Abstract
This study aimed to evaluate the representativeness of HER2-low and ultralow status in core needle biopsies (CNB) with AI assistance and investigate the clinicopathological and molecular characteristics of HER2-ultralow, HER2-null, and HER2-low breast cancer patients in the Chinese population. CNB-surgical excision biopsy (SEB) concordance was high between HER2-null and ultralow/low groups but limited in the ultralow subgroup. Univariate analyses showed that TNBC subtype, HR negative expression, high Ki-67 index, and AR negative expression were potential indicative factors for the discordance of HER2 status between CNB and SEB. Clinicopathological features showed no significant differences between low and ultralow groups. Genomic analysis revealed subtle mutation pattern differences. The HER2 copy number level did not show additional value in distinguishing subgroups. Conclusions highlight the need to address CNB-SEB diagnostic discordance, particularly in ultralow cases, emphasizing the necessity of HER2 retesting in surgical specimens.
Similar content being viewed by others
Introduction
Breast cancer is a heterogeneous disease, and the status of human epidermal growth factor receptor 2 (HER2) is crucial in determining the appropriate treatment strategy. HER2 status is typically determined using immunohistochemistry (IHC) and in situ hybridization (ISH) techniques. The groundbreaking data of DESTINY-Breast 04 trial on Trastuzumab deruxtecan (T-DXd), an antibody-drug conjugates (ADCs) of trastuzumab, in HER2-low patients underscores the importance of distinguishing between HER2 immunohistochemistry (IHC) scores of 0 and HER2-low(IHC scores of 1+ or 2+ and have a negative ISH testing result)1. Up to 60% of HER2-negative breast cancers are now classified as HER2-low2.
The DAISY trial reported that modest anti-tumor activity was also observed in patients with HER2 IHC 03. HER2 0 can be divided into HER2-null and HER2-ultralow. HER2-null is defined as the absence of staining, while HER2-ultralow is characterized by faint or barely perceptible and incomplete staining in <10% of tumor cells4,5. The DESTINY-Breast06 trial suggested the efficacy of ADCs in patients with HER2-ultralow breast cancer, and the current data suggest that there is no need to discriminate between HER2-low and HER2-ultralow disease due to the consistent benefit in both populations6. The potential to target low levels HER2 expression in clinical practice has increased the need for a more accurate classification of low levels HER2 protein expression, especially distinguishing between scores of null and ultralow/low.
However, in the real world, the accuracy of scoring HER2 IHC in the low range (0 and 1 + ) is poor. On one hand, there is subjectivity in HER2 IHC staining scores and differences among and within observers7. On the other hand, there is strong intra-tumoral heterogeneity in HER2-low expression tumors, which may lead to poor consistency in HER2 interpretation. Artificial intelligence (AI) based image analysis tools can recognize tumor cells by machine-learning HER2 immunohistochemical images, evaluate the staining pattern of tumor cells, and analyze the expression level of HER2 in tumor cells by measuring the staining intensity, completeness of the cell membrane, and positive proportion, thus assisting in the interpretation of HER2 results and generating reliable, reproducible results8.
Core needle biopsy (CNB) is a commonly used method for initial diagnosis and HER2 assessment in breast cancer. Evaluating HER2 status in preoperative specimens is crucial for neoadjuvant therapy selection and is the only sample for assessing tumors with complete pathological response or in metastatic patients ineligible for surgery. However, previous studies have shown variable consistency rates of HER2 status between CNBs and surgically excised specimens9,10,11,12. In Rossi C’s study, the concordance between biopsy and surgical specimen for HER2 was 68%, and the concordance was especially low (37%) in the HER2 1 + category10. Chen R’s study demonstrated that the diagnostic value of CNB for determining HER2 status in breast cancer was limited, especially in the HER2-low population13. The diagnostic value of CNB in determining HER2-null, HER2-ultralow and HER2-low status have not been adequately investigated. It is urgent for us to further verify the concordance between CNB and surgical excision biopsy (SEB), especially to evaluate the diagnostic value of CNB among patients with HER2-ultralow and HER2-low breast cancer. A more precise HER2 status assessment, combined with comprehensive characterization of the clinical and molecular features of HER2-null and HER2-ultralow, is required.
The aims of this study were to evaluate whether the HER2-low and ultralow status in CNB can represent the whole tumor with the assistance of AI and to investigate the clinicopathological features and molecular characteristics of patients with HER2-ultralow, HER2-null, and HER2-low expression in the Chinese breast cancer population.
