Abstract
Aberrant cyclin-dependent kinase 2 (CDK2) activity is implicated as a resistance mechanism to CDK4/6 inhibitors (CDK4/6i) in hormone receptor–positive (HR+)/human epidermal growth factor receptor 2-negative (HER2−) breast cancer. Using preclinical patient-derived xenograft models, the CDK2i + CDK4/6i combination was active broadly across CDK4/6i-resistant and -naïve HR+ and triple-negative breast cancer models. A novel, weighted mRNA expression signature involving CCND1, CCNE1, RB1, and CDKN2A (p16) predicted response to combined inhibition of CDK2 and CDK4/6. Addition of endocrine therapy significantly enhanced antitumor activity in HR+ models, providing preclinical proof-of-concept for the broad antitumor activity of the triple combination. Early clinical data demonstrated activity of BLU-222, a potent and selective CDK2 inhibitor, both as monotherapy (CCNE1 amplified) and in combination with ribociclib and fulvestrant in patients with HR+/HER2− breast cancer. These findings provide evidence that CDK2i combined with CDK4/6i can address multiple known mechanisms of resistance to CDK4/6i, enhancing antitumor responses in preclinical breast cancer models.
Similar content being viewed by others
Introduction
Cyclin-dependent kinase 4/6 inhibitors (CDK4/6i) have significantly improved patient outcomes for hormone receptor-positive (HR+)/human epidermal growth factor receptor 2-negative (HER2−) breast cancers. Currently used in combination with endocrine therapy (ET) as adjuvant therapy and first-line standard of care for patients with metastatic breast cancer, the CDK4/6i+ET combination enhances antitumor responses and improves overall survival1. Approximately 20% of patients with breast cancer exhibit primary resistance to CDK4/6i+ET, but even patients who initially respond eventually develop resistance2,3. The mechanism of resistance to CDK4/6i can vary4 but converge on parallel pathways to promote cell cycle progression in the absence of CDK4/6 activity5. Palbociclib resistance is associated with high levels of cyclin E1 (CCNE1) expression, and poorer overall survival was observed in patients with high versus lower CCNE1 levels6. Implicated as both an intrinsic and acquired mechanism of resistance, high CCNE1 leads to aberrant activation of CDK2, which could promote G1/S phase transition in the absence of CDK4/6 activity. Therefore, selective CDK2 inhibition has been proposed as a promising therapeutic strategy in this setting5,7,8,9,10. Given the plasticity between CDK4/6 and CDK2 in regulating the G1/S phase transition11, it is possible that the benefit of a CDK2 inhibitor (CDK2i) could extend beyond the CCNE1 high setting when combined with CDK4/6i. Identification of predictive biomarkers of response for CDK2i combination strategies would enable treatment to be tailored for optimal therapeutic benefit.
Emerging preclinical data support the combination of CDK2i with CDK4/6i to overcome or delay resistance to the CDK4/6i+ET combination in HR+/HER2- breast cancer models12,13. Previous reports examining the activity of the CDK2i + CDK4/6i combination have been limited by both the use of a single CDK2 small molecule inhibitor and small numbers of tumor models or cell lines in vitro14,15. Here, we demonstrated consistent in vitro activity profiles in breast cancer cell lines, regardless of the CDK2 small molecule inhibitor tested. Further, we evaluated antitumor activity in a panel of patient-derived xenograft (PDX) models to enhance the translatability of the preclinical findings using BLU-222 as a benchmark for potent and selective CDK2i. BLU-222 demonstrated tolerability in patients when dosed in combination with ribociclib and fulvestrant in patients with HR+/HER2− breast cancer in a second-line treatment setting16. We elucidated functional determinants of response to CDK2i as a single agent and in combination with CDK4/6i (e.g., ribociclib) in models of intrinsic (HR+/HER2− and triple-negative breast cancer [TNBC]) and acquired (HR+/HER2−) CDK4/6i resistance, as well as CDK4/6i-naïve models. Further, we demonstrated that BLU‑222 + CDK4/6i combined with fulvestrant enhances antitumor response across HR+ breast cancer PDX models, addressing multiple known mechanisms of CDK4/6i resistance. Notably, early clinical observations demonstrated BLU-222 activity as monotherapy and in combination with CDK4/6i+ET in patients.
Results
Cyclin E1, Rb, and p16 are determinants of CDK2 pathway dependence in breast cancer cell lines
The G1/S phase transition relies on the cooperative action of CDK4/6-cyclin D and CDK2-cyclin E to inactivate retinoblastoma protein (Rb) and promote E2F signaling, committing the cell to initiate DNA synthesis17. In CDK4/6i-responsive cells, the G1/S phase transition is dependent on CDK4/6, and CDK2 cannot compensate for the loss of CDK4/6 catalytic activity in this context15. We therefore hypothesized that a differential expression of CDK-Rb-E2F pathway regulators could determine cell cycle dependence on CDK2 versus CDK4/6.
We first tested the antiproliferative effect of BLU-222 and palbociclib across a panel of breast cancer cell lines (n = 38) (Supplementary Data S1). BLU-222 is a potent and selective CDK2i with strong selectivity over other CDK family members18. While there was no statistically significant difference in CDK2i sensitivity based on receptor status, HR+/HER2− and HER2+ cell lines were largely sensitive to palbociclib, while TNBC cell lines were largely refractory (Supplementary Fig. S1A). In general, strong responders to BLU-222 tended to be palbociclib resistant (P < 0.01), while BLU-222 nonresponders were palbociclib sensitive (P < 0.05; Fig. 1a; Supplementary Fig. S1B). Previously, a composite biomarker signature of high CCNE1 expression, low phosphorylated-Rb (pRb), and high p16 expression was identified as a signature for CDK4/6i resistance in breast cancer PDX models19. To elucidate biomarkers that predict sensitivity to CDK2i (BLU-222) versus CDK4/6i (palbociclib), we analyzed the gene expression data from the Cancer Cell Line Encyclopedia database20. Compared with palbociclib responders, BLU-222 responders had lower expression of RB1, cyclin D1 (CCND1), and ESR1, and higher levels of CCNE1 and the endogenous CDK4/6 inhibitors, CDKN2A (p16INK4a), and CDKN2B (p15INK4b) (Fig. 1b). Although overexpressed CCNE1 or high CCND1 (defined as the 75th percentile or above using mRNA expression data from the Cancer Cell Line Encyclopedia; Western blot data in Supplementary Fig. S1C) was sufficient to enrich for palbociclib resistance, consistent with previous reports6,21, selection of cell lines based on any single biomarker was insufficient to predict single agent CDK2i sensitivity (Supplementary Fig. S1D and S1E), suggesting that predicted response can be improved using a multiple biomarker approach.
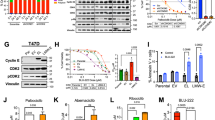
a BLU-222 and palbociclib proliferative GI50 determined by CyQUANT direct proliferation assay, 5-day compound treatment. BLU-222 responder threshold was set at 200 nM as previously described18. Dashed lines connect the BLU-222 and palbociclib IC50s from an individual cell line. P-value determined by paired t-test. b Volcano plot assessing differential gene expression in BLU-222 responders versus palbociclib responders from cell lines in (a). Points above the horizontal dashed line have an adjusted P < 0.05. The left vertical dashed line and the right vertical dashed line represent log2(FC) = -1.5 and log2(FC) = 1.5, respectively. c Kinome-selectivity of CDK2 inhibitors, as measured by KINOMEscan at 3 μM compound. S(10) is the number of kinases inhibited at <10% of control, divided by the total number of human wild-type kinases. Kinome illustrations reproduced courtesy of Cell Signaling Technology, Inc. (www.cellsignal.com) (CSTI). The foregoing website is maintained by CSTI, and Blueprint Medicines is not responsible for its content. d Antiproliferative response to CDK2 inhibitors in breast cancer cell lines as in (a). Cell lines were categorized by RB1 (Rb) expression (+: intact, −: <25th percentile expression with copy number loss or mutation), CDKN2A (p16) expression (+: >75th percentile of mRNA expression, -: ≤25th percentile of mRNA expression), and CCNE1 (cyclin E1) expression (overexpression defined as above the 75th percentile of mRNA expression). CCNE1 copy number status is noted by symbol color. Statistical significance was evaluated using the Wilcoxon rank sum test. e–l CyQuant, 5-day compound treatment, in T-47D isogenic cell lines overexpressing CCNE1, CDKN2A, or both. Mean and standard error of the mean (SEM) of two biological replicates are plotted. m Cell cycle distribution in T-47D isogenic cell lines in response to 24-h treatment BLU-222 (100 nM) or palbociclib (250 nM), ±siRB1. Mean and standard deviation (SD) of two biological replicates are plotted. n Antitumor activity of BLU-222 (60 mg/kg twice a day [BID]), ribociclib (50 mg/kg once a day [QD]), and BLU-222+ribociclib (60 mg/kg BID + 50 mg/kg QD) in BCX.022, a TNBC PDX model. 2-way ANOVA statistical deviation in tumor volume at study end is noted, *P < 0.05, **P < 0.01. Mean tumor volume and SEM are plotted, n = 6 per group.
Previously, we identified a CCNE1-high/Rb-intact/p16-high biomarker signature to predict CDK2i single-agent activity in ovarian and endometrial cancer models18. Here, we profiled a collection of highly selective CDK2 inhibitors, including three inhibitors that have been dosed in patients (BLU-222 [NCT05252416]18, INX-315 [NCT05735080]13, PF-4091 [NCT04553133])22, and two novel CDK2 inhibitors (BLU-225623 and BLU-1954 [Cpd 7724]). All molecules display favorable selectivity over CDK family members but vary in overall kinome selectivity as tested by KINOMEscan at 3 μM compound (Fig. 1c; Supplementary Data S2. In breast cancer cell lines (n = 38), the CCNE1 high, Rb-intact, p16 high biomarker signature enriched for CDK2i single agent responders (Fig. 1d). These results demonstrate both that the predictive biomarker signature can be extended to breast14,18,25 and the high degree of similarity between the selective CDK2 inhibitors in the in vitro cellular context. The biomarker signature enriches for CDK2i single-agent responder cell lines, whether using RB1 mRNA (Fig. 1d) or protein levels from immunoblotting (Supplementary Fig. S2A, S2F).
To further examine CDK sensitivity in the context of CCNE1, Rb, and p16, we developed an isogenic T-47D cell line system. T-47D is HR+ sensitive to CDK4/6i, and expresses wild-type Rb and low levels of CCNE1 and CDKN2A (p16). We overexpressed CCNE1, CDKN2A, or both (Supplementary Fig. S2A) and found that co-overexpression of CCNE1 and p16 is required to sensitize to single-agent CDK2 inhibitors (Fig. 1e–g) and lead to resistance to palbociclib (Fig. 1h) and ribociclib (Supplementary Fig. S2B), in agreement with the cell line panel results. Basal cell cycle distribution in the T-47D + CCNE1, +CDKN2A, or +CCNE1 + CDKNA was not significantly altered from the control (Supplementary Fig. S2C). The CDK2i single agent response was specific to the CDK2-CDK4/6 axis, as CCNE1/p16 overexpression was not sufficient to shift the antiproliferative response to a CDK1 inhibitor (RO-3306) (Fig. 1i), a pan-CDK inhibitor (roniciclib) (Fig. 1j), or a Wee1 inhibitor (azenosertib/Zn-c3) (Fig. 1k). This response was also not a factor of increased replication stress, as sensitivity to the ATM/ATR inhibitor berzosertib (VE-822) was not shifted (Fig. 1l). Cell cycle analysis demonstrated that in the control, palbociclib treatment led to G1 accumulation, while BLU-222 treatment had no effect (Fig. 1m; Supplementary Fig. S2C). In the CCNE1 + CDKN2A co-overexpressed T-47D cells, the effect on cell cycle distribution was reversed: BLU-222 treatment led to accumulation in G1, while palbociclib had no effect (Fig. 1m). Palbociclib induced a G1 arrest in the +CCNE1 only and +CDKN2A only cell lines (Supplementary Fig. S2C), and thus overexpression of the individual biomarkers was not sufficient to alter the cell cycle effects of CDK4/6i in this system. Furthermore, the BLU-222–induced G1 accumulation in the T-47D CCNE1 + CDKN2A cell line is Rb-dependent, as small interfering RB1 (siRB1) led to loss of G1 accumulation upon BLU-222 treatment (Fig. 1m; Supplementary Fig. S2D, S2E).
