Abstract
It is widely acknowledged that articular cartilage lacks the ability to regenerate. However, if such regeneration were possible, which cell type would generate new tissue? The p21−/− mouse provides an excellent platform to explore this question, hence, we conducted lineage tracing on Paired related homeobox 1 (Prrx1/Prx1) cells post-injury to determine whether endogenous Prx1+ cells contribute to regenerated tissues post-injury. p21−/− mice displayed enhanced endogenous cartilage regeneration, accompanied by notable differences in the number and kinetics of Prx1+ cells within and around the injury site. In p21−/− mice, Prx1+ cells underwent chondrogenesis, ultimately contributing to the regenerated articular cartilage layer. These findings underscore the impact of tissue-resident cells on cartilage regeneration, albeit under abnormal conditions. If the conditions within the joint could be manipulated to favor such a regenerative environment, these endogenous cell types might be recruited to facilitate the formation of a new articular cartilage surface post-injury.
Similar content being viewed by others
Introduction
Articular cartilage regeneration has been known to lack intrinsic regenerative capacity since at least the 1700 s1. Unlike more vascular tissues, cartilage lacks a blood supply, limiting its healing potential in response to injury2. Even small defects in cartilage fail to restore their low-friction, Proteoglycan-4/lubricin-rich surface, leading to further deterioration over time, and culminating in osteoarthritis (OA)3,4.
While there’s a growing interest in utilizing exogenously delivered mesenchymal progenitor cells (MPCs) for cartilage injury and OA treatment5, the MRL/MpJ (MRL, “super-healer”) mouse strain has shown remarkable endogenous cartilage regeneration6,7,8. This phenomenon, previously thought to be limited to amphibians, has sparked significant interest9. Studies in MRL mice suggest that their superior healing ability may be due to differences in MPC quantity and/or quality. Naviaux et al. noted the presence of undifferentiated stem cells in uninjured MRL mice, retaining stemness markers even in differentiated tissues like the heart10. Additionally, exogenously delivered MRL synovial MPCs have shown homing capabilities to cartilage injury sites, with increased numbers observed within full-thickness cartilage defects (FTCD)11.
Earlier research linking amphibian regeneration to cell accumulation in G2/M phase prompted further investigation by Bedelbaeva et al. who connected a single gene deletion in p21 to the MRL regeneration phenotype using p21−/− mice, which displayed complete ear hole closure12. Supporting this, our lab group demonstrated that p21−/− mice can regenerate FTCD, unlike age-matched C57BL/6 mice13. However, identifying the specific cell type(s) responsible for articular cartilage regeneration remains a challenge.
To address this, we utilized the p21−/− mouse model, known for robust spontaneous cartilage regeneration, to investigate the cells driving this abnormal repair response. Our study focuses on Prx1+ cells as potential candidates for articular cartilage regeneration. Prx1 is a transcription factor highly expressed in all mesenchymal cells of the developing limb14. Previous in vivo studies have shown that Prx1+ cells give rise to various mesenchymal lineages, including bone, cartilage, and fat, in postnatal mice15,16,17,18. Additionally, cartilaginous callus formation post-fracture is predominantly derived from chondrocytes differentiated from Prx1+ cells19. These findings provide a strong rationale for investigating Prx1+ cells’ involvement in bone and cartilage repair in vivo.
Results
Deletion of p21 enhances spontaneous articular cartilage regeneration
To validate the positive impact of p21 absence on the endogenous healing capacity of articular cartilage post-injury (PI), histological samples from p21−/− mice were compared to those from C57BL/6 (p21+/+) mice before and after an FTCD at various time points: 1 and 3 days post-injury (DPI), 1, 2, 4, and 8 weeks post-injury (WPI) (Fig. 1A–C). It is also important to note, that this full thickness defect model extends to the subchondral bone8.
The experimental outline (A) is depicted, followed by representative Safranin O stained sections depicting uninjured articular cartilage as the injury site post-FTCD in C57BL/6 (B) and p21−/− (C) mice. Fluorescent images (D) show the localization of Prg4 and Col2 and Acan within the uninjured cartilage and the FTCD area at 4 and 8 weeks post-injury (WPI) for C57BL/6 and p21−/− mice (n = 5 mice for each group at each timepoint). Cartilage healing was assessed using a grading scale (E) in sagittal tissue sections from both C57BL/6 and p21−/− mice (n = 5 mice per strain per timepoint). A T-test was used to determine significance (p < 0.05) between strains at each timepoint. Scale bars are equal to 50 µm.
In both C57BL/6 and p21−/− mice, a visible disruption to the Safranin O-positive (red staining) articular cartilage surface was observed post-FTCD at all examined time points. While the surface irregularity appeared to qualitatively improve in C57BL/6 mice at 2 weeks post-injury (Fig. 1B), proteoglycan matrix staining (Safranin O) within the defect area remained absent and decreased over time post-FTCD. Conversely, p21−/− mice displayed increased Safranin O staining within the injury site by 2 weeks post-FTCD, and this increased staining persisted at 4 and 8 WPI (Fig. 1C).
A significant difference in cartilage healing/repair score was observed as early as 3 DPI (Fig. 1E), with p21−/− mice demonstrating enhanced cartilage regeneration compared to C57BL/6 mice. This enhanced cartilage regeneration ability of p21−/− mice was observed at all time points examined except for 1 DPI.
To determine if the repaired/regenerated tissue was morphologically/molecularly consistent with articular cartilage, sections from C57BL/6 and p21−/− mice were stained for Prg4 (also known as Lubricin), which is required for normal cartilage lubrication and homeostasis, Collagen 2 (Col2), the primary extracellular matrix (ECM) molecule in articular cartilage and Aggrecan (Acan), the primary proteoglycan present in the ECM (Fig. 1D). Within uninjured murine articular cartilage, Prg4, Col2 and Acan staining was observed within both strains. Minimal Prg4 staining was observed in the FTCD within C57BL/6 mice over time, while staining was evident in the FTCD area in p21−/− mice. However, it should be noted that the Prg4 staining in p21−/− mice was not enriched in the superficial layer, where it is normally found in normal cartilage. Minimal Col2 staining was observed in C57BL/6 mice post-injury, in contrast to p21−/− mice, where robust staining was observed throughout the injury site (Fig. 1D). The same trend was observed with Acan staining in where pronounced Acan staining was observed in the FTCD area of p21−/− mice at 8 WPI. The mean fluorescence intensity (MFI) for Prg4, Col2 and Acan staining was quantified, and increased MFI for all markers was observed in p21−/− mice at 8 WPI (Fig. S1).
These results indicate that p21−/− mice not only demonstrate the capacity to regenerate a morphologically consistent articular-like cartilage layer post-injury but also express proteins found in native articular cartilage that are critical for its normal function.
Presence of tissue resident MPCs and macrophages post-FTCD in C57BL/6 and p21−/− mice
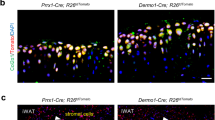
In Figs. 2–6, fluorescence imaging was utilized to assess the presence and distribution of endogenous Prx1+ MPCs (lineage traced (LT)/tdTomato+, and Prx1promoter-driven expression (PD)/GFP+) in knee joints of both C57BL/6 and p21−/− mice across time points within the cartilage injury site (FTCD), adjacent synovium (SYN), and fat pad (FP). Characterization was performed in uninjured joints and injured ones at 1-day and 1-, 2-, 4-, 8-weeks post injury.