Results
Concordance between historical and re-scoring results of CNB
Regarding the historical results of CNB, 84 cases were HER2 0, 398 cases were HER2 1 + , 24 cases were HER2 2 + . After re-scoring, 54 cases were classified as HER2-null, 12 as HER2-ultralow and 440 as HER2-low (HER2 1 + , Fig. 1). Among the 84 HER2 0 cases, 54 cases were re-classified as HER2-null, 9 as HER2-ultralow and 21 as HER2-low. Among the 398 HER2 1+ cases, none were re-classified as HER2-null, 3 as HER2-ultralow and 395 as HER2-low. The concordance of HER2 0 and HER2-low between historical and re-scoring results was 95.26% (Kappa = 0.678, p < 0.001). This indicates a relatively high agreement between the initial scoring and the pathologists’ re-scoring. When analyzed by AI, 32 were HER2-ultralow and 474 were HER2-low (Fig. 1). Among the 54 HER2-null cases determined by manual re-scoring, 21 were classified as HER2-ultralow and 33 as HER2-low. Among the 12 HER2-ultralow cases determined by manual re-scoring, 4 were classified as HER2-ultralow and 8 as HER2-low. Among the 440 HER2-low cases determined by manual re-scoring, 7 were classified as HER2-ultralow and 433 as HER2-low. The concordance of HER2 0 and HER2-low between the AI and manual re-scoring results was 90.51% (Kappa = 0.465, p < 0.001). The concordance of IHC null, ultralow and low between AI and manual re-scoring results was 86.36% (Kappa = 0.029, p < 0.001). In 32 HER2-ultralow cases identified by AI, the percentage of positive tumor cells was slightly lower in patients in HER2-null cases than HER2-ultralow and low patients determined by manual re-scoring (Fig. S1).
Sankey diagrams demonstrating concordance of HER2 status among historical score, manual re-scoring and AI based score in core needle biopsy (CNB).
These results suggest that the AI algorithm shows a good concordance with manual scoring, although some differences still exist. Additionally, with the assistance of AI, it was found that all HER2-null slides exhibited a small amount of faint HER2 membrane staining and were re-classified as ultralow by AI. This indicates that quantitative analysis based on AI may be more sensitive in detecting this subgroup.
Concordance between CNB and SEB
Based on the manual re-score of SEB, 89(17.59%) cases were classified as HER2-null, 51(10.08%) cases as HER2-ultralow and 366(72.34%) cases as HER2-low. The discordance rate (for HER2-null, ultralow and low) between CNB and SEB was 23.71%. Among the 12 HER2-ultralow cases in CNB, 34% changed to HER2-null and 33% changed to HER2-low in SEB. Among the 54 HER2-null cases in CNB, 4 cases changed to HER2-ultralow and 14 changed to HER2-low in SEB. Among the 440 HER2-low cases in CNB, 44 cases changed to HER2-ultralow and 49 changed to HER2-null in SEB (Fig. 2A).
A Sankey diagrams depicting the changes in HER2 manual re-score from core needle biopsy (CNB) to surgical excision specimens (SEB). B Sankey diagrams depicting the changes in HER2 AI-based score from core needle biopsy (CNB) to surgical excision specimens (SEB).
The discordance rate (for HER2 0 and low) between manual re-scoring CNB and SEB was 22.13% (Kappa=0.208, p < 0.001). Regarding the conversion of HER2 status from CNB to SEB, 28.79% (19/66) patients experienced a conversion from HER2-null/ultralow to HER2-low status in the SEB sample (manual scoring). 21.14% (93/440) showed a conversion from HER2-low to HER2-null/ultralow.
The discordance rate (for HER2-null and ultralow/low) between re-scoring CNB and SEB was 14.03% (71/506, Kappa=0.427, p < 0.001).33.33% (18/54) patients experienced a conversion from HER2-null to HER2-low/ultralow status in the SEB sample. 11.73% (53/452) showed a conversion from HER2-low/ultralow to HER2-null.
The discordance rate (for HER2 0 and low) between AI-based CNB and SEB was 24.11% (Kappa = 0.125, p < 0.001). The discordance rate (for HER2-null, ultralow and low) between AI-based CNB and SEB was 28.85%. The discordance rate (for HER2-null and ultralow/low) between AI-based CNB and SEB was 17.59%. Using the AI-based scoring, 21.88% (7/32) patients experienced a conversion from HER2-ultralow to HER2-low status in the SEB sample and 24.26% (115/474) showed a conversion from HER2-low to HER2-null/ultralow (Fig. 2B, Fig. 3A–F). In the 32 HER2-ultralow cases identified by AI in CNB, the percentage of positive tumor cells in CNB slides was slightly lower in patients who converted to HER2-null/ ultralow status than in HER2-low patients (p = 0.31, Fig. 3G). Among the 474 HER2 1+ cases, the percentage of positive tumor cells in CNB slides was lower in patients who converted to HER2-null/ ultralow status than in HER2-low patients (p < 0.001, Fig. 3H).
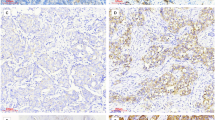
A–F Reprehensive images showing the conversion of HER2 status from CNB to SEB. G Density plot illustrating the percentage of HER2 positive tumor cells in 32 CNB slides with HER2-ultralow status, categorized by the HER2 immunohistochemistry (IHC) status of the corresponding SEB. H Density plot showing the percentage of HER2 - positive tumor cells in 474 CNB slides with HER2-low status, classified according to the HER2 IHC status of the respective SEB. All *p < 0.05, **p < 0.01, ***p < 0.001, n.s. not significant.
Overall, the concordance of HER2-null, ultralow, and low status between CNB and SEB was limited. The agreement could be improved by clustering IHC categories, and the highest concordance between CNB and SEB was observed when comparing HER2-null with the ultralow/low grouped cluster. However, in the HER2-ultralow category, the concordance remained limited.
