Abstract
Stevioside (ST) and rebaudioside A (Reb A) are 250 times sweeter than sucrose, but their bitterness and low water solubility limit their application. Rebaudioside Aα (Reb Aα) was synthesized using dextransucrase. Reb Aα exhibited 80-fold greater water solubility and lower thermal mass loss than Reb A and had melting point >180 °C. Reb Aα was stable within pH range 1.4–10 and exhibited excellent storage stability in aqueous solution. In vitro digestion revealed that Reb Aα was hydrolyzed into ST and glucose. In vivo, Reb Aα administration did not affect body weight gain, glucose tolerance, energy homeostasis, or liver histology. Also, in the Reb Aα group, inflammatory crown-like structures were not observed in white adipose tissue and uncoupling protein 1 was significantly upregulated in brown adipose tissue. Gut microbiota analysis revealed no significant differences in microbial composition between the control and Reb A or Reb Aα groups.
Similar content being viewed by others
Introduction
The overconsumption of added sugars is a well-documented contributor to metabolic disorders, including obesity, cardiovascular disease, and type 2 diabetes1,2. In response, global interest in sugar alternatives has surged, particularly those that reduce caloric intake without compromising sweetness3. Natural sweeteners, such as steviol glycosides derived from Stevia rebaudiana, have attracted significant attention due to their plant origin, high sweetness intensity (200–300 times sweeter than sucrose), and functional properties including antioxidant, antihypertensive, and anti-inflammatory activities4,5,6.
Among the steviol glycosides, stevioside (ST) and rebaudioside A (Reb A) are the most abundant and commonly used. However, their application in food systems is limited by their bitter aftertaste and poor aqueous solubility5. To overcome these limitations, enzymatic glycosylation has been explored as a strategy to improve their physicochemical and sensory properties. Cyclodextrin glucanotransferase (CGTase) is frequently used for this purpose, but it tends to randomly attach excessive glucose residues (up to 20 units), leading to heterogeneous products with increased caloric content7.
To achieve more selective glycosylation, dextransucrase has emerged as a promising alternative. This enzyme transfers D-glucopyranosyl residues from sucrose to hydroxyl groups of carbohydrate or non-carbohydrate molecules via α-glucosidic linkages, producing more structurally defined glycosides8. Previous studies have demonstrated that dextransucrase derived from Leuconostoc citreum can efficiently glycosylate steviol glycosides with high regioselectivity and conversion efficiency9. However, while α-glucosylated steviol glycosides have been shown to improve taste and solubility, most research has focused on glycosylation efficiency or sensory improvement without fully elucidating their molecular structures or biological behavior10. Moreover, although α-glucosylation is generally understood to enhance water solubility through the introduction of hydrophilic glucose moieties11, the precise physicochemical and in vivo properties of such derivatives remain largely unexplored. In particular, the metabolic fate and safety of glucosylated steviol glycosides are of interest. According to the European Food Safety Authority (EFSA), these compounds are expected to be hydrolyzed to steviol by intestinal microbiota, following metabolic pathways similar to those of native steviol glycosides, suggesting comparable safety profiles12.
In this context, we developed a novel α-glucosylated derivative of Reb A, designated Rebaudioside Aα (Reb Aα), through dextransucrase-mediated bioconversion. Despite its promising features, no prior study has structurally identified this compound or systematically compared it with Reb A in terms of solubility, stability, and bioavailability under both in vitro and in vivo conditions. Therefore, the aim of this study is to structurally characterize Reb Aα and to evaluate its physicochemical properties, solubility, and stability in comparison with Reb A. Furthermore, this study investigates the in vivo behavior of Reb Aα, including its absorption, metabolism, and safety, to determine its potential as a next-generation natural sweetener.
Results
Enzymatic synthesis of Reb Aα
Reb Aα was produced using L. citreum KM20-derived dextransucrase via glucosylation. Thereafter, it was purified using an MPLC C18 column at an ACN concentration of 33–34% under linear gradient conditions. Finally, Reb Aα was obtained via freeze-drying at –85 °C, yielding 39 g at an approximate conversion rate of 60% (Supplementary Fig. 1). This yield was comparable to that reported in a previous study¹⁷, indicating the consistency of the dextransucrase-mediated conversion process.
Reb Aα NMR spectra
The 1H- and 13C- NMR spectra of Reb Aα were compared with those of ST according to Jaitak et al., (2009)13. The ¹H- NMR spectrum of Reb Aα exhibited additional anomeric proton signals (δ 5.52, 4.24–4.26, 4.64, 4.20–4.24, 4.54–4.57, 4.47–4.50, and 4.35–4.38). Additionally, the corresponding ¹³C- NMR signals (δ 100.96, 72.67, 75.76, 74.43, 74.64, and 63.19) were confirmed through 1H-1H COSY and HSQC experiments (Table 1). These results indicate that Reb Aα contained one additional glucose moiety compared with ST, displaying consistency with the [M+Na]+ at m/z 989.5 observed in a previous liquid chromatography–mass spectrometry analysis14.
Furthermore, the anomeric proton (H-1”“) at δ 5.52 yielded a coupling constant (J) of 3.5 Hz in the 1H-NMR spectra. HMBC spectra confirmed the correlation between H-1”“ and C-6”, indicating that the α-D-glucose moiety was coupled to C-6” of ST. Therefore, Reb Aα was structurally interpreted as O-α-D-glucosyl-(1”“ → 6”)-stevioside (Fig. 1).
A reabudioside A. B rebaudioside Aα.
Water solubility evaluation
The aqueous solubilities of Reb A and Reb Aα were evaluated. Those of ST and Reb A (with only β-glycosidic linkages) were 10.97 ± 3.11 and 5.10 ± 2.93 mg/mL, respectively. However, the aqueous solubility of Reb Aα (with an α-glycosidic linkage) was 406.28 ± 20.91 mg/mL (Table 2), exhibiting a significant 80-fold improvement compared with that of Reb A, its isomer with the same molecular weight. This study found varying solubilities among steviol glycosides depending on the glycosidic linkage type (alpha or beta).
FT-IR analysis
The FT-IR spectra of Reb A and Reb Aα were compared (Fig. 2A). The Reb A spectrum displayed vibrations at 1729, 1072, 3336, and 2920 cm−¹. In a previous study, Reb A’s –C=O vibrations and C–O–C fragment were observed at 1,727 and 1076 cm−¹, respectively. Additionally, –OH vibrations appeared at 3387 cm−¹, and the –CH₃/–CH₂/–CH fragment was detected at 2939 cm−¹, aligning with the peaks observed in this study15. Similarly, Reb Aα yielded characteristic peaks at 1728, 1071, 3325, and 2920 cm−¹. Overall, Reb A and Reb Aα exhibited similar FT-IR spectra, suggesting that differences in glucose linkage type did not significantly affect the overall FT-IR spectra. Nevertheless, subtle peak shifts indicate structural differences between Reb A and Reb Aα.