The top section provides a diagrammatic representation of the experimental design and time-points. Safranin O stained sections shown the injury site in both C57BL/6 (A–C) and p21−/− mice (F–H). Fluorescent images depict the localization of Prx1 (non-lineage traced/promoter driven GFP expression in green; lineage traced in red) progenitors and Ki67/proliferating cells (blue) within the cartilage defect area (B, C, G, H), adjacent synovium (D, I), and fat pad (E, J) in both C57BL/6 and p21−/− mice. In images A-E, C57BL/6 mice are shown, while images F-J represent p21−/− mice. Notably, arrows in C’ and H’ (lacking the Nuc channel) highlight tdTomato+ cells within the articular cartilage at the margin of the FTCD (arrows). Cell quantification was performed within three regions: FTCD, SYN, and FP, utilizing a tissue cytometry approach (K). A T-test was used to determine significance (p < 0.05) between strains at each timepoint for each marker. An n = 5 mice per strain per timepoint was used in this experiment. All values are expressed as mean ± standard deviation (SD). Scale bars are equal to 50 µm. The white dotted lines outline the cartilage injury site, the yellow dotted lines mark the boundary between the articular cartilage and subchondral bone.
The top panel illustrates the experimental design and time-points. Safranin O stained sections shown the injury site in both C57BL/6 (A) and p21−/− mice (F). Fluorescent images depict the localization of Prx1 progenitors (non-lineage traced/promoter driven GFP expression in green; lineage traced in red) and Ki67/proliferating cells (blue) within the cartilage defect area (B, C, G, H), adjacent synovium (D, I), and fat pad (E, J) in both C57BL/6 (A–E) and p21−/− mice (F–J), with arrows in H’ indicating GFP+ cells lining the articular cartilage at the surface of the FTCD. Images C’ and H’ have the Nuc channel removed. Cell quantification was performed within three regions: FTCD, SYN, and FP using tissue cytometry (K). A T-test was used to determine significance (p < 0.05) between strains at each timepoint for each marker. An n = 5 mice per strain per timepoint was used in this experiment. All values are expressed as mean ± standard deviation (SD). Scale bars are equal to 50 µm. The white dotted lines outline the cartilage injury site, the yellow dotted lines mark the boundary between the articular cartilage and subchondral bone.
The top panel illustrates the experimental design and time-points. Safranin O stained sections shown the injury site in both C57BL/6 (A) and p21−/− mice (F). Fluorescent images depict the localization of Prx1 progenitors (non-lineage traced/promoter driven GFP expression in green; lineage traced in red) and Ki67/proliferating cells (blue) within the cartilage defect area (B, C, G, H), adjacent synovium (D, I), and fat pad (E, J) in both C57BL/6 and p21−/− mice. Images C’ and H’ have the Nuc channel removed. Arrows highlight tdTomato+ cells within the articular cartilage surface of the full-thickness cartilage defect (FTCD). Cell quantification was performed within three regions: FTCD, SYN, and FP using tissue cytometry (K). A T-test was used to determine significance (p < 0.05) between strains at each timepoint for each marker. An n = 5 mice per strain per timepoint was used in this experiment. All values are expressed as mean ± standard deviation (SD). Scale bars are equal to 50 µm. The white dotted lines outline the cartilage injury site, the yellow dotted lines mark the boundary between the articular cartilage and subchondral bone.
The diagram illustrates the experimental design and time-points. Safranin O stained sections shown the injury site in both C57BL/6 (A) and p21−/− mice (F). Fluorescent images depict the distribution of Prx1 progenitors (non-lineage traced/promoter driven GFP expression in green; lineage traced in red) and Prg4 expression (blue) in the cartilage defect area (B, C, G, H), adjacent synovium (D, I), and fat pad (E, J) of both C57BL/6 and p21−/− mice. GFP+ and tdTomato+ cells within the articular cartilage surface of the full-thickness cartilage defect (FTCD) are highlighted with arrows (C’H’). Images C’ and H’ have the Nuc channel removed. Cell quantification was performed in three regions (FTCD, SYN, and FP) using tissue cytometry (K). A T-test was used to determine significance (p < 0.05) between strains at each timepoint for each marker. An n = 5 mice per strain per timepoint was used in this experiment. All values are expressed as mean ± standard deviation (SD). Scale bars are equal to 50 µm. The white dotted lines outline the cartilage injury site, the yellow dotted lines mark the boundary between the articular cartilage and subchondral bone.
The diagram illustrates the experimental design and time-points. Safranin O stained sections shown the injury site in both C57BL/6 (A) and p21−/− mice (F). Fluorescent images depict the distribution of Prx1 progenitors (non-lineage traced/promoter driven GFP expression in green; lineage traced in red) and Prg4 expression (blue) in the cartilage defect area (B, C, G, H), adjacent synovium (D, I), and fat pad (E, J) of both C57BL/6 and p21−/− mice. GFP+ and tdTomato+ cells within the articular cartilage surface of the full-thickness cartilage defect (FTCD) are highlighted with white arrows (C’H’), while Prg4 expression is identified by the yellow arrows. Images C’ and H’ have the Nuc channel removed. Cell quantification was performed in three regions (FTCD, SYN, and FP) using tissue cytometry (K). A T-test was used to determine significance (p < 0.05) between strains at each timepoint for each marker. An n = 5 mice per strain per timepoint was used in this experiment. All values are expressed as mean ± standard deviation (SD). Scale bars are equal to 50 µm. The white dotted lines outline the cartilage injury site, the yellow dotted lines mark the boundary between the articular cartilage and subchondral bone.
Uninjured Joints
To determine the localization of the Prx1+ and Prx1 lineage-traced cells in the uninjured joints, mice were injected with 4-OHT for 5 consecutive days followed by a 7 day washout period. At the conclusion of this washout period, C57BL/6 and p21−/− mice were sacrificed and their joints examined with histology and immunofluorescence (IF) (Fig. S2). Staining for GFP and tdTomato showed that most labelled cells in both strains were present in the SYN and FP, with no positive cells observed in the articular cartilage (CART). In C57BL/6 mice, that majority of labelled cells in the synovium and fat pad co-expressed GFP and tdTomato. In p21−/− mice, few double positive cells were observed, with GFP+Tomato- cells being observed in the intimal layer and GFP-tdTomato+ cells being observed deeper in the synovium – which also appeared to be taking on a chondrocyte-like morphology and SafO staining (Fig. S2). This suggests that while Prx1+ cells are maintained in synovium and fat pad of C57BL/6 joints, even over a relatively short pulse/chase period in uninjured p21−/− mice there is a differentiation of Prx1+ cells and concurrent appearance of a new (non-lineage traced) population of Prx1+ cells (GFP+Tomato-).
1 Day Post Injury
To understand how the how the dynamics of the Prx1+ cell population wound change immediately after a joint injury, knee joints were harvested from both mouse strains at 1 DPI. GFP+ (Prx1+) and tdTomato+ (Prx1+ lineage traced) cells were observed within the FTCD of both C57BL/6 (Fig. 2A–C) and p21−/− mice (Fig. 2F–H). In both mouse strains, GFP+ cells were distributed throughout the defect and adjacent subchondral bone. However, while tdTomato+ cells were present in the subchondral bone of p21−/− mice, in C57BL/6 animals, the majority of tdTomato+ cells were observed in the FTCD. In both mouse strains, tdTomato+ cells were observed associated with the articular cartilage lining the margins of the FTCD (Fig. 2, arrows C’H’). This indicates a near immediate (~24 h) response of these cell populations to the injury in both strains - as no GFP+ nor tdTomato+ cells were observed in the cartilage of the uninjured controls (Fig. S2). A tissue cytometry workflow was developed and employed to identify cells based on their nuclear marker expression and then quantify the different populations in each region (Fig. S3), including proliferating cells (Ki67+) given the established role of p21 as a critical cell cycle regulator.
Within the FTCD, no difference in the abundance of tdTomato+ cells was observed, but an increase in GFP+ cells was noted in p21−/− mice (Fig. 2K). Furthermore, in p21−/− mice, there was an increase in proliferating cells, including progenitor populations (Fig. 2K). At 1 DPI, there were no differences in progenitor abundance or proliferating cells within the adjacent synovium. However, the proliferative trends observed in the fat pad mirrored those of the defect site (Fig. 2).
