Abstract
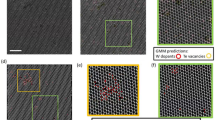
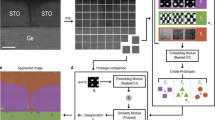
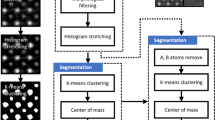
Atomic-scale defects govern many functional properties of materials, yet their systematic identification and quantification remain challenging because supervised learning approaches require extensive labeled datasets, which are scarce in atomic-resolution microscopy due to the complexity and diversity of defect structures. To overcome this limitation, we introduce a fully unsupervised machine learning framework capable of discovering and clustering defect structures without prior labeling or predefined defect classes. The framework employs a convolutional variational autoencoder (CVAE) to reconstruct ideal, defect-free images, enabling the generation of difference images that isolate local structural anomalies. From these, 47 features are extracted and refined through a three-tier feature selection process to minimize redundancy and noise. Dimensionality reduction via principal component analysis (PCA), combined with silhouette score optimization, guides the determination of the optimal cluster number prior to applying k-means clustering, which yields well-separated groups corresponding to distinct defect types. Validated on CdTe and SrTiO3 datasets, this unsupervised, label-free approach enables high-throughput defect discovery and clustering in scanning transmission electron microscopy (STEM) and related imaging modalities.
Similar content being viewed by others
Data availability
The data and Python codes supporting the findings of this study are openly available on GitHub at https://github.com/RAW-Ayyubi/defect-classification-in-stem-images.
References
Liu, E. et al. High Responsivity Phototransistors Based on Few-Layer ReS2 for Weak Signal Detection. Adv. Funct. Mater. 26, 1938–1944 (2016).
Shim, J. et al. High-Performance 2D Rhenium Disulfide (ReS2) Transistors and Photodetectors by Oxygen Plasma Treatment. Adv. Mater. 28, 6985–6992 (2016).
Lagunas, F.et al. In Situ Formation of Ripplocations in Hybrid Organic–Inorganic MXenes. Adv. Mater. 37, https://doi.org/10.1002/adma.202411669 (2025).
Rhodes, D., Chae, S. H., Ribeiro-Palau, R. & Hone, J. Disorder in van der Waals heterostructures of 2D materials. Nat. Mater. 18, 541–549 (2019).
Wu, Z. & Ni, Z. Spectroscopic investigation of defects in two-dimensional materials. Nanophotonics 6, 1219–1237 (2017).
Li, C. et al. Column-by-column observation of dislocation motion in CdTe: Dynamic scanning transmission electron microscopy. Appl. Phys. Lett. 109, 143107 (2016).
Li, C., Poplawsky, J., Yan, Y. & Pennycook, S. J. Understanding individual defects in CdTe thin-film solar cells via STEM: From atomic structure to electrical activity. Mater. Sci. Semiconductor Process.65 64–76 https://doi.org/10.1016/j.mssp.2016.06.017 (2017).
Jiang, J., Xu, T., Lu, J., Sun, L. & Ni, Z. Defect Engineering in 2D Materials: Precise Manipulation and Improved Functionalities. Research 2019, 4641739 (2019).
Hong, J. et al. Exploring atomic defects in molybdenum disulphide monolayers. Nat. Commun. 6, 6293 (2015).
Qiu, H. et al. Hopping transport through defect-induced localized states in molybdenum disulphide. Nat. Commun. 4, 2642 (2013).
Guan, X. et al. Targeted elimination of tetravalent-Sn-induced defects for enhanced efficiency and stability in lead-free NIR-II perovskite LEDs. Nat. Commun. 15, 9913 (2024).
Vaibhav, V. et al. Experimental identification of topological defects in 2D colloidal glass. Nat. Commun. 16, 55 (2025).
Bielinski, N. et al. Floquet–Bloch manipulation of the Dirac gap in a topological antiferromagnet. Nat. Phys. 21, 458–463 (2025).