Clinicopathological characteristics comparison between HER2-null and ultralow groups
There were no significant difference in clinicopathological parameters between HER2-low and ultralow groups. Compared with HER2-null cases, patients with HER2-ultralow expression were more likely to have positive AR expression (p = 0.001). There was no significant difference in other clinicopathological parameters such as age, histological grade, tumor size, lymph node status, and molecular subtype between HER2-null and ultralow groups. Compared with HER2-null cases, patients with HER2-low expression had a lower proportion of grade 3 tumors (p = 0.005), a lower Ki-67 index (p = 0.003), a higher frequency of positive AR expression (p < 0.001) and hormone receptor (HR) positive expression (p < 0.001). Additionally, there was a significant difference in histological types between the HER2-null and low groups (p = 0.011, Table 1).
In the HR positive subgroup (Table S1), no significant difference in clinicopathological parameters were observed between HER2-low and ultralow groups, nor between HER2-null and ultralow groups. However, patients with HER2-low expression were more likely to have positive AR expression than those with HER2-null expression (p = 0.034). In the HR-negative subgroup (Table 2), there were no significant differences in clinicopathological parameters between the HER2-low and ultralow groups. Patients with HER2-low expression had a lower proportion of T3 tumors than those with HER2-null expression (p = 0.024). Compared with HER2-null cases, HER2-ultralow cases were more likely to belong to the luminal androgen receptor (LAR) subgroup (p = 0.007) and to have lymph node metastasis (p = 0.048). In both the HR-positive and HR-negative groups, there were no statistically significant differences in disease-free survival and overall survival among the three groups, probably due to the relatively short follow-up period (Fig. S2A–D).
A total of 71 (14.03%) patients showed discordance between CNB and SEB based on clinical classification (for HER2-null and ultralow/low). Univariate analyses indicated that TNBC subtype, HR negative expression, high Ki-67 index, and AR negative expression were potential indicative factors for the discordance of HER2 status (Table 3).
Overall, there were no significant differences in clinicopathological parameters between the HER2-low and ultralow groups. In TNBC, HER2-ultralow cases were more likely to belong to the LAR subgroup and to have lymph node metastasis than HER2-null cases. These results showed that HER2-ultralow cases demonstrated overlapping clinicopathological characteristics with HER2-low cases.
Genomic landscape of the HER2-null, HER2-ultralow and HER2-low breast cancers
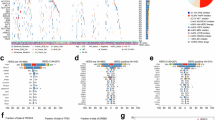
In all 506 cases, the most prevalent mutations were TP53, PIK3CA, MAPK3K1, and GATA3. In HER2-null breast cancer patients, the most commonly mutated genes were TP53 (60.7%), PIK3CA (27.0%), PIK3R1(9.0%) and KMT2D (9.0%). However, in HER2-ultralow cases, the most commonly mutated genes were TP53 (51.0%), PIK3CA (39.2%), MAP3K1(11.8%) and PDGFRB (9.8%). In HER2-low cases, the most commonly mutated genes were PIK3CA (42.1%), TP53 (35.8%), MAP3K1(11.5%) and GATA3 (9.01%). There was some variation in the frequency of the same genes among the three groups. Given the distribution of HR status was different between HER2-null and low cases, we investigated the differences of mutation landscape in HR positive and negative subgroup separately (Fig. 4). In the HR positive subgroup, the mutation frequencies of MAP3K1 (2.0% vs. 14.1%; p = 0.016), KMT2D (12.0% vs. 3.0%; p = 0.012) and KMT2B (10.0% vs. 2.0%; p = 0.012) were significantly different between HER2-null and HER2-low cases. Additionally, the mutation frequency of IRS1 (8.3% vs. 1.7%; p = 0.044) differed significantly between HER2-ultralow and HER2-low cases (Figs. 4A, 5A). To comprehensively understand the functional implications of somatic mutations, we constructed landscapes depicting the altered genes within oncogenic signaling pathways (Fig. S3A). In HER2-ultralow cases, the most commonly mutated oncogenic signaling pathways were PIK3CA/AKT/mTOR (63.9%) TP53 (38.9%), and RTK-RAS (33.3%). In HER2-low cases, the most commonly mutated oncogenic signaling pathways were PIK3CA/AKT/mTOR (63.8%) TP53 (30.5%), and RTK-RAS (27.9%). In HER2-null cases, the most commonly mutated oncogenic signaling pathways were also PIK3CA/AKT/mTOR (52.0%) TP53 (40.0%), and RTK-RAS (36.0%). The distinctions among HER2-low, HER2-ultralow, and HER2-null cases in HR-positive breast cancer were subtle and did not show statistically significant differences.
Molecular landscape of HER2-null, ultralow and low breast cancers in hormone receptor (HR) positive (A) and negative cases (B). Genes marked with * represent those with differential mutation frequencies among HER2-null, HER2-ultralow and HER2-low patients.
A Bar plots showing the differentially mutated somatic genes rate of the top 30 genes within 506 breast cancers. B Density plot displaying the HER2 copy number in 506 CNB slides, grouped by the HER2 IHC status determined by manual scoring in CNB. All *p < 0.05, **p < 0.01, ***p < 0.001, n.s. not significant. C Association between the percentage of positive tumor cells as determined by AI and HER2 copy number per cell quantified by next -generation sequencing (NGS).