Given that spontaneous regenerating mouse strains such as MRL demonstrate a blunted macrophage response post-spinal cord injury20, and neutrophil depletion in C57BL/6 mice reduced inflammation and resulted in an accelerated corneal healing phenotype similar to MRL mice21, we also examined macrophage populations post-FTCD. F4/80 was employed as a pan macrophage marker, while CD38 was employed as a pro-inflammatory macrophage marker and CD206 as an anti-inflammatory macrophage marker (Fig. S4). These markers are routinely employed to identify these macrophage (sub)populations in mice22,23,24,25. There was an overall trend of increased macrophage abundance (of all sub-types) in C57BL/6 mice vs. p21−/− mice within the defect, synovium, and fat pad (Fig. S4I) at this timepoint.
1 Week Post-Injury
At 1 WPI (Fig. 3), cells expressing GFP and/or tdTomato were observed both throughout and adjacent to the injury site (Fig. 3A–H) in both strains. Additionally, GFP+ cells formed a continuous lining on the surface of the defect in p21−/− mice (Fig. 3, H’arrow). Notably, there was a stark contrast in the localization of tdTomato+ cells between strains. In p21−/− mice, tdTomato+ cells were distributed throughout the defect and adjacent subchondral bone, whereas in C57BL/6 mice, these cells were largely confined to a non-cartilaginous-like tissue that developed adjacent to the cartilage surface (Fig. 3A–H). Although significant differences in the abundance of the examined cell populations were observed, these differences between strains were not as pronounced as those seen at 1 DPI. It is noteworthy that a higher percentage of proliferating GFP+ and/or tdTomato+ cells were observed in the synovium of p21−/− mice, while these populations were more proliferative in the fat pad of C57BL/6 mice (Fig. 3K)
Upon examining macrophage populations at 1 WPI, it was found that all populations were increased within the FTCD of p21−/− mice, while the numbers of CD206+ and F4/80+ positive cells were elevated in the synovium and fat pad of C57BL/6 mice (Fig. S5I).
2 Weeks Post-Injury
At 2 weeks post-injury (WPI), a notable shift in the distribution of GFP+ and tdTomato+ cells within the injury site occurred (Fig. 4). GFP+ cells were observed both lining and dispersed throughout the defect site in both strains (Fig. 4C’, H’arrows). However, distinct differences were noted in the localization of tdTomato+ cells within the defect between strains. In p21−/− mice, tdTomato+ cells were prominently concentrated in the center of the defect, particularly within the regenerating articular cartilage surface (Fig. 4, H’arrow). Conversely, in C57BL/6 mice, TdTomato+ cells were observed throughout the depth of the defect and on its surface, albeit mainly localized to the injury margins similar to what was observed at 1 DPI in this strain (Fig. 4, C’arrow).
Regarding cell proliferation, for the most part, higher counts of proliferating cells, including those double positive for Ki67 and either GFP or tdTomato, were noted in FTCD, synovium and fat pad of p21−/− mice (Fig. 4, K).
Examination of macrophage populations at 2 WPI in the FTCD found no differences in CD206+ cells between strains, while CD38+ cells were increased in p21−/−mice and F4/80+ cells were increased in C57BL/6 mice (Fig. S6). In the synovium and fat pad, no differences were observed with the CD38+ population between strains, however, both CD206+ and F4/80+ cells were increased in C57BL/6 mice (Fig. S6I).
4 Weeks Post-Injury
At 4 WPI (Fig. 5), there was a dramatic change in the localization tdTomato+ cells throughout the injury site (Fig. 5A–H). Nearly the entire newly regenerated articular cartilage surface (Fig. 5, H’) in p21−/− mice contained tdTomato+ cells signifying that this new tissue layer is derived primarily from the Prx1+ MPCs present at the time of 4-OHT delivery (Fig. 5H’arrows). In C57BL/6 mice, there were GFP+ and tdTomato+ cells lining the surface of the injury site (Fig. 5C’arrows), however, the tissue present in this area did not resemble articular cartilage histologically (Fig. 5A), nor did it stain for Safranin O. To determine if superficial chondrocytes had been reconstituted at this point in the healing process, Prg4 was used as marker26. While there was Prg4 expression observed in both strains, minimal staining was observed in either strain at the cartilage surface. However, there was very intense Prg4 staining observed in the marrow cavity of the subchondral bone in p21−/− mice that was not observed in C57BL/6 mice (Fig. 5B, G). Since Prg4 is a secreted protein that is found throughout the body and blood, we cannot comment if this Prg4 was produced from cells in the marrow cavity or elsewhere, therefore, we performed in situ hybridization to investigate the localization of Prg4 mRNA (Fig. S7). In the bone marrow cavity of C57BL/6 and p21−/− mice, Prg4 was enriched in cells adjacent to blood vessels (Fig. S7, A’,B’arrows). However, in p21−/− mice Prg4 was also found throughout the bone marrow cavity similar to what was observed with the Prg4 protein expression. Furthermore, this phenomenon of increased Prg4 protein in the marrow cavity was only observed at this timepoint in the p21−− mice.
When macrophage populations were examined at 4 WPI, it was found that there was an increase in all populations within the injury site of C57BL/6 mice, while there were only minimal differences observed in the different macrophage populations between strains in the synovium and fat pad (Fig. S8).
8 weeks post-injury
To examine if the regenerated cartilage-like tissue in p21−/− mice was stable at later timepoints, animals were also examined at 8 WPI (Fig. 6). In p21−/− mice, tdTomato+ cells were still observed throughout the surface of the regenerated articular cartilage (Fig. 6, H’white arrow). In C57BL/6 only minimal lineage traced cells were observed in the injury site, primarily at the margin (Fig. 6C’white arrow) similar to earlier timepoints. Prg4 expression was observed throughout the surface of the newly regenerated articular cartilage in p21−/− mice (Fig. 6H’yellow arrow). Prg4 surface staining was also observed in C57BL/6 mice, however, it was not observed as a continuous layer in this strain (Fig. 6C’yellow arrow).
When macrophage populations were examined at 8 WPI, it was found that there was an increase in CD38+ (pro-inflammatory) macrophages in the FTCD, synovium and fat pad in C57BL/6 mice (Fig. S9I).
Trends in populations over time
When the quantification of Prx1+ MPCs and macrophage populations were examined as a whole over-time (Fig. S10), several distinct patterns between the mouse strains were observed. Overall, there appeared to be a quicker response of MPCs to enter the defect site in p21−/− mice vs. C57BL/6 mice. This was not observed in the synovium or fat pad, as both strains demonstrate a rapid increase in MPCs post-injury. In terms of macrophage response, there was a heightened CD38+ cell response in C57BL/6 mice immediately after the cartilage injury which decreased overtime, with a slight increase in these populations again at 8 weeks post-injury in the C57BL/6 mice (Fig. S10). Overall, we believe this suggests that there is some level of coordination of cell responses in the cartilage and adjacent tissues post-injury and that this is partially dysregulated with the loss of p21. Furthermore, there appears to be a greater increase in MPCs and their progeny across all tissues examined vs. macrophages, even in vascularized tissues such as the synovium and fat pad.
Differential regulation of the inflammatory response post-injury
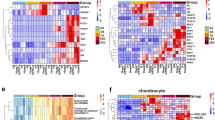
Since inflammation plays a prominent role in cartilage degeneration and it has previously been shown that healer strains of mice7,21,27,28 demonstrate a number of differences in systemic cytokine expression including lower levels of IL-1α and higher levels of IL-4 and IL-10 post-injury29, we next decided to examine if the cartilage regeneration in p21−/− mice was also associated with differential regulation of systemic inflammation post-FTCD (Fig. 7).While a number of differences within individual cytokine expression was observed over time, the two timepoints that stood out were 3DPI and 8WPI (Fig. 7A). Specifically, an increase in expression of most of the cytokines examined was observed within p21−/− mice at 3DPI. These trends are even more evident when directly compared to the baseline levels (Fig. S11). Specifically, at 3DPI nearly all cytokines are increased in the p21−/− mice compared to baseline and C57BL/6 post-injury except IL-10. At 8WPI, there are very few similarities between the C57BL/6 and p21−/− mice and there remain several differences within strains at 8WPI compared to the baseline levels suggesting that the systemic respond triggered by the cartilage injury has not resolved.