A FT-IR, B XRD, C DSC, D TGA, E pH stability, F storage stability, G glucose generation rate, and H in vitro digestion rate.
XRD analysis
XRD analysis revealed that Reb A and Reb Aα yielded a strong diffraction peak around 16°. Additionally, several weak yet distinguishable peaks were identified. Reb A peaks were observed at 14.16, 16.47, 19.74, and 22.76°, while those of Reb Aα appeared at 14, 16, 19, and 22° (Fig. 2B). The abovementioned diffraction peak for Reb A (16°) was observed at the same position in both Reb A and Reb Aα in this study. Furthermore, the other peaks (14.08, 16.92, 19.34, and 22.64°) also appeared at comparable positions15. Reb A and Reb Aα exhibited similar diffraction patterns, confirming their amorphous and slight crystalline structures.
DSC analysis
This study compared the DSC thermograms of Reb A and Reb Aα (Fig. 2C), which exhibited two thermal endothermic peaks. Their first peak was confirmed at approximately 100 °C and attributed to sample dehydration. The amount of dehydration appeared not to differ between Reb A and Reb Aα. Reb A second peak was confirmed at 203.83 °C (ΔH = 4.32 J/g) and observed over a broad range, diverging from crystalline compounds, which are characterized by distinct peaks. A study found crystalline Reb A’s melting peak to occur at 238 °C, aligning with the endothermic peak in this study16. Nonetheless, Reb Aα second endothermic peak occurred at a lower temperature [184.83 °C (ΔH = 19.15 J/g)]. Similar to that of Reb A, this peak was also observed over a broad range. Reb Aα endothermic peak appeared 19 °C lower than that of Reb A, suggesting a structural difference between the two.
TGA analysis
This study compared the TGA thermograms of Reb A and Reb Aα (Fig. 2D), which exhibited third-stage mass loss owing to thermal decomposition. The first mass loss occurred at approximately 100 °C, where Reb A and Reb Aα retained 92.65 ± 2.34% and 92.29 ± 1.15% of their initial mass, respectively. This mass-loss temperature is identical to that previously reported (100 °C) for Reb A17. Therefore, this peak corresponds to the dehydration peak observed in the DSC thermograms, confirming that the mass loss emanates from dehydration.
Second-stage mass loss for Reb A and Reb Aα commenced at approximately 280–290 °C, during which significant mass loss occurred. During this stage, Reb A and Reb Aα retained 19.89 ± 1.67% and 25.64 ± 1.88% of their initial mass, respectively (Table S2). This temperature range is similar to that previously reported for Reb A (284.71 °C), further validating the results18. At 597 °C, Reb A retained 9.92 ± 0.45% of its initial mass, while Reb Aα retained 12.57 ± 1.08%, indicating that Reb Aα experienced less mass loss than Reb A.
pH stability
The stability of Reb A and Reb Aα was evaluated within a 1.4–10 pH range (Fig. 2E). At pH 10, Reb A and Reb Aα retained 84.56 ± 1.68% and 85.16 ± 1.04% of their initial concentrations, respectively, indicating no significant stability difference under alkaline conditions. However, significant differences occurred under acidic conditions. At pH 2, Reb A retained 82.67 ± 2.39% of its initial concentration, while Reb Aα retained 94.06 ± 0.98%. The difference increased at pH 1.4, where Reb A retained only 25.26 ± 3.19% of its initial concentration, while Reb Aα retained 63.17 ± 1.09%, demonstrating that Reb Aα was approximately 2.5-fold more stable than Reb A at pH 1.4. Despite minimal stability differences between Reb A and Reb Aα under alkaline conditions, Reb Aα exhibited significantly greater stability under acidic conditions.
Storage stability in aqueous solution
Significant differences in 1-month storage stability at 25 °C were observed between Reb A and Reb Aα (Fig. 2F). In 1% and 2% (w/v) aqueous solutions, Reb A exhibited considerable precipitation, with only 52.61 ± 1.69% and 32.14 ± 3.78% of the initial concentration remaining, respectively. In contrast, Reb Aα retained >90% of its initial concentration in most aqueous solutions. Notably, in 1% (w/v) aqueous solutions, >95% of the initial concentration remained, confirming its high stability in aqueous solutions.
GGR measurement
α-Glucosidase primarily hydrolyzes α-glycosidic bonds. The amount of glucose released from diverse steviol glycosides was measured using RIAP-derived α-glucosidase. Reb Aα steadily released glucose, reaching 2.3 ± 0.46 mg/mL in 2 h. Conversely, ST and Reb A exhibited almost no glucose release, indicating that Reb Aα with the α-glycosidic linkage had been hydrolyzed by α-glucosidase (Fig. 2G). Additionally, after α-glycosidic linkage hydrolysis of Reb A, the compound converted to ST (Supplementary Fig. 2). ST and Reb A are recognizably stable compounds that do not interfere with the intestinal metabolic pathway19, suggesting that, like ST and Reb A, Reb Aα remains stable during digestion, further supporting the in vitro digestion simulation findings.
Reb Aα bioaccessibility under in vitro digestion
The most prevalent, widely accepted in vitro method for simulating digestion is the InfoGest protocol, which employs simulated gastrointestinal fluids containing enzymes20. Reb A and Reb Aα were evaluated via in vitro digestion simulation of the human digestive system. In vitro simulated digestion resulted in ~99% Reb Aα hydrolysis, while only 2% of ST and Reb A were hydrolyzed (Fig. 2H). No significant differences in the ST and Reb A degradation rates were observed during oral, gastric, and small intestinal digestion. Carbohydrates containing an α-glycosidic linkage are reportedly hydrolyzed during digestion simulations21. Steviol glycoside α-glycosidic linkages were degraded by the digestive enzyme solution.
Reb Aα’s effect on HFD-induced obesity
In vitro examination revealed that Reb Aα’s α-glycosidic linkage is degradable under digestive conditions. If the α-glycosidic linkage is digested, the truncated glucose is presumably absorbed in the body. As excess glucose is potentially converted into lipids through de novo lipogenesis (DNL)22, individuals with obesity may be more susceptible to Reb Aα than lean individuals. Therefore, we investigated Reb Aα’s in vivo effect under diet-induced obesity (DIO) conditions.