In the HR negative subgroup, PIK3R1(17.9% vs. 2.9%; p = 0.011) and SMARCA4(0.0% vs. 11.8%; p = 0.026) showed significantly different frequencies between HER2-null and HER2-low cases, and PREX2 (0.0% vs. 20.0%; p = 0.018) had a significantly different frequency between HER2-ultralow and HER2-null (Figs. 4B, 5A). In HER2-ultralow cases, the most commonly mutated oncogenic signaling pathways were TP53 (86.7%), NOTCH (46.7%) and PIK3CA/AKT/mTOR (46.7%). In HER2-low cases, the most commonly mutated oncogenic signaling pathways were TP53 (73.5%), RTK-RAS (35.3%) and PIK3CA/AKT/mTOR (30.9%). In HER2-null cases, the most commonly mutated oncogenic signaling pathways were TP53 (89.7%), RTK-RAS (53.8%) and PIK3CA/AKT/mTOR (35.9%, Fig. S3B). The distinctions were not statistically different.
We compared the HER2 copy number levels with the IHC scores of the following HER2 subcategories: HER2-null, HER2-ultralow and HER2-low. The HER2 copy number level was significantly higher in HER2-low cases than in the HER2-null group (P < 0.001), but there was substantial overlap in the copy number levels between IHC null and IHC low groups (Fig. 5B). The HER2 copy number level was slightly higher in HER2-low cases than in the HER2-ultralow group (p = 0.870), and it was also slightly higher in HER2-ultralow cases than in the HER2-null group (p = 0.427). The percentage of positive tumor cells in CNB slides was positively correlated with HER2 copy number levels (p < 0.001, Fig. 5C).
Taken together, HER2 copy number level does not appear to provide additional value in distinguishing HER2-null from HER2-low and ultralow tumors. Comparing the frequencies of somatic mutations among HER2-null, HER2-ultralow and HER2-low subgroup across the most frequently mutated genes in these cases, we found that the HER2-null and HER2-low subgroup had moderately different mutation patterns, the HER2-null and HER2-ultralow subgroups had slightly different mutation patterns, and the HER2-low and HER2-ultralow subgroup also had slightly different mutation patterns.
Discussion
The recent research on HER2-low levels breast cancer has expanded the spectrum of HER2 positivity in breast cancer and the concepts of HER2-low and HER2-ultralow have emerged as crucial areas of study. HER2-ultralow is characterized by faint/barely perceptible and incomplete staining in <10% of tumor cells.
The advantages of IHC testing include its wide availability, relatively low cost, easy preservation of stained slides, and the use of a familiar routine microscope14. It should be noted that the current standard HER2 assays (IHC and ISH) were developed to identify patients with HER2 overexpression who could benefit from trastuzumab, and they are likely to be inaccurate in distinguishing low levels of HER2 expression (ultralow and low) from HER2-null. The detection of low levels of HER2 expression by IHC might be affected by several pre-analytical and analytical issues, such as the ischemia time, formalin fixation and artifacts, storage time of wax blocks and white pieces, and technical and biological heterogeneity15. This makes it extremely difficult to accurately identify low levels of HER2 status, whether through manual scoring or AI algorithms. Previous studies have evaluated the interobserver reproducibility of HER2 IHC and demonstrated that the interobserver agreement was limited. Baez-Navarro X’s study indicated that the interobserver agreement could be improved by clustering IHC categories, and the highest reproducibility was observed when comparing IHC no staining with the ultralow/1 + /2+ grouped cluster (74.3% of cases with 100% agreement)16. In this study, we found that the highest concordance between CNB and SEB was observed when comparing HER2-null with the combined ultralow/low clustering group.
An in vitro study established that cells containing 0–20,000, 20,000–100,000, 100,000–500,000, and more than 2,000,000 receptors corresponded to IHC null, IHC ultralow, IHC 1+ or 2 + , and IHC 3 + , respectively. The slight difference in receptor number between IHC null and ultralow may explain the difficulty in distinguishing IHC null from ultralow with the current diagnostic methods. Inaccurate HER2 evaluation can lead to the misassignment of many patients for treatment with T-DXd. There is an urgent need for novel quantitative assays to improve our ability to identify patients with HER2-low breast cancer. Determining the threshold of HER2 expression in the beneficial population is an issue worthy of future attention. The AI algorithm developed in this study has shown potential in assisting with HER2 assessment. Compared with manual scoring, it can provide a more objective and consistent assessment, reducing the impact of interobserver variability. In our study, with the assistance of AI, all HER2-null slides were found to have a small amount of faint HER2 membrane staining and were classified as ultralow by AI, suggesting that AI-based quantitative analysis may be more sensitive in detecting this subgroup. Among the 32 HER2-ultralow cases identified by AI, the percentage of positive tumor cells was lower in patients with HER2-null cases than in those with HER2-ultralow and low cases determined by manual re-scoring.
