Abstract
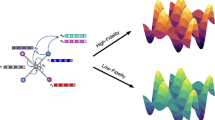
The rapid development of foundation potentials (FPs) in machine learning interatomic potentials demonstrates the possibility for generalizable learning of the universal potential energy surface. The accuracy of FPs can be further improved by bridging the model from lower-fidelity datasets to high-fidelity ones. In this work, we analyze the challenge of this transfer learning (TL) problem within the CHGNet framework. We show that significant energy scale shifts and poor correlations between GGA and r2SCAN hinder cross-functional transferability. By benchmarking different TL approaches on the MP-r2SCAN dataset, we demonstrate the importance of elemental energy referencing in the TL of FPs. By comparing the scaling law with and without the pre-training on a low-fidelity dataset, we show that significant data efficiency can still be achieved through TL, even with a target dataset of sub-million structures. We highlight the importance of proper TL and multi-fidelity learning in creating next-generation FPs on high-fidelity data.
Similar content being viewed by others
Introduction
Atomistic simulations provide a powerful framework for predicting and virtually screening material properties and have led to multiple predictions of interesting functional materials1,2,3. These simulations are enabled by accurate determination of the potential energy surface (PES) as a function of atomic positions, permitting prediction of stability properties, reaction mechanisms, and dynamic behavior4,5,6,7.
Historically, two main approaches have been used to model the PES: ab-initio quantum chemical calculations and empirical force fields8. Ab-initio quantum chemical calculations such as density functional theory (DFT) provide high accuracy and generality, but at a substantial computational cost. Furthermore, this cost scales rapidly with system size, typically, \(\sim {\mathcal{O}}({N}_{e}^{3})\) or \({\mathcal{O}}({N}_{e}\log {N}_{e})\) with Ne the number of electrons9,10, thereby limiting the achievable length and time scales. In contrast, empirical force fields typically trade accuracy and generality for speed, limiting their applicability in complex systems.
To address these limitations, surrogate energy models such as machine learning interatomic potentials (MLIPs) have been developed to accelerate atomistic simulations. MLIP methods aim to disrupt the long-standing trade-off between accuracy and efficiency, achieving near training set accuracy while offering a computational cost reduction of several orders of magnitude compared to traditional DFT methods11 and maintaining \({\mathcal{O}}(N)\) computational efficiency, where N is the number of atoms12.
MLIPs are parametrized to reproduce energies from ab-initio quantum mechanical calculations, such as DFT. The total energy of a material system is decomposed and predicted through a learnable mapping of atomic positions and chemical species, where each atom’s contribution is determined by its surrounding local atomic configuration within a defined cutoff radius:
The learnable function ϕ maps the position vectors \({\{{\overrightarrow{r}}_{j}\}}_{i}\) and chemical species \({\{{C}_{j}\}}_{i}\) of neighboring atoms j to the energy contribution of atom i. Forces \(\{{\hat{{\boldsymbol{f}}}}_{i}\}\) are derived as the negative gradient of the total energy with respect to atomic coordinates. The choice of design features ϕ is crucial for MLIPs to encode the system’s physical and chemical properties, such as using equivariant feature encoding13,14 and including atomic charge information15,16.
Recently, foundation potentials (FPs) trained on millions of DFT calculations demonstrate promising transferability in atomic simulations across diverse chemical spaces. The FPs such as M3GNet17, CHGNet15, MACE-MP-018, SevenNet-MF-019, and Orb20 have been developed from open-source materials databases such as the Materials Project21,22 and Alexandria23. Industry FPs such as GNoME24, MatterSim25, and EquiformerV2-OMAT26 demonstrate improved PES predictability with larger data and model sizes in various downstream materials modeling tasks such as phonon spectra prediction, phase diagram construction, catalyst screening, and molecular dynamics simulations27,28,29,30,31.
Despite these successes in improving models and data, there remain challenges for further improvements of FPs. One significant issue reported by Deng et al.27 shows a consistent underprediction of energies and forces in FPs27, which calls for improved sampling in FP training datasets. The predominant approach to generate FP datasets relies on DFT calculations using generalized gradient approximations (GGAs), limiting FPs to GGA-level accuracy and posing potential challenges for migrating to higher-accuracy functionals like meta-GGAs. Recently, Kaplan et al.32 released the MatPES dataset that incorporates regularized strongly constrained and appropriately normed (r2SCAN) meta-GGA functional calculations, which opens the possibility for FPs to migrate to high level of theory. See ref. 33 for a definition of GGAs and meta-GGAs and Ref. 34 for an overview of their well-established limitations in describing crystalline and molecular systems.
In this work, we discuss the challenges and practical approaches that help better understand the fine-tuning process in FPs, particularly when dealing with multi-fidelity data transferability across different functionals. Successfully achieving this multi-fidelity data transferability is crucial, as it can enable more accurate and efficient atomistic simulations without the substantial computational costs typically associated with high-fidelity calculations. By showing the correlation between the labels from different levels of theory, we emphasize the importance of training at the right scale through energy referencing when conducting transfer learning.
Results
Data challenges in existing foundation potentials
An essential component in building improved FPs comes from reliable datasets. The current FP datasets applicable to crystalline materials are predominantly composed of GGA and GGA + U-level DFT calculations15,17,18,20. While GGA-based training data is widely available and computationally efficient to generate, several limitations of GGA are known35,36,37 and other functionals are now available38,39,40. A widely used method to alleviate some of the self-interaction in GGA is the Hubbard U correction41, which adds an energy correction to localized electron states (e.g., d or f orbitals). The use of + U is particularly important when dealing with metal oxidation/reduction in formation enthalpies, reaction energies, or electrochemical potentials36,42. At the same time, the application of + U is not appropriate for metallic systems where electron delocalization is appropriate. Because of these conflicting requirements, compatibility schemes between GGA and GGA + U have been designed43 and some datasets contain a mixture of GGA and GGA + U calculations. We call attention to three data challenges in existing FPs, which were primarily trained with a mixture of GGA/GGA + U DFT calculations.
Firstly, GGA/GGA + U exhibit lower transferability across chemical bonding environments37. The Perdew-Burke-Ernzerhof (PBE) GGA44 is found to have a mean absolute error (MAE) of 194 meV/atom dominated by the large error in oxides and strongly bound systems, in a large-scale test on the formation energy of 987 compounds45. In contrast, the SCAN meta-GGA functional developed by Sun et al.38 predicts formation energies with an MAE of 84 meV/atom. Isaacs and Wolverton46 also demonstrate that SCAN is more accurate in predicting formation energy for strongly bound compounds, crystal volumes, magnetism, and band gaps, as compared to the PBE GGA. The r2SCAN39 revision of the SCAN meta-GGA balances numerical stability with high general accuracy45 and has therefore become the preferred method to evaluate thermophysical properties of materials45,47,48. While the demonstrated prediction errors in Ref. 45 are high, it is worth noting that many of the compounds included have formation reactions from molecular species such as H2, N2, O2, and thereby are more similar to cohesive energies. When evaluating only solid-state reactions, energy errors are typically smaller for GGA49.