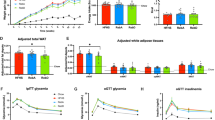
Mice were fed a HFD with either Reb A- or Reb Aα-supplemented water, or water alone (control), ad libitum for 16 weeks (Fig. 3A). Reb A and Reb Aα were supplied at 0.2 g/L, aligning with previous studies23. Reb A/Reb Aα administration had no effect on body weight (Fig. 3B); moreover, water intake and food consumption were also comparable across the groups (Fig. 3C). The estimated average daily intake of Reb A or Reb Aα, calculated based on the drinking volume, was approximately 0.675 mg per mouse. Considering that the no-observed-adverse-effect level (NOAEL) of Reb A in rats is 4100 mg/kg/day, the intake of 0.675 mg per mouse falls within a safe range24. Additionally, Reb A/Reb Aα did not increase gWAT weight (Fig. 3D). Although Reb Aα slightly decreased the number of small-sized adipocytes compared with the control and Reb A, Reb A significantly increased the crown-like structure (CLS), an inflammatory indicator in adipose tissue (Fig. 3E–G). These results suggest that Reb A, but not Reb Aα, exacerbates HFD-induced adipose tissue inflammation. We analyzed the protein levels related to lipogenesis (ACC and FAS) and lipolysis (p-HSL and p-PLIN)25. Interestingly, p-PLIN levels substantially decreased in the Reb A and Reb Aα groups compared with that in the control group (Fig. 3H). However, it should be noted that the number of large adipocytes and levels of serum cholesterol and TG remained unchanged following Reb A/Reb Aα administration (Fig. 3F, I). Collectively, Reb Aα, unlike Reb A, neither worsened HFD-induced obesity nor caused inflammation.
A Schematic description of the experiment. Analysis of mice after 16-week HFD feeding and treatment. B Body weights of the mice (n = 10 per group). C Water consumption and food intake volumes (n = 10 per group). D Gonadal white adipose tissue (gWAT) weight (n = 10 per group). E Hematoxylin and eosin (H&E) staining of gWAT (n = 5 per group). Representative images are shown. Scale bar = 100 μm. F Frequency of adipocyte size (n = 5 per group). G Percentage of crown-like structures (n = 5 per group). H Immunoblot of total gWAT lysates (left) and quantification of protein levels (right) (n = 4 per group). I Serum cholesterol and triglyceride levels (n = 5 per group). All values are presented as the mean ± SEM. Statistical analysis was conducted using repeated two-way analysis of variance (ANOVA) (B, F) and one-way ANOVA (C, D, G, H, I). Asterisks exhibit significant differences: *p < 0.05, ***p < 0.001, and ****p < 0.0001; N.S. not significant.
Reb Aα’s effect on HFD-induced steatosis and thermogenesis
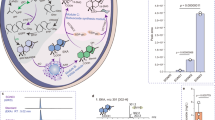
Individuals with obesity are more susceptible to diabetes owing to their impaired glucose metabolism26. The liver is the primary organ for glucose homeostatic control27, and obesity-induced insulin resistance compromises the inhibition of hepatic glucose production28. Moreover, excess glucose promotes hepatic lipid accumulation via DNL, potentially progressing to hepatic steatosis29. Therefore, we analyzed DIO mouse livers to determine whether Reb Aα administration disrupts glucose homeostasis and induces hepatic steatosis. To ascertain whether Reb Aα affects gluconeogenesis, we used qPCR to assess the expression of gluconeogenic markers, G6pc and Pck1, whose levels displayed no differences among the groups (Fig. 4A). Impaired glucose tolerance and fasting glucose differences remained unobserved among the groups (Fig. 4B, C). Nonetheless, feeding-state glucose levels tended to decrease in the Reb Aα group (Fig. 4C). These results suggest that Reb Aα consumption does not promote gluconeogenesis or glucose intolerance. Subsequently, we investigated Reb Aα’s impact on hepatic steatosis. HFD-induced hepatic steatosis is characterized by increased lipid accumulation and decreased hepatocytic OXPHOS30. Therefore, we analyzed lipogenic and OXPHOS-related gene and protein expressions. No significant gene-expression differences occurred among the groups (Fig. 4D). Conversely, the protein expression levels of ACC and FAS, both involved in lipogenesis, exhibited distinct alterations wherein Reb A increased ACC and Reb Aα decreased FAS (Fig. 4E). OXPHOS protein levels were comparable among the groups (Fig. 4E). Additionally, liver weight and histology did not differ among the groups (Fig. 4F, G). ALT and AST, liver impairment indicators, tended to decrease in the Reb Aα group, albeit not statistically significant (Fig. 4H). These results suggest that Reb Aα does not negatively affect hepatic glucose metabolism or HFD-induced steatosis.
A mRNA levels of G6pc (left) and Pck1 (right) in the liver (n = 6 per group). B Glucose tolerance test (left) and AUC (right) (n = 10 per group). C Fasting glucose level (left, n = 10 per group) and glucose level under feeding state (right, n = 5 per group). (D) mRNA levels of the indicated genes in the liver (n = 6 per group). E Immunoblot of total liver lysates (left) and quantification of protein levels (right) (n = 4 per group). F Liver weight (n = 10 per group). G Hematoxylin and eosin (H&E) staining of the liver (n = 5 per group). Representative images are shown. Scale bar = 100 μm. H Serum ALT and AST levels (n = 5 per group). I Brown adipose tissue (BAT) weight (n = 10 per group). J Hematoxylin and eosin (H&E) staining of BAT (n = 5 per group). Representative images are shown. Scale bar = 100 μm. K mRNA levels of the indicated genes in BAT (n = 6 per group). L Immunoblot of total BAT lysates (left) and quantification of protein levels (right) (n = 4 per group). All values are presented as the mean ± SEM. Statistical analysis was conducted using repeated two-way ANOVA (B [left]) and one-way ANOVA (A, B [right], C–L). Asterisks exhibit significant differences: *p < 0.05; N.S., not significant.
Energy homeostasis disruption potentially leads to obesity and diabetes, and BAT is the primary regulator of energy expenditure via non-shivering thermogenesis31. Therefore, we determined whether Reb Aα consumption influences BAT function. Tissue weight and histology did not differ among the groups (Fig. 4I, J). However, the expression of Fgf21, a thermogenic gene, and Cox5b, an OXPHOS-related gene, increased in the Reb Aα group’s BAT (Fig. 4K). Moreover, the protein expression of UCP1, which is pivotal for thermogenesis, significantly increased in the Reb Aα group compared with that in the control (Fig. 4L). These results indicate that Reb Aα, but not Reb A, may help activate adipocyte thermogenesis, although no body weight change was observed (Fig. 3B).
