Abstract
Energetic Materials (EMs) play important roles in military, civilian and aerospace fields. Energy and stability are the two most important but contradictory properties in practical application, thus leading to difficult challenges in developing new EMs with high comprehensive performance. Motivated by the challenge, we exploit a de novo design framework targeting multiple objectives by integrating deep learning generator, machine learning prediction models, Pareto front optimization and quantum mechanics (QM) validation. First, heat of explosion (Q) and bond dissociation energy (BDE) are calculated by high-precision QM for 778 explosives experimentally reported. With the reliable dataset, RNN coupled with transfer learning is exploited to generate a new massive search space with 2 × 105 potential energetic molecules. Q and BDE prediction models with high accuracy are further developed by data augmentation and improvements in feature representation and model architectures, to quickly and accurately evaluate these new energetic molecules. The modified 3D-GNN achieves an R2 = 0.95 for the Q prediction, while the XGBoost coupled with the feature complementarity and PADRE data augmentation performs best for the BDE prediction (R2 = 0.98). To screen energetic compounds with trade-off energy and stability from the vast new molecule space, the predicted values and uncertainties are simultaneously considered, and Pareto front-based multi-objective screening is conducted by using 2D P[I] metric. QM calculation confirms the superior performance of the top 60 candidates to CL-20 in Q. 25 promising energetic molecules with high energy and desired stability, as well as synthesis feasibility provide valuable candidates for experimental development. Also, the design strategy can be extended to other material fields.

Similar content being viewed by others
Introduction
Energetic materials (EMs), such as explosives, pyrotechnics, and propellants, have played important roles in military and civilian applications1,2,3. However, it is a long process for developing new EMs with high performance4,5 due to the extremely complex multi-parameter nature and high safety risk for experimental investigation. Energy and stability are the two most important but contradictory properties of EMs. Nowadays, EMs have entered into a new phase of high-energy-density materials (HEDMs)6. Unlike conventional energetic compounds such as HMX, RDX, TNT, etc., HEDMs are composed of nitrogen-rich skeletons and energetic substituents, which can contribute to higher energy7,8. However, the high energy often accompanies low stability with high safety risk, limiting its application in practice. Thus, the contradiction between the energy and the stability makes the development of novel EMs face difficult challenges.
To accelerate the development of EMs, computational simulation methods like quantum mechanics (QM) calculation and molecular dynamics (MD) simulation were introduced to assist experiments9,10, yet high computational cost limits their exploration to vast and unknown chemical spaces. With the advent of artificial intelligence (AI), machine learning (ML) has emerged as a powerful tool for accelerating material development11,12, as it can intelligently capture the causality underlying complex data. MLs already exhibit great success in medicine, chemistry and material fields13,14,15,16. Currently, the data-driven ML techniques have been increasingly exploited to accelerate the development of EMs. These works mainly focused on property prediction tasks for EMs, involving density17, decomposition temperature18,19, heat of formation20,21, detonation parameters22,23,24, thermal stability, and impact sensitivity25,26,27, etc. The size of datasets used in the property prediction varies from a few dozen to several thousands. For real explosives, the data set is typically limited to a few hundreds while some studies incorporated organic molecules to expand the data set to several thousand samples. With the predictive models, researchers have screened potential energetic molecules from existing large chemical databases. For example, Chen et al.20 used 451 molecules selected from the Cambridge Structural Database (CSD) to establish machine learning models for EM’s crystal density and solid phase enthalpy of formation, through which they screened out 56 candidates with high detonation performance from the 150 million-sized PubChem database. In contrast to property prediction tasks, the molecular design of new EMs has been very limited, which were mainly based on the combination of existing energetic fragments and then screened target candidates by some computational metrics or ML models. For example, Liu et al.19 combined -NO2, -NH2, -NHNO2, and -N=N-NO2 functional groups with scaffolds from the CCDC database to generate 70,188 structures and then utilized ML-based decomposition temperature and density models to one by one screen heat-resistant energetic molecules. Cheng et al.28 generated 35,322 bistetrazole-based molecules by combining 20 bridgeheads with 29 substituents, and then applied sequential single-objective filtering based on calculated oxygen balance, SYBA scores, and density, to identify high-energy and low-sensitivity candidates. Gou et al.29 constructed the ML model of the detonation velocity to screen four promising energetic molecules from 1053 prismane derivatives generated by group substitution at the prismane backbone. Similarly, Song et al.23 constructed ML models of several properties involving denotation and stability to successfully screen some potential energetic molecules from a large molecule space generated by substitution numeration on several energetic backbones. Wen et al.30 constructed a candidate space of 107 molecules based on 71 cage scaffolds extracted from the ZINC15 database, and theoretically discovered a new potential ZN-OB9 with high energy and good safety. More details regarding the methodologies and performance of these molecular design works are summarized in Table S3.
Despite the promising potential of ML exhibited in the EM field, there remain many challenges in applying machine learning to EMs31. Firstly, compared to other fields, high risk in safety and long development cycle have led to the scarcity of EM data, which limits robustness and generalization of machine learning to unseen samples. Secondly, energetic molecules in the search space were mainly constructed by a combinatorial chemistry way under some prior rules, which limit the structure diversity and rationality. In addition, most of the related works mainly relied on ML-predicted values and a single or single-similar objective (one-by-one for several properties) to screen target molecules. The screen way does not consider the prediction uncertainty of MLs, which easily induces a high risk with large bias to unseen samples, in particular for the ML model trained on the small-sized data. Furthermore, using the single-objective design, the improvement in the single target property most probably compromises the others32, in particular for mutually contradicting properties like the energy and the stability of EMs. However, materials with highly comprehensive properties are more desired in practical applications, which need to simultaneously consider multiple objectives in designing new molecules, thus presenting greater challenges than the single objective design. Practically, the aforementioned issues not only exist in the field of EMs but also in many other chemistry and material fields with low data available, thus being a common question in the real world. Consequently, there is an urgent need to develop more effective methods to accelerate the development of new materials with desired comprehensive properties.
Motivated by the urgent need, we developed a de novo multi-objective optimization design framework for new energetic molecules with trading off energy and stability. Herein, we selected two calculated metrics (heat of explosion (Q) and bond dissociation energy (BDE) of the weakest bond) to separately represent the energy and stability of EMs. Practically, BDE has been considered as an important metric for evaluating stability and reactivity across various molecules33,34,35. To achieve the de novo multi-objective design, the design framework integrates de novo deep learning molecular generation coupled with transfer learning, uncertainty-aware ML property prediction, Pareto front-based multi-objective screening, and high precision QM validation. Firstly, we extensively searched literature and collected energetic molecules synthesized to conduct a real explosive data set. With the aid of a large chemical molecule database and the explosive data set, a deep learning generative model coupled with transfer learning strategy is exploited to de novo generate massive new energetic candidates with diverse and rational structures, which can circumvent the scarcity of EM data and limitations of combinatorial chemistry. To rapidly and reliably evaluate the two important properties of new energetic molecules in the unknown space, we constructed accurate property prediction models by exploiting multiple traditional ML architectures coupled with new feature representations and data augmentation as well as introducing deep learning pre-trained models. The comparison between the different ML models and the different feature representations also provides methodological guidelines for the application of machine learning. To effectively and quickly screen new potential EM molecules that can trade off the two contradictory properties (high energy and low stability), we adopted a 2D P[I] multi-objective optimization strategy, which simultaneously considers the predicted value and the uncertainty of the ML model to obtain Pareto front solutions based on two-dimensional P[I] decisions36. High-precision QM calculations and comprehensive synthetic accessibility evaluation were further performed to identify 25 energetic candidates with good performance of energy and stability. A detailed comparison between our de novo multi-objective design framework and prior studies on designing energetic molecules is provided in Table S3.
