Abstract
The development of fast ionic conductors to improve the performance of electrochemical devices relies on expensive high-throughput (HT) density functional theory (DFT) calculations of transport properties. Machine learning (ML) can accelerate HT workflows but requires high-quality data to ensure accurate predictions from trained models. In this study, we introduce the LiTraj dataset, which comprises 13,000 percolation and 122,000 migration barriers, and 1700 migration trajectories, calculated for Li-ion in diverse crystal structures using empirical force fields and DFT, respectively. With LiTraj, we demonstrate that classical ML models and graph neural networks (GNNs) for structure-to-property prediction of percolation and migration barriers can distinguish between “fast” and “poor” ionic conductors. Furthermore, we evaluate the capability of GNN-based universal ML interatomic potentials (uMLIPs) to identify optimal Li-ion migration trajectories. Fine-tuned uMLIPs achieve near-DFT accuracy in predicting migration barriers, significantly accelerating HT screenings of new ionic conductors.
Similar content being viewed by others
Introduction
Li-ion conductors are used in a wide range of electrochemical devices, including batteries1, sensors2, memristors3, and synaptic transistors4. Improving the energy density, charge rate, and stability of a solid-state battery, the response time of a sensor, or the switching time of a memristor can be achieved by innovations in the architecture of devices and/or material components3. Thus, understanding of ionic transport mechanisms and their characteristics is essential for designing advanced ionic conductors.
Ionic conductivity corresponds to the processes of mobile ions transport and, as a consequence, charge transport. This process can be characterized by the migration barrier (Em), a key factor governing ionic transport, which corresponds to the energy required for an ion to pass through a transition (intermediate) state between two stable sites in the crystal lattice5.
In the last few decades, density functional theory (DFT) has become the main workhorse in computational materials science6. This approach to atomistic modeling is routinely used to study ionic conductors and accelerate the development of the materials with improved ionic mobility, moving away from inefficient trial-and-error approaches. In particular, the method is widely used for modeling battery materials7 including intercalation electrodes8,9,10,11,12, solid electrolytes5,13,14,15, and coating materials16,17. A typical DFT study of an ionic conductor involves expensive calculations of Em with nudged elastic band (NEB) method18, which allows evaluating the energy profile of a mobile ion migration. Alternatively, ab initio molecular dynamics (AIMD), which additionally takes into account lattice dynamics effects at finite temperature, can be used for direct evaluation of ionic mobility7.
Each approach has its own limitations. DFT-NEB converges slowly if the initial trajectory is poorly chosen or may even converge to a wrong pathway19, and must be performed for numerous ionic hops to construct the percolating network that represents macroscopic diffusion. Also, DFT-NEB tends to underestimate Em by 0.1–0.3 eV, as the effects of lattice dynamics are not included20. A comprehensive benchmark of DFT-NEB accuracy in calculating migration barriers with different exchange-correlation (XC) functionals compared to experimental data was conducted by Devi et al., who demonstrated that the error can reach up to 0.3 eV21.
On the other hand, AIMD does not require prior knowledge of migration pathways, offers more insights into diffusion mechanisms and correlation effects, and generally predicts Em values that are closer to experiment due to more realistic modeling conditions20. However, AIMD simulations are more computationally demanding7 and are typically performed for systems containing a few hundred atoms over sub-nanosecond timescales. This limitation impacts the accuracy of the extracted properties due to the finite-size effects22 and the limited number of diffusion events captured23.
Given the high computational cost of DFT, the research effort has been focused on developing approximate models for accelerated discovery of fast ionic conductors in a high-throughput (HT) manner. For example, Kahle et al. introduced the “pinball” model, which reduces the cost of AIMD diffusion study by keeping the host lattice “frozen”24. This model was subsequently applied to screen Li-ion conductors25. Rong et al. invented an efficient algorithm for finding the optimal migration trajectory using a DFT charge density static potential26. The method was later used by Rutt et al. for screening Mg-ion conductors27. Finally, Adams et al. developed an empirical force field, the Bond Valence Site Energy (BVSE) method28, for calculating the minimum energy barrier required for an ion to percolate through the host framework, known as the percolation barrier. This characteristic correlates well with DFT-derived and experimentally measured energy barriers29 establishing it as a viable tool for rapid screening of ionic conductors30,31,32.
With the growing volume of experimentally and theoretically derived materials33,34,35,36, driven by advances in tools for predicting new structures37,38,39,40, the task of tuning the ionic conductivity through doping15,41 and performing HT calculations of ionic transport properties demand substantial time and computational resources. In this context, the development of fast and accurate machine learning (ML) models will allow replacing conventional techniques for calculating transport properties in a HT manner.
Various data-driven solutions have already been applied to predict the properties of ionic conductors, leveraging both experimental and theoretically calculated data through developing crystal structure descriptors and using deep neural networks and classical ML models42,43,44. In the case of neural networks, a large amount of data is needed for training the model effectively, whereas classical ML models require less data but necessitate careful development of features tailored to a specific task45. The development of ML models for structure-to-property prediction of a mobile ion energy barrier has been actively pursued in recent years, albeit with several caveats. First, most of the datasets used to develop the models have been generated using the empirical BVSE method46,47, while the DFT-derived data is lacking. Second, numerous studies have focused on specific structural types, such as NASICONs and argyrodites48, or a limited range of chemical compositions including Li-Zn-X-O49, K-Fe-Ti-O50, LiMPO4X51). Third, relatively small datasets with limited ranges of calculated migration barriers have been utilized. However, the overarching question of whether it is possible to develop a model capable of predicting Li-ion migration barriers for any material with DFT-level accuracy remains largely unanswered.
The development of interatomic ML potentials has been a major breakthrough in modeling ionic conductors52,53,54, which are reported to achieve DFT-level accuracy in predicting Em and ionic conductivity at much lower computational cost. Despite their accuracy and efficiency, these potentials are typically tailored to specific chemistries and are not suitable for HT exploration across diverse chemical spaces. As a solution, current research focuses on developing universal machine learning interatomic potentials (uMLIPs). State-of-the-art uMLIPs, such as M3GNet55, CHGNet56, SevenNet57, and MACE58, represent graph neural networks (GNNs)59 trained on the Materials project’s (MP) data33 spanning a wide range of chemical compositions. The authors assert that these uMLIPs can be applied to downstream tasks, such as modeling adsorption and diffusion processes, potentially accelerating the discovery of novel materials with superior properties. However, despite their application to model Li-ion diffusion60, no benchmark scheme has yet been proposed to assess their robustness and accuracy in predicting Li-ion transport properties across the entire periodic table.
The main objective of our work is to evaluate the accuracy and applicability of existing ML models for HT screening of Li-ion conductors. For this purpose, we collect the LiTraj dataset, which consists of four parts, namely BVEL13k, nebBVSE122k, nebDFT2k, and MPLiTrj datasets. The BVEL13k dataset is designed with the purpose of benchmarking classical ML models and GNNs in structure-to-property prediction of Li-ion percolation barrier derived with BVSE method. We use the nebBVSE122k and nebDFT2k datasets to evaluate surrogate models for structure-to-property prediction of Li-ion migration barrier calculated at the BVSE and DFT level of theory, respectively. Finally, with the nebDFT2k and MPLiTrj datasets, we evaluate the ability of uMLIPs to reproduce the optimal Li-ion migration trajectory and corresponding energetics predicted by DFT-NEB.