Gut microbiota composition following Reb Aα consumption
Reb Aα neither raised blood glucose levels nor induced glucose intolerance, despite releasing glucose. Therefore, we inferred that the hydrolysis of Reb Aα began under gastric conditions and was rapidly accelerated in the intestinal phase, suggesting that Reb Aα-derived glucose was not significantly absorbed, likely due to microbial utilization in the gut. Moreover, certain studies have reported that rebaudioside can negatively impact host health through gut microbiota alterations18,32,33,34,35,36. Therefore, we analyzed the fecal microbiota of Reb A-/Reb Aα-administered HFD mice over 14 weeks. Alpha diversity, as measured by species richness (observed feature, Chao1) and evenness (Shannon, Simpson) indices, exhibited no significant differences within groups (Fig. 5A–D). Furthermore, multiple comparison analysis revealed no statistical differences among the control, Reb A, and Reb Aα groups across all alpha diversity indices (Table S3). In beta diversity, principal coordinate analysis plots based on weighted and unweighted UniFrac distances indicated no clear separation among the groups in microbial community structure (Fig. 5E–F). Additionally, ANOSIM-based and multiple comparison analyses revealed no statistically significant differences within and among the groups, respectively (Table S4). These findings suggest that Reb Aα did not influence the gut microbiota.
Species richness was measured using A observed features and B Chao1 diversity indices. Species evenness was measured using C Shannon and D Simpson diversity indices. Data are presented as the mean ± SD (n = 4). Statistical significance for alpha diversity was calculated using the Kruskal–Wallis test. Principal coordinate analysis (PCoA) plots of different mouse groups based on weighted (E) and unweighted (F) UniFrac distance metrics. Beta diversity significance was assessed using ANOSIM, with p-values of 0.054 and 0.058 for unweighted and weighted UniFrac distances, respectively.
To identify each group’s gut microbiota composition, the relative abundance of taxa in Reb A-/Reb Aα-treated mice was determined using the RDP classifier. Bacillota and Pseudomonadota were the dominant phyla across all mouse groups (Fig. 6A), while 124 genera were identified from feces, with Ligilactobacillus, Lactobacillus, Clostridium sensu stricto, Faecalibaculum, Limosilactobacillus, and Allobaculum dominating all samples; moreover, the relative abundance of each taxon represented >1% of all sequences (Fig. 6B). These data demonstrate that Reb Aα supplementation in HFD mice does not induce any significant changes in gut microbial diversity, suggesting that Reb Aα consumption is safe for gut microbiota.
The bar plots show the relative abundance of each microbial taxon across mouse groups at the phylum (A) and genus (B) levels.
Discussion
Reb Aα was synthesized to address the limitations of conventional compounds (e.g., ST and Reb A). It was produced using ST and L. citreum KM20, and its molecular weight was identical to that of Reb A14. Compared with ST’s previously reported NMR spectra14, 1H- and 13C-NMR spectra revealed one additional glucose unit. Furthermore, two-dimensional NMR spectra confirmed that the additional glucose unit was attached to ST via a 1,6-α-glycosidic linkage.
Reb Aα (containing an α-glycosidic linkage) exhibited significantly improved water solubility compared with Reb A and ST (containing only β-glycosidic linkages), suggesting that water solubility depends on linkage type (alpha or beta). Flavonoids display a similar phenomenon. Flavonoids with α-glycosidic linkages are more soluble than those with β-glycosidic linkages because α-glycosidic linkages disrupt intramolecular hydrogen bonds and increase structural flexibility, thereby improving solubility. Reb Aα (containing an α-glycosidic linkage) exhibited improved solubility owing to its increased ability to disrupt intramolecular hydrogen linkages and more flexible structure compared with that of steviol glycosides (containing only β-glycosidic linkages)36.
FT-IR and XRD analyses demonstrated that Reb Aα and Reb A exhibited similar amorphous structures with low crystallinity and no significant differences. The additional glucose unit did not significantly alter the overall FT-IR and XRD patterns. However, subtle differences in the patterns indicated that the two compounds were structurally distinct. These subtle structural differences potentially influence their properties (e.g., stability and thermal properties). DSC thermograms confirmed that Reb Aα’s melting point was 184.83 °C, 19 °C lower than that of Reb A. Nevertheless, Reb Aα retained 2.71% more mass at elevated temperatures. These results suggest that Reb Aα is sufficiently stable for use in hot beverages or thermally processed foods.
Reb Aα demonstrated approximately 2.5-fold greater stability than Reb A at pH 1.4. Steviol glycosides contain ester and ether linkages, which undergo chemical degradation under acidic conditions37. However, Reb Aα’s α-glycosidic linkage conferred greater stability against hydrolysis under acidic conditions than Reb A’s ester and ether linkages. Hence, Reb Aα was expected to be more stable than Reb A in acidic beverages. Furthermore, Reb Aα exhibited superior long-term stability to Reb A over 1 month. Steviol glycosides with α-glycosidic linkages have already proven to exhibit excellent stability in aqueous solutions, and this study further corroborates these findings9,38, suggesting Reb Aα’s suitability for use in food products without significant degradation during long-term storage.
Reb Aα was hydrolyzed during in vitro digestion, as evidenced by the GGR and InfoGest protocols, confirming that its α-glycosidic linkage was hydrolyzed during digestion. Although Reb Aα was hydrolyzed, it was eventually converted to ST, recognized for its stable structure. Additionally, Reb Aα exhibits high water solubility, thermal stability, and pH stability, suggesting its potential as a sweetener. Therefore, systemically evaluating its impact on the body was imperative. Sugar substitutes are used not only by individuals striving to reduce caloric intake but also by those with diabetes. Therefore, we examined Reb Aα’s effect in HFD-induced obese mice. Because DIO mice exhibit obesity, insulin resistance, and diabetes, they constitute a suitable model for mimicking individuals requiring sugar substitutes. In this study, Reb Aα-administered DIO mice did not exhibit weight gain, glucose intolerance, or energy homeostasis disturbance. Moreover, they displayed reduced hepatic FAS levels and increased BAT UCP1 levels. Although we observed no significant improvements in steatosis or reductions in body weight, longer-term Reb Aα administration may improve metabolic health by inhibiting lipogenesis and activating thermogenesis. Notably, the Reb Aα group exhibited fewer CLSs than the Reb A group in gWAT. These results indicate that Reb Aα not only demonstrates safety but also possesses potential beneficial effects.
High doses of rebaudioside A have been associated with increased relative abundances of Lactobacilli in mice, as well as elevated levels of Bacteroides goldsteinii and Bacteroides thetaiotaomicron in rats32. However, in our study, 14-week Reb A/Reb Aα administration in HFD-fed mice did not significantly alter microbial diversity. This discrepancy is attributable to dietary variations. Unlike studies that administered a chow diet, we fed mice an HFD. HFD feeding significantly alters the microbiota profile39. Accordingly, in our study, microbial diversity remained consistent, possibly because of the HFD’s predominant influence, which might have attenuated the effects of Reb A/Reb Aα. Nonetheless, although partial Reb Aα degradation might have occurred, it might not have influenced its functional impact upon intake, indicating its retention of stability in terms of biological activity.
