Abstract
Moyamoya disease (MMD) is a progressive cerebrovascular disorder with intracranial arterial stenosis and collateralization. Over 70% of sporadic cases lack known genetic drivers; RNF213 variants explain only 23% of Chinese cases, highlighting unmet diagnostic and therapeutic needs. Trio-based whole-exome sequencing (WES) of 126 Chinese sporadic MMD patients (30 pediatric-parent trios) underwent cross-platform validation, cohort screening (n = 268), and functional analysis in human brain microvascular endothelial cells (HBMECs). Variants were prioritized by population frequency, pathogenicity, and case-control comparisons. Endothelial function and oxidative stress were assessed via proliferation, migration, tube formation, and molecular markers. WES identified 42 rare sporadic and 15 de novo mutations; Sanger validation confirmed 11 sporadic/11 de novo variants including de novo NF1 p.Q2002X and recurrent CARS1 p.E712V. CARS1 p.E712V carriers showed early-onset stenosis (mean age 7.5 ± 4.4 years) and right-dominant Suzuki stage ≥4, while NF1 p.Q2002X correlated with severe bilateral stenosis in a child (onset age 3). CARS1 p.E712V showed significant patient enrichment (P = 0.004). Silencing NF1/CARS1 in HBMECs enhanced proliferation, migration, tube formation, and reduced GPX4 (CARS1-specific). CARS1 mutation augmented angiogenesis, indicating functional alteration. This study identifies NF1 and CARS1 as novel susceptibility genes for sporadic MMD in Chinese, expanding the genetic landscape beyond RNF213.
Similar content being viewed by others
Introduction
Moyamoya disease (MMD) is a chronic, progressive cerebrovascular disorder characterized by bilateral stenosis or occlusion of the terminal internal carotid arteries (ICAs) and/or the proximal segments of the anterior and middle cerebral arteries (MCAs), accompanied by the compensatory formation of abnormal basal collateral vascular networks1. The disease has been reported worldwide, with a distinct east-west gradient in prevalence, the highest incidence being observed in East Asian populations2. In China, the disease ranks third globally, with an estimated prevalence of 3.92 per 100,000 population3. MMD demonstrates a bimodal age distribution, presenting as pediatric-onset (ages 5–10) and adult-onset (ages 30–40) forms4. It is a leading cause of ischemic stroke in both pediatric and adult populations, accounting for approximately 80% of childhood ischemic strokes associated with cerebrovascular anomalies5. MMD frequently leads to neurological deficits, with high rates of disability and mortality, severely impairing patients’ quality of life and imposing substantial socioeconomic burdens. The incomplete understanding of the molecular mechanisms underlying MMD initiation and progression limits the development of targeted preventive, diagnostic, and therapeutic strategies.
Genetic factors play a significant role in the pathogenesis of MMD: approximately 10% and 6% of Japanese and US patients, respectively, have affected first-degree relatives6,7. The RNF213 p.R4810K variant has been established as a significant contributor to MMD susceptibility8,9, clinical severity10,11, and prognostic progression12,13. However, this variant is found in only ~23% of Chinese patients with MMD, suggesting its limited explanatory power for disease etiology in this population. While familial cases indicate a genetic predisposition, the genetic architecture of sporadic MMD remains poorly understood, necessitating comprehensive approaches to identify high-confidence pathogenic variants. Recent advances in trio-based whole-exome sequencing (WES) have led to the identification of de novo mutations (DNMs), which are genetic alterations arising during parental gametogenesis and transmitted to offspring, as critical drivers of sporadic neurodevelopmental disorders14,15,16,17. Pinard et al.18 identified three RNF213 DNMs in 28 MMD trios, associated with early disease onset and multi-arterial occlusion. Similarly, Kundishora et al.19 reported DIAPH1 DNMs in non-East Asian cohorts (both familial and sporadic cases), demonstrating their disruptive effects on vascular actin cytoskeleton remodeling, with significant diagnostic and therapeutic implications. DNMs, which typically arise during meiosis in parental germ cells and are absent from parental somatic tissues, play pivotal roles in sporadic diseases due to their absence in parental genomes and the lack of inheritance patterns20.
While previous studies have advanced our understanding of DNMs in MMD, several critical gaps remain. Firstly, most studies have been limited by small sample sizes, reducing statistical power to detect rare DNMs and their phenotypic correlations. Secondly, population-specific biases persist, yet the relevance of such variants to East Asian populations remains unclear. Thirdly, prior study designs have often conflated familial and sporadic cases, or relied solely on single-platform sequencing, which may introduce platform-specific technical artifacts. Finally, many studies have prioritized DNMs without systematically integrating inherited variants or validating findings across independent cohorts, potentially overlooking polygenic or oligogenic contributions.
In this study, our dual-cohort design with cross-platform validation and ethnically matched controls may address these limitations by minimizing both technical and population biases. The integration of pediatric trios and case-control analysis allowed for a systematic differentiation between de novo and inherited contributions. This multi-layered approach not only strengthened the prioritization of causal genes but also elucidated population-specific genetic mechanisms in Chinese patients with sporadic MMD. Collectively, our framework may enhance the precision of genetic discovery in heterogeneous disorders and provide a scalable model for future studies.
Results
Clinical description of study population
A total of 126 patients with sporadic MMD from two independent cohorts were enrolled in this study. DNA was isolated, and WES was performed. The baseline characteristics of the cohort are summarized in Table 1. Among these patients, 64 (50.8%) were male, and the median age at onset was 8.5 years. A majority (70.7%) presented with cerebral ischemia as the initial symptom, and 30% exhibited posterior cerebral artery involvement. All diagnoses were confirmed via magnetic resonance angiography (MRA), digital subtraction angiography (DSA), or non-invasive ultrasonography, with the confirmed absence of a family history of MMD. The geographical distribution of the 394 patients in the final combined analysis cohort, as detailed in the Methods, demonstrated broad representation across major Han Chinese subgroups, supporting the genetic comparability with the ChinaMAP reference population.
The clinical baseline characteristics of the 30 trios are summarized in Table 2. All 30 pediatric patients had an age at onset under 16 years, with the majority presenting ischemic manifestations as initial symptoms. None of the parents of these patients were diagnosed with MMD following MRA and DSA screenings.
Identification of pathogenic gene mutation
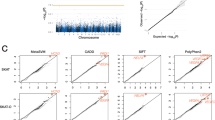
In the WES analysis aimed at identifying pathogenic variants in patients with sporadic MMD, cohort 1 (74 patients) initially revealed 401,387 mutation sites, while cohort 2 (52 patients) contained 105,468 variants. Following the filtering pipeline outlined in Fig. 1A, which incorporated criteria such as minor allele frequency (MAF) from public databases, genomic regions and functional annotations of mutations, and pathogenicity predictions from computational tools, 17,954 variants remained in cohort 1 and 8869 in cohort 2. These variants were subsequently matched with SNPs from the ChinaMAP database, and case-control allele frequency analysis was performed to exclude loci lacking statistical significance. The intersection of remaining variants between both cohorts yielded 42 shared mutations, consisting of 38 non-synonymous mutations (all missense variants) and four stop-gain mutations. Detailed characteristics of these candidate variants are presented in Table S3. Among the 42 rare mutations, 11 loci with functional relevance to the pathological mechanisms of MMD were selected for Sanger sequencing in all mutation carriers based on gene function annotations. These results demonstrated complete concordance with the WES data. Detailed characteristics of these prioritized loci are summarized in Table S4.