A Luminex assay measuring 32 cytokines was undertaken on serum in both strains (A). The 2 columns that are highlighted in red dashed boxes indicate timepoints where a number of differences were observed. The white vertical dashed box represents differences in Ccl2 expression between strains. Graphical representation of the experimental design (B). Changes in serum Ccl2 levels over time and between strains (C). Quantification of Ccl2 mRNA expression in CD38+, CD206+ and GFP+ cells isolated from the synovium of both strains over time (D). *=p < 0.05, **=p < 0.01, ***=p < 0.001.
We would like to highlight Ccl2 as a cytokine of interest (Fig. 7A), as we and others have previously impacted Ccl2/Mcp1 in cartilage degeneration/regeneration23,30,31 as well as the chondrogenic potential of synovial MPCs32. While Ccl2 levels dramatically increased in both strains at 2 WPI, by 4 WPI, levels in p21−/− mice were approximately double those observed in C57BL/6 mice. At 8 WPI, Ccl2 levels dropped off sharply in p21−/− mice, almost to baseline; yet remained dramatically elevated in C57BL/6 (Fig. 7C). Because of the prominent increase in Ccl2 levels post-FTCD in both strains and since we have previously demonstrated Ccl2 is required for cartilage repair in vivo23, we decided to investigate if cells in the synovium of these mice were producing Ccl2 post-FTCD. CD38+ and CD206+ macrophages in addition to GFP+ Prx1 MPCs were isolated from the joints of mice without joint injuries or at 2, 4 or 8 WPI and examined for Ccl2 mRNA expression (Fig. 7D). In CD38+ cells, there was a significant upregulation of Ccl2 in both strains at injury, with expression returning to baseline levels in p21−/− mice at 8 WPI. There was minimal Ccl2 expression in CD206+ cells in either strain. Interestingly, there was a dramatic increase in Ccl2 expression in GFP+ cells in C57BL/6 mice, and the levels increased over time. This trend was not observed in p21−/− mice.
Cellular mechanisms
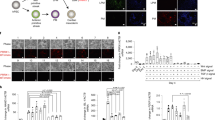
To gain a better understanding of the potential cellular mechanism(s) behind the observation that Prx1+ MPCs reconstitute the articular cartilage surface in p21−/− mice, GFP+ cells were isolated from the synovium of both p21−/− and C57BL/6 mice and differentiated into chondrocytes in vitro (Fig. 8). Three primary variables were examined in the chondrogenic protocol; the initial cell pellet number, Bmp2 concentration, and Tgfβ3 concentration. These three variables were examined over a continuum and Col2a1 mRNA expression was used as the primary outcome of chondrogenesis. It was observed that MPCs derived from p21−/− mice had the ability to effectively undergo chondrogenesis over a wider range of initial cell pellet number, and Bmp2 or Tgfβ3 concentrations vs. MPCs derived from C57BL/6 mice (Fig. 8A). This observation was validated by differentiating MPCs using sub-optimal levels of all three variables (50,000 starting cells, 250 µg/ml Bmp2 and 5 ng/ml Tgfβ3) (Fig. 8B). Under these sub-optimal conditions, MPCs from p21−/− mice maintained robust expression of Aggrecan and Sox9, while MPCs derived from C57BL/6 mice demonstrated only minimal expression of Aggrecan (Fig. 8C). These results may suggest that the MPCs present in the joints of p21−/− mice are more sensitive to pro-chondrogenic signals post-injury vs. their wild-type counterparts.
GFP+ cells isolated from the synovium of p21−/− and C57BL/6 mice were induced to undergo chondrogenesis in vitro. The initial start density of cells/pellet, Bmp2 concentration and Tgfβ3 concentration were all used as input variables while the expression of Col2a1 mRNA was employed as the outcome measure (A). Immunofluorescence of Aggrecan and Sox2 was used to validate chondrogenesis (B) and the expression of these markers were quantified using a tissue cytometry approach (C).
Discussion
We have previously shown that p21 deficiency in mice permits endogenous articular cartilage regeneration post-FTCD and that, compared to C57BL/6 wild-type mice, the newly regenerated cartilage tissue in p21−/− mice resembled hyaline cartilage at the histological level13. Therefore, the objective of the current study was to identify the cell type(s) (e.g., stem/progenitor and/or immune) involved in this spontaneous endogenous articular cartilage regeneration in p21-/- mice.
In support of our previous findings13, and those of an independent group33, we have confirmed that p21−/− mice regenerate a hyaline-like cartilage tissue after injury unlike C57BL/6 mice. Furthermore, we have now shown that the resultant cartilage tissue in p21−/− mice is positive for the articular cartilage ECM proteins Col2 and Prg4. While previous studies have shown that MRL6 and DBA/134 mice can spontaneously regenerate articular cartilage after injury, to our knowledge no study has examined the cell type(s) involved in the healing phenotype observed in these mouse strains. In the current study, we compared the numbers of Prx1+ cells (GFP+) and their differentiated progeny (tdTomato+) in uninjured knee joints from C57BL/6 and p21−/− mice and found that the Prx1+ lineage participates in reconstituting the new articular cartilage in this regenerating strain. We also identified several differences in the kinetics/timing of the Prx1+ lineage in the injury site, synovium and fat pad between the strains. Generalizing the results from all three tissues, there is an increase in Prx1+ cells immediately post-injury within the FTCD adjacent synovium and fat pad in both strains, which decreases over time. However, in C57BL/6 mice it appears that the numbers are beginning to increase again at 8 weeks post-injury, which may suggest the restarting of a second attempt at a repair response. It would be interesting to see if this cycle occurs continuously in the C57BL/6 over time with the degeneration of the cartilage and later the joint7, resulting from an ongoing ineffective repair response (Fig. 9). More specifically, p21−/− mice exhibit superior articular cartilage regeneration by healing from both the wound margin and the center. This enhanced repair is attributed to increased chondrocyte proliferation, migration, and reduced cellular senescence, fostering robust tissue regeneration13,33,35. In contrast, wild-type C57BL/6 mice demonstrate stagnant wound healing, with repair limited to the wound margins and little to no central integration (Fig. 9). This suggests that p21 deletion enhances intrinsic cartilage regenerative capacity, allowing for complete tissue restoration. Understanding these mechanisms provides insights into potential therapeutic strategies for improving cartilage repair in OA and joint injuries, where regeneration is typically poor.
p21⁻/⁻ mice exhibit superior articular cartilage regeneration compared to C57BL/6 mice, forming hyaline-like cartilage with increased Prx1⁺ cell activity and proliferation. In p21⁻/⁻ mice, Prx1⁺ MPCs migrate immediately into defect site and differentiate more effectively, contributing to improved cartilage restoration from both the wound margin and center. In contrast, C57BL/6 mice show limited repair and delayed or ineffective progenitor response. Additionally, p21⁻/⁻ mice demonstrate transient, tightly regulated inflammatory cytokine expression (e.g., Ccl2), while C57BL/6 mice exhibit persistent inflammation, hindering regeneration and favoring joint degeneration over time. This image was made with Biorender.
The differential response of the Prx1+ lineage to the cartilage injury in the synovium and fat pad (and between strains) suggests that a difference in MPC kinetics within the injury site and the origin of the MPCs (synovium vs. fat pad and even subchondral bone) may play role in determining the balance between repair (fibrocartilage) vs. regeneration (articular cartilage). This hypothesis agrees with a growing body of evidence suggesting that synovial-derived MPCs (endogenous or exogenous) migrate to areas of cartilage injury36,37,38,39,40. Yet, little evidence exists on what factors regulate the change in behaviour to favour regeneration. Our results suggest that the loss of p21 expression is sufficient to drive regeneration, and that the Prx1+ lineage in these mice is more sensitive to pro-chondrogenic cues (in vitro) than their wild-type counterparts. Yet, it remains unknown if this is what is occurring in vivo and what is the exact mechanism that is driving the reconstitution of the damaged articular cartilage, in part, by the Prx1+ lineage. There is supporting evidence of the observation that the Prx1+ lineage can directly differentiate into chondrocytes in vivo since these cells directly differentiate into the cartilage callus during the fracture repair process15,41.