Secondly, the application of the Hubbard U correction to mitigate self-interaction errors in GGA is inherently semi-empirical and non-universal. GGA + U fails to predict accurate energy differences between some compounds with localized electronic states and those with delocalized electronic states43. There is also no precise definition of an “optimal” U, and approaches such as the linear response method50 suggest that such an optimal U would be system-dependent. However, the GGA/GGA + U FP datasets were generated using the same U value for each element regardless of the local environment or formal valence state, calibrated to minimize discrepancies between DFT-calculated oxidation energies and experimental measurements for a limited number of 3d transition metal oxides42,43.
Thirdly, to correct for some of the self-interaction error in GGA which is particularly large when calculating the energy of reactions that reflect charge transfer such as oxide formation enthalpies, an ad hoc scheme of mixing GGA and GGA + U calculations is typically used to bridge the gap between GGA and GGA + U43,51. Such coarse-grained, non-universal adjustments can potentially cause issues when fitting a FP, such as sudden jumps of potential energy at the scale of a few hundred meV per atom when moving between training data computed with these mixing schemes. Last, there is no corresponding mixing scheme applied to the GGA/GGA + U interatomic forces and stresses. This may be less of an issue as both are derivative properties of a given functional, and thus should be independent of the energy scale of the underlying DFT approximation. However, this has not been formally verified.
Overall, the use of approximate exchange-correlation functionals, combined with the non-universality of Hubbard U corrections and compatibility adjustments, leads to less accurate and somewhat noisy data within the GGA/GGA + U framework. Such data noise makes it challenging for graph neural network models (GNNs) to accurately learn and capture the underlying interactions within materials.
Cross-functional transferability challenges in foundation potentials
One possible solution to overcome the challenges of GGA and GGA + U is to shift the FP training and benchmarking dataset to DFT calculations performed with higher-fidelity functionals. These higher-fidelity calculations come with higher computational costs, leading to challenges in constructing datasets on a substantial scale. One possible solution is to leverage existing lower-fidelity GGA and GGA + U calculations and existing pre-trained FPs as a starting point.
There are three main strategies to achieve explicit or implicit transferability between multi-fidelity DFT datasets: transfer learning, multi-fidelity learning, and mixed multi-fidelity training.
Transfer learning (TL) involves pre-training a large neural network on extensive lower-fidelity datasets. The pretrained weights from this network are then transferred to initialize machine-learning tasks on smaller, higher-fidelity datasets. This approach is both computationally efficient and data-efficient52,53. However, if the correlation between the two different fidelity datasets is not strong enough, TL is not effective and can even deteriorate the learning performance, known as negative transfer54.
Multi-fidelity learning can be conducted either at the feature (input) level or at the label (output) level55, i.e., low-fidelity data is utilized as input features to predict high-fidelity data, or the task of learning high-fidelity data can be transformed into learning the difference between high-fidelity and low-fidelity data, an approach known as Δ-machine learning56,57. Multi-fidelity learning tends to be more computationally expensive than TL58. When applying multi-fidelity learned models to make real predictions for unknown cases, one must first calculate low-fidelity data to obtain input features (input level) or use it to add the predicted difference to get the final high-fidelity prediction (output level).
Mixed multi-fidelity training aims to simultaneously learn and predict datasets of varying fidelity levels. Chen et al.59 encoded the fidelity of each dataset and embedded the dataset type as a vector in the global state feature input to the M3GNet model for band gap prediction. Ko and Ong60 adopted this method to construct highly accurate GNN-based interatomic potentials for two model systems—silicon and water. Allen et al.61 used meta-learning techniques to build pre-trained potentials that simultaneously incorporate information from multiple large organic datasets, calculated at different levels of theory. Kim et al.19 developed a high-fidelity MLIP by one-hot encoding each fidelity, concatenating it to the scalar part of the input node feature at each linear layer, and adding different atomic energy shift scale blocks for each fidelity database to the SevenNet model. Similar to TL, mixed-fidelity training tends to be computationally expensive when additional poorly correlated data are added to the trained model.
Each of the three strategies presents its own advantages and challenges. So far, no clear evidence exists that TL consistently outperforms multi-fidelity learning or mixed multi-fidelity approaches, or vice versa. In this work, we focus on how to tackle the transferability challenges of efficient TL across GGA/GGA + U mixed data and r2SCAN data in the CHGNet model, though our conclusion should hold more generally for other FPs.
MP-r2SCAN dataset
We use a r2SCAN dataset, MP-r2SCAN, parsed from Materials Project21,22 r2SCAN relaxation trajectories, for high-fidelity training tasks. Following the data parsing criteria described in Data preparation, we obtain 34,927 material IDs with 238,247 structures. Compared to the MPtrj Dataset15, which has 145,923 materials IDs with 1,580,395 structures, the MP-r2SCAN is significantly smaller in size.
Figure 1 a presents the element distribution in the MP-r2SCAN dataset with a total of 238,247 structures. The color of each element indicates the total number of times each element is present in the MP-r2SCAN dataset, with a lower cutoff of 1000. Elements with 1000 or fewer occurrences all share the same color. The MP-r2SCAN dataset covers 88 elements in the periodic table.
a Element distribution of the MP-r2SCAN dataset of 238,247 structures. The color indicates the total number of occurrences of an element in the MP-r2SCAN dataset with a lower cutoff of 1000. b Total Energy of materials computed from GGA/GGA+U vs. r2SCAN functionals. Each point represents a material with a materials ID that has r2SCAN calculations in Materials Project, with the x-axis showing the total energy after r2SCAN structure relaxation and the y-axis showing the total energy after GGA/GGA+U structure relaxation. The marginal histograms on the top and right illustrate the distributions of total energies for the same collection of materials, as calculated by r2SCAN and GGA/GGA + U, respectively. c, d Feature importance in the formation energy differences between GGA/GGA + U mixing and r2SCAN. Each element is treated as a feature, with its importance indicated by colors on the periodic table. Higher values correspond to greater importance and therefore larger energy difference between GGA/GGA + U and r2SCAN. Panel c presents the feature importance when anion and compatibility corrections are included in the mixed GGA/GGA + U data, and d presents the feature importance without these adjustments. Compositional corrections are applied primarily to pnictogens, chalcogens, and halogens.
Energy differences across two functionals
Machine learning transferability can be quantified by assessing the correlations between the source and target datasets62. To investigate the feasibility and effectiveness of TL between DFT functionals, we analyze the scale of the total energy differences between r2SCAN and GGA/GGA + U.
Figure 1 b presents the comparison of the relaxed total energies calculated using r2SCAN (x-axis) and GGA/GGA + U (y-axis), which represent the training label of most FPs. In Fig. 1b, each point represents a single compound from the Materials Project, and the corresponding GGA/GGA + U energies have applied anion and compatibility corrections63. The marginal histograms on the top and right side show the distributions of energies calculated using r2SCAN and GGA/GGA + U, respectively, for all r2SCAN materials IDs in Materials Project. As depicted in Fig. 1b, the total energy of r2SCAN and GGA/GGA + U are distributed on different scales. The shift from the GGA/GGA + U to r2SCAN is at the scale of 0–70 eV/atom, which is significantly larger than the energy accuracy of FPs (~ 30meV/atom), indicating these r2SCAN energy labels are not directly transferrable without proper reference or normalization.