We confirmed CDK2i (BLU-222) single-agent antitumor activity in a biomarker-positive TNBC PDX model, BCX.022 (CCNE1 high [CN = 3; CN, copy number; mRNA overexpressed], Rb-intact, p16-high) (Fig. 1n; Supplementary Fig. S3A), and a TNBC cell-line derived xenograft (CDX) model, HCC1569 (CCNE1 amplified/overexpression, CN > 30, Rb-intact, p16-high) (Supplementary Fig. S3B–D). Notably, the antitumor effect of BLU-222 was not further enhanced in combination with ribociclib in the BCX.022 model, suggesting that the cells have adapted to high basal levels of p16 and therefore strong, endogenous inhibition of CDK4/6.
Intact Rb and low p16 expression enrich for CDK2i + CDK4/6i synergistic response
Our previous findings demonstrated that response to the BLU-222 + CDK4/6i combination could be predicted in CCNE1-high, Rb-intact cell lines with low p16 levels in preclinical ovarian and endometrial cancer models18, and others have demonstrated in select breast cancer cell lines that low p16 is associated with a combination response in vitro14,25. To assess the penetrance of these biomarkers in breast cancer models, we tested selective CDK2i+ribociclib in a dosing matrix to evaluate antiproliferative synergy in a high-throughput breast cancer cell line screen (n = 37). CCNE1 status alone was insufficient to predict a CDK2i+ribociclib synergistic response (Supplementary Fig. S4A). In contrast, CCND1 overexpression was significantly correlated with the greatest synergistic area scores based on the highest single-agent synergy model in 2/5 CDK2 inhibitors tested (Supplementary Fig. S4B). When stratified by RB1 and CDKN2A status, the synergistic potential was significantly greater in RB1+/CDKN2A− cells than in RB1+/CDKN2A+ cells for all selective CDK2 inhibitors, including both the investigational and novel CDK2i (BLU-2256, BLU-222, BLU-1954, PF-4091, INX-315) (Fig. 2a). Within the RB1+/CDKN2A− population, selection by CCND1 status did not further enrich for synergistic potential whereas the two CCNE1 overexpressed cell lines had high synergistic potential (Supplementary Fig. S4C and S4D). Further, the RB1+/CDKN2A− biomarker signature may predict CDK2i + CDK4/6i combination response across other cancer types (colorectal, non-small cell lung, prostate, and sarcoma), suggesting a broad potential benefit for the combination when selecting by biomarkers (Supplementary Fig. S5A–F).
a and b In breast cancer cell lines, CDK2i + ribociclib synergy was assessed by dosing matrix using CyQUANT Direct Proliferation Assay. Percent inhibition on day 5 was used to determine HSA MSA scores by SynergyFinder. MSA was evaluated by a cell line RB1 and CDKN2A status, and b cell line ribociclib response, where ribociclib-resistant cell lines were defined as a GI50 > 1 µM. Statistical significance was evaluated using the Wilcoxon rank sum test. c and d The T-47D-engineered CCNE1 and/or CDKN2A OE cell line system (as described in Fig. 1) was developed into in vivo CDX models. c Protein expression in tumors was measured by Western blot. d Tumor volume was measured over 50 days in the EV + EV (empty vector), CCNE1 + EV, and CCNE1 + CDKN2A models in BALB/c nude female mice, n = 8 per group. Mean tumor volume and SEM are plotted. Statistical deviation between treatment groups was assessed by two-way ANOVA at study end: **P < 0.01, ***P < 0.001, ****P < 0.0001.
We evaluated the importance of these biomarkers in predicting CDK2i + CDK4/6i combination potential in the context of baseline CDK4/6i response. In ribociclib-resistant cell lines (GI50 > 1 μM), synergistic potential was significantly higher in the p16-low cell lines compared with p16-high cell lines for all CDK2i tested (Fig. 2b). Notably, ribociclib-sensitive cell lines (GI50 < 1 μM) were almost exclusively p16 low, and their average synergistic potential was comparable with the ribociclib-resistant, p16-low cell lines (Fig. 2b). Together, these results demonstrate that in vitro, the BLU-222+ribociclib combination achieves the greatest synergy effects in cell lines with intact Rb and low p16 expression, with a potential role for CCNE1 and CCND1 expression, regardless of ribociclib sensitivity. This suggests that CDK2i+ribociclib will have the most active combination benefit in breast cancer cells in which there is no intrinsic, basal inhibition of CDK4/6 activity by high levels of p16.
Next, we examined the contribution of p16 levels to the BLU-222+ribociclib response using the T-47D isogenic system as CDX models in mice (Fig. 2c; Supplementary Fig. S6A–F). In agreement with the in vitro data, CCNE1 or p16 overexpression alone was insufficient to induce single-agent antitumor activity or significantly shift ribociclib response (Fig. 2d). Strikingly, the T-47DCCNE1+CDKN2A model was refractory to ribociclib and sensitized to BLU-222 single agent (Fig. 2d), confirming the importance of these proteins in predicting response to CDK2i and CDK4/6i. In combination, BLU-222+ribociclib induced significantly greater tumor growth inhibition (TGI) compared with either single agent in the p16 low models, T-47DEV+EV and T-47DCCNE1, but no additive benefit in the CDKN2A overexpression models (Fig. 2d). Together, these data provide mechanistic insight into determinants of the CDK2 versus CDK4/6 sensitivity in breast cancer cell lines, with p16 levels being a critical determinant of CDK2i single agent activity or CDK2i + CDK4/6i combination activity.
BLU-222 in combination with ribociclib enhances antitumor activity in CDK4/6i-resistant models of breast cancer
The reciprocal response to CDK2i and palbociclib demonstrated here (Fig. 1a) and by others7,9,14,15,26 suggested that the CDK2i single agent may be efficacious in the CDK4/6i progressed/resistant setting. To evaluate this, we first used an isogenic MCF-7 CDX system in which the resistant model (R) was generated by chronic exposure to palbociclib in vitro, implantation into mice, and passaging once to improve growth kinetics (WuXi AppTec). Differential gene expression analysis demonstrated RB1 downregulation and CCNE1 upregulation, as well as changes in other known markers of CDK4/6i resistance (Supplementary Fig. S7A). Compared with the parental MCF-7 model (Fig. 3a), the MCF-7 palbociclibR model was strongly sensitized to both BLU-222 (Fig. 3b) and BLU-2256 (Fig. 3c) as single agents, leading to tumor stasis and decreased pRb in tumors (Supplementary Fig. S7B–S7J). Similar results have been demonstrated in engineered CDX models and select in vitro cell lines13,14,25. TGI in both the MCF-7 parental and palbociclibR models correlated with inhibition of thymidine kinase 1 (TK1), indicating suppression of the CDK-Rb-E2F pathway leading to antitumor response (Supplementary Fig. S7K–S8L).
a MCF-7 parental or b, c palbociclib-resistant CDX models in BALB/c nude female mice treated with BLU-222 (100 mg/kg BID), BLU-2256 (1, 5, or 10 mg/kg BID), ribociclib (75 mg/kg QD), or the combination (BLU-222 100 mg/kg BID + ribociclib 75 mg/kg QD or BLU-2256 5 mg/kg BID or 10 mg/kg QD + ribociclib 75 mg/kg QD) for 28 days. Tumors were monitored for an additional 21-day, dosing-free observation period to assess treatment durability. Mean tumor volume and SEM are plotted, n = 8 per group. Statistical deviation of tumor volume between groups at study end is indicated in the figure, 2-way ANOVA: **P < 0.01, ***P < 0.001, ****P < 0.0001. TGI and statistical deviation of tumor volume of treatment groups versus vehicle was assessed at dosing-end (28 days) by 2-way ANOVA, **P < 0.01, ***P < 0.001, ****P < 0.0001: MCF-7 parental (a): BLU-222 TGI = 45%****; ribociclib TGI = 97%****; BLU-222+ribociclib TGI = 108%****; MCF-7 palbociclib-resistant (b): BLU-222 TGI = 83%***; ribociclib TGI = 23%; BLU-222+ribociclib TGI = 111%****; (c) BLU-2256 1 mg/kg BID TGI = 12%, BLU-2256 5 mg/kg BID TGI = 72%, BLU-2256 10 mg/kg QD TGI = 75%*, ribociclib TGI = 29%, BLU-2256 5 mg/kg BID + ribociclib TGI = 113%****, BLU-2256 10 mg/kg QD + ribociclib TGI = 123%****. d Protein expression by Western blot in T-47D and MCF-7 parental and CDK4/6i-resistant cell lines. e–h Antiproliferative effect determined by CyQUANT after 5 d treatment in T-47D and MCF-7 parental and CDK4/6i-resistant cell lines; e ribociclib single agent (IC50 mean ± SEM, n ≥ 2); f BLU-222 single agent (IC50 mean ± SEM, n ≥ 2); g and h BLU-222+ribociclib highest single agent synergy plots.
We next examined if the CDK2i + CDK4/6i combination induced additional antitumor activity in the MCF-7 parental and palbociclibR models. While ribociclib induced a strong antitumor effect when on treatment in the parental model (97% TGI), the response was not durable and tumors regrew after treatment cessation (Fig. 3a). The BLU-222+ribociclib combination had strong on-treatment antitumor activity, and this response was durable, with continued stasis after treatment was removed (Fig. 3a). In the MCF-7 palbociclibR model, the CDK2i combination with ribociclib only modestly improved upon the strong single-agent activity induced by CDK2i (Fig. 3b, c).
To explore CDK2i activity in additional models of CDK4/6i resistance, T-47D and MCF‑7 cells were rendered resistant (R) to ribociclib, abemaciclib, or atirmociclib by chronic exposure in vitro over ~6 months, producing a polyclonal population of resistant cells. T-47D–resistance cell lines trended toward increased CDK6 expression and lower Rb expression, with total loss in the atirmociclibR cell line (Fig. 3d). MCF-7–resistance cell lines trended toward increased cyclin E1 and CDK6 expression (Fig. 3d). Other markers of known CDK4/6i resistance were also found to be modulated in these cell lines (Supplementary Fig. S8A, S8B). While these cell lines were resistant to ribociclib (Fig. 3e) and other CDK4/6i (Fig. 3e; Supplementary Fig. S9A–C), they were not sensitized to CDK2i single agent (Fig. 3f), in contrast to the MCF-7 palbociclibR CDX model (Fig. 3a–c). These cell lines were also not sensitized to INX-315 (Fig. S9D, S9E), an investigational CDK2i that had previously been demonstrated to have enhanced activity in abemaciclib-resistant MCF-7 and T-47D cell lines13. This suggests that different mechanisms of CDK4/6i resistance may emerge during model development, which is a polyclonal and stochastic process, and that not all will be addressable with CDK2i as a single agent. Instead, convergence of multi-biomarker signatures will predict CDK2 essentiality in CDK4/6i-resistant models.