Results
Percolation barrier prediction with the BVEL13k dataset
In this section, we benchmark classical ML and GNN models for structure-to-property prediction using the BVEL13k dataset. The BVEL13k dataset comprises 12,807 Li-ion 1D, 2D, and 3D percolation barriers (\({E}_{a}^{1D}\), \({E}_{a}^{2D}\), \({E}_{a}^{3D}\)) calculated for Li-containing crystal structures retrieved from the MP database33. The dataset overview is given in Fig. 1a–d.
a The counter of the chemical elements present in the collected crystal structures, b The distribution of the Li-ion 1, 2, and 3D percolation barriers in the BVEL13k dataset, c An exemplary Li-containing structure (mp-2878) and d corresponding Li-ion percolation (diffusion) map calculated using the BVSE method, e Parity plot for the prediction of the 1, 2, and 3D percolation barriers from the test data using the most accurate GNN models, f The top7 most important features for predicting 1D percolation barrier according to the impurity-based method, g The negative mean squared error (MSE) of predicting 1D percolation barrier vs. number of features selected by the forward feature selection method with the KNN regression model.
The percolation barrier, calculated using the BVSE force fields, represents the minimal energy barriers required for infinite diffusion through the crystal. To evaluate this value, a single Li-ion is iteratively placed at each point of a grid spanning the unit cell of the host framework. The corresponding Li-ion energy is evaluated using BVSE. The resulting spatial energy distribution is analyzed to identify the percolating regions (Fig. 1c, d) and the corresponding percolation barriers.
We train random forest (RF), extreme gradient boosted RF (XGBRF), k-nearest neighbor (KNN), kernel ridge (KR), and support vector machine (SVR) regression models on the calculated percolation barriers. To employ the models, we perform a Voronoi partitioning of the crystal structure and use aggregated values of geometric and elemental characteristics over equilibrium Li sublattice and over approximate Li transition states. This is in accordance with the previously reported studies on the correlation between the mobile ion energy barrier and crystal structure attributes43,47,49,61,62. The full list of the features designed for predicting percolation and migration barriers along with detailed description of them is given in Supplementary Section S1 of the Supplementary Information (SI). In addition, we utilize the Matminer’s63 general purpose featurization scheme developed by Ward et al.64.
Among the GNNs, we train three structure-to-property prediction models, namely Nequip65, Allegro66, and M3GNet55.
The performance metrics calculated for the test subset of BVEL13k data are shown in Table 1. According to the results, the proposed featurization scheme performs better than that of Matminer for 1D and 2D percolation for most of the models tested. On contrary, the 3D barriers are better predicted with the XGBRF model trained on the Matminer-derived descriptors. Combining two feature sets provides no significant improvement in quality of target predictions (see Supplementary Table S3 of the SI), whereas the computational complexity becomes much higher due to the larger number of features being computed and used within the fit.
The group of GNN models shows better performance. Interestingly, the M3GNet model with approximately 0.3 million parameters performs as well as Nequip with 2 million parameters in predicting 1D percolation barriers. Considering a smaller Nequip version, which is about the same size as M3GNet, we obtain the prediction quality that is worse compared to classical ML models trained on the developed feature set. Regardless of the trained model, mean absolute error (MAE) and root mean squared error (RMSE) tend to increase with the dimensionality of the percolation map, which can be due to the increase in variance of the target property and its range for higher dimensionalities (Fig. 1b, e). In any case, the GNN models perform much better compared to the classical ML models in predicting 2D and 3D percolation barriers.
We determine the impact of each descriptor on the model prediction for 1D barrier using the impurity-based feature importance method. According to this, the fraction of the covalent free space in the structure has the highest impact on the RF model (Fig. 1f). The average volume of the Voronoi polyhedra constructed around the Li sites within the anion sublattice, the effective coordination number of Li, the average oxidation state of the Li-ion nearest anion neighbors, the average volume of Voronoi polyhedra constructed around approximate transition states, and volume per atom of the host structure are in top 7 most important features.
We use the KNN regressor for forward feature selection to estimate the optimal number of features required for fast and accurate predictions. According to the results, the optimal number of crystal structure descriptors for predicting the 1D percolation barrier with the KNN regressor is 30-35, since the negative mean squared error of prediction converges within this range of feature numbers (Fig. 1g). The model refitted on this subset is slightly better (MAE = 0.14 eV, RMSE = 0.27 eV, R2 = 0.82) compared to the KNN regressor trained on the full set of features.
Migration barrier prediction with nebBVSE122k and nebDFT2k datasets
In this section, we benchmark the ML models for the structure-to-property prediction of the Li-ion migration barrier, Em, using the nebBVSE122k and nebDFT2k datasets.
The nebBVSE122k dataset represents energy barriers, obtained by the BVSE-NEB optimization of the linearly interpolated migration trajectory of Li vacancy from i-th starting to j-th final position (Fig. 2a–c). Overall, there are 122,421 barriers calculated for 11,290 distinct structures representing a subset of the crystal structure from the BVEL13k dataset. We use the nebBVSE122k data to sample Li-ion migration trajectories in a stratified manner for further DFT-NEB optimization. The resulting 1681 BVSE-NEB and DFT-NEB optimized trajectories obtained for 876 distinct crystal structures and 45 chemical elements form the nebDFT2k dataset. Fig. 2d shows the Em distribution in the nebBVSE122k and nebDFT2k datasets.
a, b An illustrative example of linearly interpolated and BVSE-NEB optimized Li-ion migration trajectory, respectively, and c corresponding energy profile, d Distributions of the BVSE- and DFT-NEB calculated Li-ion migration barriers, e Parity plot for the test data predictions obtained with the M3GNet model trained on the nebBVSE122k, f The correlation between the weight of the edge connected to the centroid and its length extracted from the last layer of the Allegro model trained on the nebBVSE122k dataset, g An illustrative example of the structure with the centroid dummy element, h Parity plots for the test data predictions obtained with the SVR model trained on the nebDFT2k datasets, i The top-5 most important features according to the impurity-based method (f1 – minimum volume of the Voronoi polyhedra constructed around the initial, final, and centroid Li sites, f2 – edge length, f3 – maximum weighted average of oxidation states of the nearest neighbors collected for initial, final, and centroid Li sites, f4 – maximum volume of the Voronoi polyhedra constructed around the initial, final, and centroid Li sites, f5 – minimal average covalent radius of nearest neighbors collected for initial, final, and centroid sites.
Apparently, the target property considered in these datasets is an attribute of the edge, eij, connecting the i-th initial and j-th final positions of the migrating ion (vacancy). Therefore, we use the Voronoi-tessellation-derived descriptors associated with this edge to develop models. In the case of GNNs, we modify the input crystal structures by introducing a dummy element ’X’ at the midpoint (centroid) of the eij edge, enabling the models to focus on the vicinity of a specific Li-ion migration pathway (Fig. 2g).
The benchmarking results of the selected ML models on the nebBVSE122k and nebDFT2k datasets are presented in Tables 2 and 3, respectively.
In the case of classical models, the RF and XGBRF models provide more robust predictions compared to the KNN model. The strongest impact on predictions of both the XGBRF and RF models is from the minimum volume of the Voronoi polyhedra among the starting, final, and midpoint sites (Fig. 2i).
According to the scores obtained, the GNN models possess better performance in predicting the BVSE-NEB calculated migration barrier compared to that of the classical models, with M3GNet being the most accurate model (MAE = 0.21 eV, R2 = 0.95).
To identify the graph edges that have the greatest impact on the GNNs’ predictions, we explore embeddings and weights at the last layer of Allegro. We find that 98% of the graph edges with the largest weights in the graph represent the edges connected to the centroid. In addition to this, we observe a negative non-linear correlation between the weights of these graph edges and their lengths in the crystal structures (Fig. 2f).