Results
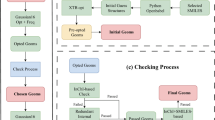
As illustrated by Fig. 1, our computation framework mainly includes five modules: dataset construction, deep learning-based molecular generation, machine learning property prediction, 2D P[I] multi-objective optimization based on the Pareto front, and molecule recommendation based on both quantum mechanics calculation and synthetic accessibility evaluation.
The workflow for multi-objective design of EMs, including five modules: (1) the construction of EMs dataset containing Q and BDE; (2) the generation of a new search space for energetic molecules based on a deep molecular generator coupled with transfer learning strategy; (3) the construction of machine learning prediction models for Q and BDE; (4) the combination of ML models and Pareto front-based multi-objective optimization to select high-performance energetic molecules balancing Q and BDE; (5) validation based on high-precision DFT calculations and comprehensive synthetic accessibility for recommendation of experiments.
Construction of a representative energetic data set
Data is the foundation of ML, including the data size, data representativeness, and data quality. To construct a reliable and representative EM dataset, we manually collected 778 synthesized CHON-containing energetic molecules from 427 published literature. Given the scarcity of experimental data with high quality for EMs, the calculated Q and BDE were adopted as theoretical indicators of the energy and stability, respectively. Practically, many studies indicated that BDE is associated with the stability of energetic molecules, including thermal stability and impact sensitivity as well as shock sensitivity37,38,39. Figure S2 shows correlation between the calculated BDEs of our dataset and two types of experimental stability metrics (Td19 and H5040), showing moderate linear correlation, in turn supporting the utility of BDE as a theoretical indicator of the molecular stability for initial assessment. However, it is noted that when the initial decomposition steps go through a more complex transition state rather than simple bond dissociation, increasing deviation may exist for the stability estimation solely based on the BDE value of the weakest bond. The 778 energetic molecules, including their SMILES, calculated Q and BDE values at the level of CBS-4M and B3LYP/6-31 G**, respectively, and related 427 references are listed in Supplementary Data 1. Table S4 summarizes the types and counts of trigger bonds corresponding to the calculated BDEs.
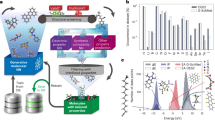
To evaluate the representativeness of the EM dataset, we performed statistical analyses on the distribution of Q and BDE, the total number of atoms, the number of C, H, O, and N atoms, and the mass percent of nitrogen. As shown in Fig. 2a, the Q values of all the energetic molecules are distributed between 300 cal·g-1 and 1900 cal·g-1, implying that there remains space to increase the Q values. The BDE values range from 50 to 550 kJ·mol-1. As shown in Fig. 2b, c, the number of total atoms in 778 molecules ranges from 7 to 87, and the mass percent of nitrogen in each molecule is in the range of 4.17–72.73%. The counts of C, H, O, and N atoms in each molecule are presented in Fig. S3. Additionally, as reflected by Fig. 2, the 778 energetic molecules include diverse categories involving different energetic substituents and skeletons41, also including classic explosives like TNT, TATB, TNB, RDX and CL-20, rather than only containing -NO2 group compounds derived from computational combination or collected from Cambridge Structural Database (CSD)26,42, These statistical results indicate the diversity of EMs for either the structures or the property labels, thus being representative.
a The distribution of Q and BDE values. The 778 energetic compounds are classified into six categories following Mathieu’s work40, of which are highlighted by different colors. Unst denotes a generic category including unstable explosophores like azides, difluoroamines, diazo phenols, tetrazoles, and triazole compounds. O-NO2, N-NO2, and C-NO2 represent categories containing O-NO2 group, N-NO2 group, and aliphatic C-NO2 groups, respectively. 5mAr denotes a category containing five-membered aromatics rings. NACs denote all remaining compounds only containing a single kind of explosophore-like nitro groups bonded to carbon atoms in an aromatic 6-membered ring. The statistical distribution of (b) atom number, and c mass percent of nitrogen for the 778 energetic molecules. d Chemical structures of representative molecules from the six categories of the dataset.
Deep generative model to construct a massive search space for energetic candidates with structure diversity and rationality
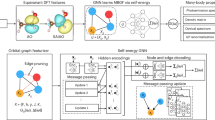
To circumvent the limitation of combinatorial chemistry in structure diversity and rationality, we introduced deep learning-based molecule generation to achieve a de novo design. However, the limited number of energetic molecules (778) collected cannot support deep learning. To address the technical obstacle, we explored a deep learning-based generative model. It includes a pre-trained RNN and a generative RNN, which are connected by a transfer learning strategy to target the energetic molecule generation, as shown in Fig. 3. The pre-trained RNN is first trained on 1.27 million molecules from the ChEMBL database43 to provide sufficient structural and grammatical knowledge for downstream energetic molecule generation tasks. In the transfer learning stage, we initialized the LSTM layers in the generative RNN with the same parameters in the pre-trained RNN. Then, we fine-tuned the fully connected layer on the EM dataset to target the energetic molecule generation. After training the generative model, large-scale sampling was performed to generate molecular structures.
a The framework consists of a pre-trained RNN and a generated RNN. Each RNN includes three LSTM layers followed by a fully connected layer with a Softmax function. A transfer learning strategy is adopted to generate the energetic molecular space. b The Tanimoto similarity distribution of the Murcko skeleton (left) and the Tanimoto similarity distribution (right) between the generated new molecules and the energetic molecules in the training set.
To evaluate the performance of the generation model, we generated 20,000 samples and calculated the proportion of valid SMILES (91.47%), demonstrating that the model can successfully learn essential molecular structures and grammar information. Subsequently, we calculated similarities of full compound and Murcko scaffold44 between the generated molecules and those in the training set to estimate the novelty of the generated molecules. As illustrated by Fig. 3b, there are around 82% of scaffolds in the generated compounds having low similarity (<0.1) to the training EMs. For the full compound similarity, approximately 90% of the compounds exhibit similarity below 0.2. The similarity value of less than 0.3 was considered as molecular dissimilarity45,46. Thus, lower similarity values (0.1 and 0.2) in the work indicate that the generative model can produce molecules with significant differences from those in the training set. Additionally, we calculated the IntDiv index, a common metric for measuring the diversity of generated molecules. The larger the IntDiv index, the higher the diversity. The results show an IntDiv index of 0.8 for the generated molecules, further supporting the high diversity of molecules generated by the RNN generative model.
To construct a massive search space for exploring new energetic molecules, we further generated 280k molecules as an initial search space. For the final search space, we conducted a preliminary filtering in terms of the following rules.
-
1.
Only molecules composed of C, H, O, and N elements were retained.
-
2.
Molecules containing three-membered or four-membered ring structures were removed.
-
3.
Electrically neutral molecules were retained, while charged molecules were removed.
Given that the most widely used energetic molecules typically consist of C, H, O, N elements, only molecules containing these elements were considered here. Additionally, molecules containing three-membered or four-membered ring structures were excluded due to their instability. Finally, electrically neutral molecules were retained due to higher stability. After these filtering steps, the remaining 201,547 molecules were taken as the final search space for energetic molecules.
Construction of optimal prediction models for Q and BDE
To rapidly assess the heat of explosions and bond dissociation energies of unknown molecules in the vast search space, it is necessary to construct property prediction models with high accuracy and robustness. Given the limited data size, we preferred to attempt some traditional machine learning algorithms. Herein, we used six traditional ML algorithms that were reported to perform well in small-size datasets47,48,49, including a least absolute shrinkage and selection operator regression model (LASSO)50, a kernel ridge regression model (KRR)51, a support vector regression model (SVR)52, a Gaussian process regression model (GPR)53, a random forest regression model (RF)54, and an extreme gradient boosting regression model (XGBoost)55.