We then assessed the BLU-222+ribociclib combination by synergy analysis in the CDK4/6i-induced-resistance cell lines. Compared with parental cell lines, the ribociclibR, abemaciclibR, and atirmociclibR cell lines had increased synergistic potential (Fig. 3g, h). Evaluating the in vitro antiproliferative effect of the BLU-222+ribociclib combination, a strong combination benefit was seen in the parental cell lines (Supplementary Fig. S9F, S9G). This combination benefit was sustained in the resistant cell lines, with the exception of the T-47D atirmociclibR cell line (Supplementary Fig. S9F), which had lost Rb expression (Fig. 3d). Consistent with this, pRb, TK1, and cyclin A2 were downregulated upon combination treatment in the ribociclibR and abemaciclibR cell lines, while no modulation in the Rb-null T‑47D atirmociclibR cell line was observed (Supplementary Fig. S10A, S10B). These results reveal that the BLU-222+ribociclib combination can overcome resistance to multiple CDK4/6i in vitro, even when combined with the CDK4/6i used to generate primary resistance.
BLU-222 in combination with ribociclib overcomes CDK4/6i resistance in breast cancer PDX models
To better assess the translatability of the BLU-222+ribociclib combination to in vivo models and patients, we measured the antiproliferative response in a panel of HR+ breast PDX models using an ex vivo spheroid platform (KIYA-Predict). In CDK4/6i-progressed PDX models selected from the XenoSTART PDX library, BLU-222 + ribociclib improved the ex vivo antiproliferative effect compared with single agents in the majority (6/9) of models tested (Fig. 4a). Interestingly, a model (STM188) derived from a patient who had progressed on both palbociclib + fulvestrant (2 years on-treatment), as well as an investigational CDK2i, was responsive to the BLU-222 + ribociclib combination (Fig. 4a). Consistent with the CDK4/6i-resistant cell line panel, these data support that BLU-222 + ribociclib combination may address resistance to multiple CDK4/6i and overcome various mechanisms of resistance.
a Ex vivo PDX drug response screen using KIYA-PREDICT dosing matrix assay platform. Viability was assessed by CTG and percent inhibition was calculated relative to DMSO control. IC50 for BLU-222 + ribociclib was calculated by generating a percent inhibition curve at equi-potent doses of the test agents across the dosing matrix, n = 2. b Antitumor activity of BLU-222 (60 mg/kg BID) and ribociclib (50 mg/kg BID) dosed as monotherapy or in combination in the TNBC PDX model, BCX.051. Mean tumor volume and SEM are plotted, n = 6 per group. Statistical deviation in tumor volume between groups was tested using a 2-way ANOVA: *P < 0.05, ***P < 0.001. c Tumor growth of BLU-222 60 mg/kg BID + ribociclib 50 mg/kg QD relative to vehicle in HR+ and TNBC PDX model panel. d CDK4/6i resistance mechanisms in the PDX models from (c) were evaluated using baseline tumor WES and RNA-seq data. Gene expression was corroborated with IHC %N for RB1, CCND1, CDKN2A (high: ≥75%, low ≤10%) and CCNE1 (high: ≥50%, low: ≤10%). Nonresponders had TGI < 50%, partial responders had TGI between 50% and 70%, and responders had TGI ≥ 70%. XC5172013 did not have transcriptomic or genomic data available. e BLU-222+ribociclib responders (%TGI > 70%) and nonresponders (%TGI < 70%) were evaluated relative to known CDK4/6i resistance mechanisms. Numbers above bars represent the number of models. f–h Proportions of known CDK4/6i resistance mechanisms shown in (e) represented within the PDX models. Models with 3+ resistance mechanisms had three or more identified markers, and those with two resistance mechanisms harbored two identified markers. PDX model names are abbreviated in (c and d); see Supplementary Data S8-8 for full model information.
Given the response seen with BLU-222+ribociclib in CDK4/6i-experienced HR+ breast cancer models, we investigated whether the combination was active in models of intrinsic (CDK4/6i-naïve) CDK4/6i resistance. The TNBC subtype is generally considered to be resistant to CDK4/6i due to high rates of Rb loss27, but both in vitro and in vivo data demonstrate that some models are sensitive to palbociclib21,28,29. BCX.051 is an Rb-intact, CCNE1 nonamplified–TNBC PDX model derived from a patient with no previous exposure to CDK4/6i or ET. In this model, single-agent BLU-222 or ribociclib induced some antitumor activity (63% TGI and 50% TGI, respectively), while the BLU-222 + ribociclib combination achieved stasis (88% TGI; Fig. 4b). This result highlights the potential utility of the BLU-222 + CDK4/6i combination in TNBC, consistent with recent reports30.
In accordance with in vitro models of CDK4/6i-resistance (Fig. 3f), BLU-222 monotherapy did not induce strong antitumor activity in the majority of CDK4/6i-resistant PDX models (Supplementary Figs. S12 and S13). To further assess the BLU-222 + ribociclib therapeutic potential and identify potential predictive biomarkers, we screened the combination in 13 HR+/HER2− and 14 TNBC PDX models in mice. The HR+/HER2− models included both CDK4/6i-experienced and -naïve models. The combination was well tolerated in mice (Supplementary Fig. S11) and induced a strong antitumor effect, eliciting >70% TGI in 69% of the HR+/HER2− and 50% of the TNBC models (Fig. 4c; Supplementary Data S3; Supplementary Figs. S12 and S13). Among the 27 models screened, 25 were also treated with BLU-222 and CDK4/6i monotherapy. Of CDK4/6i monotherapy nonresponders (n = 22), 55% (12/22) responded to the BLU-222 + ribociclib combination, irrespective of prior CDK4/6i exposure (Supplementary Figs. S12 and S13). The combination elicited superior TGI response in 100% (25/25) of the models compared with CDK4/6i monotherapy and in 81% (22/27) compared with BLU-222 monotherapy (Supplementary Data S3; Supplementary Figs. S12A and S13A).
Genomic or transcriptomic analyses were performed on all 27 models to assess known intrinsic and acquired markers of CDK4/6i resistance (markers described in Asghar et al.4) as well as other cell cycle-related markers (Fig. 4d). Resistance mechanisms were identified in 78% (21/27) of the models, with the most common being RAS activation, CCNE1 high, and FGFR2 activation (Fig. 4e and f). Of responders to the combination, 81% (13/16) harbored one to two known CDK4/6i resistance markers (Fig. 4g), while 73% (8/11) nonresponders to the combination harbored ≥3 known markers of resistance (Fig. 4h). These data indicate that the BLU-222 + ribociclib combination can overcome a broad spectrum of CDK4/6i resistance mechanisms, with the notable exception of Rb loss or when ≥3 resistance mechanisms are present. Importantly, these findings demonstrate that targeting CDK2 extends therapeutic benefit beyond the CCNE1 overexpressed setting when combined with a CDK4/6i.
Cyclin E1/D1, p16, and Rb expression can be used to predict BLU-222 + ribociclib response in HR+ and triple-negative breast PDX models
In breast cell lines, we demonstrated that RB1 and CDKN2A (p16) mRNA expression levels were associated with a CDK2i+ribociclib combination benefit, both in the CCNE1 normal and CCNE1 overexpression setting (Fig. 2a; Supplementary Fig. S4). Notably, breast cancer cell lines with CCND1 overexpression exhibited significant synergistic potential, and high CCND1 levels have been paradoxically associated preclinically with both increased sensitivity to CDK4/6 inhibition21 and resistance to CDK4/6i19. To further investigate, we assessed whether CCND1, CCNE1, RB1, and/or CDKN2A baseline mRNA expression could predict antitumor response to the BLU-222 + CDK4/6i combination. PDX models were treated with BLU-222, ribociclib, or palbociclib as monotherapy, or the BLU-222 + CDK4/6i combination (Supplementary Data S3; Supplementary Figs. S12 and S13). Among the biomarkers analyzed, CCNE1 expression as a single biomarker was not significantly correlated with response to BLU-222 + CDK4/6i, while higher CCND1 expression, higher RB1 expression, and lower CDKN2A expression were each significantly correlated with response (Fig. 5a), consistent with directional trends observed in breast cell lines (Fig. 2a; Supplementary Fig. S4). mRNA expression significantly correlated with protein expression by IHC in these PDX models (Supplementary Fig. S14).
a Kendall’s tau correlation between BLU-222 + ribociclib-induced TGI in PDX models and gene expression for a single biomarker of interest. b Pearson’s correlation between observed BLU-222+ribociclib-induced TGI and predicted BLU-222 + ribociclib TGI estimated from a weighted, multivariate biomarker signature developed from a generalized linear model. c Differential observed TGI between predicted BLU-222 + ribociclib responder (predicted TGI ≥ 70%) and predicted BLU-222+ribociclib nonresponder (predicted TGI < 70%). d Receiver operator curves for the prediction models for each single biomarker and combinations of CCND1, CCNE1, RB1, and CDKN2A. e CDK2i, ribociclib, or the combination in a predicted CDK2i + CDK4/6i responder PDX model. PDX.003.014 is an ER+ breast cancer model derived from a patient who had progressed on CDK4/6i. f Antitumor activity in a predicted CDK2i + CDK4/6i combination nonresponder PDX model. PDX.003.234 is an ER+ breast cancer model derived from a CDK4/6i-naïve patient previously treated with chemotherapy. BLU-222 studies dosed as follows: BLU-222 60 mg/kg BID, ribociclib 50 mg/kg QD, BLU-222+ribociclib 60 mg/kg BID/50 mg/kg QD. BLU-2256 studies dosed as follows: BLU-2256 10 mg/kg QD, ribociclib 75 mg/kg QD, BLU-2256+ribociclib 10 mg/kg QD/75 mg/kg QD. Mean tumor volume and SEM are plotted, n = 6-8 per group. TGI from vehicle is noted: 2-way ANOVA, **P < 0.01, ***P < 0.001, ns = not significant.
Although individual markers were correlated with response, prediction models built on any single marker only achieved a maximum validation accuracy of 62.5%, highlighting the need for multivariate biomarker approaches (Supplementary Data S4). To address this, we developed a multivariate biomarker signature based on the weighted mRNA expression of CCND1, CCNE1, RB1, and CDKN2A to predict response to the combination of BLU-222 and CDK4/6i in both HR+ and TNBC models. The multigene signature predicted response with 75% accuracy, 100% sensitivity, and 67% positive predictive value (PPV) in its validation set (n = 8) (Supplementary Data S4). Across the entire data set (n = 27), the model predicted response with 89% accuracy, 94% sensitivity, and 88% PPV (Fig. 5b, c; Supplementary Data S4). Furthermore, we modeled potential combinations of these four genes to predict TGI as a binary outcome (responder ≥ 70% TGI, nonresponder < 70% TGI), and confirmed that the CCND1/CCNE1/RB1/CDKN2A combination provided the most robust predictive power compared with any single biomarker or other combination (Fig. 5d). Therefore, this novel, weighted biomarker signature predicts that breast tumors with intact Rb, high CCND1, low CDKN2A, with or without high CCNE1 expression, would be the strongest responders to the BLU-222 + CDK4/6i combination, regardless of receptor subtype.
Although the signature was developed using BLU-222 data, we hypothesized that its predictive utility could extend to other selective CDK2 inhibitors. To test this, we evaluated the signature in two ER+ models treated with BLU-2256, a CDK2i that is structurally diverse from BLU-222 and that was not included in the training set. PDX.003.014 was correctly predicted as a responder to CDK2i+ribociclib (predicted TGI = 88%), with BLU-222+ribociclib achieving 108% TGI (P < 0.001) and BLU-2256+ribociclib achieving 104% TGI (P < 0.001) (Fig. 5e; Supplementary Data S4). In contrast, PDX.003.234 was correctly predicted as a nonresponder to CDK2i + CDK4/6i (predicted TGI = 56%), with BLU-222+ribociclib achieving 55% TGI (P > 0.05) and BLU-2256+ribociclib achieving 62% TGI (P > 0.05) (Fig. 5f; Supplementary Data S3). These results validate the predictive power of the multivariate biomarker signature and demonstrate its generalizability across structurally diverse CDK2i + CDK4/6i combinations. Further, these data support the use of this biomarker signature to identify tumors most likely to benefit from dual CDK2i, CDK4/6 inhibition in breast cancer.