A Flowchart illustrating the screening process for identifying rare candidate genes in 126 Chinese sporadic MMD cases from two independent cohorts through WES. B Workflow depicting the screening of de novo mutation candidates identified via WES in 30 MMD probands from core families. MMD Moyamoya Disease, WES, whole-exome sequencing, MAF minor allele frequency, SIFT sorting intolerant from tolerant, assessing mutation impact on protein structure/function, PolyPhen Polymorphism Phenotyping, predicting structural/functional alterations of coding nonsynonymous variants; MutationTaster Tool for evaluating mutation pathogenicity, CADD combined annotation dependent depletion, a scoring system for assessing the deleterious effects of SNPs/indels, predicting functional impacts of coding and non-coding variants.
Following the DNM identification strategy, WES of 30 trios initially detected 2250 DNMs, which were then filtered according to the workflow outlined in Fig. 1B. After applying criteria including genomic regions, functional annotations, MAF thresholds in public databases, and computational pathogenicity predictions, 95 mutations remained. Post-filtering, 15 high-confidence DNMs were retained, comprising 13 non-synonymous variants, one variant predicted to introduce a stop-gain (p.Q2002X in NF1), and one mutation of unknown functional consequence. Detailed information for these candidate loci is provided in Table S5. The 15 identified DNMs were subjected to Sanger sequencing validation, and 11 mutations were successfully validated (Table S5). These included 10 non-synonymous mutations and one de novo variant resulting in a p.Q2002X substitution in NF1 (c.C6004T). The p.E712V (c.A2135T) variant in the CARS1 gene was confirmed in both the de novo and rare sporadic mutation cohorts.
Targeted multiplex PCR amplicon sequencing validates identified pathogenic genes in an independent patient cohort
To further validate the frequency of mutations confirmed by Sanger sequencing, targeted multiplex PCR amplicon sequencing was performed on an independent cohort of 268 patients with disease onset aged ≤10 years. The baseline characteristics of this cohort are summarized in Table 1. A total of 42 SNPs were analyzed, including 11 rare sporadic mutations and 11 DNMs validated by Sanger sequencing in this study (with the CARS1 p.E712V shared between both categories), as well as 21 rare mutations in three genes (RNF21318, DIAPH119, and ANO121) previously identified in non-Asian populations. Detailed information for these loci is provided in Table S6.
The results indicated that none of the 21 mutations in RNF213, DIAPH1, or ANO1 were detected in this cohort, suggesting potential ethnic specificity and limited pathogenicity relevance to Asian populations. However, larger-scale validation studies are still required. Among the 21 validated rare sporadic and DNMs, 11 were identified in the expanded cohort, including 10 rare sporadic mutations and 1 DNM. Notably, the mutation in the DDX53 gene was observed as homozygous in two patients, while all other mutations were heterozygous. The detection profiles for these mutations are summarized in Table S7.
Validation of the CARS1 p.E712V (c.A2135T) mutation as a susceptibility gene in sporadic MMD
Based on the multiplex PCR results, the frequency of each identified mutation was calculated across the two-phase cohort, which included a total of 394 patients. A case-control allele frequency analysis was performed by comparing the frequencies of these mutations with SNP carrier rates in non-MMD individuals from the ChinaMAP database to assess differences between patients and controls. Nine SNPs from the ChinaMAP database overlapped with genes identified in this study. The CARS1 p.E712V variant (rs139920420) is a low-frequency polymorphism in East Asian populations, with an allele frequency of 0.00737 in the ChinaMAP control database, consistent with other public genomic databases (Table S8). The allele was significantly enriched in the MMD patient cohort compared with ChinaMAP controls (χ² = 8.295, P = 0.004), with an odds ratio of 2.260 (95% CI: 1.278–3.998) (Table 3).
Clinical presentation of CARS1 c.A2135T and NF1 c.C6004T carriers and cerebral angiography
The CARS1 p.E712V mutation was identified in 13 patients (3.30%) (Table S9), with the age of onset ranging from 3 to 18 years (mean 7.5 ± 4.4 years), showing a female predominance (53.8%). Most patients presented with transient ischemic attacks (TIA) (69.2%). Advanced Suzuki stages (≥4) were predominantly observed in the right hemisphere (69.2% vs. 38.5% in the left hemisphere). Cerebral angiography confirmed stenosis/occlusion at the terminal ICA and proximal MCA in all patients, further supporting the pathogenic role of the CARS1 p.E712V mutation in driving cerebrovascular pathology. A pediatric case with a de novo NF1 p.Q2002X variant exhibited early-onset MMD (age 3), presenting with ischemic stroke-induced aphasia and Suzuki stage Ⅱ bilateral ICA/anterior cerebral artery (ACA)/MCA stenosis. This case underscored the phenotypic overlap between sporadic MMD and NF1-related cerebrovasculopathy, suggesting that truncating mutations in NF1 may contribute to pediatric-onset vascular remodeling and the development of MMD.
Bioinformatic characterization of NF1 and CARS1 mutations reveals structural and functional perturbations linked to MMD pathogenesis
Bioinformatic analyses were performed to assess the pathogenic potential of the CARS1 p.E712V (c.A2135T) and NF1 p.Q2002X (c.C6004T) mutations identified in sporadic MMD patients and MMD trios. Evolutionary conservation analysis revealed both mutated residues to be highly conserved across species (Fig. 1A, E). In CARS1, the p.E712V (c.A2135T) missense mutation substitutes a negatively charged glutamic acid with a hydrophobic valine, altering hydrogen-bonding distances and disrupting local charge distribution (Fig. 2B). Structural domain analysis identified critical motifs in CARS1, including the HIGH, KIIK, and KMSKS regions, which are vital for tRNA synthetase activity (Fig. 2C). PPI network analysis in sporadic MMD revealed a novel physical association between the newly identified susceptibility gene CARS1 and GPX4 (Fig. 2D), a key regulator of ferroptosis. For the NF1 p.Q2002X variant, bioinformatic analysis predicts the loss of the bipartite nuclear localization signal (residues 2555–2571) and disordered polar regions (residues 2787–2839), which are essential for protein regulation (Fig. 2F). Homology modeling suggested that the NF1 mutation introduces a premature termination codon, which could result either in nonsense-mediated mRNA decay or, should the transcript escape degradation, in the production of a truncated protein lacking critical functional domains (Fig. 2G). PPI analysis revealed a close association between the newly identified de novo NF1 mutation and RNF213 in MMD trios (Fig. 2H). This interaction suggests potential functional synergy or shared pathogenic mechanisms between these two genes.