This study systematically compared the physicochemical properties, digestive behavior, and in vivo metabolic effects of Reb A and its glucosylated derivative, Reb Aα. Both compounds exhibited considerably similar physicochemical characteristics, including thermal behavior and structural features, as evidenced by FT-IR, DSC, and TGA analyses. Reb Aα demonstrated markedly enhanced water solubility and greater stability under acidic and long-term aqueous conditions, validating its suitability for multiple industrial applications. Although Reb Aα was partially hydrolyzed during digestion, its metabolic impact was negligible, exerting no adverse effects on weight gain, glucose metabolism, lipid accumulation, or gut microbiota composition under HFD conditions. These findings indicate that Reb A and Reb Aα possess comparable physicochemical profiles and can be considered safe for consumption, with Reb Aα offering additional advantages in solubility and stability.
Methods
Materials and enzymes
ST and Reb A (purity: >90%) were purchased from MSC Co., Ltd. (Yangsan, Korea). Sucrose was procured from Bio Basic Inc. (Ontario, Canada) and used as an enzyme reaction substrate. The solvents used in high-performance liquid chromatography (HPLC) were of HPLC grade, and those used in medium-pressure liquid chromatography (MPLC) were of extra-pure grade. The HPLC and MPLC solvents were acquired from Duksan General Science Co., Ltd. (Seoul, Korea). L. citreum KM20-derived recombinant dextransucrase was synthesized in Escherichia coli BL21 (DE3) according to previous study14.
Dextransucrase activity
Enzyme activity was measured at 30 °C in a 1 mL reaction mixture containing 400 mM sucrose, 40 mM sodium acetate buffer (pH 5.5), and crude L. citreum KM20-derived dextransucrase. After 15 min of reaction, the reaction was heat-inactivated at 100 °C for 5 min and centrifuged (8000 × g, 4 °C, 20 min) to obtain the supernatant. The amount of fructose produced from sucrose in the supernatant was analyzed using the d-Fructose Assay Kit (Megazyme, Bray, Ireland) and measured at 340 nm using an ultraviolet–visible (UV/Vis) spectrophotometer (Hidex, Turku, Finland). One unit (U) of dextransucrase activity was defined as the enzyme quantity releasing 1 μmol of fructose per min.
Enzymatic synthesis of Reb Aα
Reb Aα was enzymatically synthesized using a modified enzymatic method of previous sutdy14. The reaction mixture (1 L) containing 40 mM sodium acetate buffer (pH 5.5), 80 mM ST, 400 mM sucrose, and 5.1 U/mg of dextransucrase was incubated at 30°C and 180 rpm. Thereafter, twice the volume of ethanol was added and the resulting mixture stored at –80 °C for 1 h. Dextran was removed via centrifugation (8000 ×g, 10 min) and the supernatant evaporated under vacuum rotary concentration (Eylea N-1300, Eylea, Tokyo, Japan).
The sample was adsorbed onto Diaion HP-20 resin and washed with deionized water to remove sugar compounds, and the adsorbed compounds were eluted with ethanol. The eluate was concentrated and Reb Aα purified on an ODS-MPLC column (240 g; Biotage, Uppsala, Sweden) using a linear acetonitrile gradient (0–50%, v/v). Reb Aα synthesis was confirmed using HPLC. HPLC analysis was performed on an SCL-40 system (Shimadzu, Tokyo, Japan) equipped with a UV/vis detector (SPD-20A UV/vis), heater (CTO-40C), pump (LC-40D), and software (LabSolutions). For HPLC, an Agilent ZORBAX 300SB-C18 column (4.6 × 150 mm, 5 µM, Santa Clara, CA, USA) was used, and separation was performed under a linear acetonitrile gradient (0–50%, v/v) for over 35 min. The following parameters were employed: injection volume, 10 µL; flow rate, 1.0 mL/min; detection wavelength, 210 nm; and column temperature, 40 °C.
Physical and structural properties of Reb Aα, and Reb A
Nuclear magnetic resonance (NMR) analysis
Reb Aα (10 mg) was dissolved in pyridine-d5, transferred into 4.2-mm NMR tubes, and stored at 4 °C before analysis. NMR spectra were generated using a Unity INOVA 500 spectrometer (Varian, Palo Alto, CA, USA) at the Gwangju Center of the Korea Basic Science Institute. The associations between the steviol and sugar moieties were determined via correlation spectroscopy (COSY), heteronuclear single quantum coherence (HSQC), heteronuclear multiple bond correlation (HMBC), and total correlation spectroscopy (TOCSY).
Water solubility evaluation
ST, Reb A, and Reb Aα were each dissolved in deionized water until supersaturation and stirred for 24 h at 25 °C. The mixtures were subsequently centrifuged (8000 × g, 25 °C, 20 min) and the supernatants transferred to new conical tubes. Finally, the supernatants were freeze-dried at –80 °C and the dried residues weighed.
Fourier-transform infrared (FT-IR) spectroscopy analysis
FT-IR analyses of Reb A and Reb Aα were performed using a PerkinElmer Spectrum 400 spectrophotometer (Waltham, MA, USA). Spectral data were collected over a 400–4,000-cm⁻¹ wavenumber range, with sample measurements performed in triplicate to ensure reproducibility.
X-ray diffraction (XRD) analysis
The crystalline structures of Reb A and Reb Aα were analyzed using an X-ray diffractometer (50 kV, 50 mA, Cu-Kα radiation [λ = 0.154 nm], X’Pert PRO MPD; PANalytical, Malvern, Netherlands). XRD patterns were recorded over a 5−40° 2θ range at a scan 1°/min speed.
Thermal properties
Differential scanning calorimetry (DSC) analysis
DSC analysis was conducted using a Mettler Toledo DSC3 instrument (Greifensee, Switzerland). Reb A and Reb Aα samples (approximately 5 mg) were each placed in a hermetically sealed aluminum pan and heated from 30 °C to 300 °C at a 10 °C/min rate in a nitrogen atmosphere.
Thermogravimetric analysis (TGA)
The thermal decomposition profiles of Reb A and Reb Aα were determined using a TGA2 analyzer (Mettler Toledo, Zurich, Switzerland). Each sample’s temperature was increased from 20 °C to 600 °C at 10 °C/min under nitrogen flow.
pH stability
pH stability was analyzed using Britton–Robinson buffer (pH range: 1.4–10), with which Reb A and Reb Aα were mixed to make 0.05% (w/v) aqueous solutions. Thereafter, stability was evaluated using HPLC.