Fulvestrant enhances the antitumor effect of BLU-222 + ribociclib broadly across HR+ breast cancer PDX models
CDK4/6i efficacy is enhanced with the addition of ET31. We evaluated the activity of the BLU-222+ribociclib+fulvestrant combination in both CDK4/6i-naïve or -experienced HR+/HER2− breast cancer PDX models. The BLU-222+ribociclib+fulvestrant combination was well tolerated in mice (Supplementary Figs. S11 and S15). Combination activity in two representative HR+ breast cancer models, ST2056 and ST941, is presented. ST2056 (Fig. 6a) was derived from a patient who had received fulvestrant+abemaciclib, then fulvestrant alone, followed by investigational therapy; ST941 (Fig. 6b) was derived from a patient who had no previous treatment with CDK4/6i.
a and b In vivo antitumor activity of BLU-222 (60 mg/kg BID [twice a day]), ribociclib (50 mg/kg QD [once a day]), fulvestrant (2.5 mg Q7D [once weekly]), or the combinations thereof. Mean tumor volume and SEM are plotted, n = 8 per group. a ST2056 is a model derived from a patient who had previously received abemaciclib+fulvestrant, and b ST941 is a model derived from a patient who had not received any prior lines of CDK4/6i therapy. For ST941, treatment ceased on day 31, and tumors were monitored for an additional 29 days to assess the durability of the response. Percent TGI was calculated using tumor volume from the final day of treatment: BLU-222, 41%; BLU-222 + fulvestrant, 80%; ribociclib+fulvestrant, 71%; BLU-222+ribociclib+fulvestrant, 97%. c Tumor growth relative to vehicle in PDX models categorized as CDK4/6i-experienced (model derived from a patient who had received prior lines of CDK4/6i therapy) or CDK4/6i-naïve (derived from a patient who had received no prior lines of CDK4/6i therapy). An additional three models in the PDX panel received BLU-222 + CDK4/6i + fulvestrant but did not receive ribociclib + fulvestrant as a comparator arm and therefore are not shown here (Supplementary Fig. S12). d Top 10 most enriched pathways from gene set enrichment analysis (GSEA) of differential gene expression between BLU-222+ribociclib+fulvestrant (triplet) and ribociclib + fulvestrant (duplet) in triplet responder PDX models that did not respond to the duplet. e Enrichment score relative to gene rank of Hallmark Interferon Alpha Response. f Enrichment score relative to gene rank of E2F Targets pathway genes.
In the ST2056 model, although the patient had experienced both fulvestrant and a CDK4/6i, antitumor activity was induced with the ribociclib+fulvestrant combination (83% TGI; Fig. 6a). BLU-222 + fulvestrant demonstrated antitumor activity (85% TGI) that was equivalent to ribociclib+fulvestrant (Fig. 6a). Combining BLU-222+ribociclib+fulvestrant led to 100% TGI, and therefore induced an antitumor response that was superior to the clinical standard of ribociclib+fulvestrant (Fig. 6a).
In the ST941 model (CDK4/6i-naïve), ribociclib+fulvestrant achieved 71% TGI (Fig. 6b). BLU-222 monotherapy induced modest activity (41% TGI), while BLU-222 + fulvestrant led to 80% TGI at the end-of-treatment period (Fig. 6b), and the triple BLU-222+ribociclib+fulvestrant combination induced tumor stasis (103% TGI; Fig. 6b), similar to results from the CDK4/6i-progressed model (Fig. 6a). Importantly, the response to BLU-222+ribociclib+fulvestrant combination was sustained for a longer period upon dosing cessation compared with duplet regimens, ribociclib+fulvestrant, or BLU-222+fulvestrant (Fig. 6b).
To extend these findings, we evaluated the triple combination, BLU‑222+ribociclib+fulvestrant, in 14 HR+ breast cancer PDX models, encompassing both CDK4/6i-experienced and -naïve tumors. The triple combination demonstrated broad activity, achieving >70% TGI in 11/14 models, and consistently outperformed ribociclib+fulvestrant (Fig. 6c). Importantly, the triple combination treatment elicited responses of ≥70% TGI in 5/8 of the nonresponders to the CDK4/6i + fulvestrant combination (Fig. 6c), including in an Rb-null model (ST1799/PBR). However, the limited number of nonresponders to the triple combination precluded biomarker-based stratification, suggesting activity independent of biomarker status. Overall, CDK4/6i-experienced models demonstrated a greater proportion of responders (8/8) to the triple combination treatment than CDK4/6i-naïve models (3/6) (Fig. 6c), suggesting a potentially greater role for CDK2 in promoting tumor growth in the CDK4/6i progressed setting.
To investigate the mechanistic basis of the observed activity, we compared gene expression changes in on-treatment tumors to elucidate the added contribution of BLU-222 to a CDK4/6i+ET regimen. We compared ribociclib+fulvestrant to BLU-222+ribociclib+fulvestrant in models that did not respond to the CDK4/6i+ET double combination but were responders to the triple combination. Gene enrichment analysis demonstrated that the addition of BLU-222 significantly enhanced suppression of E2F pathway gene expression, a key downstream target of the CDK-Rb axis (Fig. 6d–f). Consistent with a previous report13, this suggests that the dual CDK2i + CDK4/6i in the presence of ET more completely inhibits the Rb-E2F signaling pathway, leading to improved antitumor activity. Additionally, the IFNα signaling pathway was significantly reduced in the triple combination (Fig. 6d–f) versus the duplet, suggesting potential involvement of the IFN or JAK/STAT pathway contributing to the antitumor effect or that the downregulation is a consequence of cell cycle arrest.
Together, these results demonstrate that strong antitumor activity can be achieved with a BLU-222 + CDK4/6i+ET regimen in HR+ breast cancer models that is deeper and acts more broadly compared with treatment with a CDK4/6i+ET alone. Furthermore, the enhanced efficacy of the triplet combination highlights the potential of a clinically meaningful benefit to patients with HR+ breast cancer.
BLU-222 monotherapy or in combination with ribociclib and fulvestrant demonstrates early clinical activity in patients
BLU-222 completed a phase I/II clinical trial (ClinicalTrials.gov. https://clinicaltrials.gov/study/NCT05252416 [2024]) investigating BLU-222 as monotherapy for adult patients with advanced solid tumors and in combination with ribociclib and fulvestrant for patients with HR+/HER2− breast cancer. BLU-222 was administered orally twice daily (BID) either as monotherapy in patients with heavily pretreated advanced cancers, or as combination therapy with ribociclib and fulvestrant in patients with HR+/HER2− breast cancers whose disease had progressed after treatment with a CDK4/6i (NCT05252416). Radiographic response was assessed per Response Evaluation Criteria in Solid Tumors, version 1.132. Two patients with confirmed partial responses are presented here as vignettes to illustrate the potential clinical activity of BLU-222 either as monotherapy or in combination with ribociclib+fulvestrant in HR+/HER2− breast cancer (data cutoff date of July 26, 2024).
BLU-222 monotherapy
A female, aged 57 years, with HR+/HER2− breast cancer and evidence of CCNE1 amplification (CCNE1 CN > 20, as detected by in situ hybridization), who had three prior lines of anticancer therapy in the metastatic setting, including palbociclib and fulvestrant, abemaciclib and anastrozole, and capecitabine, was enrolled in the monotherapy cohort at a dose level of 800 mg BID. She was dose-reduced to 400 mg BID on cycle 1 day 8 due to grade 3 nausea, which resolved with this dose adjustment. After 8 weeks of BLU-222 monotherapy treatment, a computed tomography (CT) scan revealed a 43% reduction in the measurable disease in her liver (Fig. 7a). A second scan at week 16 confirmed a partial response with 37% overall tumor reduction. The patient subsequently came off study at week 17 for increasing disease-related bone pain without objective signs of radiographic progression.
a A CT scan demonstrates a 43% reduction in measurable disease in the liver of a patient with HR+ breast cancer after 8 weeks of treatment with BLU-222 monotherapy (400 mg BID) (patient 1). b A CT scan demonstrates loss of measurable disease in the liver after 16 weeks of treatment with BLU-222 (400 mg BID) + ribociclib (400 mg QD) + fulvestrant in a patient with HR+ breast cancer (patient 2). c Representative IHC images of baseline and on-treatment tumor biopsy samples from patient two responding to treatment with BLU-222+ribociclib+fulvestrant stained with antibodies against pRb S807/811 and Ki67. d Longitudinal ctDNA tumor fraction and variant allele frequency in patient 2 responding to treatment with BLU-222+ribociclib+fulvestrant.
BLU-222 in combination
A female, aged 77 years, with HR+/HER2− breast cancer, who had six prior lines of anticancer therapy in the metastatic setting, including palbociclib and letrozole, everolimus, and exemestane, alpelisib and fulvestrant, capecitabine, gemcitabine, and trastuzumab deruxtecan, was enrolled in the combination therapy cohort at a dose level of 400 mg BID BLU-222 and 400 mg once daily (QD) ribociclib in combination with fulvestrant (500 mg). At baseline, immunohistochemistry (IHC) of a tumor biopsy showed cyclin E1 as 2% positive, H score 2, Rb as 80% positive, H score 114, and p16 as 45% positive, H score 75. After 8 weeks of combination therapy, a 30% reduction in the measurable disease in the patient’s liver was observed by computed tomography scan. At week 16, the target liver lesion was no longer detectable (Fig. 7b), confirming a partial response (given persistent nontarget bone lesions). An on-treatment biopsy on cycle 1 day 10 demonstrated an 82% reduction in pRb and 90% reduction in Ki67 compared with baseline (Fig. 7c). Interrogation of circulating tumor DNA in plasma collected at baseline and at 2 weeks after starting therapy demonstrated a rapid 68% reduction in circulating tumor DNA tumor fraction with a reduced allele burden in multiple genes of interest, including PIK3CA and ESR1 (Fig. 7d). Additionally, a significant 81% reduction in TK1 level was observed at the 2-week interval. The patient remained on the study with ongoing response for 12 cycles.
Discussion
Identifying novel treatment regimens for CDK4/6i-progressed patients is critical to meet the increasing need for efficacious therapies to address resistance in HR+ breast cancer. Currently, a combination of CDK4/6i and ET is the standard of care for HR + /HER2- advanced or metastatic breast cancer, but resistance remains an inevitable challenge for most patients. The mechanisms of CDK4/6i resistance are heterogeneous, involving varied aberrations in cell growth and cell cycle control pathways, which can be either intrinsic or acquired mechanisms of resistance4,10. Here, we reported that the combination treatment of CDK2i with CDK4/6i (±ET) has the potential to address multiple mechanisms of resistance. Furthermore, using BLU-222 as an exemplar of selective CDK2i, we identified a multivariate biomarker signature comprising Rb, p16, CCNE1, and CCND1 that reflects the underlying biology and predicts response to CDK2i + CDK4/6i combination therapy.