A Evolutionary conservation analysis of the mutated residues in CARS1 p.E712V and (E) NF1 p.Q2002X across species. B, F Domain architecture of CARS1 and NF1. C, G Homology-based three-dimensional structural models of wild-type (WT) and mutant CARS1/NF1 proteins generated using SWISS-MODEL and visualized in PyMOL (D, H). Protein-protein interaction networks (STRING database) for CARS1 and NF1.
NF1 and CARS1 promote angiogenesis in human brain microvascular endothelial cells (HBMECs)
As demonstrated in Fig. 3A, F, targeted silencing of NF1 and CARS1 significantly reduced their respective protein expression levels, whereas CARS1 overexpression resulted in a marked upregulation of its protein expression, confirming transfection efficiency. A trend toward elevated VEGF protein expression was observed in both NF1- and CARS1-silenced groups compared to the negative control (NC) group (Fig. 3B, I), although these changes did not reach statistical significance. These molecular changes were accompanied by enhanced cellular functions: silenced groups showed significantly increased cell proliferation (P < 0.05, Fig. 3C, G) and migration capacity (P < 0.05, Fig. 3E, K). Tube formation assays revealed substantial pro-angiogenic effects, with both branch node count and total capillary length showing significant increases in silenced groups (P < 0.05, Fig. 3D, J). CARS1 silencing led to a concomitant reduction in GPX4 protein expression (P < 0.05, Fig. 3H). Neither the CARS1 p.E712V point mutation nor CARS1 overexpression groups showed significant differences in protein expression compared to the empty vector controls (Fig. 3I). The CARS1 p.E712V mutant group demonstrated significantly enhanced angiogenic capacity (P < 0.05, Fig. 3J). Although aminoacylation activity was not directly measured in this study, independent biochemical evidence confirms that this variant (referred to as CARS E795V) exhibits ~20% reduction in catalytic activity22.
A ~ E results of protein and phenotype experiments of NF1 gene; F ~ K results of protein and phenotype experiments of CARS1 gene. A Representative images of western blotting for NF1 and GAPDH. The statistical results of mRNA and protein expression were calculated. n = 3, **P < 0.01, ****P < 0.0001. B Representative images of western blotting for VEGF and GAPDH. Relative grey values of the protein bands are shown, and the statistical results of protein expression were calculated. n = 3. C Effect of silencing NF1 on the proliferative capacity of HBMEC. n = 3, *P < 0.05, **P < 0.01, ****P < 0.0001. D The tubule-like structure formation of HBMEC in the NF1-silenced group and the statistical comparison of the number of branch points and total capillary length. n = 3, *P < 0.05, **P < 0.01. E Effect of silencing NF1 on HBMEC cell migration. n = 3, *P < 0.05. F Representative images of western blotting for CARS1 and GAPDH. Relative grey values of the protein bands are shown, and the statistical results of protein expression were calculated. n = 3, ***P < 0.001, ****P < 0.0001. G Effect of different treatments of CARS1 on the relative proliferative capacity of HBMEC. n = 3, ****P < 0.0001. H Representative images of western blotting for GPx4 and GAPDH. Relative grey values of the protein bands are shown, and the statistical results of protein expression were calculated. n = 3, *P < 0.05. I Representative images of western blotting for VEGF and GAPDH. Relative grey values of the protein bands are shown, and the statistical results of protein expression were calculated. n = 3. J The tubule-like structure formation of HBMEC in the CARS1 treatment group and the statistical comparison of the number of branch points and total capillary length between treatment groups. K Effect of Effect of different treatments of CARS1 on HBMEC cell migration. n = 3, **P < 0.01, *P < 0.05, ****P < 0.0001. siNF1 NF1 silencing, siCARS1 CARS1 silencing, ev empty vector, CARS1-oe, CARS1 overexpression, CARS1-mut cells expressing the CARS1 p.E712V mutant, nc negative control.
Discussion
This study ultimately prioritized two genetic variants: the DNM NF1 p.Q2002X and the dual-signature variant CARS1 p.E712V, which was identified both as a DNM and a rare sporadic mutation. In vitro studies further demonstrated that the NF1 p.Q2002X mutation likely contributes to vascular pathology, potentially involving the VEGF signaling pathway, while the CARS1 p.E712V variant may promote angiogenesis by enhancing ferroptosis mechanisms. Both genetic alterations were shown to drive excessive angiogenesis, contributing to the characteristic cerebrovascular lesions observed in MMD.
The NF1 p.Q2002X (c.C6004T) variant identified in our study represents a de novo mutation in a pediatric case of early-onset MMD. Bioinformatic analysis classifies this variant as a stop-gain mutation, predicting the introduction of a premature termination codon, however, its precise molecular consequence remains unclear. A key question is whether this mutant transcript undergoes nonsense-mediated decay (NMD) or escapes surveillance to produce a stable, truncated protein. Although direct experimental evidence from the patient’s cells is lacking, insights can be drawn from a systematic series of investigations of a highly analogous NF1 mutation (p.Q181X, c.541 C > T)23,24,25. The p.Q181X mutation was initially identified in a pedigree with NF1 and severe cerebral vasculopathy23. A mouse model harboring the Nf1-Q181X mutation subsequently demonstrated robust vascular pathology, including increased neointima formation and vessel lumen occlusion following arterial injury24. This in vivo evidence suggests that a pathogenic gene product, potentially a truncated protein resulting from NMD escape, is functionally active. Mechanistic studies further indicated that the Nf1-Q181X mutation drives M2 macrophage polarization and vascular smooth muscle cell dysfunction25, supporting the concept that C-terminal truncating NF1 mutations can act via dominant-negative or gain-of-function mechanisms. The position of the identified p.Q2002X mutation, similar to p.Q181X, in the C-terminal region of NF1 gene, raises the possibility that it might similarly evade NMD. Moreover, the severe cerebrovascular phenotype in our patient and our in vitro data showing that NF1 silencing promotes angiogenesis are consistent with a loss-of-function mechanism in the vasculature. While direct validation of the p.Q2002X mutant product remains necessary, the precedent set by the studies on the p.Q181X mutation provides a plausible framework for interpreting our finding, suggesting that de novo NF1 variants in the C-terminus, may contribute to MMD pathogenesis through aberrant protein products that disrupt vascular homeostasis.
The de novo NF1 p.Q2002X mutation identified in our study further adds to the growing evidence implicating NF1 dysfunction in moyamoya vasculopathy. Moyamoya syndrome (MMS) is the most frequently observed cerebrovascular abnormality in NF1 patients and represents a leading cause of stroke in this population26. Although our study does not experimentally validate the molecular consequence of the p.Q2002X variant, its de novo status in a patient with early-onset, severe bilateral stenosis provides compelling genetic evidence for its potential role. This finding aligns with recent studies highlighting associations between specific NF1 variants and severe cerebrovasculopathy. For example, a clinical whole-exome sequencing study by Nakamura et al.27 identified several pathogenic NF1 splicing mutations that generate premature termination codons (e.g., p.Y1292RfsX7, p.R1526SfsX54). Furthermore, case series have reported frameshift (e.g., p.Tyr1692Valfs*3) and nonsense (e.g., p.Arg440*) NF1 mutations in pediatric patients with cerebrovascular manifestations28, and studies in Chinese populations indicate that truncating NF1 variants constitute a major contributor to the disease29. A recent case report also described a pediatric patient with a de novo NF1 missense mutation (p.Arg1830Gly) presenting with MMS, epilepsy, and hydrocephalus, further emphasizing the clinical relevance of NF1 dysfunction in severe cerebrovascular complications30. Taken together, although the transcript-level fate of p.Q2002X mutation remains to be determined, its identification is consistent with emerging evidence that impaired neurofibromin function, potentially through haploinsufficiency or dominant-negative effects, may contribute to the dysregulated angiogenesis and vascular remodeling characteristic of MMD.