While p21 is known primarily as a cell cycle regulator, previous studies have shown that by genetically decreasing p21 levels (iPSCs42,43) or by inhibiting p21 (synovial MPCs44), chondrogenic potential can be enhanced. This agrees with our Prx1 data since the regenerated cartilage tissue was tdTomato+, yet it is likely that other stem/progenitor populations also contribute to the regeneration either directly or indirectly. An additional MPC population to investigate in future studies would be Growth Differentiation Factor 5 (Gdf5) positive cells, since most, if not all, joint tissues (e.g., articular cartilage, ligaments, inner synovial lining) are derived from Gdf5+ cells39,45,46.
Taking into account that p21 is a potent cell cycle inhibitor47, and previous studies showed increased cell proliferation contributed to cartilage regeneration in p21−/− mice33 and has also been implicated in MRL regeneration9,21,48,49,50; we also quantified cell proliferation (Ki67+ cells) in the joint. We observed a dramatic increase in proliferation in the defect site of and p21−/− mice immediately post-FTCD, with similar levels of cell proliferation observed in C57BL/6 mice at 1 WPI. There was also an increase in proliferation in the adjacent synovium and fat pad of p21−/− mice at most of the timepoints examined. The increased cell proliferation observed in both strains post-FTCD (albeit at different times) suggests that cartilage injury induces proliferation normally and that the absence of p21 may prime the cells in the environment to react to this insult. These results strongly suggest that, in addition to cell cycle regulation, additional mechanisms are likely involved in endogenous articular cartilage regeneration.
Several studies have implicated a decreased inflammatory response (e.g. macrophages and/or pro-inflammatory cytokines) in the superior healing ability of MRL mice21,27,28,51. Furthermore, other ‘healer’ strains such as the “macrophageless” PU.1−/− mouse demonstrate complete and scar-free wound repair52. On the other hand, macrophage depletion has been shown to inhibit regeneration in several other regenerating animals53,54,55,56, convoluting our understanding of immune-regulated tissue regeneration. In our study, deletion of p21 resulted in altered macrophage numbers post-FTCD and resulted in altered systemic inflammatory cytokine expression. We observed opposing trends in the abundance of pro-inflammatory (CD38+) and anti-inflammatory (CD206+) macrophages between the strains and joint tissues within each strain. There is still significant controversy on the role and phenotypes of macrophages involved in tissue repair vs. regeneration57,58, with our own previous work in p21−/− mice ear regeneration demonstrating that in this strain, CD206+ macrophages are increased post-injury44.
In terms of systemic inflammatory response post-injury, our data suggests that an initial increase in inflammatory cytokine expression may be required to initiate a biological process required for cartilage regeneration post-FTCD. However, if the inflammatory response is persistent (as in C57BL/6), this may lead to an abortive healing response (e.g. fibrosis/fibrocartilage) and/or induction of potent pro-inflammatory molecules such as Tnfα59 that can trigger a degenerative process; accordingly, TNFα was dramatically elevated in C57BL/6 at 8 weeks post-FTCD (Fig. 7). It is also important to discuss the levels of CCL2/MCP-1 observed in the current study. Elevated Ccl2 expression is a hallmark of synovial joint injury in humans60,61,62,63, and has also been observed in mice after surgical induction of OA64. Yet, p21−/− mice displayed increased systemic Ccl2 expression at 3 days, 2 and 4 weeks post-FTCD. This was followed by the level of Ccl2 dropping off rapidly by 8 weeks post-FTCD in p21−/− but not C57BL6 mice. This suggests that not only is Ccl2 expression differentially regulated in the absence of p21, but that strict temporal regulation of Ccl2 may be playing a critical role in cartilage regeneration. This is consistent with our finding that Ccl2 is essential for endogenous cartilage regeneration23 in mice, but prolonged exposure can negatively impact the chondrogenic ability of synovial derived MPCs32. Overall, the literature to date would suggest that Ccl2 plays a dual role in articular cartilage biology. It promotes regeneration by recruiting reparative immune cells and MPCs, aiding tissue repair. However, in chronic inflammation, Ccl2 drives degeneration by attracting pro-inflammatory macrophages and exacerbating catabolic processes, contributing to OA progression and cartilage matrix breakdown.
In conclusion, we have demonstrated that there is migration of Prx1+ MPCs to the FTCD site in p21−/− and C57BL/6 mice, and in both strains these cells can differentiate into new articular cartilage, yet reconstitution of the articular cartilage surface from these cells is only effective in p21−/− mice (Fig. 9). Furthermore, our data suggests that alterations in cell signaling and/or trophic factors, rather than an increase in absolute cell numbers (e.g., MPCs and/ or immune cells), is likely responsible for endogenous articular cartilage regeneration in p21−/− mice. Using this unique model of articular cartilage regeneration, we have shown that cells of the Prx1 lineage have the potential to regenerate the articular cartilage surface post-injury, but for some reason(s) they are incapable of performing this function in wild-type mice. The elucidation of the mechanism(s) behind this observation would have a profound impact on the development of new therapeutic treatment strategies for cartilage injuries and/or OA.
Methods
Animal strains and sample size
All mice were handled in accordance with the Canadian Council on Animal Care Guidelines and the animal care protocol was approved by the University of Calgary Animal Care Committee (AC16-0043). The method of euthanasia approved by this protocol and used in this study was CO2 asphyxiation followed by cervical dissociation. The Prx1CreERT2-GFP mouse line used in this study was kindly provided by Dr. Shunichi Murakami (Case Western Reserve, Cleveland, OH)15. C57BL/6 and p21−/− (B6;129S2-Cdkn1atm1Tyj) were purchased from The Jackson Laboratory (Bar Harbor, ME)
To examine Prx1+ cells or their differentiated progeny we created the following mice strains on both mutant and wild-type backgrounds. Prx1CreERT2-GFP;R26RTdTomato;C57BL/6, and Prx1CreERT2-GFP;R26RTdTomato;p21−/− mice were used for experiments presented. To generate the Prx1CreERT2-GFP;R26RTdTomato;C57BL/6 mice the following breeding scheme was employed: Prx1CreERT2-GFP+/+ mice (C57BL/6 background) were bred to R26RTdTomato+/+ mice (C57BL/6 background) and genotyped to identify Prx1CreERT2-GFP+/+;R26RTdTomato+/+ (Fig. S12). The breeding scheme used for generation of Prx1CreERT2-GFP;R26RTdTomato; p21−/− mice was as follows: Prx1CreERT2-GFP+/+ (C57BL/6 background) mice were bred to p21−/− (C57BL/6 background) mice and genotyped to select for Prx1CreERT2-GFP+/+; p21−/− mice. R26RTdTomato+/+ mice were bred to p21−/− mice and genotyped to select for R26RTdTomato+/+; p21−/− mice. Established Prx1CreERT2-GFP+/+;p21−/− and R26RTdTomato+/+; p21−/− mouse lines were then crossed and genotyped to generate Prx1CreERT2-GFP+/+;R26RTdTomato+/+; p21−/− mice (Fig. S12).
FTCD were surgically induced in the left knee of 8 week old mice as described below. The sample sizes were; C57BL/6 (n = 35), Prx1CreERT2-GFP;R26RTdTomato;C57BL/6 (n = 35), p21−/− (n = 35), and Prx1CreERT2-GFP;R26RTdTomato; p21−/− (n = 35). Serum cytokine expression was analyzed from C57BL/6 (n = 27) and p21−/− (n = 52) mice post-FTCD.