Developing more sensitive and specific assays would improve the diagnostic accuracy for HER2-ultralow. The findings of Baez-Navarro X et al. indicate a strong agreement between mRNA expression quantified by reverse transcription quantitative polymerase chain reaction (RT-qPCR) and HER2 IHC scores17. However, there were discrepancies in classification. Specifically, 10 of 17 HER2 0/ultralow cases by IHC (58.8%) were classified as HER2-low by MammaTyper, 2 of 71 cases (2.8%) were HER2-low by IHC and HER2 0/ultralow by MammaTyper, and 2 (2.8%) were HER2-low by IHC and HER2-positive by RP-qPCR17. Li X’s study indicated that HER2 RNA levels detected by RNAscope strongly correlated with HER2 protein levels and with HER2 IHC H-scores from the PATHWAY and HercepTest assays. RNAscope results were strongly associated with HER2 protein levels and patients’ response to T-DXd treatment. However, neither protein levels nor RNA levels significantly differed among cases scored 0, ultralow, and 1+ by PATHWAY and HercepTest18. Another study examined whether HER2 mRNA detected through the 21-gene assay, Oncotype DX, could refine the diagnosis of HER2-low and HER2-0, and demonstrated that due to the substantial overlap, HER2 gene expression was unable to properly distinguish HER2-low and HER2-0. In addition, there was no difference in HER2 gene expression between the HER2-null and HER2-ultralow subgroups19.
Moreover, breast tumors are known to be heterogeneous, and different regions of the tumor may have varying HER2 expression levels. CNB samples a relatively small portion of the tumor. This limited sampling may not fully represent the heterogeneity of the entire tumor, and this heterogeneity can lead to differences between CNB and SEB results20. Intertumoral HER2 IHC heterogeneity is more prevalent in HER2-low samples, contributing to inconsistent interpretation of HER2-low results21. It should be noted that a subset of HER2-null tumors in CNB samples might respond to targeted therapy due to intratumor heterogeneity. SEB provides a more comprehensive sample of the tumor, including a larger portion and is more likely to capture the full range of HER2 expression levels within the tumor. In this study, the low concordance between CNB and SEB in the HER2-ultralow category (only 25% in our study) highlights the challenges associated with its diagnosis. The high conversion rates between HER2-ultralow and other statuses (34% to null and 33% to low in our study) further emphasize the instability and ambiguity of this category. Additionally, the spatial and temporal evolution of HER2-low expression also affect the HER2 interpretation22. Shang J’s study investigated the evolution of HER2- low from primary breast cancer to residual breast cancer after neoadjuvant therapy and demonstrated the inconsistency rate of HER2 expression was 21.42% between primary breast cancer and residual tumor after neoadjuvant therapy, mainly manifested as the conversion of HER2-low cases to HER2-0 (10.19%) and the conversion of HER2-0 to HER2-low (6.45%). Therefore, re-detection of HER2 in breast cancer after neoadjuvant therapy should be recommended and may lead to new treatment opportunities for a certain proportion of patients23.
Currently, there is limited research on whether patients with HER2-ultralow breast cancer exhibit distinct clinicopathological features compared to those with HER2-null and HER2-low tumors. Chen Z et al. found that HER2-ultralow BC patients demonstrated distinct clinicopathological features from HER2-null and HER2-low tumors. However, HER2 status (null, ultralow, or low) had no prognostic value in this HER2-negative BC population. HER2-ultralow patients differed from HER2-low patients in terms of N stage, HR status, Ki-67 expression, and type of surgery. There were also significant differences in histologic type and postoperative endocrine therapy between HER2-ultralow and HER2-null patients24. Another study demonstrated that among HR-positive BC, HER2-null has lower Ki-67 expression compared to HER2-ultralow and HER2-low group. Among TNBC, HER2-ultralow has lower Ki-67 expression compared to HER2-low group. There are no other significant differences among these three groups25. In our study, no significant difference in clinicopathological parameters was found between HER2-low and ultralow groups. In all cases, HER2-null and low group differed in terms of histological type, histological grade, Ki-67 index, AR expression and HR expression. In TNBC, HER2-ultralow cases were more likely to belong to the LAR subgroup and to have lymph node metastasis than HER2-null cases. These results showed that HER2-ultralow cases showed overlapping clinicopathological characteristic with HER2-low cases. In both the HR-positive and HR-negative groups, there were no statistically significant differences in disease-free survival and overall survival among the three groups, probably due to the relatively short follow-up period. Analyses of large retrospective cohorts of patients with BC often reported inconsistent findings. Most studies have identified no major difference in clinicopathologic characteristics nor prognosis for HER2-low expression, few studies have instead suggested a better or worse prognosis associated with HER2-low BC2,26. A meta-analysis with a large sample of patients demonstrated that compared to patients with HER2-0 BC, those with HER2-low BC had better OS in the overall and HR-positive populations but similar OS in the HR-negative populations. HER2-low BC had better DFS in the HR-positive populations but similar DFS in the overall and HR-negative populations and the overall populations27. Guan F’s study revealed that the HER2-ultralow expression group displayed lower rates of ER, PR, and AR positivity, as well as fewer Luminal types compared to the HER2-low group, and exhibited a higher likelihood of lymph node metastasis than the HER2-null group28. HER2-low tumors tend to have more mutations in PI3K-AKT signaling pathway-related genes than HER2-0 tumors1,29,30. In our study, the distinctions among HER2-low, HER2-ultralow and HER2-null cases in both HR-positive and negative cases breast cancer were subtle and did not show statistically significant differences. Additionally, comparing the frequency of somatic mutations among the HER2-null, ultralow, and low subgroups across the most frequently mutated genes in these cases showed that the HER2-null and low subgroups had moderately different mutation patterns, the HER2-null and ultralow subgroups had slightly different mutation patterns, and the HER2-low and ultralow subgroups also had slightly different mutation patterns.