Aberrant CDK2 activation via upregulation of CCNE1 has been implicated as a bypass mechanism for CDK4/6 inhibition, suggesting a key function for CDK2 in promoting cell cycle progression in this setting15. This provides a strong rationale for targeting CDK2 to overcome resistance to CDK4/6i. In this study, we demonstrated that Rb, p16, and CCNE1 are key biomarkers predictive of CDK2i monotherapy response both in vitro (Fig. 1) and in vivo (Fig. 3). However, monotherapy with BLU-222 was not sufficient to induce strong antitumor activity broadly across CDK4/6i-resistant PDX models (Fig. 4d; Supplementary Figs. S12 and S13). To address this, we moved to a combination approach and leveraged the molecular interplay of the identified biomarkers to develop a multivariate, weighted CCND1/RB1/CDKN2A/CCNE1 biomarker signature capable of predicting the combination benefit of BLU-222 (CDK2i) + CDK4/6i across different breast cancer subtypes (Fig. 5). Our findings indicated that this combination resulted in a strong antitumor response in various breast cancer models, including those resistant to CDK4/6i. The combination effectively overcame multiple mechanisms of resistance (Fig. 4d) and delayed the emergence of CDK4/6i resistance in the CDK4/6i-naïve setting (Fig. 6b). Notably, the BLU-222+ribociclib combination was active in models of TNBC (Fig. 4c; Supplementary Fig. S13), a subtype that is largely resistant to CDK4/6i and for which CDK4/6i is not standard of care27, suggesting broad potential for clinical application.
One of the prevailing hypotheses in CDK4/6i-resistant breast cancer is that high levels of CCNE1 lead to aberrant CDK2 activation, rendering CDK2 a targetable vulnerability in this setting6,27. Our data identify that high CCNE1 expression, along with Rb and p16 levels, is an important component for predicting response to BLU-222 monotherapy. However, interestingly, high CCNE1 levels are not a prerequisite for a CDK2i + CDK4/6i combination benefit, either in cell lines (Fig. 2, Supplementary Fig. S4) or in in vivo models (Figs. 4d and 5). Instead, our results indicate that CCNE1 overexpression is one of several CDK4/6i mechanisms of resistance that can be effectively addressed with the CDK2i + CDK4/6i combination (Fig. 4d–f). In fact, CCND1 levels were more important for predicting response to combination treatment than CCNE1 levels based on the weighted biomarker signature (Fig. 5b–d). The role of cyclin D in mediating response to CDK4/6i is conflicting. While previous work in cell lines demonstrated that high levels of CCND1 correlated with responsiveness to palbociclib21, CCND1 was not identified as a predictive biomarker of clinical response to palbociclib in PALOMA-1, -2, or -3 trials33. Furthermore, CCND1 amplification and high CCND1 expression were markers of CDK4/6i resistance in a panel of PDX models19. In breast cancer models, the high levels of CCND1 could either be impacting CDK4/6-cyclin D activity or, alternatively, could be leading to noncanonical CDK2-cyclin D activity to impact cell cycle progression8, resulting in CDK2 vulnerability. The role of cyclin D in mediating response to the CDK2i + CDK4/6i combination remains to be elucidated, and its utility as a predictive biomarker of response will be informed by analysis of clinical data.
The BLU-222 + CDK4/6i biomarker signature was impactful in elucidating functional determinants of response and mechanism of action of targeting CDK2 and CDK4/6 in breast cancer. The current standard of care for metastatic breast cancer is a regimen of CDK4/6i paired with ET27, and therefore, it was important to evaluate the triple BLU-222 + CDK4/6i + ET combination. In HR+, CDK4/6i-experienced and -naïve models, we demonstrated that the triple combination was well-tolerated in preclinical mouse models and enhanced efficacy over the BLU-222+ribociclib or ribociclib+fulvestrant (ET) doublet combinations, in HR+, CDK4/6i‑experienced and -naïve models (Fig. 6a–c; Supplementary Fig. S12). The BLU‑222+ribociclib+fulvestrant combination also improved efficacy across a range of genotypes, for example, inducing a strong antitumor effect in an Rb-null model of HR+ breast cancer (ST1799/PBR) (Fig. 6c; Supplementary Fig. S12). Rb loss is relatively infrequent in patients participating in clinical trials (5–10%)4, but these patients receive no benefit upon CDK4/6i retrial34. In our newly generated CDK4/6i-resistant cell line panel, Rb loss was observed in the atirmociclibR T-47D cell line (Fig. 3d). Therefore, the frequency of Rb loss as an acquired mechanism of resistance to CDK4-selective inhibitors could evolve in this patient population and is worth monitoring. The triple combination of BLU-222 + ribociclib + ET could potentially represent a potent combination strategy to address patients who progress on CDK4/6i (or CDK4i) + ET, regardless of mechanism of resistance, capturing a broad patient population.
CDK2 inhibitors have the potential to significantly shape the breast cancer treatment landscape, offering a promising combination partner for CDK4/6 inhibitors and ET to address and/or delay resistance to CDK4/6i. The excitement for this target is reflected in the rapidly growing number of CDK2 inhibitors in clinical development35. Our findings with BLU‑222 + CDK4/6i elucidate the underlying pathway modulation that results in enhanced suppression of the CDK-Rb-E2F pathway, overcoming mechanisms of resistance to CDK4/6i and supporting clinical use in combination with ET. Preliminary, first proof-of-concept clinical data show promising activity of BLU-222 in patients with breast cancer, both as monotherapy (CCNE1 amplified) (Fig. 7a) and in combination with ribociclib and fulvestrant (Fig. 7b–d). This reinforces the translational relevance of our preclinical findings and suggests that this novel combination strategy could offer a promising therapeutic option for patients with HR+ breast cancer, particularly those who have developed resistance to CDK4/6i. As the therapeutic landscape evolves, we can expect to see novel therapeutic modalities enter the clinic that offer potential selectivity and potency advantages, such as CDK2-36 or CDK4-targeted protein degraders, as well as next-generation CDK4-selective ATP-competitive small molecule inhibitors37. The underlying mechanism of action and biomarkers of importance identified here may aid in the interpretation of emerging clinical data and inform future clinical trial patient selection strategies, ultimately leading to increased therapeutic promise of CDK2i + CDK4/6i combinations for patients.
Methods
Cell lines and culture
Cell line and culturing information are provided in Supplementary Data S1. All cell lines are regularly tested for mycoplasma and identity confirmed by short tandem repeat analysis.
CyQUANT proliferation assay
Cells were seeded in black, clear-bottom, 384-well (Corning CLS3571) or 96-well (Corning 3904) plates at densities listed in Supplementary Data S1 and allowed to adhere overnight at 37 °C/5% CO2. For single-agent treatment, cells were treated with the compound in a 10-point dose titration, top dose 25 µM, 1:4 dilutions. For combination treatment, cells were dosed with compound in a 9-point dose titration matrix, top dose 3 µM, 1:3 dilutions. Cells were treated for 120 hours (5 days), and the CyQUANT Cell Direct Proliferation Assay was performed according to the manufacturer’s instructions (Invitrogen, C35012). Experiments were run in at least duplicate. Half-maximal inhibitory concentration (IC50) values were calculated using percent inhibition relative to dimethylsulfoxide (DMSO) and staurosporine controls at 120 h. Half-maximal growth inhibition values were calculated using percentage inhibition relative to DMSO and day 0. Synergy scores were determined using SynergyFinder 3.0 (RRID:SCR_019318)38. The most synergistic area was determined as the 3 × 3 dose window with the highest average synergy score, and synergy scores > 10 had synergistic potential.
T-47D CCNE1/CDKN2A engineered cell lines
For in vitro cell line models, T-47D (ATCC HTB-133; 0.2 × 106 cells) were transduced at 1:1 multiplicity of infection with lentiviral constructs containing pLVX-ER1α-IRES-puro (GenScript), pLV-ER1α-IRES-neo, CCNE1_pLVX-ER1α-IRES-puro (GenScript), or p16_pLVX-ER1α-IRES-neo (GenScript). Transductants were selected in 1 μg/mL puromycin and 800 μg/mL G418, per construct marker, for 7 days. Expression of cyclin E1 and p16 was confirmed by Western blot. For in vivo models, T-47D-luc-BSD cells were passaged twice through mice to select for more favorable growth kinetics (named xxT47D-luc-BSD, WuXi AppTec). xxT47D-luc-BSD cells were used for stable overexpression cell line generation as above.
CDKN2A (p16) insert sequence:
ATGGATTACAAGGATGACGACGATAAGGAACCTGCCGCCGGAAGCAGCATGGAACCCAGCGCCGATTGGCTGGCTACAGCCGCTGCTAGGGGCAGAGTGGAAGAGGTGCGGGCCCTGCTGGAAGCCGGAGCTCTGCCTAACGCCCCTAACAGCTACGGCCGCAGACCTATCCAGGTGATGATGATGGGCTCCGCCAGAGTGGCCGAGCTGCTGCTGCTCCACGGCGCCGAGCCCAACTGCGCCGACCCCGCCACACTGACCCGGCCAGTTCATGATGCCGCCAGAGAGGGCTTCCTGGACACCCTGGTGGTGCTGCACCGGGCCGGCGCCCGGCTGGATGTGCGGGACGCCTGGGGCAGACTGCCAGTGGACCTGGCCGAGGAGCTGGGCCACAGAGATGTCGCCAGATACCTGAGAGCCGCTGCAGGCGGCACCAGAGGCTCTAATCACGCCAGAATCGACGCCGCTGAGGGACCTAGC GACATCCCTGACTGA
CCNE1 insert sequence:
GAATTCATGCCGAGGGAGCGCAGGGAGCGGGATGCGAAGGAGCGGGACACCATGAAGGAGGACGGCGGCGCGGAGTTCTCGGCTCGCTCCAGGAAGAGGAAGGCAAACGTGACCGTTTTTTTGCAGGATCCAGATGAAGAAATGGCCAAAATCGACAGGACGGCGAGGGACCAGTGTGGGAGCCAGCCTTGGGACAATAATGCAGTCTGTGCAGACCCCTGCTCCCTGATCCCCACACCTGACAAAGAAGATGATGACCGGGTTTACCCAAACTCAACGTGCAAGCCTCGGATTATTGCACCATCCAGAGGCTCCCCGCTGCCTGTACTGAGCTGGGCAAATAGAGAGGAAGTCTGGAAAATCATGTTAAACAAGGAAAAGACATACTTAAGGGATCAGCACTTTCTTGAGCAACACCCTCTTCTGCAGCCAAAAATGCGAGCAATTCTTCTGGATTGGTTAATGGAGGTGTGTGAAGTCTATAAACTTCACAGGGAGACCTTTTACTTGGCACAAGATTTCTTTGACCGGTATATGGCGACACAAGAAAATGTTGTAAAAACTCTTTTACAGCTTATTGGGATTTCATCTTTATTTATTGCAGCCAAACTTGAGGAAATCTATCCTCCAAAGTTGCACCAGTTTGCGTATGTGACAGATGGAGCTTGTTCAGGAGATGAAATTCTCACCATGGAATTAATGATTATGAAGGCCCTTAAGTGGCGTTTAAGTCCCCTGACTATTGTGTCCTGGCTGAATGTATACATGCAGGTTGCATATCTAAATGACTTACATGAAGTGCTACTGCCGCAGTATCCCCAGCAAATCTTTATACAGATTGCAGAGCTGTTGGATCTCTGTGTCCTGGATGTTGACTGCCTTGAATTTCCTTATGGTATACTTGCTGCTTCGGCCTTGTATCATTTCTCGTCATCTGAATTGATGCAAAAGGTTTCAGGGTATCAGTGGTGCGACATAGAGAACTGTGTCAAGTGGATGGTTCCATTTGCCATGGTTATAAGGGAGACGGGGAGCTCAAAACTGAAGCACTTCAGGGGCGTCGCTGATGAAGATGCACACAACATACAGACCCACAGAGACAGCTTGGATTTGCTGGACAAAGCCCGAGCAAAGAAAGCCATGTTGTCTGAACAAAATAGGGCTTCTCCTCTCCCCAGTGGGCTCCTCACCCCGCCACAGAGCGGTAAGAAGCAGAGCAGCGGGCCGGAAATGGCGGATTACAAGGATGACGACGATAAGTGATCTAGA.