These eV/atom scale energy shifts between functionals are related to the ambiguity in the Kohn-Sham energy levels which have an arbitrary reference energy64,65,66. These energy shifts are well understood in electronic structure theory and do not contribute to any physical quantities due to the cancellation of energy references in any physical property. The total energy itself is not a physically measurable quantity, as it is “gauge dependent” on the vacuum level, but energy differences such as the cohesive energy are measurable and gauge invariant67. Because MLIPs are typically trained on absolute total energies, these eV/atom scale energy differences from GGA/GGA + U and r2SCAN can cause significant challenges in TL.
One method to remove the significant total energy shifts is by fitting the MLIPs with physical quantities such as formation energies, which has been shown to be easier to transfer in crystal graph attention networks52,68. The formation energies describe the strengths of the interactions that form the compound from pure elemental phases and are better correlated between different functionals than the total energy labels, although small deviations can still be present due to the different levels of accuracy.
To determine which elements contribute most to the formation energy differences between r2SCAN and GGA/GGA + U calculations, we queried the formation energies from Materials Project and fitted decision tree models on the formation energy differences through scikit-learn69. The input to this model is the compositional fraction matrix of all materials with r2SCAN materials IDs in Materials Project, and the target variable is the formation energy difference between the two functionals. We calculated the feature importance (see Feature importance) for each element and plotted the strength of the importance through the color bar in the periodic table in Fig. 1c and d. The importance of a feature is computed as the normalized total reduction of the criterion brought by that feature. The higher the value the more important the feature. Figure 1c presents the feature importance with GGA/GGA + U mixing and anion corrections included, and Fig. 1d includes the same analysis but with uncorrected GGA/GGA + U formation energies.
In Fig. 1c, we observe that d-block elements such as V, Cr, Mn, Fe, Co, Ni, Mo, and W exhibit high importance, indicating they significantly contribute to the formation energy differences between GGA/GGA + U and r2SCAN. These are precisely the elements for which Hubbard U corrections and compatibility adjustments are applied in transition metal oxides and fluorides. Similarly, p-block elements with high importance—O, F, S, Cl, Se, Br, and Te—also undergo compatibility corrections when they serve as anions in compounds. Notably, Cl exhibits a very high feature importance. We can attribute the relatively higher feature importance of Cl to two sources: (i) the compatibility scheme imposed on GGA/GGA + U energies places the second largest correction ( −0.614 eV/atom in magnitude) to Cl, second only to oxides ( − 0.687 eV/atom in magnitude); (ii) PBE struggles to describe the weaker covalency and van der Waals interactions typical of ionic crystals70, whereas r2SCAN describes both covalent and ionic bonding reasonably well39 and improves the description of medium-range van der Waals interactions71,72. The differences in Fig. 1c and d show clearly that the removal of the corrections scheme almost eliminates the higher feature importance of the chalcogens and halogens seen in Fig. 1c. Without the energy correction scheme, the eight transition metals, O, and F remain a higher feature importance (see Fig. 1d).
TL with different atomic reference energies
Shifting the PES with a constant value for each element is an effective and commonly used approach in training GNN-based MLIPs. As described in Fig. 2a, in CHGNet and other models like M3GNet, NequIP13 and CACE14, the prediction of total energies (per atom) is divided into two parts: EAtomRef and EGNNs17,73. First, the composition row vector celem and atomic reference energies (AtomRef) Eelem are obtained, and their dot product gives EAtomRef. The composition vector celem represents the fraction of each element in the structure, and in CHGNet, its dimension is 1 × 94. Next, a composition model is used to fit a linear regression of total energies, where Eelem are the weights:
Here, A is the composition matrix obtained by stacking celem for all structures in the training set, and Etotal is the matrix of total energies. Subsequently, the remaining fine-grained energy is predicted by GNNs. Overall, the total energy prediction of a structure can be expressed using Etotal = celem ⋅ Eelem + EGNNs. Both AtomRef, which represent the weights of the composition model, and GNNs can be trainable.
a Schematic representation of the role and application of AtomRef in calculating total energies. The energy contribution from AtomRef is obtained by taking the dot product of the composition row vector (with LiCoO2 used here as an example) and the AtomRef vector. b The correlation between the scaled energies of GGA/GGA + U and r2SCAN (total energies with the respective AtomRefs subtracted). The marginal histograms on the top and right illustrate the distributions of r2SCAN and GGA/GGA + U scaled energies, respectively, for the same collection of materials.
For cross-functional TL on a FP with a fitted AtomRef from GGA/GGA + U total energies, one can refit the FP’s AtomRef to shift the FP’s energy to the scale of new DFT labels and, in principle, improve the correlation between pre-training and fine-tuning datasets. Refitting the AtomRef essentially replaces the fitted GGA/GGA + U AtomRef with the fitted r2SCAN AtomRef and shifts the FP’s predicted energy scale to r2SCAN. Figure 2b shows that, after replacing the AtomRef, a stronger correlation between GGA/GGA + U and r2SCAN total energies can be achieved.
Indeed, the Pearson’s correlation coefficient ρ improves from 0.0917 between the unmodified GGA/GGA + U and r2SCAN datasets to 0.9250 between the r2SCAN energies (with r2SCAN AtomRef subtracted) and the GGA/GGA + U energies (with GGA/GGA + U AtomRef subtracted).
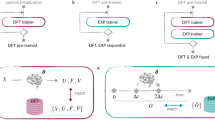
To compare in more detail how well various strategies for aligning energies from different functionals perform, we performed an ablation study using four training strategies to either pre-train or fine-tune CHGNet on the MP-r2SCAN dataset.
In Method 1 (Training from scratch), we first fitted AtomRef using the r2SCAN total energies, randomly initialized the GNN parameters of CHGNet, and then trained the GNNs on the MP-r2SCAN dataset while keeping the r2SCAN AtomRef frozen.
In Method 2 (TL with trainable AtomRef), we started from the GGA/GGA + U-pre-trained CHGNet and allowed both the GNN parameters and the AtomRef to be trainable during TL. In this manner, the AtomRef, initially set to the fitted GGA/GGA + U AtomRef, was gradually updated throughout the TL process.
In Method 3 (TL with frozen AtomRef), we again used the GGA/GGA + U-pre-trained CHGNet as the starting point, but only the GNN parameters were allowed to be trainable during TL. As a result, the AtomRef remained fixed at the fitted GGA/GGA + U AtomRef, forcing the GNNs to transfer and accommodate to the large energy differences observed in Fig. 1b.
In Method 4 (TL with r2SCAN AtomRef), we first replaced the GGA/GGA + U AtomRef in the pre-trained CHGNet model with the r2SCAN AtomRef, and then performed TL on the GNNs while keeping the r2SCAN AtomRef frozen.
Table 1 presents the MAEs on the test set for energy, force, stress, and magnetic moment (magmom) predictions (see Data preparation for details on data splitting). Methods 2 and 3 (TL with trainable and frozen AtomRef, respectively) yield similar performance across all metrics, with Method 1 (Training from scratch) achieving a comparable energy error (27 meV/atom) but reduced force (45 meV/Å) and stress error (0.239 GPa). This suggests that without properly shifting the reference energy, neither Method 2 nor Method 3 benefits from the GGA/GGA + U pre-training. In contrast, Method 4 (TL with r2SCAN AtomRef) attained the lowest MAEs for energy, force, and stress, indicating that the optimal approach to fine-tuning MLIPs is to first shift the reference energy and then train the GNNs.