In our previous study,56 a combination descriptor SEC, consisting of Sum of Bonds (SOB), Electrotopological state Fingerprints (E-state), and Custom Descriptor Set (CDS), was found to well characterize the structure knowledge associated with the heat of explosion for 88 energetic molecules. Thus, in this work, we still employed the SEC descriptor coupled with six traditional machine learning models to construct Q prediction models for the 778 energetic compounds. Table 1 summarizes their prediction performance, in which GPR exhibits the best performance for the independent test set, as highlighted in bold in Table 1, achieving an R2 value of 0.91, RMSE of 63.41 cal·g−1 and MAE of 36.75 cal·g−1.
For the BDE prediction, we adopted one model recently explored by us34, in which we proposed a hybrid feature representation by coupling the local environment of the target bond (CBD) into the global energetic characteristics (SEC) to sufficiently characterize the structure of EMs associated with BDE. In addition, to alleviate the limitation of the low data, we introduced pairwise difference regression (PADRE)56as a data augmentation with the advantages of reducing systematic errors and the improvement in the diversity. The main principle and computational details of PADRE-based data augmentation are described in Supplementary Information. In addition, a detailed pseudocode for the PADRE strategy is provided in Table S5. Benefiting from the hybrid feature and PADRE-based data augmentation, the XGBoost model achieved high precision with R2test = 0.98 and MAE = 8.8 kJ·mol−1, significantly outperforming existing competitive models.
Compared to traditional MLs, deep learning possesses a more powerful learning capacity and can avoid manual feature engineering, thus being broadly applied in various fields. However, due to the limited data of EMs available, existing ML works in the EMs mainly utilized traditional ML algorithms, less concerning deep learning methods. Practically, with the aid of the transfer learning strategy, some deep learning models have exhibited performance superior to traditional machine learning methods even on small-size datasets57,58. Thus, we also attempted to exploit deep learning models for Q and BDE. By comparing them with the traditional ML models, we hope to obtain the optimal prediction model on the one side, and provide methodological guidelines for the use of machine learning in the field of EMs on the other side. Here, we attempted two deep learning pre-training models (MG-BERT58 and 3D-GNN59) derived from large chemical databases. To obtain the prediction uncertainty for improving the reliability of subsequent screening, we modified the two DL models by replacing their last layer with a mixture density network (MDN) layer (see Methods for more details). MDN can model the conditional probability distribution of the output as a mixture of Gaussian, through which the prediction uncertainty can be obtained. Furthermore, the uncertainty quantification can enhance the reliability of the prediction by increasing confidence60,61. As evidenced by the ablation experiment (vide Table S6), the MDN replacement also to some extent improved the predictive performance for the test set compared with the original architecture without the replacement.
MG-BERT model and 3D-GNN were pre-trained on 1.7 million compounds selected from the ChEMBL database and 100 k molecules with three-dimensional conformations from the GEOM-QM9 database62, respectively. Then, we fine-tuned them with energetic compounds. As reflected by Fig. 4b, the two pre-trained models outperform the best traditional ML model (GPR) in the Q prediction task, in particular for 3D-GNN with the highest R2 of 0.95. The superior performance of the two deep learning models should benefit from rich structure knowledge learned in the pre-training stage of the large database. Practically, the high energy density of EMs mainly comes from the energetic groups and skeletons involving a large number of atoms and bonds in the molecule. For 3D-GNN, the molecular graph representation coupled with the 3D information can more accurately describe the connections and interactions between atoms, thus exhibiting the best performance for Q. However, the advantages of the two deep learning models are not extended to the BDE prediction model. As shown in Fig. 4b, the XGBoost model coupled with the PADRE data augmentation and the hybrid feature still exhibits the best performance (R2 = 0.98), better than MG-BERT (R2 = 0.94) and 3D-GNN (R2 = 0.90). The reason should be that the two deep learning frameworks equally treat the atoms and bonds in self-learning structural features of molecules while the bond dissociation energy heavily depends on a specific breaking bond. Thus, the predictive performance of the two DL models on BDE is inferior to the XGBoost model, which simultaneously emphasizes local target bond features in describing the global energetic feature and utilizes the PADRE data augmentation with the advantage of dropping systematic error to alleviate the limitation of the small size of energetic molecules. Taken together, we finally selected the XGBoost34 and the 3D-GNN models as the BDE and Q prediction tools in the subsequent screening, respectively.
a Six traditional machine learning methods and two deep learning models; b Predictive performance on the independent test set for the heat of explosion (left) and bond dissociation energy (right).
Multi-objective optimization based on Pareto front for screening energetic molecules with desired comprehensive performance
As outlined above, it is a challenge to trade off the two contradictory properties (Q and BDE) to achieve high comprehensive performance for EMs. Herein, we employed the Two-Dimensional Improved Probability-based (2D P[I]) Efficient Global Optimization, which is a Bayesian-based multi-objective optimization method63. The P[I] represents the expected probability that the new design will provide an improvement over the current Pareto front, which is calculated by using the predicted mean values and corresponding uncertainties. For the multi-objective optimization, a joint probability density function is constructed based on the predicted means and standard deviations of both objectives. The 2D P[I] is then obtained by integrating this joint probability density over the region of potential Pareto improvement. A molecule with larger 2D P[I] value is considered a more valuable candidate. Thus, 2D P[I] can evaluate the improvement probability across both Q and BDE by incorporating both predicted values and uncertainties so that it identifies the most promising candidates for subsequent experimental validation. To this end, we first need to define the optimal target area for the heat of explosion (Q) and bond dissociation energy (BDE), as illustrated by a rectangle made of red and blue dashed lines in Fig. 5a. For Q, we aim to design new EMs with Q as high as possible. Therefore, we set the highest value of heat of explosion (1896 cal·g−1) of the collected 778 EMs to be the lowest limit, and not setting an upper limit. For BDE, the decomposition reaction barrier above 120 kJ·mol−1 is generally considered as a theoretical metric of applicable stability42,64,65. In addition, 87.5% of 778 real explosives we collected exhibit higher BDE than 120 kJ·mol−1. Thus, to ensure the applicable stability for energetic molecules screened, we adopted 120 kJ·mol−1 as the lowest limit while the highest value of bond dissociation energy of the 778 EMs reported is set as the upper limit (450 kJ·mol−1).
a Illustration of Two-Dimensional Improved Probability (2D P[I]). The optimization goal is to maximize objective 1 and keep objective 2 in the target area. The inset is an illustration of the ML prediction (yellow solid dot), uncertainty (shaded area), and effective 2D P[I] area (yellow area) with respect to a target zone (rectangle made of red and blue dashed lines). b Distribution of Q and BDE of the 778 EMs and 100 candidates from the Pareto front of the multi-objective optimization. Triangles represent the 778 reported energetic molecules, and the red five-pointed stars represent the top 100 molecules in the Pareto front. Yellow dot represents CL-20 that is the most powerful explosive. c Comparison of Q and BDE between the single-objective screening and the multi-objective one for the top 100 compounds from their Pareto fronts. Green and red points denote the single-objective optimization and multi-objective optimization, respectively. d Correlation plot of predicted values and calculated values of Q and BDE for the top 60 new energetic molecules. The blue dot represents the heat of explosion and the red dot represents the bond dissociation energy.