Tamoxifen preparation and injections
The active isomer of tamoxifen ((Z)-4-Hydroxytamoxifen, Sigma) was dissolved in filter-sterilized sunflower oil to make solutions with a final concentration of 10 mg/mL. Aliquots were protected from light and stored at −20 °C. The day of each tamoxifen injection, aliquots were placed in a sonicator for 20 min at 37 °C to resuspend the tamoxifen prior to injection. Prior to FTCD, transgenic mice were injected (i.p.) with approximately 100 µL tamoxifen solution (100 mg/kg) once a day over 5 days, followed by a 1 week waiting period to allow for recombination.
Cartilage defect model
Prior to surgery, all animals received an intraperitoneal injection of buprenorphine (0.05 mg/kg) and were anaesthetized under isoflurane (Baxter) anesthesia (1.5% v/v 02) for the duration of the procedure. The FTCD injury was performed based on a protocol described by Eltawil et al.34. Briefly, a small skin incision on the medial side of the left knee was made using a scalpel and a 26 G needle was used to create a standardized cartilage defect injury13. The needle was held horizontally and the tip carefully inserted underneath the patella and into the femoral groove. After contacting the femoral groove inside the joint capsule, the needle was moved to a vertical position. A slight pressure with a twisting motion was applied at the contact with the femoral groove to make a circular wound approximately 0.6 mm in diameter that penetrated into the underlying subchondral bone7,8,13. After the FTCD was made, the needle was gently removed and the skin closed with a sterile wound clip.
Scoring of cartilage repair
Animals were sacrificed at 1 day, 3 days, 1, 2, 4, and 8 weeks post-induction of FTCD. Intact mouse knee joints were dissected and fixed in neutral buffered formalin (Sigma) for 5 days, then decalcified in EDTA for 10 days. Afterwards, samples underwent tissue processing for paraffin sectioning. To identify the defect site, every 5th serial sagittal sections (10 μm) was co-stained with Safranin-O and fast green. The defect site was landmarked as being directly under the patella. Furthermore, all slides containing the injury were viewed and the slide closeted to the center of the defect was graded for cartilage repair. There were three sections per slide per animal that were graded. The histological grading6 was undertaken for cell morphology (0–4), matrix staining (0–3), surface regularity (0–3), cartilage thickness (0–2), and integration with native cartilage (0–2). On this scale, healthy, uninjured cartilage is 14, while cartilage absence is 0. Blinded grading of all images was performed by two independent observers.
Immunofluorescence
At least 1 slide (three sections per slide) were taken from the center of the injury site per animal and stained for the respective markers. To prepare samples for immunostaining, serial sagittal paraffin sections (10 μm) were deparaffinized in CitriSolv (Fisher Scientific) and rehydrated using a series of graded ethanol to distilled water steps. Next, samples were subjected to antigen retrieval (10 mM sodium citrate, pH 6.0) and blocking (1:500 dilution; 100 μL rat serum:50 mL TRIS-buffered saline, 0.1% Tween 20 (TBST) for 1 h) steps were performed prior to going through TBST wash and antibody application steps. Antibodies conjugated to fluorophores for stem cells: Sca1 - Alexa Fluor 488 (Clone # D7), CD140a - phycoerythrin (PE) (Clone # APA5), macrophages: CD38 - Alexa Fluor 488 (Clone # 90), CD206 - R-PE (Clone # MR6F3), F4/80 – APC (Clone # BM8)(all eBioscience), cell proliferation: Ki67 – APC (Clone # SolA15), pro-inflammatory cytokine CCL2 – APC (Clone # 2H5), collagen 2 – APC (Developmental Studies Hybridoma Bank), and lubricin (PRG4, clone # 9G3, Millipore) – APC (All Thermo Fisher except where noted) were applied at 4 °C overnight. Sections were then washed three times at 10 min/wash in TBST, prior to mounting using EverBrite™ Mounting Medium with DAPI (Biotium) for nuclear counterstaining and coverslipped. Isotype controls were performed for IgG1, IgG2a, IgG2b with no positive staining observed.
Chondrogenic pellet sections were also stained with Aggrecan and Sox9. Primary antibodies conjugated to fluorophores included: Alexa Fluor® 647 Anti-Aggrecan antibody [6-B-4] (Abcam) and PE anti-Sox9 (D8G8H) (Cell Signalling). All slides were counterstained with the nucleic acid stain DAPI.
Tissue dissociation
In experiments where RNA was collected and/or single cells were required for cell sorting/flow cytometry; the synovium within the knee joints of mice was surgically removed and dissociated using the Multi Tissue Dissociation Kit 1 (Miltenyi Biotec). GFP+, CD38+ or CD206+ cells were sterilely sorted (University of Calgary, Flow Cytometry Core Facility). An aliquot of the cells was re-analyzed using flow to confirm the presence and purity of these populations.
Chondrogenesis
Prx1+ MPCs were pelleted at 50,000, 75,000 or 100,000 cells per pellet in 1.5 ml Eppendorf tubes by centrifugation. Cells were cultured in DMEM-F12 media with 10% FBS, 1% Anti-Anti, and 1% non-essential amino acids (all Thermo-Fisher) for 21 days. Chondrogenic control (chondro) was cultured in media containing 0, 100, 250 or 500 ng/mL BMP-2 (Peprotech), 0, 2.5, 5 or 10 ng/mL TGF-β3 (Peprotech), 10 M dexamethasone, 50 μg/mL ascorbic acid, 40 μg/mL proline, 100 μg/mL pyruvate and supplemented with 1× insulin, transferrin and selenium (all Sigma), and the pH of the final solution was adjusted to 7 with NaOH. Media was changed every 2–3 days, with special attention not to disturb the pellet.
Quantitative PCR
RNA from cultured cells was extracted using E.Z.N.A® Total RNA Kit I (OMEGA). RNA was converted to cDNA using the High-Capacity Reverse Transcriptase cDNA kit (Thermo-Fisher). mRNA levels were analyzed using TaqMan® Universal PCR Master Mix using TaqMan® Gene Expression Assay primers for mouse Ccl2, Col2a1 and 18S (endogenous control) on Quantstudio 6 Real-Time PCR System (all Thermo-Fisher). Samples were run in triplicate.
Histological imaging
An Axio Scan.Z1 Slide Scanner microscope (Carl Zeiss) outfitted with a Plan-Apochromat objective (10x/0.8 M27) was used to image slides. The following filters were applied: DAPI (353/465 nm), EGFP (493/517 nm), R-phycoerythrin (R-PE) (565/576 nm), DsRed (563/581 nm) and allophycocyanin (APC) (650 nm/660 nm).
Tissue cytometry
Cells of a given phenotype were identified and quantitated using the TissueQuest software (TissueGnostics), with cut-off values determined relative to the negative controls (non-stained and secondary alone controls). Gating and quantification of single/double positive cells were undertaken using these thresholds5,65. Briefly, the TissueQuest method uses algorithms to segment nuclei and the cell boundary surrounding each nucleus. Each additional fluorescent marker is then examined in the context of these segmented nuclei to determine if the marker overlaps with the boundary of the cell. It is also important to note that areas of bone marrow present in the subchondral bone region of the FTCD defect area were excluded in the tissue cytometry workflow due to the vast number of cells present in these areas and the inconsistency of the presence of regions from injury-to-injury site.