The association between NF1 and MMD is predominantly observed in the pediatric age group, affecting a minority of children with NF131,32. Our study specifically focused on patients under 16 years of age. While NF1 and RNF213 are both implicated in MMD, they participate in distinct pathways: NF1 functions as a negative regulator of the Ras pathway, controlling cellular division and proliferation33, whereas RNF213 suppresses the non-canonical Wnt signaling pathway to promote vascular regression34.
The CARS1 p.E712V mutation identified in this study represents both a risk variant in sporadic MMD and a DNM in trios, suggesting its potential role in MMD pathogenesis. Although our study did not directly assess aminoacylation activity, independent biochemical evidence confirms that this mutation impairs enzymatic function, resulting in approximately a 20% reduction in catalytic efficiency22. This finding supports that the mutation alters enzymatic activity. Subsequent research from the same group further elucidated the role of CARS in neurological disorders, showing increased CARS expression in AD brains, its involvement in neuroinflammation via the TLR2/MyD88 pathway, and its accumulation in Aβ plaques35. Collectively, these findings highlight the functional significance of CARS in neurological processes and provide a strong biological rational for its potential involvement in cerebrovascular pathology. The pleiotropic nature of CARS1 mutations is further exemplified by the report of this identical CARS1 p.E712V variant in a Chinese family presenting with parkinsonism and spinocerebellar ataxia, without documented MMD cases22. This observation suggests that CARS1 mutations may contribute to distinct neurological and vascular phenotypes, likely modulated by genetic background, environmental factors, or tissue-specific expression patterns.
It is noteworthy that the CARS1 p.E712V variant (rs139920420) occurs at a low frequency (approximately 0.6%–0.9%) in East Asian populations, as reported in the gnomAD, 1000 Genomes, and ChinaMAP databases—a frequency consistent with its identification in both our sporadic and de novo mutation screens. The significant case-control association observed in this study (P = 0.004) suggests that CARS1 p.E712V functions not as a highly penetrant rare mutation, but rather as a susceptibility allele with moderate effect size, contributing to MMD risk in a subset of genetically predisposed individuals. This interpretation is consistent with the complex, multifactorial etiology increasingly recognized in sporadic MMD. Accordingly, we consider CARS1 p.E712V a risk factor rather than a deterministic cause, whose effect on endothelial function and ferroptosis may act synergistically with other vascular risk pathways to promote disease progression. Patients carrying this mutation showed consistent neuroimaging features, including multiple cerebrovascular abnormalities predominantly characterized by stenosis or occlusion of the ICA and MCA. Evolutionary conservation analyses revealed that CARS1 p.E712V was highly conserved, underscoring its functional importance. The substitution of glutamic acid (Glu) with valine (Val) at position 712 removes the negatively charged residue, potentially disrupting the local electrostatic environment and altering interactions with enzymatic substrates or cofactors. Structural modeling further suggests that this mutation may destabilize the enzyme’s active site or impair substrate binding capacity, potentially leading to aberrant protein aggregation or altered membrane localization.
CARS1 encodes cysteinyl-tRNA synthetase, an enzyme that catalyzes the ligation of cysteine to its cognate tRNA during protein translation, ensuring accurate synthesis of cysteine-containing proteins36. Furthermore, cysteine metabolism is tightly linked to glutathione (GSH) synthesis via the System Xc- pathway, which facilitates the exchange of intracellular glutamate for extracellular cystine, enabling cystine uptake into the cell. Once inside, cystine is reduced to cysteine, a rate-limiting precursor for GSH biosynthesis. GSH, a tripeptide (γ-glutamyl-cysteinyl-glycine), is a pivotal cellular antioxidant essential for maintaining redox homeostasis and protecting cells from oxidative stress-induced ferroptosis37. The CARS1 p.E712V mutation may drive the characteristic vasculopathy of MMD through two interconnected mechanisms: impaired GPX4 synthesis and ferroptosis activation, and cysteine metabolic dysregulation with subsequent oxidative stress. The reduced enzymatic activity of mutant cysteinyl-tRNA synthetase could compromise the biosynthesis of cysteine-dependent proteins, including GPX4. Diminished GPX4 levels would impair cellular capacity to detoxify lipid peroxides, thereby promoting ferroptosis in vascular endothelial cells. This process may trigger maladaptive compensatory angiogenesis and disrupt vascular wall integrity, ultimately contributing to arterial stenosis, collateral vessel formation, and the distinctive vasculopathy observed in MMD. Additionally, the mutation may perturb cysteine metabolism, provoking compensatory overactivation of System Xc-. Chronic cystine depletion could exhaust cellular cysteine reserves, limiting the synthesis of GSH and resulting in cumulative oxidative stress. Prolonged oxidative damage to the vascular endothelium may exacerbate endothelial dysfunction and accelerate cerebrovascular remodeling, driving disease progression.
Ferroptosis is characterized by the accumulation of reactive oxygen species (ROS) and lipid peroxides38. ROS inhibit the activity of prolyl hydroxylases, thereby stabilizing hypoxia-inducible factor-1α (HIF-1α), which in turn transcriptionally regulates VEGF and other pro-angiogenic factors. Consequently, ferroptosis may modulate angiogenesis39,40,41. Building upon the observed ferroptosis induction, as indicated by the downregulation of GPX4 following CARS1 p.E712V mutation and silencing, and supported by our functional angiogenesis data, such as VEGF elevation, enhanced proliferation/migration, and excessive tube formation, we propose a mechanistic framework. In this framework, CARS1 dysfunction synergizes with ferroptosis-induced states to activate pro-angiogenic pathways. It remains to be determined whether this enhanced angiogenesis is driven by ferroptotic stress-induced potentiation of HIF-1α/VEGF signaling via lipid peroxide accumulation. This requires further mechanistic investigation. Furthermore, the known role of CARS1 in suppressing tumor cell invasion42 suggests its involvement in extracellular matrix (ECM) remodeling. Abnormal thickening of the vascular basement membrane, a pathological hallmark of MMD, may reflect dysregulated ECM dynamics linked to CARS1 dysfunction. The p.E712V mutation might impair CARS1’s regulatory capacity to regulate ECM homeostasis, exacerbating the structural disorganization of the cerebral vasculature in the progression of MMD.