After determining the optimization goal, we used the modified 3D-GNN model and the XGBoost model to separately predict Q and BDE as well as their uncertainties for the 201547 energetic molecules in the search space. The reliability of uncertainties was proved by the calibration curve, as shown in Fig. S4. Based on these values, we calculated the 2D P[I] and selected the top 100 compounds in the Pareto front of 2D P[I] as potential candidates for further QM calculations. Figure 5b shows the distribution of the predicted values for Q and BDE for the 100 candidates and the 778 EMs in the training set, where the 100 candidates in the Pareto front exhibit higher Q values than the 778 EMs reported, and their BDEs are higher than 120 kJ·mol−1, showcasing the effectiveness of the multi-objective optimization strategy.
Comparison with single-objective optimization
To further validate the advantages of the multi-objective design strategy, we conducted a comparison with single-objective optimization screening only based on the heat of explosion. Specifically, we used the optimal 3D-GNN model to predict the Q values and their uncertainties for the 201547 compounds of the search space, and calculate one-dimensional P[I] for each compound. Finally, we selected the top 100 compounds in the Pareto front of 1D P[I] as potential candidates to compare with those from the multi-objective screening.
Figure 5c displays the comparison result of Q and BDE between the single-objective and multi-objective optimizations. It can be seen that the Q screened by the single object is not inferior to the multiple objective design, yet BDEs derived from the multiple objective optimization are significantly superior to the single objective design. Most compounds screened by the single objective screening do not meet the stability required in practical applications (>120 kJ·mol−l)64, of which some are very unstable. The result clearly confirms that the optimization only considering single objective can indeed improve the target property, but at the expense of other properties, highlighting the necessity and effectiveness of our multi-objective optimization strategy.
High-precision QM verification and comprehensive evaluation for generated top candidates with experimental feasibility
Considering the inherent errors in machine learning prediction models, we selected the top 60 compounds based on 2D P[I] ranking for further high-precision QM validation. Figure 5d shows the predicted values inferred from the ML model and calculated values based on the QM method. It can be seen that our prediction models can reliably predict the heat of explosions and bond dissociation energies of unseen compounds. To comprehensively evaluate the detonation performance of 60 candidates, we also calculated other important properties related to detonation performance, such as density (ρ), detonation velocity (D), detonation pressure (P), and oxygen balance (OB), as listed in Table 2. In the work, the detonation velocity and detonation pressure were calculated by using Kamlet–Jacobs scheme, as it can provide reliable prediction for CHNO-based energetic molecules66,67,68. Although it was indicated that the K-J scheme is poorly suitable for compounds with low intrinsic oxidizability, for example, compounds with OB lower than -40%69. Practically, 59 of 60 candidates in Table 2 present higher OB values than -40%, with OB values of approximately 50% molecules > −10% (including zero). Thus, it is reasonable to use the K-J equation for evaluating the D and P performance of these candidates screened. Theoretical models and equations used for calculating the detonation performance are provided in the Supplementary Information, along with experimental data to support the computational reliability (Table S7).
As shown in Table 2, all the 60 candidates exhibit much higher Q than CL-20 (1612.50 cal·g−1 calculated at the same DFT level), in which three compounds (m1, m2 and m3) present the heat of explosion exceeding 2000 cal·g−1. Moreover, the bond dissociation energy of each candidate is greater than 120 kJ·mol−1, meeting the standard for practical application. The result demonstrates that our multi-objective optimization strategy has achieved the desired target. In addition, 52 of the 60 candidates have detonation velocities exceeding 9000 m·s−1, meeting the detonation velocity standard for new high-energy and low-sensitivity energetic compounds70, of which 15 candidates present higher detonation velocities than that of CL-20. Most of the 60 candidates also exhibit excellent performance in the detonation pressure, in which 7 candidates present higher detonation pressures than CL-20. The candidate m28 has the highest density of 1.96 g·cm−3, slightly lower than that of CL-20 (1.97 g·cm−3 calculated at the same DFT level). Practically, a density greater than 1.8 g·cm−3 was considered to meet the requirements of new EMs71. 54 of the 60 candidate EMs present greater densities than 1.8 g·cm−3. Moreover, we also estimated the OB value of the 60 candidates, as a reasonable oxygen balance is essential for achieving high detonation parameters72. In general, a value of zero denotes perfectly oxygen-balanced compositions, negative values for oxygen-poor compositions, and positive values for oxygen-rich compositions. Among the 60 candidates, 7 molecules exhibit zero oxygen balance.
In addition, we also analyzed their molecular structures, as displayed in Fig. S5. Most of the molecules belong to high-nitrogen heterocyclic compounds, which is consistent with the current design concept for high-performance energetic molecules. The skeletons include five-membered heterocycles like furan, triazole, and tetrazole, six-membered heterocycles like triazine, tetrazine, and other single-ring skeletons, and bicyclic skeletons fused with five-membered and six-membered heterocycles. The substituents include various types of -NO2 (such as C-NO2, N-NO2, and O-NO2), -NH2, and -N3). These skeletons and substituents demonstrate the advantage of the RNN-based generative model in de novo producing diverse energetic structures.
Syntheses of energetic compounds remain large challenges in practice due to safety risks. Thus, it is important to reliably evaluate a molecule’s synthetic accessibility in the design stage. To this end, we integrated three evaluation approaches (SAScore73, SCScore74 and SYBA75 score), rather than single metrics. SAScore combines calculated fragment contributions from Pubchem with molecular complexity, in which a lower SAScore implies greater synthetic accessibility. SCScore assesses synthetic complexity from a reaction perspective with the aid of a neural network model trained on 12 million reactions, in which lower scores denote fewer synthetic steps and lower difficulty. SYBA considers the contribution of molecular fragments derived from Bernoulli Naïve Bayes classifier. Higher SYBA scores correspond to greater likelihood of successful synthesis. The combination of three metrics can obtain more reliable evaluation on the synthetic accessibility. More details regarding the three synthetic accessibility are described in Supplementary Information. Table S8 lists the three type scores of the 60 candidate molecules. However, there is still a lack of a statistically evaluated threshold for EMs. To address the problem, we conducted a statistical analysis of the synthetic accessibility scores for the collected 778 real explosives (vide Fig. S6 and Fig. S7), with detailed scores provided in Supplementary Data 2. Thresholds were determined using the interquartile range (IQR) method, which can effectively avoid the influence of extreme values to give a more reasonable range of data distribution. As shown in Table S9, most of the real explosives fall within the ranges identified by Tukey’s method76. Thus, the score thresholds are determined to be smaller than 3.89 for SCScore, larger than −6.13 for SYBA score and smaller than 5.08 for SAScore. Consequently, 25 molecules that meet all three synthetic accessibility criteria were selected as candidates with experimental feasibility. Their structures with calculated Q and BDE values are presented in Fig. 6. A careful check finds that four molecules shadowed in green in Fig. 6 were already reported in experimental synthesis works. Among these, m54 (also known as DNTF) has already served as a melt-cast explosive and energetic filler of propellants77. m2977 and m3078 were considered as promising candidates for next-generation secondary EMs, as they presented high performance and acceptable sensitivity. m2679 exhibits impressive detonation velocity and pressure performance. Notably, none of these four compounds are included in our fine-tuning EM dataset. In addition, eleven generated molecules share known energetic molecular skeletons but with different substituents and substitution positions (vide structures shaded in blue in Fig. 6), expanding structural diversity. More importantly, our framework generates ten new molecular skeletons for energetic molecular design, five of which have been previously observed in non-energetic compounds (vide structures shaded in deep pink in Fig. 6), and the remaining five shaded in light pink in Fig. 6, present entirely novel scaffolds. As our generative framework is a de novo design strategy, rather than a simple combination of existing energetic fragments, the above results clearly confirm the effectiveness and application potential of our multi-objective optimization design framework in practice and provide novel candidates for experimental design and development.