Since the migration barriers from the nebBVSE122k and nebDFT2k datasets represent the same property, but computed with two different solvers: the faster but less accurate empirical force field and the more computationally expensive but more accurate DFT, we explore potential gain from the transfer learning (TL) approach. Within the TL paradigm, the weights of the GNN model trained on the BVSE-NEB data (pre-trained model) are used as a starting point for the subsequent fine-tuning the model on the DFT-NEB data. In turn, for the classical models, we include the BVSE-NEB calculated barrier as a feature for predicting the DFT-NEB barriers. We observe that TL as well as the inclusion of the target calculated with a simpler level of theory improve the quality of the predictions. The M3GNet model trained starting from randomly initialized weights has the lowest accuracy among the models built, while the RF model with the BVSE-NEB barrier included as an additional feature yields the best accuracy in predicting the DFT-NEB barrier (MAE = 0.25 eV, R2 = 0.80).
Benchmarking universal interatomic potentials with the nebDFT2k and MPLiTrj datasets
In this section, we use the nebDFT2k and MPLiTrj datasets to evaluate the ability of uMLIPs, specifically M3GNet55, CHGNet56, SevenNet57, and MACE-MP-058 (MACE, for a shorter notation), to predict minimum energy Li-ion pathway and corresponding energetics. The MPLiTrj dataset contains 929,066 configurations with calculated energies, forces, and stress tensors, obtained during the DFT-NEB optimization of 2698 migration trajectories corresponding to 1269 crystal structures. We use a subset of this data for validating and fine-tuning uMLIPs (see the “Fine-tuning” Section of Methods).
We utilize the uMLIPs trained on the MP data. However, our datasets are computed using a different computational scheme than that used by MP. We employ the PBE (XC) functional without spin polarization, while MP uses spin polarization and Hubbard DFT+U corrections for d-orbital electrons of transition metals (TMs). Devi et al.21 showed that applying Hubbard U corrections to the PBE XC functional can increase the Li-ion migration barrier by several hundred meV for TM-containing systems. To account for this effect, we split the nebDFT2k dataset into two parts – with (“with TM”) and without (“w/o TM”) TM systems – for further benchmark of uMLIPs. We also include the BVSE-NEB trajectory optimization results, since this method can be considered as a universal, although empirical, interatomic pair potential.
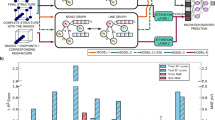
Figure 3 shows the distributions of migration barrier prediction errors for the models benchmarked. All uMLIPs show much better perfomance for the “w/o TM” part of the data compared to the “with TM” part. The MACE and SevenNet models have fairly close accuracies in predicting Em (MAEs of 0.07–0.08 and 0.20–0.21 eV for “w/o TM” and “with TM” data splits, respectively). Compared to SevenNet, the error distribution obtained for the MACE model is centered closer to 0. We observe a significant underestimation of Em by the CHGNet and M3GNet models. The M3GNet model has the most pronounced underestimations, resulting in high MAEs of 0.32 and 0.43 eV for the “w/o TM” and “with TM” parts, respectively. In contrast, the BVSE-based model greatly overestimates the migration barriers, which is due to the fixed geometry of the host framework during the Li-ion migration trajectory optimization. The calculated performance metrics for the models under consideration are shown in Table 4.
a Violin plots showing the prediction error distributions for each model studied for the “w/o TM” (left splits) and “with TM” (right splits) subsets of the nebDFT2k dataset. For better visual appearance, the error distributions are cropped to the range of −1.75 to 1.75 eV. The error distributions in the full range are given in SI, b Parity plot for the predictions of the SevenNet-ft uMLIP.
To elucidate the reasons for the much worse uMLIP performance for the “with TM” part compared to “w/o TM”, we predict forces and energies for the stratified subsample of the test split of the MPLiTrj dataset (see the “Fine-tuning” section of Methods). The results clearly show that uMLIPs completely fail to predict forces for the TM subset (force RMSE of 0.4-0.5 eV/Å, and Pearson correlation coefficient, Rp, of 0.20–0.24, see Supplementary Table S7, Supplementary Figs. S2-5 of the SI). Obviously, any potentials fitted to the MP data, which are derived using PBE+U, cannot accurately predict forces for the data calculated with just PBE. Moreover, the DFT+U corrections and spin polarization, which are not included in our calculation scheme, affect the equilibrium bond length explaining discrepancies in the minimum energy path prediction. Meanwhile, the force prediction error for the “w/o TM” part is also high (RMSE = 0.12–0.17 eV/Å, Rp = 0.68–0.76 (see Table 7, Supplementary Figs. S2–5 of the SI)
Next, we fine-tune the MACE-MP and SevenNet models on a subsample of the MPLiTrj training data split (see the “Fine-tuning” section of Methods, and Supplementary Section S2 of SI). The comparison of the accuracy metrics for the pre-trained and fine-tuned models is given in Table 5. It is worth noting that we compute the performance metrics for all trajectories from the nebDFT2k dataset regardless of the data split (training/validation/test). It is done from the perspective of solving the downstream task of optimizing the trajectory as a key target, rather than predicting forces and energies. The scores for the test split of the nebDFT2k dataset is given in Supplementary Table S7 of the SI.
The fine-tuning significantly reduces errors of the Em prediction for the TM subset. Thus, for the fine-tuned SevenNet (“SevenNet-ft”) model, the lowest MAE of Em assessments of 0.11 eV is achieved. It also greatly improves the accuracy of force and energy prediction for both parts of the dataset (RMSE = 0.07 and 0.09 eV/Å, Rp = 0.91 and 0.94 for “w/o TM” and “with TM” parts, respectively, for the SevenNet-ft model, see Supplementary Table S7, Supplementary Figs. S6-7 of the SI). Along with this, the accuracy of Em prediction of the fine-tuned MACE model on the “w/o TM” subset becomes slightly worse (MAE = 0.07 and 0.10 eV for the pre-trained and fine-tuned models, respectively). This observation may be attributed to a poorly chosen fine-tuning protocol. Despite the improvement in average accuracy, the fine-tuned models tend to underestimate Em, indicating a slight softening of the potentials (Fig. 3a).
To show that improvement in accuracy is related not only to a better fit to our computational scheme but to inclusion of the Li-ion transition states and better sampling of the potential energy surface (PES), we fine-tune the SevenNet model on the slice of the MPLiTrj dataset comprising the configurations sampled when optimizing the vacancies (endpoints of the migration trajectories). The results clearly indicate softening of the potential obtained leading to the pronounced underestimation of Em (see Supplementary Tables S7, 8, Supplementary Fig. S8–9 of the SI). We attribute the strong softening of the “SevenNet-ft-vac” model due to the training on near-equilibrium geometries, which leads to the wrong prediction of the PES curvature67.
We assess the ability of the models to predict a correct geometry of the migration pathway by calculating the path averaged MAE (PA-MAE, see Methods for the details) of the trajectories optimized with surrogate models with respect to the DFT-derived ones. The calculated distributions are given in Supplementary Fig. S12 of the SI. The results indicate that the pre-trained models have a larger error for the “with TM” data split compared to “w/o TM” (0.09–0.10 vs. 0.02–0.03 Å, respectively) trajectories. Fine-tuning the MACE and SevenNet models improves the quality of geometry prediction for both datasets, 0.02 Å and 0.03–0.04 Å for “w/o TM” and “with TM”, respectively.
We observe the highest error in predicting the optimal trajectory for the BVSE model (0.05 and 0.11 Å for “w/o TM” and “with TM” data splits). Within this approach, the host structure ions are fixed, which, generally, leads to underestimations of average distances between the migrating ion and its neighbors along the trajectory.