The uniqueness of this study lies in its dual-cohort design, which combines complementary approaches. Both cohorts underwent WES. However, distinct sequencing platforms (Illumina HiSeq X Ten and NovaSeq 6000) and bioinformatics pipelines were adopted. This design may minimize platform-specific biases and enhance the reliability of overlapping candidate genes. Intersecting variants identified in both cohorts were further prioritized through a case-control analysis against the ChinaMap database, a population-specific reference that provides ethnically matched controls to reduce genetic heterogeneity. The second cohort included 30 pediatric patients (under 16 years old) with parental trios, allowing for the exploration of DNMs, a critical yet understudied mechanism in sporadic MMD. This dual strategy, which combines cross-cohort variant filtering with family-based analysis, addresses both inherited and de novo contributions, offering a comprehensive view of the genetic etiology. By integrating multi-platform sequencing, ethnically tailored controls, and trio-based validation, this study not only refines the genetic landscape of sporadic MMD but also establishes a framework for identifying high-confidence pathogenic variants in heterogeneous disorders.
The first consideration is the population allele frequency of the prioritized CARS1 p.E712V variant. Although statistically significant in our cohort, this variant is not ultra-rare among East Asian populations. This pattern suggests a model of incomplete penetrance or a moderate effect size, common challenges in the genetic study of complex disorders. The mechanistic link between this specific polymorphism and MMD pathogenesis remains to be elucidated. Future studies involving larger, independent cohorts are needed to quantify its population-attributable risk and clarify the underlying molecular mechanisms. Secondly, to address potential population stratification, we employed geographical and ethnic matching of patients with the ChinaMAP reference cohort. Although the absence of individual-level control data precluded a formal joint PCA, the consistent allele frequency of the CARS1 variant across multiple East Asian databases supports the validity of our association. Future studies incorporating internally sequenced controls would further minimize residual confounding. Thirdly, an additional limitation arises from our conservative variant filtering strategy. While the use of two sequencing platforms and the requirement that candidate variants be present in both cohorts effectively minimized technical artifacts and false positives, this approach inherently reduced sensitivity. It likely excluded very rare, genuine pathogenic variants present in only one cohort. Future studies with larger, homogeneously sequenced cohorts will be necessary to capture these ultra-rare contributors to sporadic MMD. Fourthly, we acknowledge that our interpretation of the NF1 p.Q2002X mutation remains speculative in the absence of direct experimental validation of its transcript stability and protein product. Although we have drawn parallels to analogous mutations reported in the literature, future studies using patient-derived cellular models or genome-edited isogenic lines are essential to determine whether this variant escapes nonsense-mediated decay and produces a pathogenic truncated protein. We are committed to pursuing these mechanistic investigations in future work to fully elucidate the molecular consequences of this de novo mutation. Fifthly, the study focused exclusively on exonic regions, which may overlook non-coding regulatory variants that could contribute to MMD. Nonetheless, this approach was consistent with the hypothesis that coding mutations in genes such as CARS1 and NF1 drive pathogenic mechanisms through protein-level disruptions. Sixthly, while bioinformatics predictions linked mutations to vascular remodeling and cellular processes, experimental validation (e.g., in vitro assays or animal models) was not performed. Future studies should prioritize functional characterization to confirm these mechanistic roles. Finally, although the 30 trios provided preliminary insights into DNMs, they lack the statistical power to definitively associate rare DNMs (e.g., CARS1 p.E712V) with MMD. Larger trio cohorts would be necessary to validate the contributions of these mutations.
This study identified NF1 p.Q2002X and CARS1 p.E712V as novel risk factors associated with sporadic MMD. Using a dual-cohort design that incorporated ethnically matched controls and trio-based analysis, we provide genetic evidence implicating these variants in disease susceptibility. In vitro experiments further demonstrate that dysregulation of NF1 and CARS1 promotes pro-angiogenic responses in brain endothelial cells. Collectively, these findings broaden the genetic landscape of sporadic MMD beyond RNF213 and suggest potential involvement of Ras signaling and oxidative stress pathways. However, direct mechanistic validation remains to be established. Further studies involving larger cohorts and comprehensive functional assays are warranted to confirm the pathogenicity of these variants and to explore their potential implications for precision medicine in MMD.
Methods
Case source and ethics approval
This study was conducted in accordance with the Declaration of Helsinki. All study procedures and protocols were conducted in accordance with ethical standards approved by the Ethics Committees of China Medical University (Approval No. [2021]43) and the Chinese PLA General Hospital (Approval No. ky-2020-5-11). Written informed consent was obtained from all participating patients and their immediate family members.
The diagnostic confirmation of MMD adhered to internationally recognized guidelines established by Japanese research consortiums specializing in cerebrovascular occlusive disorders43. Exclusion criteria systematically eliminated individuals with comorbid conditions, including but not limited to atherosclerotic vascular changes, immune-mediated pathologies, intracranial tumors, neuroinflammatory processes, genetic syndromes such as trisomy 21, and other defined cerebrovascular etiologies.
This study was conducted in two phases, with all participants recruited from the Department of Neurosurgery at Chinese PLA General Hospital. Blood samples and basic demographic data were collected through outpatient clinics, hospital admissions, and follow-up visits. In the initial WES analysis phase, 126 sporadic MMD patients and 60 healthy parents were enrolled. This cohort was divided into two subgroups: Group 1, consisting of 74 patients randomly selected from those diagnosed between 2017 and 2022; and Group 2, comprising 52 patients diagnosed between 2008 and 2016, including 30 pediatric-onset cases (age < 16 years) and their healthy parents, forming 30 trios for DNM analysis. The validation phase included an additional 268 MMD patients for confirmatory studies.
To ensure ethnically matched comparisons with the China Metabolic Analytics Project (ChinaMAP) control database44, the geographical origins of all 394 MMD patients in the final combined cohort were meticulously recorded and categorized according to the established Han Chinese subpopulations. The cohort included 390 Han Chinese patients, distributed across the seven major geographical clusters (North, East, South, Northwest, Central, Lingnan, and Southeast Han), along with four patients from other ethnic groups. This geographical profiling enabled a principled approach to population matching and helped minimize potential population stratification bias. The ChinaMAP cohort represents a general population sample derived from large-scale epidemiological studies in China and includes individuals with a range of health conditions.
Exome capture and sequencing methodology
Exon capture was performed on 126 MMD patients using the SureSelectXT Human All Exon V6 probe (Agilent Technologies, CA, USA). The captured libraries were sequenced using a 150-bp paired-end strategy across two distinct platforms: Cohort 1 (n = 74) was sequenced on the Illumina NovaSeq 6000 platform, and Cohort 2 (n = 52) on the Illumina HiSeq X Ten platform. Raw sequencing data were aligned to the GRCh37/hg19 reference genome using BWA-MEM (Burrows-Wheeler Aligner, version 0.7.12), followed by variant calling with the GATK HaplotypeCaller. Single-nucleotide variants (SNVs) were filtered using GATK Variant Filtration, and annotations for single nucleotide polymorphisms (SNPs) and insertions/deletions (indels) were generated using ANNOVAR (version 2016 Feb 01). This pipeline was consistent with methodologies established in our prior work45.
In a cohort of 30 trios, DNMs were systematically identified through comprehensive genotype comparison between affected offspring and their biological parents. Mutational sites were classified as de novo when they met the following criteria: (1) the variant allele was present in the proband’s genotype, (2) neither parent carried the mutant allele at the corresponding genomic position, and (3) the proband’s genotype at the mutation site demonstrated Mendelian inconsistency with the parental genotypes.