Storage stability in aqueous solution
Reb A and Reb Aα aqueous solutions (1–2%, w/v) were prepared. The samples were incubated at 25 °C for up to 1 month. The sample amount remaining in the supernatant after 0, 7, 15, and 30 days was quantified using HPLC.
Glucose generation rate (GGR) measurement
Rat intestinal acetone powder (RIAP) was purchased from Sigma-Aldrich (St. Louis, MO, USA) and used as a mammalian mucosal α-glucosidase. The GGRs of ST, Reb A, and Reb Aα using mammalian mucosal α-glucosidases were assayed as previously described37. Briefly, RIAP (0.5 g) was mixed with 10 mL of 100 mM sodium phosphate buffer (pH 6.2) and incubated at 4 °C for 24 h. The mixture was subsequently centrifuged at 8000 × g for 20 min and the resulting supernatant was used as α-glucosidase solution. The α-glucosidase solution was freeze-dried at –80 °C until use. Before use, the freeze-dried α-glucosidase solution was mixed with 10 mL of 100 mM sodium phosphate buffer. ST, Reb A, and Reb Aα (5 mg/mL, w/v) and the mixed α-glucosidase solution were used to calculate the GGRs. Samples were aliquoted at 0, 10, 20, 30, 60, and 120 min and inactivated using a heating block for 10 min. The amount of glucose released was calculated using a glucose assay kit (Megazyme, Ireland).
Reb Aα measurement during in vitro digestion
Simulated salivary, gastric, and intestinal fluids were prepared according to a previously described method, with slight modifications38. The formulations were as follows: simulated saliva (pH 7.0; 1.5 mM CaCl₂ + 75 U/mL salivary amylase), simulated gastric juice (SGJ, pH 2.0; 0.15 mM CaCl₂ + 2000 U/mL pepsin + 60 U/mL lipase), and simulated intestinal juice (SIJ, pH 7.0; 10 mM bile salts + 0.6 mM CaCl₂ + 100 U/mL pancreatin + 25 U/mL pancreatic α-amylase).
For static in vitro digestion, Reb A/Reb Aα (50 mg) was placed in a 100-mL flask and incubated at 37 °C with continuous stirring through successive oral, gastric, and intestinal phases. The oral phase was initiated by adding simulated saliva, with samples withdrawn at 5 min, followed by the gastric phase wherein SGJ (10 mL) was added, with aliquots collected after 5 and 120 min. The intestinal phase involved SIJ (10 mL) addition, with sampling at 240 min. Collected aliquots were immediately heated at 100 °C for 5 min to terminate enzymatic activity. The samples were subsequently centrifuged at 6238 × g for 10 min and the resulting supernatants filtered using a 0.22-μm membrane. Glucose release was quantified using a glucose assay kit (Megazyme, Wicklow, Ireland).
Mice
Mouse experiments were conducted in accordance with national and institutional guidelines for the care and use of laboratory animals. The study protocol was approved by the Institutional Animal Care and Use Committee (IACUC) of [Chonnam National University], under protocol number [Approval No. CNU IACUC-YB-2024-217].
Eight-week-old C57BL/6 N mice were purchased from Damul Science (Deajeon, Korea) and housed in a stable environment (room temperature, 22–24 °C; humidity, 45–50%; and a 12/12-h light/dark cycle). The mice received a high-fat diet (HFD) containing 60% kcal fat (D12492; Research Diets, NS, USA) for 16 weeks. The experimental groups (Control, Reb A, and Reb Aα) were provided normal drinking water ad libitum as a control or with Reb A (0.2 g/L) and Reb Aα (0.2 g/L) aqueous solutions, respectively. Body weight was recorded weekly, and food and water intakes were measured every other day. Finally, the mice were euthanized using carbon dioxide at a fill rate of 50% of the chamber volume per minute, in accordance with the NIH ARAC guidelines.
Histological analysis
Adipose tissue and liver samples were obtained from individuals with the median body weight in each group. After overnight fixation in 10% neutral-buffered formalin, the samples were embedded in paraffin wax. Tissue sections were prepared on slides with a thickness of 7 μm for gonadal white adipose tissue (gWAT) and 5 μm for liver and interscapular brown adipose tissue (iBAT). The slides underwent 5-min deparaffinization in xylene three times and 2-min rehydration by immersion in a graded ethanol series (100%, 95%, 90%, 80%, 70%, and distilled water). Afterward, they underwent staining; for H&E staining, they were incubated with Harris hematoxylin (#3550; BBC Biochemical, Mount Vernon, WA, USA) for 6 min and subsequently incubated with Eosin Y Alcoholic (#3610; BBC Biochemical) for 2 min after bluing. Thereafter, the slides were dehydrated by immersion in a graded ethanol series (70%, 80%, 90%, 95%, and 100%) three times. They were mounted with xylene-based mounting medium (9990440; Thermo Fisher Scientific, Waltham, MA, USA) after clearing. All images were acquired using a Nikon ECLIPSE Ni-U microscope (Nikon Co., Tokyo, Japan). Adipocyte size was analyzed using the Adiposoft plug-in of Image J software (National Institutes of Health [NIH], Bethesda, MC, USA). For each group, approximately 3,000 cells were counted using four images per mouse from a total of five mice.
Immunoblot analysis
Tissue samples from individual mice with the median body weight per group were prepared and subjected to sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE), as previously described39. An equal amount of protein was loaded onto SDS-PAGE gels and subsequently transferred to nitrocellulose (NC) membranes (#XO-P-NC; XOGENE Co., Ltd., Seoul, Korea). To block non-specific binding, the NC membranes were incubated in 5% skim milk for 30–60 min after transfer. Briefly washed membranes with Tris-buffered saline with Tween 20 were incubated with primary antibodies overnight at 4 °C. Afterward, the NC membranes were rocked at room temperature for 2–3 h with secondary antibodies. To detect proteins, enhanced chemiluminescence solutions (#EOE001S; Enzynomics, Daejeon, Korea; HY-K1005; MedChemExpress, Suite Q Monmouth Junction, NJ, USA) were used. This study utilized antibodies against fatty acid synthase (FAS; ab22759), oxidative phosphorylation (OXPHOS; ab110413), and uncoupling protein 1 (UCP1; ab10983) from Abcam; acetyl-CoA carboxylase (ACC; 3676), phosphorylated hormone-sensitive lipase (p-HSL; 4139), and heat shock protein 90 (HSP90; 4874) from Cell Signaling Technology; and phosphorylated perilipin (p-PLIN; #4854) from Vala Sciences. Immunoreactive bands were visualized using a luminescent image analyzer (ImageQuant™ LAS 4000 mini; GE HealthCare, Chicago, IL, USA). ImageJ (NIH) was used to quantify immunoblot band intensity.