Figure 3 shows the model training gradients and training errors vs. epochs for Method 3 and Method 4 during the TL. Figure 3a illustrates the range of gradient values for several representative model layers. Gradient values are recorded every 1/10 of an epoch for these model layers during the first transfer learning epoch. We observe that Method 3 without refitting AtomRef exhibits gradient magnitudes at least one order larger than those of Method 4 with refitting. Figure 3b and c show the evolution of energy MAE during the full training process of 50 epochs, without and with AtomRef adjustments, respectively. Figure 3b displays larger initial and final energy MAE, indicating a less effective training process. In contrast, Fig. 3c demonstrates that refitting AtomRef results in a more stable and reliable training history.
a Gradient values recorded every 1/10 of an epoch for various model layers during the first transfer learning epoch, comparing models with and without AtomRef refitting. The layers include “AtomEmb” (atom embedding), “BondEmb” (bond embedding), “AngleEmb” (angle embedding), “AtomConv0_W0” and “AtomConv3_W3” (weights of the two-body atom convolution layers), “BondConv0_W0” and “BondConv2_W3” (weights of the two-body bond convolution layers), and “MLP_Layer0” (weights of the first layer in the multi-layer perceptron). b Energy training history for Method 3, showing the lowest energy MAE of 18.37 meV/atom at the last epoch. c Energy training history for Method 4, showing the lowest energy MAE of 11.82 meV/atom at the last epoch.
Stability prediction from MLIPs
As a more stringent prediction test, we evaluate relative stability of compounds through the convex hull construction. Relative stability of a compound can be measured by its decomposition energy, calculated by the total energy difference between a given compound and its competing compounds in a specific chemical space. This is a more stringent test than measuring MAE, as the scale of decomposition energy is small and relies on significant error cancellation in DFT74.
Figure 4 presents the general workflow for predicting decomposition energy. Predicting decomposition energy with FPs is particularly challenging as it depends not only on the energy of a single material but also on that of the neighboring competing phases in a phase diagram75. The physical outcome of decomposition energy is binary with negative values indicating stable compounds and positive values indicating unstable or metastable compounds. As such, small non-systematic energy errors from MLIPs will easily alter the stable entries in the phase diagram, by changing the decomposition energy from small negative values to positive values and vice versa. This issue is further exacerbated by the fact that machine learning models exhibit poorer error cancellation compared to DFT74.
The left plot shows a schematic of a convex hull energy diagram constructed using r2SCAN DFT-calculated data, providing decomposition energy values based on competing phases identified in the DFT phase diagram (e.g., for a2, the competing phases are a1 and a3; for a4, they are a3 and a6). The right plot schematically shows the convex hull constructed by CHGNet-relaxed energies. The decomposition energy and model-identified competing phases differ from DFT.
We constructed all phase diagrams in the chemical space of our dataset using r2SCAN DFT data and calculated the decomposition energy as the ground truth. A similar phase diagram can be constructed by the fine-tuned CHGNet, which allows the determination of CHGNet predicted decomposition energy. The initial configurations for all structures are sourced from Materials Project and further relaxed using the pre-trained or fine-tuned CHGNet models of corresponding methods. This process relies solely on the FP’s capability to obtain relaxed energies and relative stabilities between polymorphs, without requiring additional information from the DFT phase diagram.
Table 1 also presents benchmark results for the decomposition energy prediction MAEs of four methods on the MP-r2SCAN test set (see Data preparation for data splitting). The MAEs of Methods 2 and 3 (41.22 and 38.54 meV/atom, respectively) are slightly larger than that of Method 1 (37.44 meV/atom), again indicating no benefit from conventional TL methods. In contrast, Method 4, which uses r2SCAN-specific AtomRef, achieves an MAE of 23.66 meV/atom, at least 13.5 meV/atom lower than the others. Additionally, Table 1 shows the formation energy MAEs for the pre-trained or fine-tuned CHGNet models, where formation energy is defined as the energy difference between a compound and its constituent elements in their reference states. Method 4 again outperforms the other methods, with an MAE of 29.38 meV/atom, at least 10 meV/atom lower than the others. Method 2 has higher MAEs for both decomposition and formation energies (41.22 and 52.43 meV/atom, respectively) compared to other methods that freeze AtomRef during training, suggesting that a trainable AtomRef may lead to less accurate predictions in practice.
In the prediction of decomposition energies, we also observed that the FP trained with Method 2 and Method 3 exhibited some failed ionic relaxations. Specifically, we found that in Method 2, 40 out of 34,927 relaxations, and in Method 3, 30 out of 34,927 relaxations, resulted in at least one atom being displaced more than 6 Å away from its nearest neighbors, creating an unrealistic atomic configuration that triggered the failure of force field calculations. This is likely due to the unstable PES in the MLIP created by the large gradient updates in TL without shifting the reference energy (see Fig. 3). In contrast, Method 4 – TL with r2SCAN AtomRef, significantly improves prediction accuracy in this complex task of predicting non-intrinsic properties.
Density prediction from MLIPs
Apart from the above benchmarks that use DFT-calculated data as ground truth, we further assess the MLIPs’ ability to predict materials’ density in comparison with experimental values. r2SCAN is expected to outperform GGA in this respect, as GGA functionals are known to more significantly overestimate volumes compared to r2SCAN45. As a result, we expect that the models fine-tuned on r2SCAN data will yield more accurate density predictions than the GGA/GGA + U pretrained CHGNet.
To perform this evaluation, we randomly selected 1000 ordered structures from the Inorganic Crystal Structure Database76 (ICSD) database. For each structure, all lattice vectors were strained by −10%, followed by structure relaxations using each model with a 30 meV/Å relaxation convergence threshold. The converged densities were then compared to the experimental values.
Figure 5 presents the distributions of the ratios between experimental and predicted values for density, for the pretrained CHGNet and models trained by Method 1, Method 2, Method 3, and Method 4. The plots also report the MAE for each method in the upper left corner.
Distribution of the predicted density ratio (experimental / predicted) for 1000 Inorganic Crystal Structure Database (ICSD) structures, shown over the range 0.80–1.20. Results are given for Method 1: training from scratch, Method 2: TL with trainable AtomRef, Method 3: TL with frozen AtomRef, Method 4: TL with r2SCAN AtomRef, and the GGA/GGA + U pretrained CHGNet model. The MAE for density (in g/cm3) for each method is reported in the upper left corner. The dashed vertical line indicates perfect prediction (ratio = 1).
The results show that Method 4 achieves the lowest MAE. Furthermore, the distributions of the experimental-to-predicted ratios demonstrate that the CHGNet model pretrained on GGA tends to underestimate densities to a greater extent, whereas Method 4 yields distributions more tightly centered around the ideal value (ratio = 1) than the other methods. These findings indicate that our transfer learning approach from GGA to r2SCAN is effective, and that models trained using Method 4 with r2SCAN data have greater potential for accurate real-world volume and density predictions.