Candidate variants identification
Rare variant screening was performed on 126 sporadic MMD patients to identify potential pathogenic mutations based on allele frequency in population databases, genomic location, functional impact, predicted pathogenicity, and gene function. The screening process is described in detail below:
For Cohort 1 (74 MMD Patients), a comprehensive variant prioritization framework was established, integrating frequency thresholds (minor allele frequency [MAF] < 0.01 across 1000 Genomes Project, Exome Aggregation Consortium [ExAC] All populations, Genome Aggregation Database [gnomAD] Allele Frequency [AF], gnomAD East Asian Allele Frequency [EAS_AF], and ClinVar Allele Frequency in the Exome Sequencing Project [ClinVar_AF_ESP]), functional impact criteria (retention of missense, nonsense, and frameshift variants; splice-site alterations [3’/5’]; stop-loss/gain mutations; coding sequence indels; initiator codon disruptions; promoter modifications; spanning deletions; UTR-3/UTR-5 elements; and functionally undefined variants), and pathogenicity consensus (concordant deleterious/probably deleterious predictions from sorting intolerant from tolerant (SIFT), Polymorphism Phenotyping v2 (PolyPhen-2), and MutationTaster with ≥ 20). This was followed by genomic coordinate standardization through GRCh37 to GRCh38 conversion via LiftOver, subsequent the ChinaMAP database filtering (MAF < 0.01), and statistical validation through case-control allele frequency association analysis to eliminate variants lacking significance (P > 0.05), thereby ensuring both biological relevance and population specificity.
For Cohort 2 (52 MMD Patients), a multi-tiered variant prioritization protocol was implemented, combining frequency constraints (MAF < 0.01 in 1000 Genomes [All/EAS], ExAC [All/EAS], and ESP6500), functional impact selection (retention of exonic/splicing variants, including frameshift indels, nonsynonymous SNVs, stop-gain/loss alterations, or undefined consequences), pathogenicity consensus (concordant deleterious classifications from SIFT, PolyPhen-2, and MutationTaster with CADD ≥ 20), and clinical relevance thresholds (presence in ≥1 patient with allele count >0). This was followed by genomic coordinate standardization through GRCh38/hg38 conversion using LiftOver, ChinaMAF filtering (MAF < 0.01 in ChinaMAP database), and statistical refinement via case-control allele frequency association analysis, with exclusion of non-significant variants (P > 0.05) to ensure biological relevance.
After independent filtering of both cohorts, only shared mutations present in both groups were retained to strengthen the reliability of candidate pathogenic variants.
Identification of DNMs in 30 trios (affected probands and healthy parents)
A multi-stage filtering protocol was systematically implemented for DNM detection, starting with the prioritization of SNPs with functional impacts (exonic/splicing regions exhibiting nonsynonymous substitutions, stop-gain/loss, or undefined consequences). This was followed by stringent frequency exclusion (MAF < 0.01 across 1000 Genomes [All/EAS], ExAC [All/EAS], and ESP6500 databases), and retention of variants demonstrating consistent pathogenicity predictions (deleterious/probably deleterious classifications from SIFT, PolyPhen-2, and MutationTaster with CADD scores ≥ 20).
Sanger sequencing
To validate WES findings orthogonally, Sanger sequencing was performed on polymerase chain reaction (PCR)-amplified targets. Primer pairs were designed using NCBI’s Primer-BLAST tool (https://www.ncbi.nlm.nih.gov/tools/primer-blast/) to ensure specificity for the genomic regions of interest. The primer sequences for the candidate rare mutations identified in sporadic cases and the de novo mutation sites identified in trios are listed in Tables S1 and S2, respectively. PCR amplification was followed by DNA sequencing using BigDye Terminator v1.1 chemistry (Applied Biosystems, Foster City, CA, USA) on an ABI GeneAmp 3730xl Genetic Analyzer (Applied Biosystems).
Multiplex PCR sequencing
Mutations confirmed as positive through Sanger sequencing were analyzed in an expanded independent case cohort. Additionally, selected rare mutations previously reported in non-Asian populations were included for analysis. The case cohort was selected from patients definitively diagnosed with MMD (excluding Moyamoya syndrome [MMS]) based on stringent inclusion criteria: (1) age of onset ≤10 years, (2) absence of family history for MMD, and (3) exclusion of RNF213 p.R4810K mutation carriers, confirmed through TaqMan probe-based reverse transcription quantitative PCR.
To ensure sufficient high-quality genomic DNA extraction, quantification was conducted using a Qubit 2.0 Fluorometer (Thermo Fisher Scientific, USA). Libraries were constructed using a two-stage amplification protocol in a T100™ Thermal Cycler (Bio-Rad Laboratories). Amplification products were purified using AMPure XP magnetic bead technology (Beckman Coulter). After quality assessment, pooled libraries underwent paired-end sequencing (2 × 150 bp) on Illumina’s HiSeq XTen platform. Bioinformatic processing included: (1) trimming adapter sequences using CutAdapt (v1.2.1), (2) removing terminal low-quality bases (Q-score < 20) via PRINSEQ-lite (v0.20.3), and (3) aligning processed reads to the GRCh38 reference genome using the BWA-MEM algorithm (v0.7.13-r1126) with standard parameters. Genotype calling and variant identification were performed using Samtools (v0.1.18) and ANNOVAR, respectively, ensuring comprehensive annotation of genetic variations.
Replicate association study in expanded cases
To prioritize candidate variants, the replicated SNPs from the independent MMD cohort were integrated with those identified through WES, followed by case-control association analysis using aggregate allele frequency data from non-MMD controls in the ChinaMAP public database. In accordance with ChinaMAP data access polices, individual-level genotype counts were unavailable; therefore, association testing was performed using the provided allele frequencies. The ChinaMAP cohort represents a general population sample derived from large-scale epidemiological studies in China and includes individuals with a range of health conditions. Given the low population prevalence of MMD makes it a suitable source for control allele frequencies in this rare disease. Subsequently, statistically significant SNPs (P < 0.05) demonstrating differential allele distributions were selected for functional validation.
Bioinformatic characterization of candidate genes
To investigate the evolutionary conservation and structural-functional implications of candidate gene mutations, cross-species sequence alignment was performed using homologous protein sequences retrieved from the NCBI database (https://www.ncbi.nlm.nih.gov/), followed by multiple sequence alignment visualization with DNAMAN software (v9.0). Structural analyses included domain mapping based on secondary structure elements annotated in UniProt (https://www.uniprot.org/) using IBS software, complemented by three-dimensional structural modeling through comparative homology with SWISS-MODEL (https://swissmodel.expasy.org/) using wild-type templates from the PDB database (https://www.rcsb.org/). Mutant protein conformations were visualized and analyzed for steric clashes or domain perturbations using the PyMOL molecular graphics system. To elucidate potential pathogenic mechanisms, protein-protein interaction (PPI) networks were reconstructed using the STRING database (https://string-db.org/) with stringent confidence thresholds.