Four green shadowed molecules were already synthesized in experiments. Eleven blue-colored molecules share known energetic molecular scaffolds but with distinctions in substituents and substituted position. Ten pink shadowed molecules represent novel energetic skeletons, in which five molecular scaffolds shadowed in deep pink color have been previously observed in non-energetic compounds and five light pink-colored molecules represent entirely new molecular scaffolds.
Discussion
High energy with applicable stability has been the focus of the development of new EMs with high performance, yet there has been a lack of efficient and intelligent design strategies for improving the comprehensive performance of EMs. Inspired by the need, we exploited a de novo design framework with multi-objective optimization to trade off the contradiction between the energy and the stability such that can achieve high comprehensive performance. To overcome the technical obstacles of machine learning applications resulting from the data scarcity of EMs, our design framework integrates the deep learning molecular generator coupled with the transfer learning, the high accuracy ML-based property prediction models suitable for the small size data, the Pareto front-based multiple objective optimization screening, and molecule recommendation based on both quantum mechanics (QM) validation and synthetic accessibility evaluation.
Machine learning is a data-driven technique. Thus, we first constructed a representative and reliable dataset of EMs (778 real explosives) through extensive literature search and high-precision DFT calculations. Then, we exploited a deep learning generator to de novo produce a massive energetic molecule space, with the advantage of avoiding the limitations of combinatorial chemistry in structural diversity and rationality. Herein, we developed a RNN-based molecule generator coupled with the transfer learning strategy to address the limitation of the small size EM dataset on the deep learning, targeting energetic molecules generation with diverse structures (2 × 105 energetic molecules). To rapidly and accurately evaluate Q and BDE of new molecules in the search space, we exploited six classical machine learning models coupled with new feature representations and data augmentation strategies. In addition, we also developed two deep learning models based on pre-trained MG-BERT and 3D-GNN frameworks. In these model constructions, we revealed the relationship between the feature presentation, the model architecture and the chemical nature of the target property to provide methodological guidelines for the application of machine learning in the low data fields. The results show that 3D-GNN achieves the best performance in the Q prediction (R2 = 0.95), outperforming the traditional MLs and MG-BERT. The reason should be that the 3D molecular graph representation can better describe the connections between atoms while the explosion process involves breaking of a large number of bonds in the molecule. For the BDE prediction, the XGBoost model coupled with the hybrid feature and PADRE data augmentation strategy presents the best performance (R2 = 0.98), benefiting from the fact that the feature hybrid can consider the global energetic structure while simultaneously focusing on the breaking bond. Additionally, the PADRE data augmentation can drop to some extent systematic error. The comparison of different ML models indicates that the model selection should consider the chemical nature of the target property. Despite the high prediction accuracy achieved, it is noted that the data under this study is derived from the computational level. In general, ML can better capture causality underlying the computational data than the experimental data with inclusion of instrumental noise, as indicated by Zhang’s work21. However, when facing the scarcity of the experimental data, the computational data can serve as a good alternative.
With the obtained optimal prediction models, we used multi-objective optimization with 2D P[I] to trade off the two contradictory properties (Q and BDE), which simultaneously consider the predicted values and predicted uncertainty. Based on the Pareto front, we quickly screened out promising new energetic molecules with high comprehensive performance from the massive search space. In addition, the comparisons with the single-objective design of Q further demonstrate the advantage of our multi-objective optimization design in trading off the high energy and low sensitivity of EMs. Subsequently, high-precision DFT was used to calculate key properties of the top 60 candidates, which present diverse structures and are superior to CL-20 in the Q and BDE properties. Finally, we combined three synthetic accessibility scores coupled with score thresholds derived from our statistical analysis on the 778 real explosives to identify 25 energetic candidates with highly comprehensive performance and synthesis feasibility, which provide promising energetic candidates for experiments. Practically, four candidates not included in our initial training set were successfully synthesized in previous experimental works, confirming the effectiveness and application potential of our de novo design framework. More importantly, ten novel energetic scaffolds were generated, offering valuable skeleton candidates for further developing high-energy and low-sensitivity explosives. Also, the technical strategies embedded in our work provide methodological guidelines for the application of AI in chemistry and material fields. The codes of the molecular generation model, the best prediction models for Q and BDE, and the 2D P[I] multi-objective optimization are freely available at https://github.com/cholin01/EM_MOO. We expect that they will become useful tools for aiding the design of new energetic molecules with desired comprehensive properties.
Methods
Calculation of heat explosion
The heat of explosion (Q) is an important indicator for evaluating the detonation performance of EMs. The higher the Q, the greater the amount of heat released per unit mass. In this work, Q was calculated based on the method proposed by Kamlet and Jacobs80, which assumes that nitrogen forms N2(g), hydrogen fully converts to H2O(g), and oxygen prioritizes CO2(g) over CO. This model simplify the reaction products, not accounting for complex thermochemical equilibria like thermochemical codes, but its reliability was accepted for explosives containing C, H, O and N66,67,68. Thus, it has been widely adopted in high-throughput computation and early-stage virtual screening for energetic molecules due to its balance between computational efficiency and acceptable accuracy. Based on the K-J equation, the enthalpy of formation (Hf) of each component in the reaction first needs to be calculated. The computational method for the heat of formation is described in the Supplementary Information. After obtaining the solid-phase formation enthalpy (\(\Delta {H}_{f,\mathrm{solid}}\)), the heat of explosion of the energetic compound CaHbOcNd can be calculated using the formulas listed in Table 3.
Calculation of bond dissociation energy
The bond dissociation energy of the trigger bond at 298 K and 1 atm corresponds to the enthalpy change between the molecule and the two free radicals generated by homolysis81, as reflected by Eq. (1). Firstly, the NBO analysis was utilized to find the trigger bond based on the value of the Wiberg bond index for each molecule. A smaller Wiberg bond index indicates a higher likelihood of the chemical bond becoming a thermally triggered bond82. After identifying the thermally triggered bonds, the bond dissociation energy was calculated using Eq. (2).
Where \({\text{H}}_{298}\left({\text{A}}\cdot \right)\), \({\text{H}}_{298}\left({\text{B}}\cdot \right)\) and \({\text{H}}_{298}\left(\text{A}-\text{B}\right)\) refer to the enthalpy at 298 K of the free radicals \({\text{A}}\cdot\), \({\text{B}}\cdot\) and molecule \(\text{A}-\text{B}\), respectively. The calculations of \({\text{H}}_{298}\left({\text{A}}\cdot \right)\), \({\text{H}}_{298}\left({\text{B}}\cdot \right)\) and \({\text{H}}_{298}\left(\text{A}-\text{B}\right)\) were performed at the level of B3LYP/ 6-31 G**.
All quantum mechanics calculations were carried out using the Gaussian 09 program83. The vibration frequencies were computed at the same level of theory to confirm that the optimized structures are the minimum on the potential energy surface. The Wiberg bond index of each bond was calculated using Multiwfn software84.