Western blot
Lysates were prepared using PhosphoSafe Lysis Buffer (EMD Millipore, 71296), and Western blots were performed using standard methods. Blots were cut to target molecular weight prior to antibody hybridization and detection. Antibodies as follows: Rb1:1000 (CST 9309; RRID:AB_823629), pRbS780 1:1000 (CST 9307; RRID:AB_330015), pRbS807/811 1:1000 (CST 8516; RRID:AB_11178658), pRbT821/826 1:500 (SC-271930; RRID:AB_10655683), CDK2 1:1000 (SC-6248; RRID:AB_627238), CDK4 1:1000 (CST 12790; RRID:AB_2631166), CDK6 1:1000 (CST 13331; RRID:AB_2721897), Cyclin A2 1:1000 (CST 4656; RRID:AB_2071958), Cyclin D1 1:1000 (CST 2922; RRID:AB_2228523), Cyclin D2 1:1000 (CST 3741; RRID:AB_2070685), Cyclin D3 1:1000 (CST 2936; RRID:AB_2070801), Cyclin E1 1:1000 (CST 20808;) RRID:AB_2783554), p16 1:1000 (CST 80772; RRID:AB_2799960), p18 1:1000 (CST-2896; RRID:AB_331203), p21 1:1000 (CST 2947; RRID:AB_823586), p27 1:1000 (CST 3686; RRID:AB_2077850), TK1 1:1000 (CST 28755; RRID:AB_3668643), pNPM1 Thr199 1:1000 (CST 3541; RRID:AB_331497), pNCL1 1:500 (ab155977; RRID:AB_3075493), Estrogen Receptor 1 Alpha 1:1000 (CST-8644; RRID:AB_2617128), HER2 1:1000 (CST 2165; RRID:AB_10692490), HSP90 1:500 (SC-13119; RRID:AB_675659), CDC37 1:500 (SC-13129; RRID:AB_627232), FGFR1 1:1000 (CST 9740; RRID:AB_11178519), FGFR2 1:1000 (CST 23328; RRID:AB_2798862), FGFR3 1:1000 (CST 4574; RRID:AB_2246903), Aurora A/AIK 1:1000 (CST 4718; RRID:AB_2061482), Actin (R) 1:2000 (CST 8457; RRID:AB_10950489), Actin (M) 1:5000 (CST 3700; RRID:AB_2242334), GAPDH 1:2000 (CST 2118; RRID:AB_561053). Secondary antibodies (LI-COR 926-33212, RRID:AB_621847; 926-33213, RRID:AB_621848; 926-68070, RRID:_AB_10956588; 926-68071, RRID:AB_10956166; 926-68076, RRID:AB_10956590 were used at 1:10,000 and signals were quantified using LI-COR Odyssey CLx Imaging Studio (RRID:SCR_014579). In Fig. 3d, pRbS807 (rabbit) was probed and detected first, then reblocked before reprobing and detecting total Rb (mouse).
Cell cycle profiling
Cells at ~65% confluency were treated with BLU-222 or palbociclib for 22 hours and pulsed with 10 μM EdU for 2 h. For knockdown experiments, cells were transfected with 25 nM ON-TARGETplus Non-targeting Control Pool or ON-TARGETplus Human RB1 siRNA SMARTpool (Horizon Discovery D-001810-10/L-003296-02) 48 hours prior to cell cycle profiling. Cell cycle phase was detected by flow cytometry using the Click-it EdU Alexa Fluor 488 Flow Cytometry Assay Kit or Alexa Fluor 647 kit (Invitrogen C10420/C10634) following manufacturer instructions. Fluorescence-activated cell sorting (FACS) analysis was performed using LSRFortessa (BD Biosciences) and analyzed using FlowJo v10.8.1 software (BD Life Sciences) (RRID:SCR_008520). Data represent at least two biological replicates.
Generation of CDK4/6i-resistant cell lines
MCF-7 and T-47D cell lines were grown in the presence of chronic, increasing doses of ribociclib, abemaciclib, or atirmociclib over a period of ~6 months. Once confluent, cells were split, and dosing was increased up to 1 μM. Resistance to CDK4/6i was confirmed by CyQUANT Direct Cell Proliferation Assay, and markers of CDK4/6i resistance were assessed by Western blot.
KIYA-PREDICT ex vivo spheroid proliferation assay
The KIYA-PREDICT Assay was performed as previously described39 using breast XPDX tumors from XenoSTART (San Antonio, TX; Supplementary Data S5). Briefly, live cryopreserved PDX tissue was thawed, dissociated into single cells, and plated into 384-well spheroid microplates for 24 h. Spheroids were treated with test compounds and controls for 96 h, and viability was assessed by CellTiter-Glo 3D (Promega, G9683). BLU-222 (0.015–100 µM, 1:3 dilution) and ribociclib hydrochloride (0.41–100 µM, 1:3 dilution) were dosed as single agents and in a 9 × 6 combination matrix. Brightfield imaging was performed on all wells with an EVOS M7000 (Thermo Fisher Scientific) prior to CellTiter-Glo 3D. Cell viability at each concentration or combination was calculated as the average of seven replicates and was normalized to untreated controls. IC50 values were calculated in GraphPad Prism (RRID:SCR_00278) for single agents and in combination at equi-doses for BLU-222 + ribociclib hydrochloride.
Biomarker signature model construction
Baseline transcriptomic data from 26 models were used to determine a predictive biomarker signature for response to treatment. A 70/30 random split was performed to separate the PDX model data into training and independent testing sets. The training data set was fitted to a generalized linear regression model with 10-fold cross-validation and tested on the 30% reserved for verification.
To evaluate the prediction model, we used the confusion matrix to determine accuracy, sensitivity, specificity, PPV, and negative predictive value. We compared across these metrics to align on the best-performing model.
Differential gene expression analysis of on-treatment PDX tumors
At study end, PDX tumors were collected at 2 or 8 h post-final dose and flash frozen. Tumor fragments from three tumors per treatment group were used for RNA-Seq analysis. Library preparation and mRNA sequencing were performed at GENEWIZ (Azenta Life Sciences, RRID:SCR_003177) following the recommended protocols. Read alignment to a Homo sapiens (hg38) and Mus musculus (mm10) hybrid reference genome, and read quantification were performed using STAR (v2.7.10b) (RRID:SCR_004463). Differential gene expression analysis between each treatment group and DMSO was performed with DESeq2 (RRID:SCR_015687) using reads mapped to the human genome (hg38).
BLU-222 phase I/II clinical trial
An international, open-label, nonrandomized, phase I/II, first-in-human, dose-escalation and dose-expansion study of BLU-222 as monotherapy in patients with advanced solid tumors or in combination with ribociclib and fulvestrant in patients with HR+/HER2− breast cancer after progression on at least one prior CDK4/6i (ClinicalTrials.gov. https://clinicaltrials.gov/study/NCT05252416 [2024]) was initiated to evaluate the safety, tolerability, pharmacokinetics (PK), pharmacodynamics (PD), and anticancer activity of BLU-222. To define the safety and maximum-tolerated dose of BLU-222, all statistical analyses of safety, PK, and PD data were descriptive in nature. The study was reviewed and approved by the institutional review board at each clinical site. Written informed consent was obtained from all patients before study entry. Key eligibility criteria for the VELA study included adult patients (aged ≥ 18 years) with advanced solid tumors progressed following standard-of-care treatment, including metastatic HR+/HER2− breast cancer; Eastern Cooperative Oncology Group performance status of 0–2; and adequate bone marrow, hepatic, renal, and cardiac functions. BLU-222 was administered orally, BID, on a 4-week cycle. Adverse events per Common Terminology Criteria for Adverse Events version 5.0, PK, PD, and radiographic response per Response Evaluation Criteria in Solid Tumors, version 1.1, were assessed. The cases presented from the study are preliminary and represent a cutoff of July 26, 2024.
Animal studies
In vivo mouse studies were performed at MDACC TRACTION, XenoSTART (RRID:SCR_026236), Wuxi AppTec (RRID:SCR_026235), and Shanghai ChemPartner (RRID:SCR_026139) according to the protocols approved by the Institutional Animal Care and Use Committee at each institution. Throughout the study period, animals were monitored for irregular behavior (e.g., changes in mobility, food and water consumption, body weight, eye/hair matting, or any other abnormal condition), and any mortality and/or abnormal clinical signs were recorded. BLU-222 was prepared as a homogeneous suspension in 0.5% (w/v) carboxymethylcellulose/1% Tween-80 in water, dosed BID (12 h/12 h) orally; ribociclib was prepared in 0.5% methylcellulose, dosed QD orally, and fulvestrant was prepared in 90% castor oil, 10% ethanol, dosed once weekly subcutaneously. Ribociclib was dosed 30 minutes prior to BLU-222 administration. TGI was calculated using tumor volume (TV) on the latest day at which vehicle and/or treatment groups remained intact using the formula: TGI (%) = (1−(TVTreatment/Dn−TVTreatment/D0)/(TVControl/Dn–TVControl/D0))×100%. One-way analysis of variance was performed to compare tumor volumes among vehicle and treatment groups. No animals were excluded from analysis. PDX model receptor subtypes were confirmed by IHC. PDX model information can be found in Supplementary Data S6–S8.
CDX studies at Shanghai ChemPartner and WuXi AppTec
Cell lines (HCC1569 [Shanghai ChemPartner], MCF7, xMCF7 PalbociclibR, T47D EV + EV, T47D CCNE1 + EV, T47D CDKN2A + EV (WuXi AppTec) were maintained in appropriate media (Supplementary Data S1) and 10 × 106 cells were implanted 1:1 with Matrigel (Corning 354248) into the flanks of 6- to 8-week-old female mice (NOD SCID mice for HCC1569, BALB/c nude for all others). Except for HCC1569, 3 days prior to inoculation, 17B-estradiol (E2) pellets (0.36 mg/60-day release for MCF7 tumors, 0.18 mg/90-day release for T47D tumors) were implanted into the left back of each mouse. Tumor size was measured twice weekly using a caliper, and volume was calculated using the formula V = 0.5 × [shortest diameter] × [longest diameter]2. Dosing started when tumors reached 150–200 mm3. BLU-222 (10 mg/mL) was dissolved in 0.5% (w/v) carboxymethylcellulose, 1% (v/v) Tween-80 to make a homogeneous suspension and dosed BID (12 h/12 h), orally. Ribociclib (7.5 mg/mL) was dissolved in 0.5% methylcellulose to make a homogeneous solution, dosed QD, orally. For combination BLU-222 + ribociclib treatment groups, BLU-222 was administered 60 min prior to ribociclib. Throughout the study, mice implanted with E2 pellets were gently pressed on their bladders to promote daily urination.
Blood for plasma PK analysis was collected by serial microsampling (~100 or ~200 μL blood volume) at the indicated time points. Blood was collected into K2-EDTA tubes and centrifuged at 2000–2500×g, for 5 min at 4 °C. Tumors were collected at the indicated time points and snap-frozen. Tumor PD was assessed by Western blot from snap-frozen tumors.
In vivo PDX studies at XenoSTART
Models were selected using patient treatment history and XPDX response to CDK4/6i (Supplementary Data S1). Tumor fragments (~70 mg) were subcutaneously implanted into the flanks of 6- to 12-week-old female athymic nude, outbred homozygous (Crl:NU(NCr)-Foxn1nu) mice. Mice were given free access to estrogen-containing water throughout the study period. Tumors and body weights were measured twice weekly. Mice were randomized into treatment groups when the mean tumor volume (TV) reached 150–300 mm3 and were treated for 60 days or until tumors reached terminal TV (2000 mm3).