Cell culture and plasmid transfection
Human brain microvascular endothelial cells (HBMECs) were acquired from Qingqi Biotechnology Co., Ltd (Shanghai, China). Gene-specific siRNA targeting NF1, siRNA/overexpression constructs for CARS1, and the CARS1 p.E712V mutant plasmid were custom-designed and generated by Sangon Biotech (Shanghai, China). Transient transfection was performed using RNA transmate (Sangon Biotech) for siRNA-mediated silencing, and Lipohigh (Sangon Biotech) for overexpression constructs and the mutant plasmid, in accordance with the manufacturer’s protocols. Control siRNA (negative control, nc) and empty plasmid (vector) were provided by Sangon Biotech for normalization.
RNA isolation and RT-qPCR (reverse transcription quantitative real-time polymerase chain reaction)
Total RNA was isolated with Trizol reagent (Ambion, Shanghai, China), and purity/concentration were verified spectrophotometrically. Reverse transcription was performed using PrimeScript RT Master Mix (Takara, Japan). Quantitative PCR amplification was conducted on a QuantStudio 6 Flex System (Applied Biosystems, USA) with SYBR Green Supermix (Bio-Rad, USA). Primer sequences are provided in Table S2. Universal 18S rRNA primers (Ambion, USA) served as the endogenous control for absolute quantification. Thermocycling parameters included an initial denaturation at 95 °C (10 min), followed by 40 cycles of 95 °C (15 s) and 60 °C (60 s). Melt curve analysis was performed between 60 °C and 95 °C. Relative mRNA expression was calculated using the 2−ΔΔCt method, with all reactions performed in technical triplicates. The primers used for CARS1 and NF1 amplification in functional assays are listed in Table S10.
Western blot
Protein lysates were extracted using RIPA Lysis Buffer (Sangon, China) supplemented with protease inhibitors. Equal amounts of protein (30 μg/lane) were resolved on 10% SDS-PAGE gels (Servicebio, China) to accommodate the molecular weights of target proteins: CARS1 (~86 kDa) and NF1 (250–280 kDa). Proteins were transferred to PVDF membranes (GVS, USA) and blocked with 5% non-fat milk for 1 h. Membranes were incubated overnight at 4 °C with primary antibodies: anti-CARS1 (1:1000; Absin, China), anti-NF1 (1:1000; ProteinTech Group, USA), anti-VEGF (1:1000; Absin, China), or anti-GAPDH (1:3000; Sangon, China). Following incubation with HRP-conjugated secondary antibodies (1:5000; Sangon, China), protein bands were visualized via enhanced chemiluminescence (ThermoFisher) and quantified using ImageJ software (NIH, USA).
Cell proliferation assay
HBMECs were seeded in 96-well plates at a density of 3–5 × 103 cells/well and allowed to adhere for 24 h. Cells were then synchronized by serum starvation (0.1% FBS DMEM) for 24 h, followed by experimental interventions (e.g., transfection or drug treatment). After 48 h of intervention, cell viability was assessed using the CCK-8 kit (Dojindo, Japan). Specifically, 10 μL CCK-8 reagent was added to each well, and cells were incubated at 37 °C for 2 h. Absorbance at 450 nm was measured using a BioTek microplate reader.
Scratch wound healing assay
Confluent HBMEC monolayers in 6-well plates were scratched linearly using a sterile 200 μL pipette tip. Detached cells were removed by PBS washing, and serum-free medium was added. Wound closure was monitored at 0 h, 12 h and 48 h post-scratch using an inverted microscope (Mshot, China). Migration rates were quantified as the percentage of wound area reduction using ImageJ software.
In vitro angiogenesis assay
HBMECs (8–10 × 104 cells/well) were seeded onto 24-well plates pre-coated with 20–30 μL Matrigel (Corning, USA) per well and incubated for 12 h at 37 °C under 5% CO2. Tube formation was imaged at ×50 magnification (Leica DMi1, Germany) across three randomly selected fields per well. Network parameters including total tube length and branch points were quantified using ImageJ (NIH, USA) with the Angiogenesis Analyzer plugin.
Data availability
With the permission of the corresponding authors and their institutions, combined with relevant documents, all data used for this analysis will be shared, following ethics approval, with other investigators for the reasonable purpose of replicating the procedures and results, if requested.
References
Suzuki, J. & Takaku, A. Cerebrovascular “moyamoya” disease. Disease showing abnormal net-like vessels in base of brain. Arch. Neurol. 20, 288–299 (1969).
Ihara, M. et al. Moyamoya disease: diagnosis and interventions. Lancet Neurol. 21, 747–758 (2022).
Shang, S. et al. Progress in moyamoya disease. Neurosurg. Rev. 43, 371–382 (2020).
Pollak, L. Moyamoya disease and moyamoya syndrome. N. Engl. J. Med. 361, 98 (2009).
Torres, V. M. & Saddi, V. A. Systematic review: hereditary thrombophilia associated to pediatric strokes and cerebral palsy. J. Pediatr. 91, 22–29 (2015).
Scott, R. M. & Smith, E. R. Moyamoya disease and moyamoya syndrome. N. Engl. J. Med. 360, 1226–1237 (2009).
Fukui, M., Kono, S., Sueishi, K. & Ikezaki, K. Moyamoya disease. Neuropathology 20, S61–64 (2000).
Liu, W. et al. Identification of RNF213 as a susceptibility gene for moyamoya disease and its possible role in vascular development. PloS One 6, e22542 (2011).
Kamada, F. et al. A genome-wide association study identifies RNF213 as the first Moyamoya disease gene. J. Hum. Genet. 56, 34–40 (2011).
Wang, Y. et al. Predictive role of heterozygous p.R4810K of RNF213 in the phenotype of Chinese moyamoya disease. Neurology 94, e678–e686 (2020).
Wang, Y. et al. Meta-analysis of genotype and phenotype studies to confirm the predictive role of the RNF213 p.R4810K variant for moyamoya disease. Eur. J. Neurol. 28, 823–836 (2021).
Hao, F. et al. High-resolution MRI vessel wall enhancement in moyamoya disease: risk factors and clinical outcomes. Eur. Radiol. 34, 5179–5189 (2024).
Ge, P. et al. Association between p.R4810K variant and postoperative collateral formation in patients with Moyamoya disease. Cerebrovasc. Dis. 48, 77–84 (2019).
Jin, S. C. et al. Exome sequencing implicates genetic disruption of prenatal neuro-gliogenesis in sporadic congenital hydrocephalus. Nat. Med. 26, 1754–1765 (2020).
Zhang, M. et al. Exome sequencing of 112 trios identifies recessive genetic variants in brain arteriovenous malformations. J. Neurointerv. Surg. 13, 568–573 (2021).
Watanabe, K. et al. Cognitive impairment in a complex family with AAGGG and ACAGG repeat expansions in RFC1 detected by expansion hunter de novo. Neurol. Genet. 8, e682 (2022).
Hoya, S. et al. Whole-exome sequencing in a family with a monozygotic twin pair concordant for schizophrenia and a follow-up case-control study of identified de-novo variants. Psychiatr. Genet. 30, 60–63 (2020).