Practical application
In this section, we present an example of integrating uMLIPs into a scheme for HT screening of Li-ion conductors. We search for cathode coatings (CC) for the Li10GeP2S12 (LGPS) solid electrolyte (SE) material proposed for use in Li-ion all-solid-state batteries68. In addition, we fix cathode active material (CAM) to LiCoO2. The purpose of using the CC is to physically separate SE and CAM to prevent SE decomposition under the battery operation conditions.
Using the MP database, we collect Li-containing structures with a bandgap larger than 4.0 eV and energies above the convex hull smaller than 0.05 eV/atom. The materials with electrochemically active elements were discarded to exclude polaron migration mechanism of electron transport69.
Following the typical funnel approach, we reduce the candidate list to stable materials within 3.0–4.5 V vs. Li/Li+ anode by calculating the electrochemical stability window according to the methodology described in refs. 70,71. Subsequently, the stability of the SE∣CC and CC∣CAM interfaces is assessed according to the methodology proposed by Richards et al.70.
For the set of materials obtained, we calculate the Li-ion percolation barriers using the SevenNet-ft model. It is worth noting that the term “percolation barrier” used here is different from the one defined for the BVEL method. Here we construct a percolating (periodic) graph on Li-Li edges that are shorter than 6.0 Å and have the minimum distance to the framework ions of longer than 0.5 Å. Using SevenNet-ft, we compute Em for Li vacancy migration corresponding to the symmetrically inequivalent edges of the constructed graph and perform percolation analysis similar to that of ref. 27. A complete list of the materials under consideration and corresponding properties is given in the SI.
As a result, we identify potential coatings with Li-ion 3D percolation barriers lower than 1.0 eV. The resulting phases include both experimentally72 and theoretically studied ionic conductors73. For example, Li3AlF6 (0.45 vs. 0.53 eV for SevenNet-ft and PBE, respectively73) and Li3PO4 (0.64 vs 0.69 eV for SevenNet-ft and GGA, respectively)74, Na3Li3Al2F12 (0.31 vs 0.34 for SevenNet-ft and PBE, respectively)75. Additionally, we predict several conductors for which the migration barriers are not yet known, but they are shown to be good Li-ion conductors and/or coatings, e.g. LiYF470,76 and Li2ZnCl477.
The energy barriers calculated using the SevenNet-ft model are in reasonable agreement with the reported data suggesting that uMLIPs can be effectively incorporated in HT screening schemes.
Discussion
Both classical ML and GNN models have been used to predict Li-ion percolation barriers based on BVEL and various ML-generated data. However, no comprehensive benchmark of their accuracy has been conducted to date46,47,49. According to our benchmark, the GNN models show superior prediction accuracy. The classical ML models trained on descriptors specifically designed for predicting the percolation barrier yield accuracies comparable to those of the GNN models for the 1D percolation barrier, but fall short in performance for higher-dimensional percolation. As a result, GNN models appear to be a favorable strategy for pre-screening of “bad”/“good” ionic conductors in a HT scheme. However, due to the relatively high RMSE (0.49 eV) in the prediction of 3D percolation barriers, we recommend the direct use of the BVEL method for smaller datasets (~1000 structures), and to employ GNN only for larger datasets, where the difference in computational time becomes significantly advantageous in favor of GNN.
When predicting Li-ion migration barriers using structure-to-property models, the M3GNet GNN model achieves the highest accuracy for BVSE-derived energy barriers. However, for DFT-derived data, its accuracy is the lowest, which we attribute to the limited amount of training data, as the nebDFT2k dataset contains two orders of magnitude less data than the nebBVSE122k dataset. It appears that GNN models are sensitive to data scarcity, resulting in classical ML models demonstrating higher accuracy on the nebDFT2k dataset compared to the GNN models, even when the GNN model was pre-trained on the BVSE-derived dataset and subsequently fine-tuned on the DFT data. Overall, achieving reliable structure-to-property prediction of Em remains challenging using either classical ML or GNNs.
To explore potential solutions to this problem, we analyzed the inclusion of spatially resolved information in the input data. The examination of the graph edge weights from the last layer of the Allegro model, trained on the BVSE-derived dataset, reveals that the local coordination at the midpoint between the starting and final positions of the migrating ion is the primary focus of GNN-based models. Thus, explicit modifications of input graphs could represent a prospective direction for further improvement of structure-to-property GNN models, aimed at predictions of local properties at the bond length scale.
Finally, using the nebDFT2k dataset, we show that the SevenNet and MACE uMLIPs exhibit high predictive power for systems without TMs (MAE = 0.07–0.08 eV), which is comparable to DFT. For TM-containing systems, however, the errors are much higher (MAE = 0.20–0.21 eV). Fine-tuning these models significantly improves their accuracy for TM-containing systems, which is a result of better fit to our calculation setup (MAE = 0.10–0.11 eV) and the inclusion of crystal structures in non-equilibrium/high-energy states, enhancing the PES description by the uMLIP67.
An important advantage of uMLIPs are insights into the optimal migration trajectory, whereas this information is not accessible with structure-to-property prediction models. While determining the optimal migration trajectory using uMLIPs requires more computational resources than structure-to-property predictions, it remains significantly more computationally efficient than the DFT-NEB approach. This advantage expands the applicability of uMLIPs, as analyzing bottlenecks in migration pathways can inform strategies for improving the ionic conductivity of a given material41.
The accuracy achieved by uMLIPs is much higher compared to the BVSE-NEB method and represents a step forward in the accurate prediction of Li-ion migration trajectories and migration barriers using surrogate models. In our practical application example of screening protective layer materials for all-solid-state batteries using Li10GeP2S12 solid electrolyte and LiCoO2 cathode active material, we show that uMLIPs can be effectively incorporated into HT schemes and provide a reliable result. Ultimately, it is up to individual researchers to decide whether this level of accuracy is sufficient for the task at hand. Alternatively, since the convergence of energy and force is slow for poor initial guess within DFT, one can use uMLIPs to precondition the migration path78, followed by final optimization with DFT.
Several limitations of our work should be noted. We use BVSE-NEB to precondition the Li-ion migration trajectory for further DFT-NEB optimization during the collection of the nebDFT2k dataset. This approach accelerates the convergence of DFT calculations78. However, it results in the loss of some high energy configurations that might have emerged if the DFT-NEB optimization had started from a linearly interpolated initial guess trajectory, due to shorter Li-framework interatomic distances. Therefore, although our dataset includes high energy Li-ion transition states, it is biased toward near equilibrium structures, which leads to a slight softening of the resulting potentials.
Furthermore, we do not evaluate the effectiveness of the fine-tuning protocol in terms of data subsets (e.g., various slices of the MPLiTrj dataset) and the hyperparameters used (e.g., optimizer choice, learning rate, energy/force/stress weights in the loss function). Future work could address these gaps and explore optimal fine-tuning protocols79.
When collecting the data for evaluating the accuracy of uMLIPs, we focused exclusively on Li-vacancy migration. Other important mechanisms of ionic transport, such as interstitial migration and the concerted migration of Li-ions/Li vacancies, are beyond the scope of this study. The presence of interstitials and their migration causes much shorter Li-Li distances and, consequently, stronger cation-cation repulsion compared to vacancy migration. Therefore, their accurate description may require the inclusion of additional configurations, which are not present in our dataset. To introduce such configurations, the search for stable interstitial sites is necessary to enable adequate NEB modeling. On this track, the Voronoi tessellation-80, BVSE-81, or charge density82-based schemes can be used for finding voids suitable for interstitials.