Whole blood for plasma PK analysis was collected at the indicated time points into K2-EDTA tubes, centrifuged at 1700 RCF for 10 min at 4 °C, then flash frozen and stored at −80 °C. Bioanalytical analysis was performed at Charles River Laboratories.
PDX studies at MDACC TRACTION
At ~8 weeks old, female NSGTM NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ mice (Stock Number 005557; The Jackson Laboratory, Bar Harbor, ME) or athymic nude female mice (Stock Number/Strain Code 490; Charles River Labs) were implanted subcutaneously on the right flank with one fresh tumor fragment (4 mm3). For ER+ models, mice were given free access to water containing 8 µg/mL estrogen until compound dosing initiation. When mean TV reached 150–200 mm3, mice were randomized into treatment groups and treated for at least 28 days or until tumors reached a terminal TV of 2000 mm3. Individual tumors were measured twice per week by caliper, and TV was estimated using the formula: TV (mm3) = 0.5 ×(length × width2).
Whole blood for plasma PK analysis was collected into K2-EDTA tubes, centrifuged at 5000 RCF for 5 min at room temperature, then flash-frozen and stored at −80 °C. Bioanalytical analysis was performed at MD Anderson Cancer Center (MDACC). Tumors were collected for formalin-fixed paraffin-embedding (FFPE) or flash-frozen and stored at −80 °C. MDACC TRACTION PDX models used in the current study are listed in Supplementary Data S5.
PDX studies at MDACC Keyomarsi Group
Breast tumor tissues were obtained from patients during surgery, after written informed consent was obtained under protocols approved by the MDACC Institutional Review Board and in accordance with recognized ethical guidelines. The TNBC PDX models BCX070 and XC5173023 were previously developed40,41. The HR+ breast cancer PDX model PR-4 (originally named T141-004), which progressed on palbociclib, was provided by Dr. Funda Meric-Bernstam’s laboratory at MDACC. The generation of the 3 PDX models PR-1, PR-2, and PR-3, originally named CTG-3277, CTG-3266, and CTG-2308, respectively, was described previously42. List of PDX models and characteristics is in Supplementary Data S6.
In this study, the PDX models were re-established from the liquid nitrogen tumor bank and transplanted into the T4 mammary fat pad of 8-week-old female NSG mice from Jackson Lab (strain #005557). After transplantation, tumor size was monitored, and when the primary tumor outgrowths reached 1000–1500 mm³, 4 × 4 mm2 fragments of the outgrowths were implanted into new hosts (n = 3 per tumor) as secondary passage. As the tumor tissues can stably grow after two passages with our protocol, we considered the PDX line to have been successfully established at that point. During maintenance, HR+ breast cancer PDX models were given drinking water supplemented with 8 μg/mL E2, except for the PR-4 and the two TNBC models, which are independent of E2. The procedures performed for these studies were included in an Institutional Animal Care and Use Committee-approved protocol.
PDX tumor fragments (4 × 4 mm) were transplanted into the T4 mammary fat pad of female NCRNU-F sp/sp nude mice from Taconic Biosciences. Tumors were allowed to grow until they reached 150–200 mm³. HR+ breast cancer PDX models (except PR4 and the TNBC models) were maintained with drinking water supplemented with E2 (8 μg/mL). Once tumors reached an average volume of 200 mm³, mice were randomized to the indicated treatment arms and treated with vehicle, palbociclib (50 mg/kg QD, orally), BLU-222 (60 mg/kg BID, orally), or a combination of palbociclib (50 mg/kg) and BLU-222 (60 mg/kg) plus fulvestrant (2.5 mg/kg, once weekly, subcutaneously). The vehicle for palbociclib and ribociclib is 0.5 w/v% methylcellulose (Fujifilm, Cat#133-17815), the vehicle for BLU-222 is 0.5% carboxymethylcellulose-Na/1% Tween-80, and the vehicle for fulvestrant is 10% DMSO diluted in 90% corn oil. Once treatments started, the E2 supplementation was removed to mimic the effect of ET. The length and width of tumor xenografts were measured by caliper three times per week, and TV was calculated using the formula: volume = (length × (width)²)/2. The mice were euthanized when the average TV in the vehicle-treatment arm reached 1000–1300 mm³. Following euthanasia, all tumors were harvested, random fragments were frozen and stored at −80 °C for RNA/protein assessment, and one piece was fixed in formalin for histopathological and IHC analysis. Palbociclib isethionate and fulvestrant were purchased from MedChemExpress (Palbociclib Cat. No. HY-A0065; fulvestrant Cat. No. HY-13636).
Fine chemicals
BLU-222, BLU-2256, and BLU-1954 were synthesized at Blueprint Medicines Corporation (Cambridge, MA). Ribociclib was sourced from MedChemExpress (HY-15777) or Target Molecular Inc (T61991211441-89-3), and fulvestrant was sourced from BluePoint Laboratories (NDC# 68001-484-86), MedChemExpress (HY-13636), or XIROMED (NDC 70700-284-98). PF-07104091 (HY-137894A), INX-315 (HY-162001), palbociclib (HY-50767), abemaciclib (HY-16297A), RO-3306 (HY-12529), roniciclib (HY-13914), azenosertib (HY-132295), berzosertib (HY-13902) were sourced from MedChemExpress. Atirmociclib (E1495) was sourced from Selleckchem.
Statistical analyses
Statistical tests were performed using R v. 4.3.2 (RRID:SCR_001905). Plots were generated using R v. 4.3.2 or GraphPad Prism v10 (RRID:SCR_00278). For box plots, the center line represents the median value, the lower box limit is the 25th percentile of the distribution, the upper box limit is the 75th percentile of the distribution, and the terminus of each whisker is the minimum or maximum of the distribution. Box plots are overlaid with points representing the sample value. Wilcoxon rank-sum tests were used to assess differences between two groups. The Kruskal–Wallis test was used for multigroup comparisons. Kendall’s tau was used to assess correlation coefficients between continuous variables. All relevant tests are two-sided unless otherwise noted.
Transcriptomic data from the Cancer Cell Line Encyclopedia’s DepMap 23Q4 public release (RRID:SCR_017655)(Broad DepMap. DepMap 22Q4 public. https://figshare.com/articles/dataset/DepMap_22Q4_Public/21637199 [2022]) were used to assess differential gene expression between responders and nonresponders to treatment in cell lines. A moderated t-test implemented through linear models for microarray data (LIMMA) tested these differences.
The Benjamin–Hochberg method of multiple testing correction was performed where applicable.
Data availability
The data generated in this study are available upon request to the corresponding author. The underlying code for this study (and training/validation datasets) is not publicly available but may be made available to qualified researchers upon reasonable request to the corresponding author.
Change history
13 January 2026
Supplementary Data was originally published with the incorrect title. The original article has been corrected.
References
Lin, M., Chen, Y., Jin, Y., Hu, X. & Zhang, J. Comparative overall survival of CDK4/6 inhibitors plus endocrine therapy vs. endocrine therapy alone for hormone receptor-positive, HER2-negative metastatic breast cancer. J. Cancer 11, 7127–7136 (2020).
Nabieva, N. & Fasching, P. A. CDK4/6 inhibitors—overcoming endocrine resistance is the standard in patients with hormone receptor-positive breast cancer. Cancers (Basel) 15, 1763 (2023).
Shah, M., Nunes, M. R. & Stearns, V. CDK4/6 inhibitors: game changers in the management of hormone receptor-positive advanced breast cancer?. Oncology 32, 216–222 (2018).
Asghar, U. S., Kanani, R., Roylance, R. & Mittnacht, S. Systematic review of molecular biomarkers predictive of resistance to CDK4/6 inhibition in metastatic breast cancer. JCO Precis. Oncol. 6, e2100002 (2022).
Stanciu, I. M. et al. Mechanisms of resistance to CDK4/6 inhibitors and predictive biomarkers of response in HR+/HER2-metastatic breast cancer—a review of the literature. Diagnostics 13, 987 (2023).
Turner, N. C. et al. Cyclin E1 expression and palbociclib efficacy in previously treated hormone receptor-positive metastatic breast cancer. J. Clin. Oncol. 37, 1169–1178 (2019).
Al-Qasem, A. J. et al. Co-targeting CDK2 and CDK4/6 overcomes resistance to aromatase and CDK4/6 inhibitors in ER+ breast cancer. NPJ Precis. Oncol. 6, 68 (2022).
Herrera-Abreu, M. T. et al. Early adaptation and acquired resistance to CDK4/6 inhibition in estrogen receptor-positive breast cancer. Cancer Res. 76, 2301–2313 (2016).
Pandey, K. et al. Combined CDK2 and CDK4/6 inhibition overcomes palbociclib resistance in breast cancer by enhancing senescence. Cancers (Basel) 12, 3566 (2020).
Watt, A. C. & Goel, S. Cellular mechanisms underlying response and resistance to CDK4/6 inhibitors in the treatment of hormone receptor-positive breast cancer. Breast Cancer Res. 24, 17 (2022).
Arora, M. et al. Rapid adaptation to CDK2 inhibition exposes intrinsic cell-cycle plasticity. Cell 186, 2628–2643.e21 (2023).
Brown, V. et al. CDK2 inhibition with BLU-222 in combination with ribociclib demonstrates robust antitumor activity in pre-clinical models of CDK4/6 inhibitor-naïve and -resistant HR+/HER2− breast cancer. Cancer Res. 83, P6-10-07 (2023).
Dietrich, C. et al. INX-315, a selective CDK2 inhibitor, induces cell cycle arrest and senescence in solid tumors. Cancer Discov. 14, 446–467 (2024).
Dommer, A. P. et al. Tumor suppressors condition differential responses to the selective CDK2 inhibitor BLU-222. Cancer Res. 85, 1310–1326 (2025).
Kumarasamy, V., Vail, P., Nambiar, R., Witkiewicz, A. K. & Knudsen, E. S. Functional determinants of cell cycle plasticity and sensitivity to CDK4/6 inhibition. Cancer Res. 81, 1347–1360 (2021).
Juric, D. et al. BLU-222, an investigational, oral, potent, and highly selective CDK2 inhibitor (CDK2i), as monotherapy in patients (pts) with advanced solid tumors and in combination with ribociclib (RIBO) and fulvestrant (FUL) in HR+/HER2− breast cancer (BC). J. Clin. Oncol. 42, 1056 (2024).
Kim, S., Leong, A., Kim, M. & Yang, H. W. CDK4/6 initiates Rb inactivation and CDK2 activity coordinates cell-cycle commitment and G1/S transition. Sci. Rep. 12, 16810 (2022).
House, N. C. et al. Profiling the activity of the potent and highly selective CDK2 inhibitor BLU-222 reveals determinants of response in CCNE1-aberrant ovarian and endometrial tumors. Cancer Res. 85, 1297–1309 (2025).
Palafox, M. et al. High p16 expression and heterozygous RB1 loss are biomarkers for CDK4/6 inhibitor resistance in ER(+) breast cancer. Nat. Commun. 13, 5258 (2022).
Tsherniak, A. et al. Defining a cancer dependency map. Cell 170, 564–576e16 (2017).
Finn, R. S. et al. PD 0332991, a selective cyclin D kinase 4/6 inhibitor, preferentially inhibits proliferation of luminal estrogen receptor-positive human breast cancer cell lines in vitro. Breast Cancer Res. 11, R77 (2009).
Shen, C. et al. PF-07104091, a first-in-class CDK2-selective inhibitor for the treatment of HR+/HER2− breast cancer and CCNE1high ovarian cancer. Cancer Res. 84, https://doi.org/10.1158/1538-7445.Am2024-5709 (2024).
Brown, V. E. et al. CDK2 regulates collapsed replication fork repair in CCNE1-amplified ovarian cancer cells via homologous recombination. NAR Cancer 5, zcad039 (2023).