Scaling law on transfer learning
To evaluate the data efficiency improvement of Method 4, we analyzed its scaling behavior on the MP-r2SCAN dataset. The neural scaling laws suggest that model performance should improve steadily as the model size, dataset size, and amount of computing used for training are increased24,77,78. The performance is expected to follow a power-law relationship with each of these factors, provided the other two are not limiting. We benchmarked the energy and force MAEs on the validation set of MP-r2SCAN using either Method 1 (Scratch) or Method 4 (Transfer). The resulting validation errors vs. training sizes are shown in Fig. 6. For each curve in Fig. 6, we performed a linear regression starting from the data point corresponding to more than 1000 training points on the x-axis, yielding the coefficient of determination (R2) shown in the figures. The Linear fits demonstrate a linear scaling law behavior for both training from scratch (orange) and transfer learning (blue). The best-performing model for both energy and force predictions is obtained by Transfer, with an energy MAE of 15 meV/atom and a force MAE of 36 meV/Å.
a Energy MAE and b Force MAE on the MP-r2SCAN validation set using either Method 4, TL with r2SCAN AtomRef (Transfer, blue) or Method 1, training from scratch (Scratch, orange) methods. Zero training points in Transfer refers to the performance of the GGA/GGA + U pre-trained CHGNet with r2SCAN AtomRef. Linear fits are applied for x > 1000 to demonstrate the neural scaling law, and the coefficients of determination (R2) are shown in the figures.
The superior data-efficiency of TL over training from scratch can be found by the reduced MAE of TL in Fig. 6. For energy MAE in Fig. 6a, the Scratch curve exhibits a log-log slope of -0.615 with an R2 of 0.994, while the Transfer curve has a log-log slope of -0.301 with an R2 of 0.964. For force MAE in Fig. 6b, the Scratch curve shows a log-log slope of -0.394 with an R2 of 0.978, while the Transfer curve has a log-log slope of -0.134 with an R2 of 0.997. The results indicate TL with merely 1K high-fidelity data points can outperform training from scratch on a high-fidelity dataset with more than 10K data points, marking more than 10-fold data efficiency gained from the GGA pre-training step.
Interestingly, we observe that the superior performance of Transfer over Scratch does not saturate even given the full-sized MP-r2SCAN dataset of 0.24 million structures. Assuming the linear scaling trend of both Transfer and Scratch, the superior performance of Transfer will only be saturated after 719,996 training points for energy and 317,475 training points for force. This result indicates TL remains data-efficient even with close-to-million scale high-fidelity data points.
Discussion
The FPs enable efficient predictions of energy across diverse chemical environments, facilitating large-scale simulations with near GGA-level accuracy. As the training of FPs is migrating toward higher levels of DFT accuracy, optimal transferability strategies are needed. In this work, we investigated and benchmarked different transfer learning methods for FPs with multi-fidelity datasets. We demonstrate that the scale of atomic reference energies varies significantly across different approximate density functionals, leading to the non-trivial choice of fine-tuning and TL approaches. We rationalized the importance of refitting the atomic reference energies when fine-tuning MLIPs across multi-fidelity datasets. To further demonstrate the validity and generality of this transfer learning approach, we present two additional experiments involving the transfer learning on other datasets: a halide Van der Waals (vdW) dataset and an HSE06 dataset, which can be found in the supplementary information. The results show that Method 4 consistently outperforms the other methods, achieving the lowest test MAEs for energy, force, and stress, as well as the most robust and smooth training dynamics.
The energy quantity that matters for physical behavior is always referenced to some reference energies and not determined by total energies. For example, the cohesive energy is referenced to the energy of neutral, free atoms at infinite separation67. The formation energy is referenced to the energy of constituent elemental unaries in their reference states (solid or gas phase)79,80, and decomposition energy is referenced to the energies of competing compounds in a given chemical space74. Consequently, the eV/atom scale shifts in total energy from GGA/GGA + U to r2SCAN do not lead to any changes in the physical interaction and behavior of materials. However, as energy is the training label for a ML model, the significant difference in the energy scales leads to challenges in the convergence of the TL.
Essentially, by using energy referencing, one can modify the energy loss component in a model’s loss function during TL. For a FP with AtomRef, the general formula for the modified energy error of a structure’s data is:
where \({E}_{{\rm{label}}}^{{\rm{target}}}\) is the target energy training label, which is often obtained from high-fidelity calculations. celem is the composition row vector representing the number of each element in the structure. \({{\bf{E}}}_{{\rm{elem}}}^{{\rm{source}}}\) represents the AtomRef of the source dataset. \({E}_{{\rm{GNNs}}}^{{\rm{source}}}\) and \({{\bf{c}}}_{{\rm{elem}}}\cdot {{\bf{E}}}_{{\rm{elem}}}^{{\rm{source}}}\) are the energy predictions of the GNN and AtomRef, which sum up to the energy prediction of the source FP that has been pre-trained from a low-fidelity source dataset. \({{\bf{E}}}_{{\rm{ref}}}^{{\rm{target}}}\) and \({{\bf{E}}}_{{\rm{ref}}}^{{\rm{source}}}\) are the energy referencing parts of the two functionals, with dimensions Nelem × 1, representing the reference energies of the structures. For cohesive energy, the reference energies are the energies of neutral free atoms at rest; for formation energy, they are the energies of unaries in their reference states. In our approach, they are also coming from the fitted AtomRefs.
Energy referencing refers to replacing the AtomRef from \({{\bf{E}}}_{{\rm{elem}}}^{{\rm{source}}}\) to \(({{\bf{E}}}_{{\rm{elem}}}^{{\rm{source}}}+{{\bf{E}}}_{{\rm{ref}}}^{{\rm{target}}}-{{\bf{E}}}_{{\rm{ref}}}^{{\rm{source}}})\) before transferring a FP to the target level. After energy referencing, the remaining contribution in the energy loss represents the differences in atomic interactions approximated by the source (GGA/GGA + U) versus the target (r2SCAN), which is the relevant part of the energy that TL on GNNs aims to learn. Using AtomRef as Eref is potentially better than referencing related to cohesive or formation energy, as AtomRef obtains atomic reference energies as statistical averages from all data in the dataset that covers a vast chemical space.
We attribute the effectiveness of using AtomRef as Eref for cross-functional TL to two key factors. Firstly, the more than 10-fold improvement in correlation from 0.0917 to 0.9250 (see TL with different atomic reference energies) significantly enhances the effectiveness of TL. Secondly, refitting AtomRef ensures gradual adjustments of the model weights, and thus a more stable and reliable training process. Without refitting AtomRef, energy shifts cause substantial discrepancies between predicted and target energies, leading to very large prediction errors and high loss values initially. This, in turn, produces large gradients that cause excessive changes with the model weights, as illustrated in Fig. 3a and b.
According to Table 1, Method 4 (TL with r2SCAN AtomRef) is shown to be most effective with the lowest energy MAE, consistent with the above rationalization of this approach. The higher prediction MAEs of Methods 2 (TL with trainable AtomRef) and 3 (TL with frozen AtomRef) compared to Method 4 – which integrates energy re-referencing with GNN-based TL – highlight the challenges of conventional TL without refitting AtomRef in FPs. Methods 2 and 3 exhibit similar MAEs since they both begin with GGA/GGA + U AtomRef, and the large energy shifts between r2SCAN and GGA/GGA + U cause poor correlation and excessive weight adjustments during early fine-tuning, driving model weights to suboptimal positions where they can become trapped. Notably, their predictions for forces, stresses, and magmoms are inferior to those of Method 1 (Training from scratch), which uses r2SCAN data directly, free from GGA/GGA + U influence. This underperformance is attributed to negative transfer54, resulting from the weak correlation between source and target datasets during GNN-based TL.