As for concerted migration, this mechanism is typically observed in superionic conductors with a disordered Li-ion sublattice, such as LGPS83, LLZO84, and LATP13, posing significant challenges for data collection. The identification of correlated ionic motion requires performing high-temperature AIMD simulations, followed by the construction of the Van Hove correlation function. This process should be followed by automated analysis, identification of individual concerted events, and potentially their further NEB modeling through the most probable Li sites in a disordered sublattice13. As a simplified approach, one could model the concerted migration of two (or more) Li-ions toward a vacancy in an ordered sublattice.
We propose that for accurate modeling of concerted effects using uMLIPs, the training dataset should include DFT-NEB and AIMD simulations of concerted migration involving both vacancies and interstitials. However, given the high universality of existing models, they may already capture concerted effects to some extent, which would be an intriguing topic for future investigation. Moreover, uMLIPs can be used directly to study systems with disorder and the effect of dopants on Em, which dramatically expands the search space for new materials.
While the dataset was initially constructed to benchmark different ML approaches to predict Li-ion migration, its applications are not limited to this purpose. As we show, by fine-tuning uMLIPs to a specific computational scheme, one may obtain models with accuracy close to the desired theoretical level. For example, Devi et al. showed that DFT-NEB modeling using the SCAN XC functional yields Em that are closer to experimental values compared to those obtained with PBE, PBE+U, and SCAN+U functionals21. In this context, the trajectories from the nebDFT2k dataset can be refined using alternative computational schemes, such as SCAN, for further development of fine-tuned uMLIPs capable of providing more realistic predictions. In addition, this refinement can serve for systematic convergence studies of the XC functional impact on migration barriers. Finally, the collected data can be included into datasets for developing general purpose (foundation) models, as the structures sampled during the optimization of Li-ion migration trajectories include dynamically unstable geometries, that can help mitigate the problem of PES softening67.
As for the screening of new ionic conductors, one should remember, that information about Em alone is insufficient to evaluate an ionic conductor. Factors such as lattice dynamics, charge carriers concentration, and correlated migration also play crucial role in determining ionic conductivity. Therefore, comparing two ionic conductors based solely on Em is not meaningful20. Hence, future research efforts should prioritize the collection of AIMD-derived ionic conductivity data for structure-to-property predictions85 or for validating the MD output of uMLIPs.
In conclusion, we carried out a systematic comparative analysis of various ML models for predicting Li-ion percolation barriers, migration barriers, and migration trajectories. For this purpose, we collected the comprehensive LiTraj dataset, which includes data computed using the BVEL, BVSE-NEB, and DFT-NEB methods. Our findings demonstrate that GNN models developed for structure-to-property prediction of the BVEL-calculated percolation barrier are effective for HT screening of ionic conductors, achieving RMSE values of 0.2–0.5 eV for 1D, 2D, and 3D percolation barriers. In the case of migration barrier regression, both GNN and classical models yields comparable accuracies for the BVSE-derived Em. However, for a smaller DFT-NEB dataset comprising ~2000 samples, the classical models exhibit higher predicting accuracy. Despite this, none of the models achieved sufficient accuracy for use in HT approaches (RMSE = 0.38 eV, R2 = 0.80). Potentially, they can still be utilized for pre-screening “good” and “bad” ionic conductors. Finally, we present a novel benchmark for uMLIPs, based on the DFT-derived dataset of Li-ion minimum energy paths and corresponding energetics. Our results indicate that the fine-tuned SevenNet and MACE uMLIPs can accurately reproduce the optimal Li-ion migration trajectory, achieving a reasonable RMSE of 0.12–0.18 eV in predicting Em. This suggests that they are suitable for HT screening of Li-ion conductors. To illustrate the incorporation of uMLIPs into a HT scheme, we used the fine-tuned SevenNet model to perform a screening of protective coatings for stabilization of the interface between the Li10GeP2S12 solid electrolyte and LiCoO2 cathode active material. Our dataset can serve as a valuable resource for further development, evaluation and selection of data-driven approaches for designing Li-ion conductors, thereby accelerating the discovery of materials with outstanding transport properties.
Methods
Bond valence site energy method
Within the BVSE approach28, the interaction between a mobile (test) ion and its oppositely charged neighbors is modeled as a Morse potential (Eq. (1)). In turn, the interaction between the ion and neighbors with the same charge sign is modeled as Coulomb repulsion (Eq. (2)). The sum of these interaction energies over all neighbors within a given cutoff provides the total energy of the test ion (Eq. (3))30.
where D0, α, Rmin are tabulated constants. R is the distance between the test ion and its neighbor.
where V, Vi, n, ni, Ri, r, ri are the test ion oxidation states (valence), neighboring cation oxidation state, test ion principal quantum number, neighboring cation principal quantum number, distance between the test and neighboring ion, and their covalent radii, respectively; Conv is the conversion factor from the reciprocal length to eV, and f is a screening factor of 0.7430.
To calculate the percolation barriers using this method, the bond valence energy landscape (BVEL) must first be obtained. First, all Li ions are removed from the structure to obtain the host framework. Next, a grid with a specified resolution is constructed over the unit cell. At each grid point (xi, yi, zi) a single test ion (e.g. Li+) is placed in the host framework, followed by the BVSE calculations of the interaction energy E(xi, yi, zi) between the test ion and its framework neighbors. The resulting energy distribution E(x, y, z) is then analyzed to determine the percolation dimensionality and the corresponding energy barrier. The percolation barrier is defined as the minimum threshold value \({E}_{tr}\), at which the isosurface described by the equation \(E(x,y,z)=min(E(x,y,z))+{E}_{tr}\) percolates in 1, 2, or 3 dimensions.
The BVlain Python package (see Code availability section) was employed to calculate the Li-ion percolation barriers. A cutoff radius of 10.0 Å was used for the pair interaction between Li+1 and the framework, and the grid resolution was set to 0.2 Å. Up to k = 200 nearest neighbors were collected within the cutoff spheres. The bond valence force field parameters were sourced from80. As the BVSE approach requires formal charges of atoms, the crystal structures were decorated with oxidation states using pymatgen’s module BVAnalyzer86, with default settings applied, except that O1− and P3− species were not allowed.
Density functional theory calculations
The Vienna Ab initio Simulation Package (VASP) was used to perform density functional theory (DFT) calculations87. The GGA PBE88 exchange-correlation functional was employed, along with PAW potentials recommended by the VASP developers, except for Li, where a potential with only one valence electron was selected.
A kinetic energy cutoff of 520 eV and a single k-point at Gamma were used, with Gaussian smearing of 0.1 eV applied at the Fermi level. Atomic coordinates were optimized using the FIRE optimizer, as implemented in the ASE Python library89, until the maximum forces acting on each ion were reduced to less than 0.1 eV/Å.
Nudged elastic band calculations
The Li-ion migration trajectories were optimized using the climbing image Nudge Elastic Band (NEB) method (with an improved tangent estimate) and the FIRE optimizer, as implemented in the ASE Python library. To reduce the interaction between periodic images, the supercells with >8.0 Å along each direction were used for the calculations.
Depending on the Li-ion hop distance, three to five intermediate images were used. For BVSE-NEB calculations, the initial migration trajectory was linearly interpolated between the equilibrium positions of the ion. The geometry around vacancies were not optimized, and only the forces acting on the moving ion were considered during the band optimization with Li-Li interaction accounted for. The BVSE-NEB optimized trajectory served as the initial guess for the DFT-NEB and uMLIP-NEB calculations. For DFT-NEB and uMLIP-NEB calculations, the geometry around vacancies were optimized until the maximum force acting on each atom was less than 0.1 eV/Å.
Sampling Li-ion migration hops
The Li-Li hops (edges) used to collect the nebBVSE122k and nebDFT2k datasets were sampled using the ions Python library (see Code Availability section). The library enables the identification of the non-equivalent ionic hops that form a percolating (periodic) graph.