We also investigated whether the HER2 copy numbers could improve the discrimination between HER2-null, HER2-ultralow and HER2-low tumors. The HER2 copy number level was significantly higher in HER2-low cases than in the HER2-null group, but showed substantial overlap between IHC null and IHC low groups. We did not observe any statistical differences in HER2 copy number level between HER2-low cases and HER2-ultralow group, or between the HER2-ultralow and HER2-null group. HER2 copy number level does not seem to have additional value to separate HER2-null from HER2-low and ultralow tumors. Lambein K et al. found that the mean HER2 copy number was higher in score 1+ tumors than in score 0 tumors31, which is consistent with our study. In Baez-Navarro X’s study, they also did not observe any statistical differences in the mean HER2 copy number between IHC 0 and 1 + , regardless of the inclusion of the ultralow group16.
This study has several limitations. First, it was based on a single-center retrospective cohort, and larger-scale prospective studies need to be carried out to validate our findings. Second, although AI has shown promising results in this study, the limitations of digital image analysis techniques and other factors can still affect the accuracy of HER2 assessment. Issues such as image quality, resolution, and the ability to accurately detect and classify cells can influence the final results. Specific training and particular attention during HER2 assessment techniques and interpretation to recognize the subtle staining patterns associated with HER2-ultralow will help ensure appropriate identification of HER2-low and HER2-ultralow tumor. However, it should be noticed that even with pretrained pathologists, there may still be some differences in judgment, especially for cases with borderline HER2 staining.
In conclusion, this study demonstrated the limited diagnostic value of CNB for determining HER2-low status and ultralow status in breast cancer. The concordance between CNB and SEB was found to be limited, especially in the HER2-ultralow category. The agreement could be improved by clustering IHC categories, and the highest concordance between CNB and SEB was observed when comparing HER2-null with the ultralow/low grouped cluster. No significant difference in clinicopathological characteristics was observed between HER2-low and ultralow groups. The distinctions of genomic landscape among HER2-low, HER2-ultralow and HER2-null cases in both HR-positive and negative cases breast cancer were subtle. AI-based quantitative analysis may be more sensitive to detect HER2-ultralow. These findings indicate the necessity of retesting HER2 in excision specimens to ensure accurate diagnosis and appropriate treatment decisions. Future studies should focus on improving the accuracy of HER2 assessment in CNB and further exploring the potential of AI in breast cancer diagnosis.
Methods
HER2 AI algorithm development
The D-Path AI platform, an innovative deep-learning framework for membrane segmentation was developed to assess HER2 status in breast cancer CNB specimens. It is a three-phase AI analysis framework consisting of the following steps: (1) segmenting tumor areas while excluding ductal carcinoma in situ (DCIS); (2) detecting and classifying tumor cells based on the membrane staining pattern; and (3) grading HER2 IHC scores according to the 2018 HER2 ASCO/CAP guideline (Fig. S4). This framework leverages nuclei point-level supervision from IHC images. This framework consists of two networks: Seg-Net and Tran-Net. Seg-Net generates segmentation results for the membranes and nuclei of various tumor cells with different HER2 expression statuses. It employs a U-Net architecture with residual connections to enhance feature extraction and segmentation accuracy, and trained with a custom loss function that combines binary cross-entropy and dice coefficient to address class imbalance. Tran-Net, based on a transformer model, encodes segmented images into semantic points. This encoding process captures essential spatial relationships and contextual information for accurate cell type discrimination. This approach allowed us to accurately identify the invasive tumor cells and classified into different categories based on the intensity and completeness of the membrane staining. Weak and incomplete membrane staining, moderate and complete membrane staining, and strong and complete membrane staining were distinguished, and corresponding cell numbers and proportions were calculated.
Study samples
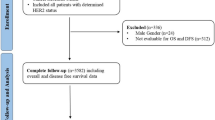
All consecutive core needle biopsies (CNB) of primary invasive breast cancers (IBCs) diagnosed at our institute in 2021—with historical HER2 IHC scores of 0 or low (1+ and 2 + /FISH-) and previously subjected to next-generation sequencing (using the FUSCC NGS panel, which detects somatic and germline mutations across 484 breast cancer-specific genes)—were rescored by a review committee blinded to the original results. SEBs were obtained following lumpectomy or mastectomy. Following rescoring, 506 CNB cases that were reclassified as HER2 0 or 1+ with paired SEB specimens were enrolled in this study. Experiments and data generation were in accordance with the ethical standards of relevant national and international rules and regulations (GCP, Declaration of Helsinki). This study was approved by the Ethics Committee of Fudan University Shanghai Cancer Center, and each participant signed an informed consent document.