Pinard, A. et al. Association of de novo RNF213 variants with childhood onset moyamoya disease and diffuse occlusive vasculopathy. Neurology 96, e1783–e1791 (2021).
Kundishora, A. J. et al. DIAPH1 variants in non-East Asian patients with sporadic Moyamoya disease. JAMA Neurol. 78, 993–1003 (2021).
Jin, Z. B. et al. Identification of de novo germline mutations and causal genes for sporadic diseases using trio-based whole-exome/genome sequencing. Biol. Rev. Camb. Philos. Soc. 93, 1014–1031 (2018).
Pinard, A. et al. Rare variants in ANO1, encoding a calcium-activated chloride channel, predispose to moyamoya disease. Brain 146, 3616–3623 (2023).
Liu, H. K. et al. A cysteinyl-tRNA synthetase mutation causes novel autosomal-dominant inheritance of a parkinsonism/spinocerebellar-ataxia complex. Neurosci. Bull. 40, 1489–1501 (2024).
Liang, J. T., Huo, L. R., Bao, Y. H., Wang, Z. Y. & Ling, F. Cerebral vasculopathy in a Chinese family with neurofibromatosis type I mutation. Neurosci. Bull. 29, 708–714 (2013).
Liang, C. et al. The Q181X point mutation in Nf1 induces cerebral vessel stenosis. Neurosci. Bull. 39, 813–816 (2023).
Yang, Y., Yao, Z. & Huo, L. The Nf1-Q181X point mutation induces M2 macrophage polarization via the AKT/STAT pathway to promote smooth muscle cell proliferation and migration. Mol. Biol. Rep. 51, 946 (2024).
Lehman, L. L. & Ullrich, N. J. Cerebral vasculopathy in children with neurofibromatosis type 1. Cancers 15, 5111 (2023).
Nakamura, A. et al. Whole-exome sequencing reveals the genetic causes and modifiers of moyamoya syndrome. Sci. Rep. 14, 22720 (2024).
Currao, P. et al. Vascular abnormalities and neurofibromatosis type 1: a paediatric case series. J. Child Neurol. 40, 49–60 (2025).
Chen, S., Cheng, H. & Zhao, G. The genetic spectrum of NF1 variants in 10 unrelated Chinese families with neurofibromatosis type 1. Neuroscience 29, 177–183 (2024).
Zhao, F., Yue, X., Wang, G. & Zhang, H. Moyamoya syndrome, epilepsy and hydrocephalus in neurofibromatosis Type 1. Int. J. Dev. Neurosci. 85, e70038 (2025).
Ghosh, P. S., Rothner, A. D., Emch, T. M., Friedman, N. R. & Moodley, M. Cerebral vasculopathy in children with neurofibromatosis type 1. J. Child Neurol. 28, 95–101 (2013).
Duat-Rodríguez, A., Carceller Lechón, F., López Pino, M., Rodríguez Fernández, C. & González-Gutiérrez-Solana, L. Neurofibromatosis type 1 associated with moyamoya syndrome in children. Pediatr. Neurol. 50, 96–98 (2014).
Li, F. et al. Neurofibromin is a novel regulator of RAS-induced signals in primary vascular smooth muscle cells. Hum. Mol. Genet. 15, 1921–1930 (2006).
Scholz, B. et al. Endothelial RSPO3 controls vascular stability and pruning through non-canonical WNT/Ca(2+)/NFAT signaling. Dev. Cell 36, 79–93 (2016).
Qi, X. H. et al. Increased cysteinyl-tRNA synthetase drives neuroinflammation in Alzheimer’s disease. Transl. Neurodegener. 13, 3 (2024).
Avalos, J., Corrochano, L. M. & Brenner, S. Cysteinyl-tRNA synthetase is a direct descendant of the first aminoacyl-tRNA synthetase. FEBS Lett. 286, 176–180 (1991).
Liu, M. et al. The critical role and molecular mechanisms of ferroptosis in antioxidant systems: a narrative review. Ann. Transl. Med. 10, 368 (2022).
Sato, M. et al. The ferroptosis inducer erastin irreversibly inhibits system x(c)- and synergizes with cisplatin to increase cisplatin’s cytotoxicity in cancer cells. Sci. Rep. 8, 968 (2018).
Prieto-Bermejo, R. & Hern ndez-Hern ndez, A. The importance of NADPH oxidases and redox signaling in angiogenesis. Antioxidants (Basel, Switzerland) 6, 32 (2017).
Kim, Y. W. & Byzova, T. V. Oxidative stress in angiogenesis and vascular disease. Blood 123, 625–631 (2014).
Xia, C. et al. Reactive oxygen species regulate angiogenesis and tumor growth through vascular endothelial growth factor. Cancer Res. 67, 10823–10830 (2007).
Zhang, W., Lin, X. & Chen, S. Cysteinyl-tRNA synthetase 1 promotes ferroptosis-induced cell death via regulating GPX4 expression. J. Oncol. 2022, 4849174 (2022).
Guidelines for diagnosis and treatment of moyamoya disease (spontaneous occlusion of the circle of Willis). Neurol. Med. Chir. 52, 245–266 (2012).
Cao, Y. et al. The ChinaMAP analytics of deep whole genome sequences in 10,588 individuals. Cell Res. 30, 717–731 (2020).
Xu, J. et al. HAPLN3 p.T34A contributes to incomplete penetrance of moyamoya disease in Chinese carrying RNF213 p.R4810K. Eur. J. Neurol. 31, e16473 (2024).
Acknowledgements
We would like to thank all the individuals who participated in this study and we also would like to express our sincere gratitude to the China Metabolic Analytics Project (ChinaMAP) for providing invaluable resources and datasets that were instrumental in our research. This research was supported by the National Natural Science Foundation of China (Grant No. 82173610); the Department of Science and Technology of Liaoning Province (Grants No. 2023JH2/20200025 and 2023-MSLH-371); and the China Postdoctoral Science Foundation (Grants No. 2024T171043, 2023MD744266, and GZC20233120).
Author information
Authors and Affiliations
Contributions
W.Y. was responsible for the identification and screening of moyamoya disease pathogenic genes, population validation, and the writing and revision of the manuscript. Z.Z.X. was responsible for the collection of clinical samples and clinical information from Moyamoya disease patients. C.G. performed molecular biology experiments to validate the pathogenic genes. Z.Q. and Z.Z.S. were responsible for the collection of clinical information. L.Y.Z. and T.X.N. performed data analysis and prepared the figures. L.S.M. acquired and analyzed imaging data from moyamoya disease patients. Z.T. conducted bioinformatics analysis and performed calibration of data analysis. Y.D. collected patient clinical samples. D.L. and L.W.Y. were responsible for the overall design and supervision of the study, manuscript writing, and revision. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, Y., Zou, Z., Chen, G. et al. Identification and validation of CARS1 p.E712V and NF1 p.Q2002X in sporadic Moyamoya disease across 30 trio pedigrees. npj Genom. Med. 11, 9 (2026). https://doi.org/10.1038/s41525-025-00539-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41525-025-00539-8