Ayyubi, R. A. W., Horing, N. J. M. & Sabeeh, K. Effect of pseudospin polarization on wave packet dynamics in graphene antidot lattices (GALs) in the presence of a normal magnetic field. J. Appl. Phys. 129, 074301 (2021).
Pennycook, S. J. & Jesson, D. E. High-resolution Z-contrast imaging of crystals. Ultramicroscopy 37, 14–38 (1991).
Guo, J. et al. Effect of selenium and chlorine co-passivation in polycrystalline CdSeTe devices. Appl. Phys. Lett. 115, 153901 (2019).
Shi, C. et al. Domain-dependent strain and stacking in two-dimensional van der Waals ferroelectrics. Nat. Commun. 14, 7168 (2023).
Lagunas, F. et al. Ion-Exchange Effects in One-Dimensional Lepidocrocite TiO2: A Cryogenic Scanning Transmission Electron Microscopy and Density Functional Theory Study. Chem. Mater. 36, 2743–2755 (2024).
Zangeneh, D., Sapkota, B., Uppuluri, R. & Klie, R. F. Atomic-Scale Tracking of Beam-Induced Phase Transitions in MgCr1.5Mn0.5O4. Chem. Mater. 37, 1491–1499 (2025).
Spurgeon, S. R. et al. Towards data-driven next-generation transmission electron microscopy. Nature Materials vol. 20 274–279 https://doi.org/10.1038/s41563-020-00833-z (2021).
Kalinin, S. V. et al. Probe microscopy is all you need. Machine Learning: Science and Technology vol. 4 https://doi.org/10.1088/2632-2153/acccd5 (2023).
Liu, Y., Checa, M. & Vasudevan, R. K. Synergizing human expertise and AI efficiency with language model for microscopy operation and automated experiment design. Mach. Learn Sci. Technol. 5, 02LT01 (2024).
Kalinin, S. V. et al. Machine learning for automated experimentation in scanning transmission electron microscopy. NPJ Comput Mater. 9, 227 (2023).
Doty, C. et al. Design of a graphical user interface for few-shot machine learning classification of electron microscopy data. Comput Mater. Sci. 203, 111121 (2022).
Shi, C. et al. Uncovering material deformations via machine learning combined with four-dimensional scanning transmission electron microscopy. NPJ Comput Mater. 8, 114 (2022).
Kalinin, S. V. et al. Machine learning in scanning transmission electron microscopy. Nat. Rev. Methods Prim. 2, 11 (2022).
Dan, J. et al. Learning Motifs and Their Hierarchies in Atomic Resolution Microscopy. Sci. Adv vol. 8 https://www.science.org (2022).
Ayyubi, R. A. W., Sultanov, S., Buban, J. P. & Klie, R. F. Unsupervised Machine Learning for Atomic-Resolution S(TEM) Image Analysis and Management. Microsc. Microanal. 31, ozaf048.1072 (2025).
Biswas, A., Vasudevan, R., Ziatdinov, M. & Kalinin, S. V. Optimizing training trajectories in variational autoencoders via latent Bayesian optimization approach. Mach. Learn Sci. Technol. 4, 015011 (2023).
Kingma, D. P. & Welling, M. Auto-Encoding Variational Bayes. arXiv:1312.6114 https://doi.org/10.48550/arXiv.1312.6114 (2014).
Prifti, E., Buban, J. P., Thind, A. S. & Klie, R. F. Variational Convolutional Autoencoders for Anomaly Detection in Scanning Transmission Electron Microscopy. Small 19, 2205977 (2023).
Ayyubi, R. A. W., Buban, J. P. & Klie, R. F. Automated Defect Detection in Atomic Resolution STEM Images: A Machine Learning Approach with Variational Convolutional Autoencoders. Microsc. Microanal. 30, ozae044.180 (2024).
Yan, X., Yang, J., Sohn, K. & Lee, H. Attribute2Image: Conditional Image Generation from Visual Attributes. arXiv:1512.00570 https://doi.org/10.48550/arXiv.1512.00570 (2015).
