotillard, A. et al. A posteriori dietary patterns better explain variations of the gut microbiome than individual markers in the American Gut Project. Am. J. Clin. Nutr. 115, 432–443 (2022).
Asnicar, F. et al. Microbiome connections with host metabolism and habitual diet from 1,098 deeply phenotyped individuals. Nat. Med. 27, 321–332 (2021).
Falony, G. et al. Population-level analysis of gut microbiome variation. Science 352, 560–564 (2016).
Aasmets, O., Krigul, K. L., Lüll, K., Metspalu, A. & Org, E. Gut metagenome associations with extensive digital health data in a volunteer-based Estonian microbiome cohort. Nat. Commun. 13, 869 (2022).
Procházková, N. et al. Gut physiology and environment explain variations in human gut microbiome composition and metabolism. Nat. Microbiol 9, 3210–3225 (2024).
Chassaing, B. et al. Randomized Controlled-Feeding Study of Dietary Emulsifier Carboxymethylcellulose Reveals Detrimental Impacts on the Gut Microbiota and Metabolome. Gastroenterology 162, 743–756 (2022).
Devolder, L. et al. Gut microbiome composition is associated with long-term disability worsening in multiple sclerosis. Gut Microbes 15, 2180316 (2023).
Tito, R. Y. et al. Population-level analysis of Blastocystis subtype prevalence and variation in the human gut microbiota. Gut 68, 1180–1189 (2019).
Nurk, S., Meleshko, D., Korobeynikov, A. & Pevzner, P. A. metaSPAdes: a new versatile metagenomic assembler. Genome Res. 27, 824–834 (2017).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinforma. 11, 119 (2010).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012).
Cantalapiedra, C. P., Hernández-Plaza, A., Letunic, I., Bork, P. & Huerta-Cepas, J. eggNOG-mapper v2: Functional Annotation, Orthology Assignments, and Domain Prediction at the Metagenomic Scale. Mol. Biol. Evolution 38, 5825–5829 (2021).
Valles-Colomer, M. et al. Variation and transmission of the human gut microbiota across multiple familial generations. Nat. Microbiol 7, 87–96 (2021).
Kultima, J. R. et al. MOCAT2: a metagenomic assembly, annotation and profiling framework. Bioinformatics 32, 2520–2523 (2016).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Perrin, A. & Rocha, E. P. C. PanACoTA: a modular tool for massive microbial comparative genomics. NAR Genom. Bioinform 3, lqaa106 (2021).
Tonkin-Hill, G. et al. Producing polished prokaryotic pangenomes with the Panaroo pipeline. Genome Biol. 21, 180 (2020).
Truong, D. T., Tett, A., Pasolli, E., Huttenhower, C. & Segata, N. Microbial strain-level population structure and genetic diversity from metagenomes. Genome Res. 27, 626–638 (2017).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Holmes, I., Harris, K. & Quince, C. Dirichlet Multinomial Mixtures: Generative Models for Microbial Metagenomics. PLoS ONE 7, e30126 (2012).
Dengler, J. Which function describes the species–area relationship best? A review and empirical evaluation. J. Biogeogr. 36, 728–744 (2009).
Zhang, X. & Yi, N. NBZIMM: negative binomial and zero-inflated mixed models, with application to microbiome/metagenomics data analysis. BMC Bioinforma. 21, 488 (2020).
Acknowledgements
We thank all study participants and the different staff members involved in the execution of this study, and Duyen Nguyen, Leen Rymenans and Chloë Verspecht for technical assistance with microbiota analyses. The clinical study was investigator-initiated study funded by an unrestricted research grant from Danone to UZ Leuven. The current study on gut microbiome was fully funded by Danone in a collaboration agreement with VIB and UZ Leuven.
Author information
Authors and Affiliations
Contributions
L.W., L.V.O., K.V., T.S., Ja.T, T.V. conceived the clinical study. L.W. collected clinical data. M.D and J.R. designed and supervised the analysis of gut microbiome. L.W. performed DNA extraction, cell count, moisture, calprotectin measurements. J.F.V.C., A.G., L.F.M. and L.W. analysed gut microbiome data. J.F.V.C. and M.D. wrote the manuscript which was critically revised by all authors.
Corresponding authors
Ethics declarations
Competing interests
T.S. and M.D. were employees of Danone at the time of this project. T.S., L.W., L.V.O., T.V. are inventors of a patent on the effect of L. rhamnosus on stress (WO2020250040A1). M.D is inventor of a patent on the effect of L. rhamnosus on gut microbiome (WO2015159125A1). L.V.O. has served as a consultant for Danone, Nestlé, and The Akkermansia Company, and received research funding from Nestlé. T.V. has served as a consultant for Danone. J.R. is inventor on the patent application PCT/EP2018/084920 in the name of VIB VZW, KAtholieke Universiteit Leuven, KU Leuven Research and Development and Vrije Universiteit Brussel. J.R. received research funding from Danone. The other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Vázquez-Castellanos, J.F., Maciel, L.F., Wauters, L. et al. Probiotic-mediated modulation of gut microbiome in students exposed to academic stress: a randomized controlled trial. npj Biofilms Microbiomes 11, 140 (2025). https://doi.org/10.1038/s41522-025-00776-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41522-025-00776-w