Deep learning generative models
The generative model comprises a pre-trained RNN and a generation RNN. Both RNN models have a three-layer LSTM architecture followed by a fully connected layer with a Softmax function. Each LSTM layer and fully connected layer contains 256 units. For some key hyperparameters, the batch size of 128 and the standard sampling temperature of 1.0 were used, referred to some generative models85,86. Other hyperparameters, such as the learning rate (0.001), dropout rate (0.2), and maximum sampling sequence length (200), were derived from optimization, as evidenced by Fig. S8 and Table S10. The forward pass for the pre-trained RNN and the generative RNN are similar. Given a sequence of tokens (\({\text{v}}_{1}\), …, \({\text{v}}_{\text{i}}\)), the pre-trained RNN or the generative RNN estimates the probability of the (i + 1)-th token \(({{\rm{P}}}_{{\rm{\theta }}}({{\rm{v}}}_{{\rm{i}}+1}))\) in terms of Eq. (3), where the model parameter θ is optimized by the Adam87 algorithm to minimize the cross entropy \({\text{l}}_{\text{i}}\) (vide Eq. (4))
Deep learning predictive models
In the work, we introduced two deep learning pre-trained models (MG-BERT58 and 3D-GNN59) derived from the large databases for predicting the Q and BDE of EMs with the aid of a transfer learning strategy. In order to enable them to output the prediction uncertainty, we did some modifications to the architectures of MG-BERT and 3D-GNN, as illustrated by Fig. 7.
We replaced the last fully connected layer of the two models with a MDN layer to obtain the uncertainty (σ) besides the prediction value (μ). The MG-BERT model architecture consists of an embedding layer, six Transformer encoder layers (orange area), a fully connected layer, and a MDN layer. The 3D-GNN model architecture is composed of an embedding layer, six message-passing layers (blue area), a mask layer, and a MDN layer.
MG-BERT58 model can learn from a large number of unlabeled molecules through the masked atom recovery task to effectively represent atomic and molecular structures and outperform existing methods on 11 ADMET datasets related to drug properties. The pre-training of the MG-BERT model was conducted on 1.7 million compounds randomly selected from the ChEMBL database43. The model architecture was composed of three components: an embedding layer, six Transformer encoder layers88, and a task-related output layer. In the embedding layer, the input word token was mapped into a continuous vector space using an embedding matrix. In the Transformer encoder layer, each word token exchanged information with others through a global attention mechanism. The embedding layer and Transformer layers were shared during the pre-training and fine-tuning stages, yet the fully connected neural network layer was not shared but fine-tuned using the target EM dataset. Dropout strategy was adopted in the output layer to minimize overfitting89. The original framework of the model only outputs prediction values, yet we need to obtain the prediction uncertainty besides the predicted value, which will be used in subsequent multi-objective screening. To this end, we replaced the final fully connected layer of the model with a Mixture Density Network (MDN) layer, as shown in Fig. 7. The MDN layer can overlay a certain distribution (usually Gaussian distribution) with specific weights to fit the final distribution, thus outputting the final predicted value and the prediction uncertainty. The input of the MG-BERT model is molecular SMILES, and the SMILES strings of the 778 explosives under study range from 10 characters to 207 characters. These molecules distribute across the five intervals of the SMILES length, such as 5.3% in the [0, 25] interval, 35.0% in [25, 50], 30.8% in [50, 75], 16.7% in [75, 100], and 12.2% in the range greater than 100. Thus, we used the five intervals for stratified sampling to ensure the representativeness of the training and testing sets. The average Tanimoto similarity based on Morgan fingerprints between the training and test sets were calculated to be 0.158, being low similarity, in turn implying low structure leakage in the data split.
The pretraining data for the 3D-GNN59 model consists of 100k molecular conformations from the large-scale molecular dataset GEOM-QM962. The model architecture includes an embedding layer, six message passing layers, and a mask layer. The embedding layer transformed the input dimension of the feature vectors into the dimension of vectors in the latent space to obtain node-level and graph-level molecular representations. The message-passing layers were used to propagate and update node information. The mask layer enhanced the model’s generalization performance by masking parts of the information in the molecules. Similarly, the last layer of 3D-GNN was replaced by MDN in order to obtain the prediction uncertainty. To obtain the 3D structures for each compound in the EMs dataset, we utilized the RDKit package to generate the 3D conformations from SMILES strings.
Model training and evaluation
Since the data size in the work is limited, traditional K-Fold cross-validation may result in quite different model performances for different folds. Therefore, we used shuffle split cross-validation to evaluate ML model performance adequately90,91. For each model, the shuffle split cross-validation was performed by randomly splitting the dataset into a training, validation, and independent test set based on an 8:1:1 ratio by 20 repeats (The MG-BERT model first performed a stratified sampling, and then the Shuffle Split cross-validation). After 20 iterations, the averaged evaluation metrics were used to evaluate the model performance, including the mean absolute error (MAE), the root mean square error (RMSE), and the coefficient of determination (R2), as shown in Eqs. (5)-(7).
Where \(\text{N}\) is the number of samples, \({\text{y}}_{\text{i}}\) is the real value, \({\hat{\text{y}}}_{\text{i}}\) is the predicted value,\(\,\bar{\text{y}}\) is the mean of the real values. The value of R2 ranges from 0 to 1, and 1 indicates a perfect fitting.
Data availability
Computational data associated with this work are provided in the Supplementary Information.
Code availability
The codes of the molecular generation model, the best models for Q and BDE, and the 2D P[I] multi-objective optimization under study are freely available at https://github.com/cholin01/EM_MOO.
References
Agrawal, J. P. High energy materials: propellants, explosives and pyrotechnics (John Wiley & Sons, 2010).
Banik, S. et al. Facile synthesis of nitroamino-1,3,4-oxadiazole with azo linkage: a new family of high-performance and biosafe energetic materials. J. Mater. Chem. A 10, 22803–22811 (2022).
Li, J. et al. Tri-explosophoric groups driven fused energetic heterocycles featuring superior energetic and safety performances outperforms HMX. Nat. Commun. 13, 5697 (2022).
Jain, A. et al. Commentary: The Materials Project: a materials genome approach to accelerating materials innovation. APL Mater. 1, 011002 (2013).
Tsyshevsky, R., Smirnov, A. S. & Kuklja, M. M. Comprehensive end-to-end design of novel high energy density materials: III. Fused heterocyclic energetic compounds. J. Phys. Chem. C. 123, 8688–8698 (2019).
Wang, Y. et al. Accelerating the discovery of insensitive high-energy-density materials by a materials genome approach. Nat. Commun. 9, 2444 (2018).
Badgujar, D. M., Talawar, M. B., Asthana, S. N. & Mahulikar, P. P. Advances in science and technology of modern energetic materials: An overview. J. Hazard. Mater. 151, 289–305 (2008).
Simpson, R. L. et al. CL-20 performance exceeds that of HMX and its sensitivity is moderate. Propellants Explos. Pyrotech. 22, 249–255 (1997).
Li, H. et al. Molecular and crystal features of thermostable energetic materials: guidelines for architecture of “Bridged” compounds. ACS Cent. Sci. 6, 54–75 (2020).
Qian, W., Xue, X., Liu, J. & Zhang, C. Molecular forcefield methods for describing energetic molecular crystals: a review. Molecules 27, 1611 (2022).
Butler, K. T., Davies, D. W., Cartwright, H., Isayev, O. & Walsh, A. Machine learning for molecular and materials science. Nature 559, 547–555 (2018).
Li, Y., Zhu, R., Wang, Y., Feng, L. & Liu, Y. Center-environment deep transfer machine learning across crystal structures: from spinel oxides to perovskite oxides. npj Comput. Mater. 9, 109 (2023).
Wang, X. et al. Multi-modal deep learning enables efficient and accurate annotation of enzymatic active sites. Nat. Commun. 15, 7348 (2024).
Zhang, H. et al. Fusion of quality evaluation metrics and convolutional neural network representations for ROI filtering in LC–MS. Anal. Chem. 95, 612–620 (2023).
Feng, J., Dong, Z., Ji, Y. & Li, Y. Accelerating the discovery of metastable IrO2 for the oxygen evolution reaction by the self-learning-input graph neural network. JACS Au 3, 1131–1140 (2023).