Wilson, D. et al. WO/2023/278326. CDK2 inhibitors. WIPO https://patentscope.wipo.int/search/en/WO2023278326 (2022).
Kumarasamy, V. et al. Discrete vulnerability to pharmacological CDK2 inhibition is governed by heterogeneity of the cancer cell cycle. Nat. Commun. 16, 1476 (2025).
Knudsen, E. S. et al. CDK/cyclin dependencies define extreme cancer cell-cycle heterogeneity and collateral vulnerabilities. Cell Rep. 38, 110448 (2022).
Gerosa, R. et al. Cyclin-dependent kinase 2 (CDK2) inhibitors and others novel CDK inhibitors (CDKi) in breast cancer: clinical trials, current impact, and future directions. Crit. Rev. Oncol. Hematol. 196, 104324 (2024).
Asghar, U. S. et al. Single-cell dynamics determines response to CDK4/6 inhibition in triple-negative breast cancer. Clin. Cancer Res. 23, 5561–5572 (2017).
Saleh, L., Wilson, C. & Holen, I. CDK4/6 inhibitors: a potential therapeutic approach for triple negative breast cancer. MedComm (2020) 2, 514–530 (2021).
Luo, L. et al. Anti-tumor activity of CDK2 inhibitor BLU-222 in combination with CDK4/6 inhibitors for overcoming resistance in HR positive and triple negative metastatic breast cancers models. Cancer Res 84, 4622 (2024).
Pu, D. et al. Efficacy of CDK4/6 inhibitors combined with endocrine therapy in HR+/HER2− breast cancer: an umbrella review. J. Cancer Res. Clin. Oncol. 150, 16 (2024).
Eisenhauer, E. A. et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur. J. Cancer 45, 228–247 (2009).
Fassl, A., Geng, Y. & Sicinski, P. CDK4 and CDK6 kinases: from basic science to cancer therapy. Science 375, eabc1495 (2022).
Mai, N. et al. Predictors of response to CDK4/6i retrial after prior CDK4/6i failure in ER+ metastatic breast cancer. NPJ Breast Cancer 10, 92 (2024).
House, I., Valore-Caplan, M., Maris, E. & Falchook, G. S. Cyclin dependent kinase 2 (CDK2) inhibitors in oncology clinical trials: a review. J. Immunother. Precis. Oncol. 8, 47–54 (2025).
Kwiatkowski, N. et al. CDK2 heterobifunctional degraders co-degrade CDK2 and cyclin E resulting in efficacy in CCNE1-amplified and overexpressed cancers. Cell Chem. Biol. 32, 556–569.e24 (2025).
Palmer, C. L. et al. CDK4 selective inhibition improves preclinical anti-tumor efficacy and safety. Cancer Cell 43, 464–481.e14 (2025).
Ianevski, A., Giri, A. K. & Aittokallio, T. SynergyFinder 3.0: an interactive analysis and consensus interpretation of multi-drug synergies across multiple samples. Nucleic Acids Res. 50, W739–W743 (2022).
Shuford, S. et al. Prospective validation of an ex vivo, patient-derived 3D spheroid model for response predictions in newly diagnosed ovarian cancer. Sci. Rep. 9, 11153 (2019).
Chen, X. et al. Cyclin E overexpression sensitizes triple-negative breast cancer to Wee1 kinase inhibition. Clin. Cancer Res. 24, 6594–6610 (2018).
Evans, K. W. et al. A population of heterogeneous breast cancer patient-derived xenografts demonstrate broad activity of PARP inhibitor in BRCA1/2 wild-type tumors. Clin. Cancer Res. 23, 6468–6477 (2017).
Navarro-Yepes, J. et al. Abemaciclib is effective in palbociclib-resistant hormone receptor-positive metastatic breast cancers. Cancer Res. 83, 3264–3283 (2023).
Acknowledgements
This study was funded by Blueprint Medicines Corporation (Cambridge, MA). We would like to thank and acknowledge Marta Murcia (Blueprint Medicines Corporation) and Christopher Vellano (MDACC TRACTION) for coordinating and managing the research collaboration between Blueprint and MDACC. Thank you to Yoon Jong Choi (Blueprint Medicines Corporation employee at the time of work) for her contribution to the discovery of BLU-222. Thank you to Chia-Cheng Li for summarizing the clinical methods, Karen Ho for cell cycle analysis and tumor processing, Kevin Keane for IHC image review, and Jian Guo for PK support (Blueprint Medicines Corporation). Thank you to Mariah Williams, Vanessa Chandler, Alejandro Hernandez Martinez, and Sanket Awate for PDX IHC processing and analysis (MDACC TRACTION). Thank you to Xiaoyan Ma and Angela Harris for PDX in vivo study support (MDACC TRACTION). Patient CT scans provided by Manish Patel (Florida Cancer Specialists) and Linda Duska (University of Virginia). Medical writing and editorial support were provided by Yvonna Fisher-Jeffes, Ph.D. (Blueprint Medicines Corporation) and Srividya Venkitachalam, Ph.D. (Healthcare Consultancy Group) with funding from Blueprint Medicines Corporation, Cambridge, MA.
Author information
Authors and Affiliations
Contributions
Conceptualization: N.C.H., M.M.C., V.B., S.R., M.L.R., L.B.M., and K.L.F. Data curation: M.M.C., FW., C.A.B., and L.B.M. Formal analysis: M.M.C,. and L.B.M. Funding acquisition: n/a. Investigation: N.C.H., M.M.C., V.B., P.R., S.M., L.Y., R.W., F.W., L.L., and N.F. Methodology: N.C.H., M.M.C., S.K., V.B., D.H.P., S.M., L.Y., R.W., F.W., L.L., N.F., C.A.B., T.A.Y., K.K., J.R.M., M.L.R., L.B.M., and K.L.F. Project administration: N.C.H., S.K., D.H.P., N.F., C.A.B., T.A.Y., K.K., J.R.M., S.R., M.L.R., L.B.M., K.L.F. Resources: n/a. Software: M.M.C., F.W., C.A.B., and L.B.M. Supervision: N.C.H., S.K., D.H.P., N.F., C.A.B., K.K., J.R.M., L.B.M., and K.L.F. Validation: N.C.H., S.K., V.B., S.M., L.Y., and L.L. Visualization: N.C.H., M.M.C., S.K., L.B.M., and K.L.F. Writing—original draft: N.C.H., M.M.C., R.W., M.L.R., L.B.M., and K.L.F. Writing—review and editing: N.C.H., M.M.C., S.K., V.B., D.H.P., S.M., L.Y., R.W., F.W., L.L., N.F., C.A.B., T.A.Y., K.K., J.R.M., S.R., M.L.R., L.B.M., and K.L.F. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
N.C.H.: Employee of Blueprint Medicines Corporation. Stockholder in Blueprint Medicines Corporation. M.M.C.: Employee of Blueprint Medicines Corporation. Stockholder in Blueprint Medicines Corporation. S.K.: No conflicts of interest. V.B.: Employee of Blueprint Medicines Corporation at the time of work. P.R.: Employee of Blueprint Medicines Corporation. Stockholder in Blueprint Medicines Corporation. D.H.P.: No conflicts of interest. S.M.: Employee of Blueprint Medicines Corporation. Stockholder in Blueprint Medicines Corporation. L.Y.: Employee of Blueprint Medicines Corporation. Stockholder in Blueprint Medicines Corporation. R.W.: Employee of Blueprint Medicines Corporation. Stockholder in Blueprint Medicines Corporation. F.W.: No conflicts of interest. L.L.: No conflicts of interest. N.F.: No conflicts of interest. C.A.B.: No conflicts of interest. T.A.Y.: Employee of University of Texas MD Anderson Cancer Center, where I am Vice President, Head of Clinical Development in the Therapeutics Discovery Division, which has a commercial interest in DDR and other inhibitors (IACS30380/ART0380 was licensed to Artios). Consultant for AbbVie, Acrivon, Adagene, Aduro, Almac, Amgen Inc., Amphista, Artios, Astex, AstraZeneca, Athena, Atrin, Avenzo, Avoro, Axiom, Baptist Health Systems, Bayer, BeiGene, BioCity Pharma, Blueprint Medicines Corporation, Boxer, BridGene Biosciences, Bristol Myers Squibb, C4 Therapeutics, Calithera, Cancer Research UK, Carrick Therapeutics, Circle Pharma, Clovis, Cybrexa, Daiichi Sankyo, Dark Blue Therapeutics, Debiopharm, Diffusion, Duke Street Bio, 858 Therapeutics, EcoR1 Capital, Ellipses Pharma, EMD Serono, Entos, FoRx Therapeutics AG, F-Star, Genesis Therapeutics, Genmab, Glenmark, GLG, Globe Life Sciences, Grey Wolf Therapeutics, GSK, Guidepoint, Ideaya Biosciences, Idience, Ignyta, I-Mab, ImmuneSensor, Impact Therapeutics, Institut Gustave Roussy, Intellisphere, Jansen, Joint Scientific Committee for Phase I Trials in Hong Kong, Kyn, Kyowa Kirin, Lumanity, MEI Pharma, Mereo, Merck, Merit, Monte Rosa Therapeutics, Natera, Nested Therapeutics, Nexys, Nimbus, Novocure, Odyssey Therapeutics, OHSU, OncoSec, Ono Pharma, Onxeo, PanAngium Therapeutics, Pegascy, PER, Pfizer, Piper-Sandler, Pliant Therapeutics, Prelude Therapeutics, Prolynx, Protai Bio, Radiopharma Theranostics, Repare, resTORbio, Roche, Ryvu Therapeutics, SAKK, Sanofi, Schrodinger, Servier, Synnovation, Synthis Therapeutics, Tango, TCG Crossover, TD2, Terremoto Biosciences, Tessellate Bio, Theragnostics, Terns Pharmaceuticals, Thryv Therapeutics, Tolremo, Tome, Trevarx Biomedical, Varian, Veeva, Versant, Vibliome, Voronoi Inc, Xinthera, Zai Labs and ZielBio. Grant/research support from Artios, AstraZeneca, Bayer, BeiGene, BioNTech, Blueprint, BMS, Boundless Bio, Clovis, Constellation, CPRIT, Cyteir, Department of Defense, Eli Lilly, EMD Serono, Exelixis, Forbius, F-Star, GlaxoSmithKline, Genentech, Gilead, Golfers Against Cancer, Haihe, Ideaya, ImmuneSensor, Insilico Medicine, Ionis, Ipsen, Jounce, Karyopharm, KSQ, Kyowa, Merck, Mirati, Novartis, NIH/NCI, Pfizer, Pliant, Prelude, Ribon Therapeutics, Regeneron, Repare, Roche, Rubius, Sanofi, Scholar Rock, Seattle Genetics, Synnovation, Tango, Tesaro, V Foundation, Vivace, Zenith and Zentalis. Stockholder in Seagen. K.K.: Funding from Apeiron, Blueprint Medicines Corporation, Repare, Schrodinger, and Novartis. J.R.M.: No conflicts of interest. S.R.: Employee of Blueprint Medicines Corporation. Stockholder in Blueprint Medicines Corporation. M.L.R.: Employee of Blueprint Medicines Corporation. Stockholder in Blueprint Medicines Corporation. L.M.: Employee of Blueprint Medicines Corporation. Stockholder in Blueprint Medicines Corporation. K.F.: Employee of Blueprint Medicines Corporation. Stockholder in Blueprint Medicines Corporation.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
House, N.C., Chen, M.M., Khazaei, S. et al. CDK2 inhibition enhances CDK4/6 inhibitor antitumor activity in comprehensive breast cancer PDX model screen. npj Breast Cancer 11, 135 (2025). https://doi.org/10.1038/s41523-025-00851-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41523-025-00851-7