As it is unlikely that one dataset will rule all of FPs, a well-founded strategy to integrate diverse datasets, such as Materials Project21,22, Alexandria23, OQMD81, AFLOWLIB82, NOMAD83, QM984, JARVIS85, OC2086, OMat2426, OCX2487, and MatPES32, will provide a promising avenue for leveraging the broad spectrum of available information and enable integration of future high quality data. Such integration will be helpful to address the data-originated issues in FPs which are otherwise challenging to solve by only model architecture improvements27. Our scaling law analysis demonstrates the superior data efficiency gained from pre-training on large-scale low-fidelity dataset when migrating to high-fidelity ones.
As FP-training is expected to transfer to higher quantum chemistry levels of theory, we also want to highlight the need to establish benchmark tests tailored to these computationally demanding quantum mechanical methods, such as r2SCAN, coupled cluster methods (e.g., CCSD), and multi-reference approaches. In this work, we exemplify this by introducing stability benchmarks based on decomposition energy and formation energy predictions, as well as the density benchmark which uses experimental values as ground truth. Current FP benchmarks such as Matbench Discovery88 are mostly limited to GGA/GGA + U tasks due to the dataset limits. We advocate for more comprehensive benchmarking frameworks that go beyond GGA/GGA + U and potentially integrate evaluations such as kinetic properties and more complex material behavior to better assess models across different functionals.
In summary, by examining how atomic reference energies influence the performance of GGA/GGA + U to r2SCAN TL, we reiterate the importance of establishing correlations between multi-fidelity datasets so that they can benefit from TL. TL with refitting atomic reference energies yields a stable and reliable MLIP for energy, interatomic forces, thermodynamic stability, and density predictions. Our benchmark results and scaling law analysis show that refitting atomic energy is data-efficient and convinces fine-tuning FPs to be a practical way for various downstream materials modeling tasks.
Methods
Data preparation
The r2SCAN Dataset, MP-r2SCAN, is parsed from the Materials Project Database in March 2024. We collected all the r2SCAN structure optimization and static task trajectories under each material ID that contain these tasks, and then followed similar criteria as those used in creating the MPtrj Dataset: (1) Final frame energies were limited to within 20 meV/atom of the primary task. (2) Structures missing energy, forces, or electronic convergence were excluded. (3) Structures with energies > 1 eV/atom or < 10 meV/atom relative to Materials Project’s ThermoDoc relaxed structures were filtered out to eliminate large energy differences resulting from variations in DFT calculation settings. (4) Duplicate structures were removed using pymatgen’s StructureMatcher and energy matcher89. For all 4 TL models, we randomly split the MP-r2SCAN dataset into training, validation, and test sets with an approximate ratio of 8:1:1 based on material IDs. The training set contains 27,943 material IDs with 190,560 structures; the validation set contains 3492 material IDs with 23,888 structures; and the test set contains 3492 material IDs with 23,799 structures. The energy, force, stress, and magmom prediction MAEs are based on the test set’s 23,799 structures. The decomposition energy prediction MAE was reported on the test set. The formation energy prediction MAE was calculated on all 34,938 r2SCAN material IDs in the Materials Project.
Training scheme
We kept most of the settings the same as the pre-trained CHGNet model, except for the following: we changed the fixed GGA/GGA + U AtomRef of the model to r2SCAN AtomRef; a Huber loss with energy, force stress and magmom loss ratio of 3:1:0.1:1 was used to train the model; we used a batch size of 64 and a learning rate of 10−3 that cosinely decays to 10−5 in 50 epochs.
Feature importance
To determine which elements contribute most to the formation energy differences between r2SCAN and GGA/GGA + U (discussed in Section Energy differences across two functionals), we used the attribute feature_importances_ in scikit-learn’s DecisionTreeRegressor.
The importance of each node on the decision tree can be calculated by (assuming only two child nodes (binary tree)):
nj represents the importance of node j, wj is the weighted number of samples reaching node j, σj denotes the impurity value (here it is variance) of node j, left(j) refers to the child node from the left split on node j, and right(j) refers to the child node from the right split on node j.
Feature importance is calculated by:
where fi represents the importance of feature i, and nj represents the importance of node j.
To obtain the normalized feature importance, each feature importance was divided by the total number of atoms of this element in the dataset and then multiplied by 9,000 for Fig. 1c and 500 for Fig. 1d to scale it back to the range of 0-1. Finally, it was visualized on the periodic table.
Data availability
The MP-r2SCAN dataset used to fine-tune CHGNet is available at https://doi.org/10.6084/m9.figshare.28245650.v290.
Code Availability
The model weights of fine-tuned CHGNet and training configuration using Method 4 are available at https://github.com/CederGroupHub/chgnet/tree/main/chgnet/pretrained/r2scan.
References
Chen, H. et al. Carbonophosphates: a new family of cathode materials for li-ion batteries identified computationally. Chem. Mater. 24, 2009–2016 (2012).
Urban, A., Seo, D.-H. & Ceder, G. Computational understanding of li-ion batteries. npj Computational Mater. 2, 1–13 (2016).
Jain, A., Shin, Y. & Persson, K. A. Computational predictions of energy materials using density functional theory. Nat. Rev. Mater. 1, 1–13 (2016).
Unke, O. T., Koner, D., Patra, S., Käser, S. & Meuwly, M. High-dimensional potential energy surfaces for molecular simulations: from empiricism to machine learning. Mach. Learn.: Sci. Technol. 1, 013001 (2020).
Li, L. et al. Representing crystal potential energy surfaces via a stationary-point network. Acta Materialia 281, 120403 (2024).
Kopp, W. A. et al. Automatic potential energy surface exploration by accelerated reactive molecular dynamics simulations: from pyrolysis to oxidation chemistry. J. Phys. Chem. A 127, 10681–10692 (2023).
Ock, J., Mollaei, P. & Barati Farimani, A. Gradnav: Accelerated exploration of potential energy surfaces with gradient-based navigation. J. Chem. Theory Comput. 20, 4088–4098 (2024).
Daw, M. S., Foiles, S. M. & Baskes, M. I. The embedded-atom method: a review of theory and applications. Mater. Sci. Rep. 9, 251–310 (1993).
Goringe, C., Hernández, E., Gillan, M. & Bush, I. Linear-scaling dft-pseudopotential calculations on parallel computers. Computer Phys. Commun. 102, 1–16 (1997).
Beck, T. L. Real-space mesh techniques in density-functional theory. Rev. Mod. Phys. 72, 1041 (2000).
Podryabinkin, E. V., Tikhonov, E. V., Shapeev, A. V. & Oganov, A. R. Accelerating crystal structure prediction by machine-learning interatomic potentials with active learning. Phys. Rev. B 99, 064114 (2019).
Zhang, L., Han, J., Wang, H., Car, R. & E, W. Deep potential molecular dynamics: A scalable model with the accuracy of quantum mechanics. Phys. Rev. Lett. 120, 143001 (2018).
Batzner, S. et al. E (3)-equivariant graph neural networks for data-efficient and accurate interatomic potentials. Nat. Commun. 13, 2453 (2022).
Cheng, B. Cartesian atomic cluster expansion for machine learning interatomic potentials. npj Computational Mater. 10, 157 (2024).
Deng, B. et al. Chgnet as a pretrained universal neural network potential for charge-informed atomistic modelling. Nat. Mach. Intell. 5, 1031–1041 (2023).