CNBs were mostly performed by ultrasound guidance with 14-gauge needles, and 4–5 pieces were obtained. The CNB specimens were fixed in 10% neutral formalin for 8–10 h, embedded in paraffin, and sectioned at a thickness of 4 um. SEBs were sliced at 5- to 10-mm intervals after operation and fixed in 10% neutral buffered formalin for 8–24 h. Then, tissue samples were in paraffin, and sectioned at a thickness of 4 um prior to analysis. Subsequently, hematoxylin and eosin(H&E) and IHC staining were carried out. The HER2 IHC staining was performed on an automated Ventana BenchmarkXT slide Stainer using a specific antibody (Ventana 4B5) against HER2. Moreover, external controls encompassing varied staining intensities and degrees of completeness were established. All samples were prepared with similar preanalytical conditions and analytical processes.
The relevant clinical-pathological characteristics, such as age, histological grade, tumor size, lymph node status, and molecular subtype, were collected. TNBCs were classified into different subtypes based on the staining results: (a) IHC-based luminal androgen receptor (IHC-LAR; AR-positive [+]), (b) IHC-based immunomodulatory (IHC-IM; AR-negative [-], CD8 + ), (c) IHC-based basal-like immune-suppressed (IHC-BLIS; AR-, CD8-, FOXC1 + ), and (d) others(including MES)32.
Evaluation of HER2 expression
Paired CNBs and SEBs were independently rescored by two pretrained expert pathologists according to the latest ASCO/CAP Guidelines Update for HER2 diagnosis33. According to the latest ASCO/CAP Guidelines Update, IHC 0 is defined as no staining observed or membrane staining that is incomplete and is faint or barely perceptible and within ≤ 10% of the invasive tumor cells. If the proportion is over 10%, tumor is defined as IHC 1 + . The definition of IHC 2+ is invasive breast cancer with weak to moderate complete membrane staining observed in > 10% of tumor cells, and when over 10% of circumferential membrane staining is complete and intense, tumor is defined as IHC 3 + . HER2 0 is further divided into HER2-ultralow and HER2-null. IHC null is defined as no observed staining, and HER2-ultralow is defined as membrane staining that is incomplete and is faint or barely perceptible and within ≤ 10% of the invasive tumor cells. In addition, HER2 low is defined as IHC 1+ or 2 + /ISH-negative. In this study, HER2 low incorporated 1+ only for CNB, while also incorporated a small number of 2 + /ISH-negative case for SEBs.
Two pathologists with approximately 5 years of experience read the cases independently. If the results between the two readers matched, it was recorded as the final result. Otherwise, the adjudicator (a third pathologist) would review the slides and each reader’s scores before making the final judgement on HER2 IHC status. All of the HER2 slides of CNBs were digitalized. The developed AI algorithm was used to score the HER2 status of the CNB samples. Then, the performance of the AI algorithm was compared with the pathologist scores to evaluate its accuracy and concordance.
Statistical analysis
The clinicopathological parameters were compared among the different subgroups using chi-square analysis. Statistical analyses were performed using SPSS 26.0 (SPSS, Chicago, IL, USA) or GraphPad Prism 9.0 (GraphPad Software, La Jolla, CA, USA). A p < 0.05 was considered statistically significant.
Data availability
The code and datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Code availability
The code used during the current study are available from the corresponding author on reasonable request.
References
Modi, S. et al. Trastuzumab deruxtecan in previously treated HER2-low advanced breast cancer. N. Engl. J. Med. 387, 9–20 (2022).
Nicolo, E., Boscolo Bielo, L., Curigliano, G. & Tarantino, P. The HER2-low revolution in breast oncology: steps forward and emerging challenges. Ther. Adv. Med. Oncol. 15, 17588359231152842 (2023).
Mosele, F. et al. Trastuzumab deruxtecan in metastatic breast cancer with variable HER2 expression: the phase 2 DAISY trial. Nat. Med. 29, 2110–2120 (2023).
Venetis, K. et al. HER2 low, ultra-low, and novel complementary biomarkers: expanding the spectrum of HER2 positivity in breast cancer. Front. Mol. Biosci. 9, 834651 (2022).
Franchina, M. et al. Low and ultra-low HER2 in human breast cancer: an effort to define new neoplastic subtypes. Int. J. Mol. Sci. 24, 12795 (2023).
Bardia, A. et al. Trastuzumab deruxtecan after endocrine therapy in metastatic breast cancer. N. Engl. J. Med. 391, 2110–2122 (2024).
Fernandez, A. I. et al. Examination of low ERBB2 protein expression in breast cancer tissue. JAMA Oncol. 8, 1–4 (2022).
Palm, C. et al. Determining HER2 status by artificial intelligence: an investigation of primary, metastatic, and HER2 low breast tumors. Diagnostics (Basel) 13, 168 (2023).
Ensani, F. et al. The core needle and surgical biopsy concordance to detect estrogen, progesterone, and HER-2 receptors in breast cancer: a comparative study. Iran. J. Pathol. 12, 202–208 (2017).
Rossi, C. et al. Concordance of immunohistochemistry for predictive and prognostic factors in breast cancer between biopsy and surgical excision: a single-centre experience and review of the literature. Breast Cancer Res. Treat. 198, 573–582 (2023).
Greer, L. T. et al. Does breast tumor heterogeneity necessitate further immunohistochemical staining on surgical specimens?. J. Am. Coll. Surg. 216, 239–251 (2013).