Luminex assay
After mice were sacrificed, whole blood was collected via cardiac puncture. Blood was allowed to form a clot before centrifugation and collection of serum. Serum inflammatory cytokine/chemokine/growth factor levels were analysis using the Luminex platform (Eve Technologies, Calgary, AB, Canada) with the mouse cytokine 32-plex discovery assay (Millipore) and prepared standards were included in all runs. The following proteins were quantified: CCL11 (Eotaxin), granulocyte-colony stimulating factor (G-CSF), granulocyte-macrophage colony-stimulating factor (GM-CSF), interferon gamma (IFNγ), IL-1α, IL-1β, IL-2, IL-3, IL-4, IL-5, IL-6, IL-7, IL-9, IL-10, IL-12 (p40), IL-12 (p70), IL-13, IL-15, IL-17, CXC chemokine ligand (CXCL) -10 (CXCL10) (IP-10), CXCL1 (KC), leukemia inhibitory factor (LIF),CXCL5 (LIX), CCL2 (MCP-1), M-CSF, CXCL9 (MIG), CCL3 (MIP-1α), CCL4 (MIP-1β), CXCL2 (MIP-2), CCL5 (RANTES), TNFα, vascular endothelial growth factor (VEGF). All samples were assayed in duplicate.
Statistical analysis
GraphPad Prism software (Version 7.0) was used to summarize the variables (mean ± SD) and perform standard descriptive statistics. Significance was reported at the level of p ≤ 0.05. Statistical analysis was by one-way or two-way ANOVA (as appropriate) followed by Tukey’s post hoc testing. SPSS v. 25 (SPSS, Inc., Chicago, IL) was used to analyze Luminex Assays. Student’s t test was used to test for cytokine differences between serum from C57BL/6 and p21−/− mice. To control for multiple comparisons a Bonferroni correction was employed. All statistical methods were reviewed by a biostatistician (Dr. Guomin Ren) at the University of Calgary.
Data availability
Data available within the article or its supplementary materials.
References
Hunter, W. VI. Of the structure and diseases of articulating cartilages. Philos. Trans. R. Soc. Lond. 42, 514–521 (1743).
Charalambous, C. P. The response of articular cartilage to mechanical injury. Class. Pap. Orthop. 64, 381–383 (2014).
Das, N. et al. Tryptase β regulation of joint lubrication and inflammation via proteoglycan-4 in osteoarthritis. Nat. Commun. 14, 1910 (2023).
Jay, G. D. et al. Prevention of cartilage degeneration and restoration of chondroprotection by lubricin tribosupplementation in the rat following anterior cruciate ligament transection. Arthritis Rheum. 62, 2382–2391 (2010).
Krawetz, R. J. et al. Mesenchymal progenitor cells from non-inflamed versus inflamed synovium post-ACL injury present with distinct phenotypes and cartilage regeneration capacity. Stem Cell Res. Ther. 14, 168 (2023).
Fitzgerald, J. et al. Evidence for articular cartilage regeneration in MRL/MpJ mice. Osteoarthr. Cartil. 16, 1319–1326 (2008).
Leonard, C. A. et al. Allogeneic bone marrow transplant from MRL/MpJ super-healer mice does not improve articular cartilage repair in the C57Bl/6 strain. PLoS One 10, e0131661 (2015).
Mak, J. et al. Evaluating endogenous repair of focal cartilage defects in C57BL/6 and MRL/MpJ mice using 9.4T magnetic resonance imaging: a pilot study. Magn. Reson. Imaging 33, 690–694 (2015).
Clark, L. D., Clark, R. K. & Heber-Katz, E. A new murine model for mammalian wound repair and regeneration. Clin. Immunol. Immunopathol. 88, 35–45 (1998).
Naviaux, R. K. et al. Retained features of embryonic metabolism in the adult MRL mouse. Mol. Genet. Metab. 96, 133–144 (2009).
Mak, J. et al. Intra-articular injection of synovial mesenchymal stem cells improves cartilage repair in a mouse injury model. Sci. Rep. 6, 23076 (2016).
Bedelbaeva, K. et al. Lack of p21 expression links cell cycle control and appendage regeneration in mice. Proc. Natl. Acad. Sci. USA107, 5845–5850 (2010).
Jablonski, C. L. C. L., Besler, B. A. B. A., Ali, J. & Krawetz, R. J. R. J. p21−/− mice exhibit spontaneous articular cartilage regeneration post-injury. Cartilage 13, 1947603519876348 (2021).
Logan, M. et al. Expression of Cre recombinase in the developing mouse limb bud driven by a Prxl enhancer. Genesis 33, 77–80 (2002).
Kawanami, A., Matsushita, T., Chan, Y. Y. & Murakami, S. Mice expressing GFP and CreER in osteochondro progenitor cells in the periosteum. Biochem. Biophys. Res. Commun. 386, 477–482 (2009).
Ouyang, Z. et al. Prx1 and 3.2kb Col1a1 promoters target distinct bone cell populations in transgenic mice. Bone 58, 136–145 (2014).
Akiyama, H. et al. Osteo-chondroprogenitor cells are derived from Sox9 expressing precursors. Proc. Natl. Acad. Sci. USA 102, 14665–14670 (2005).
Krueger, K. C., Costa, M. J., Du, H. & Feldman, B. J. Characterization of cre recombinase activity for in vivo targeting of adipocyte precursor cells. Stem Cell Rep. 3, 1147–1158 (2014).
Esposito, A., Wang, L., Li, T., Miranda, M. & Spagnoli, A. Role of Prx1-expressing skeletal cells and Prx1-expression in fracture repair. Bone 139, 115521 (2020).
Kostyk, S. K., Popovich, P. G., Stokes, B. T., Wei, P. & Jakeman, L. B. Robust axonal growth and a blunted macrophage response are associated with impaired functional recovery after spinal cord injury in the MRL/MpJ mouse. Neuroscience 156, 498–514 (2008).
Ueno, M. et al. Accelerated wound healing of alkali-burned corneas in MRL mice is associated with a reduced inflammatory signature. Invest. Ophthalmol. Vis. Sci. 46, 4097–4106 (2005).
Jablonski, K. A. et al. Novel markers to delineate murine M1 and M2 macrophages. PLoS One 10, e145342 (2015).
Jablonski, C. L., Leonard, C., Salo, P. & Krawetz, R. J. CCL2 but not CCR2 Is required for spontaneous articular cartilage regeneration post-injury. J. Orthop. Res. 37, 2561–2574 (2019).
Li, W. et al. CD38: a significant regulator of macrophage function. Front. Oncol. 12, 775649 (2022).
Ray, A. et al. Targeting CD206+ macrophages disrupts the establishment of a key antitumor immune axis. J. Exp. Med. 222, 40957 (2025).
Schmidt, T. A., Schumacher, B. L., Klein, T. J., Voegtline, M. S. & Sah, R. L. Synthesis of proteoglycan 4 by chondrocyte subpopulations in cartilage explants, monolayer cultures, and resurfaced cartilage cultures. Arthritis Rheum. 50, 2849–2857 (2004).
Heber-Katz, E. & Gourevitch, D. The relationship between inflammation and regeneration in the MRL mouse: Potential relevance for putative human regenerative (scarless wound healing) capacities?. Ann. N. Y. Acad. Sci. 1172, 110–114 (2009).
Lewis, J. S. et al. Genetic and cellular evidence of decreased inflammation associated with reduced incidence of posttraumatic arthritis in MRL/MpJ mice. Arthritis Rheum. 65, 660–670 (2013).
Ward, B. D. et al. Absence of posttraumatic arthritis following intraarticular fracture in the MRL/MpJ mouse. Arthritis Rheum. 58, 744–753 (2008).
Miotla Zarebska, J. et al. CCL2 and CCR2 regulate pain-related behaviour and early gene expression in post-traumatic murine osteoarthritis but contribute little to chondropathy. Osteoarthr. Cartil. 25, 406–412 (2017).
Appleton, C. T. G. et al. Reduction in disease progression by inhibition of transforming growth factor α-CCL2 signaling in experimental posttraumatic osteoarthritis. Arthritis Rheumatol. 67, 2691–2701 (2015).
Harris, Q. et al. Monocyte chemotactic protein-1 inhibits chondrogenesis of synovial mesenchymal progenitor cells: an in vitro study. Stem Cells 31, 2253–2265 (2013).
Ibaraki, K. et al. Deletion of p21 expression accelerates cartilage tissue repair via chondrocyte proliferation. Mol. Med. Rep. 21, 2236–2242 (2020).