To construct the percolation graph on the Li-ion sublattice, we collect all neighbors in the unit cell and its 26 nearest periodic replicas within a pre-defined cutoff. The edges connecting the starting and final positions of the migrating ion are rejected if the distance between the migrating ion and any other ion in the structure is less than 0.5 Å. Next, we find the shortest Li-ion hop distance (edge length) ensuring that at least one Li-ion in the unit cell can percolate through the constructed graph to its periodic replica sites in 1, 2, or 3 dimensions. The resulting symmetrically non-equivalent edges are selected for BVSE-NEB calculations.
The cutoff for the Li-Li hop distance was set to 8.0 Å in accordance with the minimum size of the supercell. For the nebDFT2k dataset, Li-Li edges were randomly subsampled from the nebBVSE122k dataset using the following criteria: i) the edge length is less than 6.0 Å, ii) edges are sampled for the chemical systems represented by at least 10 data entries in the nebBVSE122k dataset, iii) the chemical space was restricted to 45 chemical elements deemed most suitable for practical applications.
Data split
Each of the introduced datasets was split into training, validation, and test subsets in an 8:1:1 ratio. The training, validation, and test data points were propagated from the nebDFT2k dataset to the other datasets (nebBVSE122k, BVEL13k, and MPLiTrj), ensuring that no crystal structures from the test subset of the nebDFT2k dataset are in the training or validation subsets of other datasets. This approach provides consistent data splits and prevents data leakage within the transfer learning (TL) approach. In addition, the compositional space of each subset was controlled such that all chemical elements present in the test subset also appeared in the validation subset, and all elements from the validation set were included in the training subset.
Models training and evaluation
For the classical ML models, the training and validation data splits were combined within a five-fold cross-validation scheme for model fitting and hyperparameters tuning. The hyperparameters were optimized using grid search technique. We used classical ML models as implemented in the scikit-learn90 and XGBoost91 Python libraries.
All GNN models were trained by minimizing the mean squared error (MSE) loss function. When selecting hyperparameters for the Allegro and Nequip models, we followed the recommendations of the model’s authors and from our previous study92. The Allegro and Nequip models were trained with r_max = 7.0 and 6.0 Å for the BVEL13k and nebBVSE122k datasets, respectively, using the ReduceOnPlato scheduler with early stopping based on validation loss and learning rate. The default M3GNet model with cutoff = 5 Å, threebody_cutoff = 4 Å, and is_intensive=False was used. The CosineAnnealing scheduler was employed for the M3GNet model training. Full configuration files and learning curves are available upon request.
The mean absolute error (MAE), root mean squared error (RMSE), and coefficient of determination (R2) metrics were calculated on the test data splits to evaluate the accuracy of the models.
MLIPs description and evaluation
For the benchmark, we used CHGNet (v0.3.0), SevenNet(2024.11.07), MACE-MP-0 (2023.12.03, the “small” model of 2.08 million trainable parameters), and M3GNet (2021.2.8) uMLIPs. The first three models were trained on the MPtrj dataset55 comprising ~1,580,000 structures, while M3GNet was trained on the smaller MPF55 dataset (~188,000 structures).
The MAE, RMSE, R2, and Pearson correlation coefficient (Rp) were used to evaluate the accuracy of models in predicting Em. In addition, the softening of the potentials was assessed by the slope of the linear regression between the target property and prediction.
The ability of models to predict a correct geometry of the migration pathway was assessed by calculating the path averaged MAE (PA-MAE) of the trajectory optimized with surrogate models with respect to the DFT-NEB-derived pathways. The formula for calculating PA-MAE is given in Eq. (4):
where M is the number of images in the trajectory, N is the number of atoms in the supercell, ri and \(\hat{{r}_{i}}\) corresponds to the positions of the i-th atom optimized with DFT and surrogate model, respectively.
Fine-tuning
The pre-trained MACE-MP and SevenNet uMLIPs were fine-tuned on a stratified subsample of the MPLiTrj training and validation data splits (see details in Supplementary Section S2). The performance metrics were calculated using the test subset.
The uMLIPs were fine-tuned by minimizing the mean squared error loss function. The model with the lowest loss on the validation subset was used for the benchmark. For the MACE model, the loss function includes the energy and force terms, weighted by 1.0 and 10.0, respectively. For the SevenNet model, the loss function includes the energy, force, and stress tensor terms, weighted by 1.0, 10.0, and 10−6, respectively. Both models were trained with a learning rate of 0.001. The batch size of 16 and 32 samples was used for the MACE and SevenNet models, respectively.
Crystal structures
The ionic transport properties were calculated for Li-containing crystal structures retrieved from the Materials Project (MP) database33. The selection criteria included a non-zero bandgap, the presence of lithium, a unit cell volume less than 1500 Å3, up to 100 atoms per unit cell, and the absence of hydrogen. The crystal structures were visualized using VESTA 3.093.
Data availability
The obtained in this work LiTraj dataset: https://github.com/AIRI-Institute/LiTraj.
Code availability
The Python library for calculating percolation barriers of mobile species in crystal structures can be found at: https://github.com/dembart/BVlain. The Python library used for the BVSE-NEB calculations is available at: https://github.com/dembart/ions.
References
Famprikis, T., Canepa, P., Dawson, J. A., Islam, M. S. & Masquelier, C. Fundamentals of inorganic solid-state electrolytes for batteries. Nat. Mater. 18, 1278–1291 (2019).
Zhang, L. et al. A database of ionic transport characteristics for over 29 000 inorganic compounds. Adv. Funct. Mater. 30, 2003087 (2020).
Zhu, Y. et al. Lithium-film ceramics for solid-state lithionic devices. Nat. Rev. Mater. 6, 313–331 (2021).
Nikam, R. D. et al. Near ideal synaptic functionalities in Li ion synaptic transistor using Li3POxSex electrolyte with high ionic conductivity. Sci. Rep. 9, 18883 (2019).
Sau, K. et al. Unlocking the secrets of ideal fast ion conductors for all-solid-state batteries. Commun. Mater. 5, 122 (2024).
Teale, A. M. et al. DFT exchange: sharing perspectives on the workhorse of quantum chemistry and materials science. Phys. Chem. Chem. Phys. 24, 28700–28781 (2022).
Urban, A., Seo, D.-H. & Ceder, G. Computational understanding of Li-ion batteries. npj Computational Mater. 2, 1–13 (2016).
Aksyonov, D. A., Fedotov, S. S., Stevenson, K. J. & Zhugayevych, A. Understanding migration barriers for monovalent ion insertion in transition metal oxide and phosphate based cathode materials: A DFT study. Comput. Mater. Sci. 154, 449–458 (2018).
Fedotov, S. S. et al. Reversible facile Rb+ and K+ ions de/insertion in a KTiOPO4-type RbVPO4F cathode material. J. Mater. Chem. 6, 14420–14430 (2018).
Drozhzhin, O. A. et al. β-NaVP2O7 as a superior electrode material for Na-ion batteries. Chem. Mater. 31, 7463–7469 (2019).
Fedotov, S. S. et al. Titanium-based potassium-ion battery positive electrode with extraordinarily high redox potential. Nat. Commun. 11, 1–11 (2020).
Shraer, S. D. et al. Development of vanadium-based polyanion positive electrode active materials for high-voltage sodium-based batteries. Nat. Commun. 13, 4097 (2022).
He, X., Zhu, Y. & Mo, Y. Origin of fast ion diffusion in super-ionic conductors. Nat. Commun. 8, 15893 (2017).