Paulauskas, T. et al. Atomic scale study of polar Lomer-Cottrell and Hirth lock dislocation cores in CdTe. Acta Crystallogr A Found. Adv. 70, 524–531 (2014).
Sultanov, S., Ayyubi, R. A. W., Buban, J. P. & Klie, R. F. Robust Spectral Anomaly Detection in EELS Spectral Images via 3D Convolutional Variational Autoencoders. Small 21, 2503019 (2025).
Mitić, N. S. et al. Correlation-based feature selection of single cell transcriptomics data from multiple sources. J. Big Data 12, 4 (2025).
Ranjan, B. et al. DUBStepR is a scalable correlation-based feature selection method for accurately clustering single-cell data. Nat. Commun. 12, 5849 (2021).
Pudjihartono, N., Fadason, T., Kempa-Liehr, A. W. & O’Sullivan, J. M. A Review of Feature Selection Methods for Machine Learning-Based Disease Risk Prediction. Front. Bioinform. 2 https://doi.org/10.3389/fbinf.2022.927312 (2022).
Zeb, M. H. et al. Machine Learning-Enhanced Prediction of Inorganic Semiconductor Bandgaps for Advancing Optoelectronic Technologies. Adv. Theory Simul. 7, 2400190 (2024).
Kaoungku, N., Suksut, K., Chanklan, R., Kerdprasop, K. & Kerdprasop, N. The silhouette width criterion for clustering and association mining to select image features. Int J. Mach. Learn Comput 8, 69–73 (2018).
Rousseeuw, P. J. Silhouettes: A Graphical Aid to the Interpretation and Validation of Cluster Analysis. J. Computational Appl. Math. 20, 53–65 (1987).
Kamalov, F. et al. Mathematical Methods in Feature Selection: A Review. Mathematics 13 https://doi.org/10.3390/math13060996 (2025).
Vogelstein, J. T. et al. Supervised dimensionality reduction for big data. Nat. Commun. 12, 2872 (2021).
Jolliffe, I. T. Principal Component Analysis. (Springer New York, New York, NY, 1986). https://doi.org/10.1007/978-1-4757-1904-8.
Abadi, M. et al. TensorFlow: Large-Scale Machine Learning on Heterogeneous Distributed Systems. www.tensorflow.org.
Pedregosa FABIANPEDREGOSA, F. et al. Scikit-Learn: Machine Learning in Python Gaël Varoquaux Bertrand Thirion Vincent Dubourg Alexandre Passos PEDREGOSA, VAROQUAUX, GRAMFORT ET AL. Matthieu Perrot. Journal of Machine Learning Research vol. 12 http://scikit-learn.sourceforge.net. (2011).
Acknowledgements
This work was supported by a grant from the National Science Foundation, NSF-DMR 2309396 and was in part based on research sponsored by the U.S. Department of Energy Office of Energy Efficiency and Renewable Energy Solar Energy Technologies Office agreement number 37989 through National Laboratory of the Rockies, operated under Contract No. DE-AC36-08GO28308. The views expressed in the article do not necessarily represent the views of the DOE or the U.S. Government. The U.S. Government retains and the publisher, by accepting the article for publication, acknowledges that the U.S. Government retains a nonexclusive, paid-up, irrevocable, worldwide license to publish or reproduce the published form of this work, or allow others to do so, for U.S. Government purposes.
Author information
Authors and Affiliations
Contributions
R.F.K. and J.P.B. conceived the project and supervised the research. R.A.W.A. conducted the investigation, developed the software, generated the visualizations, and prepared the original draft of the manuscript. S.S. contributed to formal analysis and validation. J.P.B. and R.A.W.A. carried out methodology development. R.F.K. and J.P.B. contributed to conceptualization. R.F.K. led funding acquisition, project administration, and resource provision. R.F.K. further shared supervision responsibilities. All authors reviewed and commented on the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ayyubi, R.A.W., Sultanov, S., Buban, J.P. et al. Unsupervised defect clustering in atomic-resolution microscopy using a convolutional variational autoencoder. npj Comput Mater (2026). https://doi.org/10.1038/s41524-026-02024-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-026-02024-x