Quantitative polymerase chain reaction (qPCR) analysis
Following the standard protocol, each tissue sample’s RNA was isolated using TRI-Solution (TS200-001; Bio Science Technology, Daegu, Korea). A NanoDrop spectrophotometer was used to measure the final RNA concentration. A 2.5-µg mRNA sample was reverse-transcribed into cDNA using Moloney murine leukemia virus reverse transcriptase (28025021; Thermo Fisher Scientific) following the manufacturer’s protocol. qPCR was conducted using SYBR Green (PB20.17-51; PCR Biosystems, London, UK) on a QuantStudio 6 Flex 384-well qPCR System (Thermo Fisher Scientific), with the following thermal cycling conditions: 95 °C for 3 min, followed by 40 cycles at 95 °C for 10 s and 60 °C for 30 s. As an internal control, TATA-box binding protein mRNA was used. The primer sequences are provided in Table S1.
Serum analysis
Whole blood was obtained via cardiac puncture after mouse euthanization. Mouse serum samples were analyzed using an auto chemistry analyzer (Dotto 2000; MTD Diagnostics Srl, Italy) to quantify cholesterol, triglyceride (TG), alanine aminotransferase (ALT), aspartate aminotransferase (AST), and glucose serum levels.
Glucose tolerance test
At treatment week 13, mice underwent 16-h fasting before receiving a 20% filtered glucose solution intraperitoneal injection (2 g/kg body weight). Blood glucose levels were measured 0, 30, 60, and 120 min using an Accu-Check glucometer (Performa; Roche, Basel, Switzerland). Blood was sampled by pricking the distal tail vein with a syringe needle.
Gut microbiota analysis
Total DNA was extracted from mouse feces using the QIAamp Fast DNA Stool Mini Kit (Qiagen, Hilden, Germany) following the manufacturer’s instructions. To prepare the 16S rRNA amplicon, a universal primer set targeting the V3–V4 regions (Bakt 341 F: 5′-CCTACGGGNGGCWGCAG-3′, Bakt 805 R: 5′-GACTACHVGGGTATCTAATCC-3′) was used. The PCR conditions were as follows: one cycle at 98 °C for 3 min, followed by 30 cycles of 98 °C for 10 s, 57 °C for 5 s, and 68 °C for 1 s, with a final extension at 72 °C for 5 min. After the first PCR cycle, the amplicons were purified using AMPure beads (Agencourt Bioscience, Beverly, MA). Following purification, the first PCR product (10 μL) was amplified for 10 cycles using NexteraXT Indexed Primers, purified with AMPure beads, quantified via qPCR, and qualified using TapeStation D1000 ScreenTape. Thereafter, the amplicons were sequenced using the MiSeq platform (Illumina, San Diego, CA, USA). Raw data were processed using QIIME2 (version 2024.5) for demultiplexing and primer sequence removal with Cutadapt, followed by denoising using the DADA2 algorithm. Filtered sequences were used to construct a phylogenetic tree with FastTree, and taxonomic assignment was performed using the naïve Bayesian classifier trained on the Ribosomal Database Project (RDP) database (version 2.14). Microbial species diversity and evenness were evaluated using the Shannon and Inverse Simpson indices. Alpha diversity was assessed via rarefaction curves and Chao1 values, while beta diversity, based on weighted and unweighted UniFrac distances, was analyzed to identify variations between comparative groups. Using the microbiomeMarker package (version 1.2.1) in R (version 4.0.1), “linear discriminant analysis effect size” analysis was conducted to identify biomarkers with significant differential abundance across groups.
Statistical analysis
Data were obtained from biologically independent samples and are presented as the mean ± standard error of the mean. One-way analysis of variance (ANOVA), followed by Dunnett’s or Tukey’s post-hoc test, and/or two-way ANOVA, with Bonferroni’s post-hoc test for repeated measurements over time, were used to evaluate statistical differences between groups. GraphPad Prism (version 9.3.1; GraphPad Software, San Diego, CA, USA) was used for statistical analysis and data visualization.
Data availability
No datasets were generated or analysed during the current study.
References
Malik, V. S., Popkin, B. M., Bray, G. A., Després, J. P. & Hu, F. B. Sugar-sweetened beverages, obesity, type 2 diabetes mellitus, and cardiovascular disease risk. Circulation 121, 1356–1364 (2010).
Hlyan, N. P. et al. From sugar spikes to pressure peaks: Navigating the world of diabetes, hypertension, obesity, and kidney health. Cureus 16, e57241 (2024).
Castro-Muñoz, R. et al. Natural sweeteners: Sources, extraction and current uses in foods and food industries. Food Chem. 370, 130991 (2022).
Rafiq, M. T., Sajid, Z. A. & Khilji, S. A. Graphene oxide nanoparticle-assisted promotion of stevioside, rebaudioside A, and selected biochemical attributes in Stevia rebaudiana Bertoni. Science 2024, 1–10 (2024).
Yang, Y., Xu, M., Wan, Z. & Yang, X. Novel functional properties and applications of steviol glycosides in foods. Curr. Opin. Food Sci. 43, 91–98 (2022).
Libik-Konieczny, M., Capecka, E., Tuleja, M. & Konieczny, R. Synthesis and production of steviol glycosides: recent research trends and perspectives. Appl. Microbiol. Biotechnol. 105, 3883–3900 (2021).
Zhang, R. et al. Efficient bioconversion of stevioside and rebaudioside A to glucosylated steviol glycosides using an Alkalihalobacillus oshimesis-derived cyclodextrin glucanotransferase. Molecules 28, 1245 (2023).
Demuth, K., Jö, H.-J. & Buchholz, K. Oligosaccharide synthesis by dextransucrase: new unconventional acceptors. Carbohydr. Res. 337, 1811–1820 (2002).
Lee, S. H. et al. Enzymatic synthesis of glucosyl rebaudioside A and its characterization as a sweetener. J. Food Sci. 84, 3186–3193 (2019).
Ji, Y. et al. Advances on the in vivo and in vitro glycosylations of flavonoids. Appl. Microbiol. Biot. 104, 6587–6600 (2020).
Jiang, J. R. et al. Conversion of puerarin into its 7-O-glycoside derivatives by Microbacterium oxydans (CGMCC1788) to improve its water solubility and pharmacokinetic properties. Appl. Microbiol. Biot. 81, 647–657 (2008).
Aquilina et al. Safety evaluation of glucosylated steviol glycosides as a food additive in different food categories. EFSA J. 20, e7066 (2022).
Jaitak, V., Bandna, Singh, B. & Kaul, V. K. An efficient microwave-assisted extraction process of stevioside and rebaudioside-A from Stevia rebaudiana (Bertoni). Phytochem. Anal. 20, 240–245 (2009).