Wu, Z. et al. Leveraging language model for advanced multiproperty molecular optimization via prompt engineering. Nat. Mach. Intell. 6, 1359−1369(2024).
Nguyen, P. et al. Predicting energetics materials’ crystalline density from chemical structure by machine learning. J. Chem. Inf. Model. 61, 2147–2158 (2021).
Huang, X. et al. Applying machine learning to balance performance and stability of high energy density materials. Iscience 24, 102240 (2021).
Liu, J. et al. Screening heat-resistant energetic molecules via deep learning and high-throughput computation. Chem. Eng. J. 507, 160218 (2025).
Chen, C. et al. Accurate machine learning models based on small dataset of energetic materials through spatial matrix featurization methods. J. Energy Chem. 63, 364–375 (2021).
Zhang, Z. et al. “Unraveling the tangle” of calculated solid-state enthalpies of formation of C-H-N-O energetic materials with a high-precision approach based on the predicted asymmetric cell total energy evaluation. Fuel 404, 136307 (2026).
Wang, X. et al. Fast explosive performance prediction via small-dose energetic materials based on time-resolved imaging combined with machine learning. J. Mater. Chem. A 10, 13114–13123 (2022).
Song, S. et al. Accelerating the discovery of energetic melt-castable materials by a high-throughput virtual screening and experimental approach. J. Mater. Chem. A 9, 21723–21731 (2021).
Wu, Q. et al. Reverse design of high-detonation-velocity organic energetic compounds based on an accurate BPNN with wide applicability. J. Mater. Chem. A 13, 1470–1477 (2025).
Lansford, J. L., Barnes, B. C., Rice, B. M. & Jensen, K. F. Building chemical property models for energetic materials from small datasets using a transfer learning approach. J. Chem. Inf. Model. 62, 5397–5410 (2022).
Wang, R., Liu, J., He, X., Xie, W. & Zhang, C. Decoding hexanitrobenzene (HNB) and 1,3,5-triamino-2,4,6-trinitrobenzene (TATB) as two distinctive energetic nitrobenzene compounds by machine learning. Phys. Chem. Chem. Phys. 24, 9875–9884 (2022).
Liu, W.-H., Liu, Q.-J., Liu, F.-S. & Liu, Z.-T. Machine learning approaches for predicting impact sensitivity and detonation performances of energetic materials. J. Energy Chem. 102, 161–171 (2025).
Cheng, P., Jin, Y., Wang, D. & Tao, S. Design and computational screening of high-energy, low-sensitivity bistetrazole-based energetic molecules. RSC Adv. 15, 11645–11654 (2025).
Guo, S. et al. Discovery of high energy and stable prismane derivatives by the high-throughput computation and machine learning combined strategy. Firephyschem 4, 55–62 (2024).
Wen, L. et al. Accelerating molecular design of cage energetic materials with zero oxygen balance through large-scale database search. J. Phys. Chem. Lett. 12, 11591–11597 (2021).
Zang, X. et al. Prediction and construction of energetic materials based on machine learning methods. Molecules 28, 322 (2023).
Jiao, F., Xiong, Y., Li, H. & Zhang, C. Alleviating the energy & safety contradiction to construct new low sensitivity and highly energetic materials through crystal engineering. CrystEngComm 20, 1757–1768 (2018).
St. John, P. C., Guan, Y., Kim, Y., Kim, S. & Paton, R. S. Prediction of organic homolytic bond dissociation enthalpies at near chemical accuracy with sub-second computational cost. Nat. Commun. 11, 2328 (2020).
Gou, Q. et al. Exploring an accurate machine learning model to quickly estimate stability of diverse energetic materials. Iscience 27, 109452 (2024).
Wen, M., Blau, S. M., Spotte-Smith, E. W. C., Dwaraknath, S. & Persson, K. A. BonDNet: a graph neural network for the prediction of bond dissociation energies for charged molecules. Chem. Sci. 12, 1858–1868 (2021).
Duan, C., Nandy, A., Terrones, G. G., Kastner, D. W. & Kulik, H. J. Active learning exploration of transition-metal complexes to discover method-insensitive and synthetically accessible chromophores. JACS Au 3, 391–401 (2022).
Zeman, S. & Jungová, M. Sensitivity and performance of energetic materials. Propellants Explos. Pyrotech. 41, 426–451 (2016).
Yan, Q. L. & Zeman, S. Theoretical evaluation of sensitivity and thermal stability for high explosives based on quantum chemistry methods: a brief review. Int. J. Quantum Chem. 113, 1049–1061 (2013).
Tan, B. S. et al. Two important factors influencing shock sensitivity of nitro compounds: bond dissociation energy of X-NO2 (X = C, N, O) and Mulliken charges of nitro group. J. Hazard. Mater. 183, 908–912 (2010).
Mathieu, D. Sensitivity of energetic materials: theoretical relationships to detonation performance and molecular structure. Ind. Eng. Chem. Res. 56, 8191–8201 (2017).
Li, C. et al. Correlated RNN framework to quickly generate molecules with desired properties for energetic materials in the low data regime. J. Chem. Inf. Model. 62, 4873–4887 (2022).
Liu, J. et al. High-throughput design of energetic molecules. J. Mater. Chem. A 11, 25031–25044 (2023).
Mendez, D. et al. ChEMBL: towards direct deposition of bioassay data. Nucleic Acids Res. 47, D930–D940 (2018).
Lancet, D. & Pecht, I. Spectroscopic and immunochemical studies with nitrobenzoxadiazolealanine, a fluorescent dinitrophenyl analog. Biochem 16, 5150–5157 (1977).
Mellor, C. L. et al. Molecular fingerprint-derived similarity measures for toxicological read-across: recommendations for optimal use. Regul. Toxicol. Pharm. 101, 121–134 (2019).
Béquignon, O. J. M. et al. Collaborative SAR modeling and prospective in vitro validation of oxidative stress activation in human HepG2 Cells. J. Chem. Inf. Model. 63, 5433–5445 (2023).
Gu, G. H., Plechac, P. & Vlachos, D. G. Thermochemistry of gas-phase and surface species via LASSO-assisted subgraph selection. React. Chem. Eng. 3, 454–466 (2018).
Dou, B. et al. Machine learning methods for small data challenges in molecular science. Chem. Rev. 123, 8736–8780 (2023).
Xu, P., Ji, X., Li, M. & Lu, W. Small data machine learning in materials science. npj Comput. Mater. 9, 42 (2023).
Mani-Varnosfaderani, A., Soleimani, M. & Alizadeh, N. Least absolute shrinkage and selection operator as a multivariate calibration tool for simultaneous determination of diphenylamine and its nitro derivatives in propellants. Propellants Explos. Pyrotech. 43, 379–389 (2018).
Shi, Z. et al. Machine-learning-assisted high-throughput computational screening of high performance metal–organic frameworks. Mol. Syst. Des. Eng. 5, 725–742 (2020).
Gu, B. et al. Incremental learning for ν-support vector regression. Neural Netw. 67, 140–150 (2015).
Higgins, K., Valleti, S. M., Ziatdinov, M., Kalinin, S. V. & Ahmadi, M. Chemical robotics enabled exploration of stability in multicomponent lead halide perovskites via machine learning. ACS Energy Lett. 5, 3426–3436 (2020).
Meyer, J. G., Liu, S., Miller, I. J., Coon, J. J. & Gitter, A. Learning drug functions from chemical structures with convolutional neural networks and random forests. J. Chem. Inf. Model. 59, 4438–4449 (2019).
Zhu, X.-Y. et al. Prediction of multicomponent reaction yields using machine learning. Chin. J. Chem. 39, 3231–3237 (2021).