Eltawil, N. M., De Bari, C., Achan, P., Pitzalis, C. & Dell’accio, F. A novel in vivo murine model of cartilage regeneration. Age and strain-dependent outcome after joint surface injury. Osteoarthr. Cartil. 17, 695–704 (2009).
Arthur, L. M. et al. Epimorphic regeneration in mice is p53-independent. Cell Cycle 9, 3667–3673 (2010).
Hunziker, E. B. Growth-factor-induced healing of partial-thickness defects in adult articular cartilage. Osteoarthr. Cartil. 9, 22–32 (2001).
Hunziker, E. B. & Rosenberg, L. C. Repair of partial-thickness defects in articular cartilage: cell recruitment from the synovial membrane. J. Bone Jt. Surg. Am. 78, 721–733 (1996).
Kurth, T. B. et al. Functional mesenchymal stem cell niches in adult mouse knee joint synovium in vivo. Arthritis Rheum. 63, 1289–1300 (2011).
Roelofs, A. J. et al. Joint morphogenetic cells in the adult mammalian synovium. Nat. Commun. 8, 15040 (2017).
Karystinou, A. et al. Distinct mesenchymal progenitor cell subsets in the adult human synovium. Rheumatology48, 1057–1064 (2009).
Murao, H., Yamamoto, K., Matsuda, S. & Akiyama, H. Periosteal cells are a major source of soft callus in bone fracture. J. Bone Miner. Metab. 31, 390–398 (2013).
Diekman, B. O. et al. Intra-articular delivery of purified mesenchymal stem cells from C57BL/6 or MRL/MpJ superhealer mice prevents posttraumatic arthritis. Cell Transpl. 22, 1395–1408 (2013).
D’Costa, S., Rich, M. J. & Diekman, B. O. Engineered cartilage from human chondrocytes with homozygous knockout of cell cycle inhibitor p21. Tissue Eng. - Part A 26, 441–449 (2020).
Bertram, K. L. et al. 17-DMAG regulates p21 expression to induce chondrogenesis in vitro and in vivo. DMM Dis. Model. Mech. 11, dmm033662 (2018).
Koyama, E. et al. A distinct cohort of progenitor cells participates in synovial joint and articular cartilage formation during mouse limb skeletogenesis. Dev. Biol. 316, 62–73 (2008).
Roelofs, A. J. et al. Identification of the skeletal progenitor cells forming osteophytes in osteoarthritis. Ann. Rheum. Dis. 79, 1625–1634 (2020).
Cazzalini, O., Scovassi, A. I., Savio, M., Stivala, L. A. & Prosperi, E. Multiple roles of the cell cycle inhibitor p21CDKN1A in the DNA damage response. Mutat. Res. - Rev. Mutat. Res. 704, 12–20 (2010).
Heber-Katz, E. The regenerating mouse ear. Semin. Cell Dev. Biol. 10, 415–419 (1999).
Heber-Katz, E., Leferovich, J., Bedelbaeva, K., Gourevitch, D. & Clark, L. The scarless heart and the MRL mouse. Philos. Trans. R. Soc. B Biol. Sci. 359, 785–793 (2004).
Chadwick, R. B. et al. Digit tip regrowth and differential gene expression in MRL/Mpj, DBA/2, and C57BL/6 mice. Wound Repair Regen. 15, 275–284 (2007).
Kench, J. A. et al. Aberrant wound healing and TGF-β production in the autoimmune-prone MRL/+ mouse. Clin. Immunol. 92, 300–310 (1999).
Martin, P. et al. Wound healing in the PU.1 null mouse - tissue repair is not dependent on inflammatory cells. Curr. Biol. 13, 1122–1128 (2003).
Godwin, J. W., Pinto, A. R. & Rosenthal, N. A. Macrophages are required for adult salamander limb regeneration. Proc. Natl. Acad. Sci. USA110, 9415–9420 (2013).
Petrie, T. A., Strand, N. S., Tsung-Yang, C., Rabinowitz, J. S. & Moon, R. T. Macrophages modulate adult zebrafish tail fin regeneration. Dev 141, 2581–2591 (2014).
Aurora, A. B. et al. Macrophages are required for neonatal heart regeneration. J. Clin. Invest. 124, 1382–1392 (2014).
Simkin, J., Gawriluk, T. R., Gensel, J. C. & Seifert, A. W. Macrophages are necessary for epimorphic regeneration in African spiny mice. Elife 6, e24623 (2017).
Wu, C. L., Harasymowicz, N. S., Klimak, M. A., Collins, K. H. & Guilak, F. The role of macrophages in osteoarthritis and cartilage repair. Osteoarthr. Cartil. 28, 544–554 (2020).
van der Kraan, P. M. The interaction between joint inflammation and cartilage repair. Tissue Eng. Regener. Med. 16, 327–334 (2019).
Kapoor, M., Martel-Pelletier, J., Lajeunesse, D., Pelletier, J.-P. & Fahmi, H. Role of proinflammatory cytokines in the pathophysiology of osteoarthritis. Nat. Rev. Rheumatol. 7, 33–42 (2011).
Furman, B. D. et al. Brief report: articular ankle fracture results in increased synovitis, synovial macrophage infiltration, and synovial fluid concentrations of inflammatory cytokines and chemokines. Arthritis Rheumatol. 67, 1234–1239 (2015).
Cuellar, V. G., Cuellar, J. M., Golish, S. R., Yeomans, D. C. & Scuderi, G. J. Cytokine profiling in acute anterior cruciate ligament injury. Arthrosc. - J. Arthrosc. Relat. Surg. 26, 1296–1301 (2010).
Watt, F. E. et al. Acute molecular changes in synovial fluid following human knee injury: association with early clinical outcomes. Arthritis Rheumatol. 68, 2129–2140 (2016).
Li, L. & Jiang, B.-E. Serum and synovial fluid chemokine ligand 2/monocyte chemoattractant protein 1 concentrations correlates with symptomatic severity in patients with knee osteoarthritis. Ann. Clin. Biochem. 52, 276–282 (2015).
Raghu, H. et al. CCL2/CCR2, but not CCL5/CCR5, mediates monocyte recruitment, inflammation and cartilage destruction in osteoarthritis. Ann. Rheum. Dis. 76, 914–922 (2017).
Mudigonda, S. et al. Proteoglycan 4 is present within the dura mater and produced by mesenchymal progenitor cells. Cell Tissue Res. 389, 483–499 (2022).
Acknowledgements
The authors would like to acknowledge the assistance of Guomin Ren for statistical advice, Catherine Leonard for assistance with histology and Anand Masson for assistance with grading the cartilage defect. Grants through Natural Sciences and Engineering Research Council (NSERC) of Canada (RJK # RGPIN-2014-04586), Canada Foundation for Innovation (RJK), Grace Glaum Professorship (RJK), and Canadian Institutes for Health Research (RJK # 201309MOP-311542-DIB-CBBA-170939) supported the work. CLJ was supported by NSERC Create and Biomedical Engineering Studentships. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Conception and design: C.L.J. and R.J.K. Analysis and interpretation of the data: C.L.J., D.M., J.C., D.M.M., P.S., and R.J.K. Drafting of the article: C.L.J. and R.J.K. Critical revision of the article for important intellectual content: C.L.J., D.M., J.C., D.M.M., P.S., and R.J.K. Final approval of the article: C.L.J., D.M., J.C., D.M.M., P.S., and R.J.K. Provision of study materials: C.L.J. and R.J.K. Obtaining of funding: P.S. and R.J.K. Collection and assembly of data: C.L.J., D.M., and R.J.K.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Jablonski, C.L., Modrcin, D., Cobb, J. et al. Prx1+ progenitors give rise to new articular cartilage when conditions are permissive for endogenous regeneration. npj Regen Med 10, 38 (2025). https://doi.org/10.1038/s41536-025-00425-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41536-025-00425-y