Ko, J. A. et al. Synthesis and characterization of glucosyl stevioside using Leuconostoc dextransucrase. Food Chem. 211, 577–582 (2016).
Hou, Y. et al. Novel self-nanomicellizing solid dispersion based on rebaudioside A: A potential nanoplatform for oral delivery of curcumin. Int. J. Nanomed. 14, 557–571 (2019).
Upreti, M., Strassburger, K., Chen, Y. L., Wu, S. & Prakash, I. Solubility enhancement of steviol glycosides and characterization of their inclusion complexes with gamma-cyclodextrin. Int J. Mol. Sci. 12, 7529–7553 (2011).
Asadzadeh, B., Zhong, J. & Yan, W. Solid-liquid equilibrium of rebaudioside A in pure and binary mixed solvents at T = (288.15 to 328.15). K. J. Chem. Eng. Data 63, 4269–4276 (2018).
Kang, H. J. et al. Synthesis and characteristics of a rebaudioside-A like compound as a potential non-caloric natural sweetener by Leuconostoc kimchii dextransucrase. Food Chem. 366, 130623 (2022).
Gardana, C., Simonetti, P., Canzi, E., Zanchi, R. & Pietta, P. Metabolism of stevioside and rebaudioside A from Stevia rebaudiana extracts by human microflora. J. Agric. Food Chem. 51, 6618–6622 (2003).
Brodkorb, A. et al. INFOGEST static in vitro simulation of gastrointestinal food digestion. Nat. Protoc. 14, 991–1014 (2019).
Torp Nielsen, M., Roman, L. & Corredig, M. In vitro gastric digestion of polysaccharides in mixed dispersions: Evaluating the contribution of human salivary α-amylase on starch molecular breakdown. Curr. Res. Food Sci. 8, 100759 (2024).
Kersten, S. Mechanisms of nutritional and hormonal regulation of lipogenesis. EMBO Rep. 2, 282–286 (2001).
Park, S. et al. Maternal low-calorie sweetener consumption rewires hypothalamic melanocortin circuits via a gut microbial co-metabolite pathway. JCI Insight 8, e156397 (2023).
Curry, L. L. & Roberts, A. Subchronic toxicity of rebaudioside A. Food Chem. Toxicol. 46, S11–S20 (2008).
Wong, R. H. F. & Sul, H. S. Insulin signaling in fatty acid and fat synthesis: a transcriptional perspective. Curr. Opin. Pharmacol. 10, 684–691 (2010).
Klein, S., Gastaldelli, A., Yki-Järvinen, H. & Scherer, P. E. Why does obesity cause diabetes?. Cell Metab. 34, 11–20 (2022).
Han, H. S., Kang, G., Kim, J. S., Choi, B. H. & Koo, S. H. Regulation of glucose metabolism from a liver-centric perspective. Exp. Mol. Med. 48, e218 (2016).
Leclercq, I. A., Da Silva Morais, A., Schroyen, B., Van Hul, N. & Geerts, A. Insulin resistance in hepatocytes and sinusoidal liver cells: mechanisms and consequences. J. Hepatol. 47, 142–156 (2007).
Sanders, F. W. B. & Griffin, J. L. De novo lipogenesis in the liver in health and disease: More than just a shunting yard for glucose. Biol. Rev. 91, 452–468 (2016).
Tarantino, G., Savastano, S. & Colao, A. Hepatic steatosis, low-grade chronic inflammation and hormone/growth factor/adipokine imbalance. World J. Gastroenterol. 16, 4773–4783 (2010).
Jung, S. M., Sanchez-Gurmaches, J. & Guertin, D. A. Brown adipose tissue development and metabolism. Handb. Exp. Pharmacol. 251, 3–36 (2019).
Nettleton, J. E. et al. Low-dose stevia (Rebaudioside A) consumption perturbs gut microbiota and the mesolimbic dopamine reward system. Nutrients 11, 1248 (2019).
Ramos-García, M. et al. Gut microbiota response to sucralose or rebaudioside A in rats under two dietary conditions. Res. Square. 1, 1–18 (2023).
Roy, J. K. et al. Production of highly water-soluble genistein α-diglucoside using an engineered O-α-glycoligase with enhanced transglycosylation activity and altered substrate specificity. Food Chem. 437, 137898 (2024).
Kroyer, G. Stevioside and stevia-sweetener in food: Application, stability and interaction with food ingredients. J. Consum. Prot. Food Saf. 5, 225–229 (2010).
Murphy, E. A., Velazquez, K. T. & Herbert, K. M. Influence of high-fat diet on gut microbiota: A driving force for chronic disease risk. Curr. Opin. Clin. Nutr. Metab. Care. 18, 515–520 (2015).
Lee, B. H. & Hamaker, B. R. Number of branch points in α-limit dextrins impact glucose generation rates by mammalian mucosal α-glucosidases. Carbohydr. Polym. 157, 207–213 (2017).
Hong, S. J. et al. A comparative study on the characteristics and properties of cyclodextrans, cyclodextrins, and their complexes with biologically active compounds. Food Hydrocoll. 168, 111587 (2025).
Park, M. J., Lee, C. M. & Kim, D. I. An easy method for the clear detection of beige fat UCP1 by Western blotting. Adipocyte 8, 357–361 (2019).
Acknowledgements
This work was supported by the National Research Foundation (NRF), Korea, under project BK21 FOUR, Graduate Program for Integrative Food, Bioscience and Biotechnology (No. 2024-1044-02). This research was supported by the Bio-Industrial Strategic Technology Development Program (No. 20026226) funded by the Ministry of Trade, Industry & Energy (MOTIE, Korea).
Author information
Authors and Affiliations
Contributions
Hye-Jin Kim: Writing – original draft, Data curation. Hye-Jin Kang: Writing – original draft, Investigation, Validation. Sangyi Lim: Writing – original draft, Investigation, Validation. Ok Cheol Kim: Methodology, Validation. Ki-Nam Yoon: Methodology, Validation. Junhyeong Lee: Methodology, Validation. Merc Emil Matienzo: Investigation. Min-Jung Park: Formal analysis, Conceptualization. Bu-Soo Park: Conceptualization, Supervision. Jeong-Yong Cho: Supervision, Data curation. Seong-Jin Hong: Writing – review & editing, Writing – original draft. Dong-il Kim: Writing – review & editing, Writing – original draft, Supervision. Young-Min Kim: Writing – review & editing, Writing – original draft, Project administration, Conceptualization.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kim, HJ., Kang, HJ., Lim, S. et al. Comparative study of rebaudioside A and rebaudioside Aα: physicochemical properties and in vivo metabolism. npj Sci Food 9, 225 (2025). https://doi.org/10.1038/s41538-025-00585-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41538-025-00585-7