Mo, Y., Ong, S. P. & Ceder, G. First principles study of the Li10GeP2S12 lithium super ionic conductor material. Chem. Mater. 24, 15–17 (2012).
Deng, Z. et al. Fundamental investigations on the sodium-ion transport properties of mixed polyanion solid-state battery electrolytes. Nat. Commun. 13, 4470 (2022).
Xiao, Y., Miara, L. J., Wang, Y. & Ceder, G. Computational screening of cathode coatings for solid-state batteries. Joule 3, 1252–1275 (2019).
Liu, C., Früchtl, H., Irvine, J. T. & Bühl, M. Computational screening of anode coatings for garnet-type solid-state batteries. Batteries Supercaps 5, e202100357 (2022).
Jónsson, H., Mills, G. & Jacobsen, K. W. Nudged elastic band method for finding minimum energy paths of transitions. In: Classical and quantum dynamics in condensed phase simulations, 385–404 (World Scientific, 1998).
Golov, A. & Carrasco, J. Enhancing first-principles simulations of complex solid-state ion conductors using topological analysis of procrystal electron density. npj Computational Mater. 8, 187 (2022).
Sagotra, A. K., Chu, D. & Cazorla, C. Influence of lattice dynamics on lithium-ion conductivity: A first-principles study. Phys. Rev. Mater. 3, 035405 (2019).
Devi, R., Singh, B., Canepa, P. & Sai Gautam, G. Effect of exchange-correlation functionals on the estimation of migration barriers in battery materials. npj Computational Mater. 8, 160 (2022).
Grasselli, F. Investigating finite-size effects in molecular dynamics simulations of ion diffusion, heat transport, and thermal motion in superionic materials. J. Chem. Phys. 156, 134705 (2022).
He, X., Zhu, Y., Epstein, A. & Mo, Y. Statistical variances of diffusional properties from ab initio molecular dynamics simulations. npj Computational Mater. 4, 18 (2018).
Kahle, L., Marcolongo, A. & Marzari, N. Modeling lithium-ion solid-state electrolytes with a pinball model. Phys. Rev. Mater. 2, 065405 (2018).
Kahle, L., Marcolongo, A. & Marzari, N. High-throughput computational screening for solid-state Li-ion conductors. Energy Environ. Sci. 13, 928–948 (2020).
Rong, Z., Kitchaev, D., Canepa, P., Huang, W. & Ceder, G. An efficient algorithm for finding the minimum energy path for cation migration in ionic materials. J. Chem. Phys. 145, 074112 (2016).
Rutt, A. et al. Expanding the material search space for multivalent cathodes. ACS Appl. Mater. Interfaces 14, 44367–44376 (2022).
Adams, S. & Rao, R. P. High power lithium ion battery materials by computational design. Phys. Status Solidi (a) 208, 1746–1753 (2011).
Chen, H., Wong, L. L. & Adams, S. SoftBV–a software tool for screening the materials genome of inorganic fast ion conductors. Acta Crystallogr. Sect. B Struct. Sci. Cryst. Eng. Mater. 75, 18–33 (2019).
Avdeev, M., Sale, M., Adams, S. & Rao, R. P. Screening of the alkali-metal ion containing materials from the inorganic crystal structure database (ICSD) for high ionic conductivity pathways using the bond valence method. Solid State Ion. 225, 43–46 (2012).
Fedotov, S. S. et al. Crystallochemical tools in the search for cathode materials of rechargeable Na-ion batteries and analysis of their transport properties. Solid State Ion. 314, 129–140 (2018).
Dembitskiy, A. D., Aksyonov, D. A., Abakumov, A. M. & Fedotov, S. S. NH4+-based frameworks as a platform for designing electrodes and solid electrolytes for Na-ion batteries: A screening approach. Solid State Ion. 374, 115810 (2022).
Jain, A. et al. Commentary: The materials project: A materials genome approach to accelerating materials innovation. APL Mater. 1, 011002 (2013).
Curtarolo, S. et al. Aflow: An automatic framework for high-throughput materials discovery. Computational Mater. Sci. 58, 218–226 (2012).
Kirklin, S. et al. The open quantum materials database (OQMD): assessing the accuracy of DFT formation energies. npj Computational Mater. 1, 1–15 (2015).
Merchant, A. et al. Scaling deep learning for materials discovery. Nature 624, 80–85 (2023).
Glass, C. W., Oganov, A. R. & Hansen, N. USPEX-evolutionary crystal structure prediction. Comput. Phys. Commun. 175, 713–720 (2006).
Wang, Y., Lv, J., Zhu, L. & Ma, Y. Crystal structure prediction via particle-swarm optimization. Phys. Rev. B 82, 094116 (2010).
Kim, S., Noh, J., Gu, G. H., Aspuru-Guzik, A. & Jung, Y. Generative adversarial networks for crystal structure prediction. ACS Cent. Sci. 6, 1412–1420 (2020).
Goodall, R. E., Parackal, A. S., Faber, F. A., Armiento, R. & Lee, A. A. Rapid discovery of stable materials by coordinate-free coarse graining. Sci. Adv. 8, eabn4117 (2022).
Wang, J. et al. Design principles for nasicon super-ionic conductors. Nat. Commun. 14, 5210 (2023).
Ling, C. A review of the recent progress in battery informatics. npj Computational Mater. 8, 33 (2022).
Sendek, A. D. et al. Holistic computational structure screening of more than 12000 candidates for solid lithium-ion conductor materials. Energy Environ. Sci. 10, 306–320 (2017).
Hargreaves, C. J. et al. A database of experimentally measured lithium solid electrolyte conductivities evaluated with machine learning. npj Computational Mater. 9, 9 (2023).
Artrith, N. et al. Best practices in machine learning for chemistry. Nat. Chem. 13, 505–508 (2021).
Wang, Z., Han, Y., Cai, J., Chen, A. & Li, J. An end-to-end artificial intelligence platform enables real-time assessment of superionic conductors. SmartMat 4, e1183 (2023).
Katcho, N. A. et al. An investigation of the structural properties of Li and Na fast ion conductors using high-throughput bond-valence calculations and machine learning. J. Appl. Crystallogr. 52, 148–157 (2019).
Zhao, Q., Avdeev, M., Chen, L. & Shi, S. Machine learning prediction of activation energy in cubic Li-argyrodites with hierarchically encoding crystal structure-based (HECS) descriptors. Sci. Bull. 66, 1401–1408 (2021).
Nakayama, M. et al. Data-driven materials exploration for Li-ion conductive ceramics by exhaustive and informatics-aided computations. Chem. Rec. 19, 771–778 (2019).
Eremin, R. A., Zolotarev, P. N., Golov, A. A., Nekrasova, N. A. & Leisegang, T. Ionic transport in doped solid electrolytes by means of DFT modeling and ML approaches: a case study of Ti-doped KFeO2. J. Phys. Chem. C. 123, 29533–29542 (2019).
Jalem, R., Nakayama, M. & Kasuga, T. An efficient rule-based screening approach for discovering fast lithium ion conductors using density functional theory and artificial neural networks. J. Mater. Chem. A 2, 720–734 (2014).
Shapeev, A. V. Moment tensor potentials: A class of systematically improvable interatomic potentials. Multiscale Model. Simul. 14, 1153–1173 (2016).
Bartók, A. P., Kondor, R. & Csányi, G. On representing chemical environments. Phys. Rev. B Condens. Matter Mater. Phys. 87, 184115 (2013).
Behler, J. & Parrinello, M. Generalized neural-network representation of high-dimensional potential-energy surfaces. Phys. Rev. Lett. 98, 146401 (2007).
Chen, C. & Ong, S. P. A universal graph deep learning interatomic potential for the periodic table. Nat. Computational Sci. 2, 718–728 (2022).