Kim, D., King, D. S., Zhong, P. & Cheng, B. Learning charges and long-range interactions from energies and forces. arXiv preprint arXiv:2412.15455 (2024).
Chen, C. & Ong, S. P. A universal graph deep learning interatomic potential for the periodic table. Nat. Computational Sci. 2, 718–728 (2022).
Batatia, I. et al. A foundation model for atomistic materials chemistry. arXiv preprint arXiv:2401.00096 (2023).
Kim, J. et al. Data-efficient multifidelity training for high-fidelity machine learning interatomic potentials. J. Am. Chem. Soc. 147, 1042–1054 (2024).
Neumann, M. et al. Orb: A fast, scalable neural network potential. arXiv preprint arXiv:2410.22570 (2024).
Jain, A. et al. Commentary: The materials project: A materials genome approach to accelerating materials innovation. APL Mater. 1, 011002 (2013).
Horton, M. K. et al. Accelerated data-driven materials science with the materials project. Nat. Mater. 24, 1–11 (2025).
Ghahremanpour, M. M., Van Maaren, P. J. & Van Der Spoel, D. The alexandria library, a quantum-chemical database of molecular properties for force field development. Sci. data 5, 1–10 (2018).
Merchant, A. et al. Scaling deep learning for materials discovery. Nature 624, 80–85 (2023).
Yang, H. et al. Mattersim: A deep learning atomistic model across elements, temperatures and pressures. arXiv preprint arXiv:2405.04967 (2024).
Barroso-Luque, L. et al. Open materials 2024 (omat24) inorganic materials dataset and models. arXiv preprint arXiv:2410.12771 (2024).
Deng, B. et al. Systematic softening in universal machine learning interatomic potentials. npj Computational Mater. 11, 1–9 (2025).
Yu, H., Giantomassi, M., Materzanini, G., Wang, J. & Rignanese, G.-M. Systematic assessment of various universal machine-learning interatomic potentials. Mater. Genome Eng. Adv. 2, e58 (2024).
Lan, J. et al. Adsorbml: a leap in efficiency for adsorption energy calculations using generalizable machine learning potentials. npj Computational Mater. 9, 172 (2023).
Chen, J., Huang, X., Hua, C., He, Y. & Schwaller, P. A multi-modal transformer for predicting global minimum adsorption energy. Nat. Commun. 16, 3232 (2025).
Sivak, J. T. et al. Discovering high-entropy oxides with a machine-learning interatomic potential. Phys. Rev. Lett. 134, 216101 (2025).
Kaplan, A. D. et al. A foundational potential energy surface dataset for materials. arXiv preprint arXiv:2503.04070 (2025).
Perdew, J. P. & Schmidt, K. vol. 577, 1 (American Institute of Physics, 2001).
Kaplan, A. D., Levy, M. & Perdew, J. P. Predictive power of the exact constraints and approximate norms in density functional theory. Annu. Rev. Phys. Chem. 74, 193–218 (2023).
Perdew, J. P. & Zunger, A. Self–interaction correction to density–functional approximations for many–electron systems. Phys. Rev. B 23, 5048–5079 (1981).
Zhou, F., Cococcioni, M., Marianetti, C. A., Morgan, D. & Ceder, G. First-principles prediction of redox potentials in transition-metal compounds with lda+ u. Phys. Rev. B-Condens. Matter Mater. Phys. 70, 235121 (2004).
Goerigk, L. et al. A look at the density functional theory zoo with the advanced GMTKN55 database for general main group thermochemistry, kinetics and noncovalent interactions. Phys. Chem. Chem. Phys. 19, 32184–32215 (2017).
Sun, J., Ruzsinszky, A. & Perdew, J. P. Strongly constrained and appropriately normed semilocal density functional. Phys. Rev. Lett. 115, 036402 (2015).
Furness, J. W., Kaplan, A. D., Ning, J., Perdew, J. P. & Sun, J. Accurate and numerically efficient r2scan meta-generalized gradient approximation. J. Phys. Chem. Lett. 11, 8208–8215 (2020).
Heyd, J., Scuseria, G. E. & Ernzerhof, M. Hybrid functionals based on a screened coulomb potential. J. Chem. Phys. 118, 8207–8215 (2003).
Anisimov, V. I., Zaanen, J. & Andersen, O. K. Band theory and mott insulators: Hubbard u instead of stoner i. Phys. Rev. B 44, 943 (1991).
Wang, L., Maxisch, T. & Ceder, G. Oxidation energies of transition metal oxides within the gga+ u framework. Phys. Rev. B-Condens. Matter Mater. Phys. 73, 195107 (2006).
Jain, A. et al. Formation enthalpies by mixing gga and gga+ u calculations. Phys. Rev. B-Condens. Matter Mater. Phys. 84, 045115 (2011).
Perdew, J. P., Burke, K. & Ernzerhof, M. Generalized gradient approximation made simple. Phys. Rev. Lett. 77, 3865 (1996).
Kothakonda, M. et al. Testing the r2scan density functional for the thermodynamic stability of solids with and without a van der waals correction. ACS Mater. Au 3, 102–111 (2022).
Isaacs, E. B. & Wolverton, C. Performance of the strongly constrained and appropriately normed density functional for solid-state materials. Phys. Rev. Mater. 2, 063801 (2018).
Kingsbury, R. et al. Performance comparison of r 2 scan and scan metagga density functionals for solid materials via an automated, high-throughput computational workflow. Phys. Rev. Mater. 6, 013801 (2022).
Liu, H. et al. Assessing r2scan meta-gga functional for structural parameters, cohesive energy, mechanical modulus, and thermophysical properties of 3d, 4d, and 5d transition metals. J. Chem. Phys. 160, 024102 (2024).
Hautier, G., Ong, S. P., Jain, A., Moore, C. J. & Ceder, G. Accuracy of density functional theory in predicting formation energies of ternary oxides from binary oxides and its implication on phase stability. Phys. Rev. B-Condens. Matter Mater. Phys. 85, 155208 (2012).
Cococcioni, M. & de Gironcoli, S. Linear response approach to the calculation of the effective interaction parameters in the LDA+U method. Phys. Rev. B 71, 035105 (2005).
Kingsbury, R. S. et al. A flexible and scalable scheme for mixing computed formation energies from different levels of theory. npj Computational Mater. 8, 195 (2022).
Hoffmann, N., Schmidt, J., Botti, S. & Marques, M. A. Transfer learning on large datasets for the accurate prediction of material properties. Digital Discov. 2, 1368–1379 (2023).
Chen, M. S. et al. Data-efficient machine learning potentials from transfer learning of periodic correlated electronic structure methods: Liquid water at afqmc, ccsd, and ccsd (t) accuracy. J. Chem. Theory Comput. 19, 4510–4519 (2023).
Wang, Z., Dai, Z., Póczos, B. & Carbonell, J. In Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, 11293–11302 (2019).
Gong, S. et al. Calibrating dft formation enthalpy calculations by multifidelity machine learning. JACS Au 2, 1964–1977 (2022).
Ramakrishnan, R., Dral, P. O., Rupp, M. & Von Lilienfeld, O. A. Big data meets quantum chemistry approximations: the δ-machine learning approach. J. Chem. theory Comput. 11, 2087–2096 (2015).