Ge, W. K. et al. Evaluation of hormone receptor, human epidermal growth factor receptor-2 and Ki-67 with core needle biopsy and neoadjuvant chemotherapy effects in breast cancer patients. Thorac. Cancer 6, 64–69 (2015).
Chen, R. et al. Diagnostic value of core needle biopsy for determining HER2 status in breast cancer, especially in the HER2-low population. Breast Cancer Res. Treat. 197, 189–200 (2023).
Ross, J. S. et al. The Her-2/neu gene and protein in breast cancer 2003: biomarker and target of therapy. Oncologist 8, 307–325 (2003).
Bussolati, G., Annaratone, L. & Maletta, F. The pre-analytical phase in surgical pathology. Recent Results Cancer Res. 199, 1–13 (2015).
Baez-Navarro, X. et al. Interobserver variation in the assessment of immunohistochemistry expression levels in HER2-negative breast cancer: can we improve the identification of low levels of HER2 expression by adjusting the criteria? An international interobserver study. Mod. Pathol. 36, 100009 (2023).
Baez-Navarro, X., van Bockstal, M. R., van der Made, A. & van Deurzen, C. H. M. A comparison between immunohistochemistry and mRNA expression to identify human epidermal growth factor receptor 2-low breast cancer. Arch. Pathol. Lab Med. 149, 635–642 (2025).
Li, X. et al. Correlation of HER2 protein level with mRNA level quantified by RNAscope in breast cancer. Mod. Pathol. 37, 100408 (2024).
Gaudio, M. et al. Unveiling the HER2-low phenomenon: exploring immunohistochemistry and gene expression to characterise HR-positive HER2-negative early breast cancer. Breast Cancer Res. Treat. 203, 487–495 (2024).
Potts, S. J. et al. Evaluating tumor heterogeneity in immunohistochemistry-stained breast cancer tissue. Lab Invest. 92, 1342–1357 (2012).
Wu, S. et al. The role of artificial intelligence in accurate interpretation of HER2 immunohistochemical scores 0 and 1+ in breast cancer. Mod. Pathol. 36, 100054 (2023).
Geukens, T. et al. Intra-patient and inter-metastasis heterogeneity of HER2-low status in metastatic breast cancer. Eur. J. Cancer 188, 152–160 (2023).
Shang, J. et al. Evolution and clinical significance of HER2-low status after neoadjuvant therapy for breast cancer. Front Oncol. 13, 1086480 (2023).
Chen, Z. et al. Is HER2 ultra-low breast cancer different from HER2 null or HER2 low breast cancer? A study of 1363 patients. Breast Cancer Res. Treat. 202, 313–323 (2023).
Tuluhong, D. et al. Molecular characteristics and prognosis of breast cancer patients with different level of HER2 positivity after adjuvant and neoadjuvant chemotherapy. Eur. J. Cancer Prev. 32, 377–387 (2023).
Yang, C. et al. Survival differences between HER2-0 and HER2-low-expressing breast cancer—a meta-analysis of early breast cancer patients. Crit. Rev. Oncol. Hematol. 185, 103962 (2023).
Tang, Y. et al. The association between HER2-low expression and prognosis of breast cancer: a systematic review and meta-analysis. Ther. Adv. Med. Oncol. 15, 17588359231156669 (2023).
Guan, F. et al. Comparison of clinicopathological characteristics, efficacy of neoadjuvant therapy, and prognosis in HER2-low and HER2-ultralow breast cancer. Diagn. Pathol. 19, 131 (2024).
Zhang, G. et al. Distinct clinical and somatic mutational features of breast tumors with high-, low-, or non-expressing human epidermal growth factor receptor 2 status. BMC Med. 20, 142 (2022).
Dai, L. J. et al. Molecular features and clinical implications of the heterogeneity in Chinese patients with HER2-low breast cancer. Nat. Commun. 14, 5112 (2023).
Lambein, K. et al. Distinguishing score 0 from score 1+ in HER2 immunohistochemistry-negative breast cancer: clinical and pathobiological relevance. Am. J. Clin. Pathol. 140, 561–566 (2013).
Zhao, S. et al. Molecular subtyping of triple-negative breast cancers by immunohistochemistry: molecular basis and clinical relevance. Oncologist 25, e1481–e1491 (2020).
Wolff, A. C. et al. Human epidermal growth factor receptor 2 testing in breast cancer: ASCO-College of American Pathologists guideline update. J. Clin. Oncol. 41, 3867–3872 (2023).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: M.L., M.-Y.C., R.-H.S. and W.-T.Y. Collected the data: M.L., M.-Y.C., H.L., and S.-L.Z. Analyzed the data and prepared figures: M.L., M.-Y.C., and Y.-F.Z. Wrote the paper: M.L., M.-Y.C. and W.-T.Y.; M.L. and M.-Y.C. contributed equally to this work. All authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, M., Cai, MY., Lv, H. et al. Illuminating the clinicopathological and genomic landscape of HER2-null, ultralow, and low breast cancers: insights into diagnostic discordance between biopsy and surgical excision. npj Breast Cancer 12, 17 (2026). https://doi.org/10.1038/s41523-025-00882-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41523-025-00882-0