Deng, B. et al. CHGNet as a pretrained universal neural network potential for charge-informed atomistic modelling. Nat. Mach. Intell. 5, 1031–1041 (2023).
Park, Y., Kim, J., Hwang, S. & Han, S. Scalable parallel algorithm for graph neural network interatomic potentials in molecular dynamics simulations. J. Chem. Theory Comput. 20, 4857–4868 (2024).
Batatia, I. et al. A foundation model for atomistic materials chemistry. Preprint at https://arxiv.org/abs/2401.00096 (2023).
Duval, A. et al. A hitchhiker’s guide to geometric GNNs for 3D atomic systems. Preprint at https://arxiv.org/abs/2312.07511 (2023).
Xie, S. R., Honrao, S. J. & Lawson, J. W. High-throughput screening of Li solid-state electrolytes with bond valence methods and machine learning. Chem. Mater. 36, 9320–9329 (2024).
Aksyonov, D., Boev, A., Fedotov, S. & Abakumov, A. Computational insights into ionic conductivity of transition metal electrode materials for metal-ion batteries-a review. Solid State Ion. 393, 116170 (2023).
Cherkashchenko, I. R. et al. Reversible multielectron redox activity of the anti-NASICON-type phosphate LiNbV(PO4)3 towards lithium and sodium intercalation. Dalton Trans. 53, 16918–16928 (2024).
Ward, L. et al. Matminer: An open source toolkit for materials data mining. Computational Mater. Sci. 152, 60–69 (2018).
Ward, L. et al. Including crystal structure attributes in machine learning models of formation energies via voronoi tessellations. Phys. Rev. B 96, 024104 (2017).
Batzner, S. et al. E (3)-equivariant graph neural networks for data-efficient and accurate interatomic potentials. Nat. Commun. 13, 2453 (2022).
Musaelian, A. et al. Learning local equivariant representations for large-scale atomistic dynamics. Nat. Commun. 14, 579 (2023).
Deng, B., Choi, Y. & Zhong, P. et al. Systematic softening in universal machine learning interatomic potentials. npj Comput Mater. 11, 9 (2025).
Kato, Y., Hori, S. & Kanno, R. Li10GeP2S12-type superionic conductors: synthesis, structure, and ionic transportation. Adv. Energy Mater. 10, 2002153 (2020).
Franchini, C., Reticcioli, M., Setvin, M. & Diebold, U. Polarons in materials. Nat. Rev. Mater. 6, 560–586 (2021).
Richards, W. D., Miara, L. J., Wang, Y., Kim, J. C. & Ceder, G. Interface stability in solid-state batteries. Chem. Mater. 28, 266–273 (2016).
Zhu, Y., He, X. & Mo, Y. First principles study on electrochemical and chemical stability of solid electrolyte–electrode interfaces in all-solid-state Li-ion batteries. J. Mater. Chem. A 4, 3253–3266 (2016).
Kobayashi, H., Yuan, G., Gambe, Y. & Honma, I. Effective Li3AlF6 surface coating for high-voltage lithium-ion battery operation. ACS Appl. Energy Mater. 4, 9866–9870 (2021).
Liu, B. et al. High-throughput computational screening of Li-containing fluorides for battery cathode coatings. ACS Sustain. Chem. Eng. 8, 948–957 (2020).
Du, Y. A. & Holzwarth, N. Li ion diffusion mechanisms in the crystalline electrolyte γ-Li3PO4. J. Electrochem. Soc. 154, A999 (2007).
Laskowski, F. A., McHaffie, D. B. & See, K. A. Identification of potential solid-state Li-ion conductors with semi-supervised learning. Energy Environ. Sci. 16, 1264–1276 (2023).
Choi, B. et al. Dielectric and ionic conduction properties in LiYF4 single crystals. Mater. Des. 21, 567–570 (2000).
Panchal, A. A. et al. Compatibility of halide bilayer separators for all-solid-state batteries. ACS Energy Lett. 9, 5935–5944 (2024).
Smidstrup, S., Pedersen, A., Stokbro, K. & Jónsson, H. Improved initial guess for minimum energy path calculations. J. Chem. Phys. 140, 214106 (2014).
Devi, R., Butler, K. T. & Sai Gautam, G. Optimal pre-train/fine-tune strategies for accurate material property predictions. npj Computational Mater. 10, 300 (2024).
He, B. et al. CAVD, towards better characterization of void space for ionic transport analysis. Sci. Data 7, 153 (2020).
Wong, L. L. et al. Bond valence pathway analyzer-an automatic rapid screening tool for fast ion conductors within softBV. Chem. Mater. 33, 625–641 (2021).
Shen, J.-X., Horton, M. & Persson, K. A. A charge-density-based general cation insertion algorithm for generating new Li-ion cathode materials. npj Computational Mater. 6, 161 (2020).
Xu, M., Ding, J. & Ma, E. One-dimensional stringlike cooperative migration of lithium ions in an ultrafast ionic conductor. Appl. Phys. Lett. 101, 031901 (2012).
Jalem, R. et al. Concerted migration mechanism in the Li ion dynamics of garnet-type Li7La3Zr2O12. Chem. Mater. 25, 425–430 (2013).
Jang, S.-H., Jalem, R. & Tateyama, Y. Predicting room-temperature conductivity of Na-ion super ionic conductors with the minimal number of easily-accessible descriptors. Adv. Energy Sustain. Res. 5, 2400158 (2024).
Ong, S. P. et al. Python materials genomics (pymatgen): A robust, open-source python library for materials analysis. Comput. Mater. Sci. 68, 314–319 (2013).
Kresse, G. & Furthmüller, J. Efficient iterative schemes for ab initio total-energy calculations using a plane-wave basis set. Phys. Rev. B 54, 11169 (1996).
Perdew, J. P., Burke, K. & Ernzerhof, M. Generalized gradient approximation made simple. Phys. Rev. Lett. 77, 3865 (1996).
Larsen, A. H. et al. The atomic simulation environment-a python library for working with atoms. J. Phys. Condens. Matter 29, 273002 (2017).
Pedregosa, F. et al. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Chen, T. & Guestrin, C. XGBoost: A scalable tree boosting system. In: Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, KDD ’16, 785–794 (ACM, 2016). https://doi.org/10.1145/2939672.2939785.
Eremin, R. A. et al. Graph neural networks for predicting structural stability of Cd- and Zn-doped γ-CsPbI3. Comput. Mater. Sci. 232, 112672 (2024).
Momma, K. & Izumi, F. Vesta: a three-dimensional visualization system for electronic and structural analysis. J. Appl. Crystallogr. 41, 653–658 (2008).
Acknowledgements
A.D., D.A., and S.F. acknowledge the financial support of Russian Science Foundation project No. 24-73-10204.
Author information
Authors and Affiliations
Contributions
A.D. proposed the original design of the study and developed the BVlain and ions Python libraries (Skoltech). A.D. collected the dataset under the supervision of R.E. (AIRI), A.D. fitted and evaluated the models under the supervision of I.H. (AIRI), A.D. wrote the original draft with significant input from R.E. and D.A. D.A. suggested the idea of calculating the DFT part of the dataset and validated the calculation results, D.A. and R.E. curated the revision of the manuscript, D.A., R.E., and S.F. made substantial contributions to the discussion of the results. D.A., S.F., and S.B. curated the research. All the authors discussed and critically revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dembitskiy, A.D., Humonen, I.S., Eremin, R.A. et al. Benchmarking machine learning models for predicting lithium ion migration. npj Comput Mater 11, 131 (2025). https://doi.org/10.1038/s41524-025-01571-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41524-025-01571-z