Tynes, M. et al. Pairwise difference regression: a machine learning meta-algorithm for improved prediction and uncertainty quantification in chemical search. J. Chem. Inf. Model. 61, 3846–3857 (2021).
Gupta, V. et al. Cross-property deep transfer learning framework for enhanced predictive analytics on small materials data. Nat. Commun. 12, 6595 (2021).
Zhang, X.-C. et al. MG-BERT: leveraging unsupervised atomic representation learning for molecular property prediction. Brief. Bioinform. 22, bbab152 (2021).
Jiao, R., Han, J., Huang, W., Rong, Y. & Liu, Y. Energy-motivated equivariant pretraining for 3D molecular graphs. Proc. AAAI Conf. Artif. Intell. 37, 8096–8104 (2023).
Fan, W., Zeng, L. & Wang, T. Uncertainty quantification in molecular property prediction through spherical mixture density networks. Eng. Appl. Artif. Intell. 123, 106180 (2023).
Li, D. et al. Enhancing probabilistic hydrological predictions with mixture density Networks: accounting for heteroscedasticity and Non-Gaussianity. J. Hydrol. 641, 131737 (2024).
Axelrod, S. & Gómez-Bombarelli, R. GEOM, energy-annotated molecular conformations for property prediction and molecular generation. Sci. Data 9, 185 (2022).
Keane, A. J. Statistical improvement criteria for use in multiobjective design optimization. AIAA J. 44, 879–891 (2006).
Ma, Q. et al. Theoretical investigations on 4,4′,5,5′-tetranitro-2,2′-1H,1′H-2,2′-biimidazole derivatives as potential nitrogen-rich high energy materials. J. Phys. Org. Chem. 28, 31–39 (2015).
Liu, J., He, X., Xiong, Y., Nie, F. & Zhang, C. Benchmark calculations and error cancelations for bond dissociation enthalpies of X-NO2. Def. Technol. 22, 144–155 (2023).
Wang, Y. et al. A simple method for the prediction of the detonation performances of metal-containing explosives. J. Phys. Chem. A 118, 4575–4581 (2014).
Gao, T. Y., Ji, Y. J., Liu, C. & Li, Y. Y. Molecular descriptor-enhanced graph neural network for energetic molecular property prediction. Sci. China Mater. 67, 1243–1252 (2024).
Xi, H.-W., Dev, S. P. & Lim, K. H. Advances in the modelling and simulation of high-energy density materials. J. Mol. Model. 31, 120 (2025).
Kimber, P. et al. Machine learning densities, detonation velocities, and formation enthalpies of energetic materials using quantum chemistry descriptors. J. Chem. Theory Comput. 21, 8406–8419 (2025).
Klapötke, Sabaté, T. M. & Bistetrazoles, C. M. Nitrogen-rich, high-performing, insensitive energetic compounds. Chem. Mater. 20, 3629–3637 (2008).
Zhang, J., Zhang, Q., Vo, T. T., Parrish, D. A. & Shreeve, J.nM. Energetic salts with π-stacking and hydrogen-bonding interactions lead the way to future energetic materials. J. Am. Chem. Soc. 137, 1697–1704 (2015).
Saraf, S. R., Rogers, W. J. & Mannan, M. S. Prediction of reactive hazards based on molecular structure. J. Hazard. Mater. 98, 15–29 (2003).
Ertl, P. & Schuffenhauer, A. Estimation of synthetic accessibility score of drug-like molecules based on molecular complexity and fragment contributions. J. Cheminform. 1, 8 (2009).
Coley, C. W., Rogers, L., Green, W. H. & Jensen, K. F. SCScore: Synthetic complexity learned from a reaction corpus. J. Chem. Inf. Model. 58, 252–261 (2018).
Vorsilak, M., Kolar, M., Cmelo, I. & Svozil, D. SYBA: Bayesian estimation of synthetic accessibility of organic compounds. J. Cheminform. 12, 35 (2020).
Páez, A. & Boisjoly, G. In Discrete Choice Analysis with R 25-64 (Springer International Publishing, 2022).
Fershtat, L. L. et al. Assembly of nitrofurazan and nitrofuroxan frameworks for high-performance energetic materials. Chempluschem 82, 1315–1319 (2017).
Zhang, G. et al. Thermal stability assessment of 3,4-bis(3-nitrofurazan-4-yl)furoxan (DNTF) by accelerating rate calorimeter (ARC). J. Therm. Anal. Calorim. 126, 1185–1190 (2016).
Larin, A. A. et al. Pushing the energy-sensitivity balance with high-performance Bifuroxans. ACS Appl. Energy Mater. 3, 7764–7771 (2020).
Kamlet, M. J. & Jacobs, S. J. Chemistry of Detonations. I. A Simple method for calculating detonation properties of C–H–N–O Explosives. J. Chem. Phys. 48, 23–2 (1968).
Politzer, P. & Murray, J. S. Relationships between dissociation energies and electrostatic potentials of C–NO2 bonds: applications to impact sensitivities. J. Mol. Struct. 376, 419–424 (1996).
Zhang, Y. L., Fan, L., Su, C., Shu, Z. Y. & Zhang, H. J. Application of machine learning in developing a quantitative structure-property relationship model for predicting the thermal decomposition temperature of nitrogen-rich energetic ionic salts. RSC Adv. 14, 37737–37751 (2024).
Frisch, M. et al. Gaussian 09, Revision D. 01, Gaussian, Inc., Wallingford CT. (2009).
Lu, T. & Chen, F. Multiwfn: a multifunctional wavefunction analyzer. J. Comput. Chem. 33, 580–592 (2012).
Mahmood, O., Mansimov, E., Bonneau, R. & Cho, K. Masked graph modeling for molecule generation. Nat. Commun. 12, 3156 (2021).
Bagal, V., Aggarwal, R., Vinod, P. K. & Priyakumar, U. D. MolGPT: molecular generation using a transformer-decoder model. J. Chem. Inf. Model. 62, 2064–2076 (2022).
Kingma, D. P. & Ba, J. Adam: a method for stochastic optimization. Preprint at https://doi.org/10.48550/arXiv.1412.6980 (2014).
Vaswani, A. et al. Attention is all you need. Adv. Neural Inf. Process. Syst. 30, 6000–6010 (2017).
Srivastava, N., Hinton, G., Krizhevsky, A., Sutskever, I. & Salakhutdinov, R. J. T. Dropout: a simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15, 1929–1958 (2014).
Haffiez, N. et al. Exploration of machine learning algorithms for predicting the changes in abundance of antibiotic resistance genes in anaerobic digestion. Sci. Total Environ. 839, 156211 (2022).
Xie, Y. et al. A property-oriented adaptive design framework for rapid discovery of energetic molecules based on small-scale labeled datasets. RSC Adv. 11, 25764–25776 (2021).
Acknowledgements
This project was supported by the Advanced Materials-National Science and Technology Major Project (Grant No. 2024ZD0607000), the National Natural Science Foundation of China (No. 62475177) and the Sichuan International Science and Technology Innovation Cooperation Project (No. 2024YFHZ0328).
Author information
Authors and Affiliations
Contributions
J.L. and Q.G. contributed equally to this work. J.L., Q.G.: conceptualization, investigation, methodology, model, formal analysis, writing—original draft; S.L., Y.G.: formal analysis, investigation; Y.H.: data curation, investigation; Y.L.: methodology, data curation; X.P.: funding acquisition, project administration, writing – review & editing, conceptualization, supervision.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, J., Gou, Q., Li, S. et al. De novo multi-objective generation framework for energetic materials with trading off energy and stability. npj Comput Mater 11, 360 (2025). https://doi.org/10.1038/s41524-025-01845-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41524-025-01845-6