Pitfield, J. et al. Augmentation of universal potentials for broad applications. Phys. Rev. Lett. 134, 056201 (2025).
Dral, P. O., Zubatiuk, T. & Xue, B.-X. Learning from multiple quantum chemical methods: δ-learning, transfer learning, co-kriging, and beyond. In Quantum Chemistry in the Age of Machine Learning, 491–507 (Elsevier, 2023).
Chen, C., Zuo, Y., Ye, W., Li, X. & Ong, S. P. Learning properties of ordered and disordered materials from multi-fidelity data. Nat. Computational Sci. 1, 46–53 (2021).
Ko, T. W. & Ong, S. P. Data-efficient construction of high-fidelity graph deep learning interatomic potentials. npj Computational Mater. 11, 65 (2025).
Allen, A. E. et al. Learning together: Towards foundation models for machine learning interatomic potentials with meta-learning. npj Computational Mater. 10, 154 (2024).
Gerace, F., Saglietti, L., Mannelli, S. S., Saxe, A. & Zdeborová, L. Probing transfer learning with a model of synthetic correlated datasets. Mach. Learn.: Sci. Technol. 3, 015030 (2022).
Wang, A. et al. A framework for quantifying uncertainty in dft energy corrections. Sci. Rep. 11, 15496 (2021).
Choe, D.-H., West, D. & Zhang, S. Revealing the vacuum level in an infinite solid by real-space potential unfolding. Phys. Rev. B 103, 235202 (2021).
Ihm, J., Zunger, A. & Cohen, M. L. Momentum-space formalism for the total energy of solids. J. Phys. C: Solid State Phys. 12, 4409 (2001).
How, W. B., Chong, S., Grasselli, F., Huguenin-Dumittan, K. K. & Ceriotti, M. Adaptive energy reference for machine-learning models of the electronic density of states. Phys. Rev. Mater. 9, 013802 (2025).
Kittel, C. & McEuen, P.Introduction to solid state physics (John Wiley & Sons, 2018).
Schmidt, J., Pettersson, L., Verdozzi, C., Botti, S. & Marques, M. A. Crystal graph attention networks for the prediction of stable materials. Sci. Adv. 7, eabi7948 (2021).
Quinlan, J. R. Induction of decision trees. Mach. Learn. 1, 81–106 (1986).
Sun, J. et al. Density functionals that recognize covalent, metallic, and weak bonds. Phys. Rev. Lett. 111, 106401 (2013).
Yang, J. H., Kitchaev, D. A. & Ceder, G. Rationalizing accurate structure prediction in the meta-gga scan functional. Phys. Rev. B 100, 035132 (2019).
Ning, J. et al. Workhorse minimally empirical dispersion-corrected density functional with tests for weakly bound systems: r2SCAN+rVV10. Phys. Rev. B 106, 075422 (2022).
Lupo Pasini, M. et al. Scalable training of trustworthy and energy-efficient predictive graph foundation models for atomistic materials modeling: a case study with hydragnn. J. Supercomputing 81, 618 (2025).
Bartel, C. J. et al. A critical examination of compound stability predictions from machine-learned formation energies. npj computational Mater. 6, 97 (2020).
Bartel, C. J. Review of computational approaches to predict the thermodynamic stability of inorganic solids. J. Mater. Sci. 57, 10475–10498 (2022).
Bergerhoff, G., Hundt, R., Sievers, R. & Brown, I. D. The inorganic crystal structure data base. J. Chem. Inf. Computer Sci. 23, 66–69 (1983).
Bahri, Y., Dyer, E., Kaplan, J., Lee, J. & Sharma, U. Explaining neural scaling laws. Proc. Natl Acad. Sci. 121, e2311878121 (2024).
Frey, N. C. et al. Neural scaling of deep chemical models. Nat. Mach. Intelligence 5, 1297–1305 (2023).
Xin, X., Lai, W. & Liu, B. Point defect properties in hcp and bcc zr with trace solute nb revealed by ab initio calculations. J. Nucl. Mater. 393, 197–202 (2009).
Stevanović, V., Lany, S., Zhang, X. & Zunger, A. Correcting density functional theory for accurate predictions of compound enthalpies of formation: Fitted elemental-phase reference energies. Phys. Rev. B-Condens. Matter Mater. Phys. 85, 115104 (2012).
Saal, J. E., Kirklin, S., Aykol, M., Meredig, B. & Wolverton, C. Materials design and discovery with high-throughput density functional theory: the open quantum materials database (oqmd). Jom 65, 1501–1509 (2013).
Curtarolo, S. et al. Aflow: An automatic framework for high-throughput materials discovery. Computational Mater. Sci. 58, 218–226 (2012).
Draxl, C. & Scheffler, M. The nomad laboratory: from data sharing to artificial intelligence. J. Phys.: Mater. 2, 036001 (2019).
Ramakrishnan, R., Dral, P. O., Rupp, M. & Von Lilienfeld, O. A. Quantum chemistry structures and properties of 134 kilo molecules. Sci. data 1, 1–7 (2014).
Choudhary, K. et al. The joint automated repository for various integrated simulations (jarvis) for data-driven materials design. npj computational Mater. 6, 173 (2020).
Chanussot, L. et al. Open catalyst 2020 (oc20) dataset and community challenges. Acs Catal. 11, 6059–6072 (2021).
Abed, J. et al. Open catalyst experiments 2024 (ocx24): Bridging experiments and computational models. arXiv preprint arXiv:2411.11783 (2024).
Riebesell, J. et al. A framework to evaluate machine learning crystal stability predictions, Nat. Mach. Intell. 7, 836–847 (2025).
Ong, S. P. et al. Python materials genomics (pymatgen): A robust, open-source python library for materials analysis. Computational Mater. Sci. 68, 314–319 (2013).
Huang, X. et al. Materials Project Trajectory Dataset of r2SCAN (MP-r2SCAN) (2025). https://figshare.com/articles/dataset/Materials_Project_Trajectory_Dataset_of_r_sup_2_sup_SCAN_MPr_sup_2_sup_SCAN_/28245650.
Acknowledgements
This work was funded by the U.S. Department of Energy, Office of Science, Office of Basic Energy Sciences, Materials Sciences and Engineering Division under Contract No. DE-AC0205CH11231 (Materials Project program KC23MP). The work was also supported by the computational resources provided by the Extreme Science and Engineering Discovery Environment (XSEDE), supported by National Science Foundation grant number ACI1053575; the National Energy Research Scientific Computing Center (NERSC), a U.S. Department of Energy Office of Science User Facility located at Lawrence Berkeley National Laboratory; and the Swift Cluster resource provided by the National Renewable Energy Laboratory (NREL). The authors thank Luca Binci and Lauren N. Walters for valuable discussions.
Author information
Authors and Affiliations
Contributions
B.D. and G.C. conceived the initial idea. X.H. performed Dataset Collection. X.H. benchmarked all the transfer learning methods. X.H. performed experiments on scaling law analysis. P.Z. and A.K. offered insights into the discussion of DFT functionals. B.D., K.P. and G.C. offered insights and guidance throughout the project. All authors contributed to discussions and approved the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Huang, X., Deng, B., Zhong, P. et al. Cross-functional transferability in foundation machine learning interatomic potentials. npj Comput Mater 11, 313 (2025). https://doi.org/10.1038/s41524-025-01796-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41524-025-01796-y