Abstract
Bronchiolitis is the leading cause of infant hospitalization. However, the molecular networks driving bronchiolitis pathobiology remain unknown. Integrative molecular networks, including the transcriptome and metabolome, can identify functional and regulatory pathways contributing to disease severity. Here, we integrated nasopharyngeal transcriptome and metabolome data of 397 infants hospitalized with bronchiolitis in a 17-center prospective cohort study. Using an explainable deep network model, we identified an omics-cluster comprising 401 transcripts and 38 metabolites that distinguishes bronchiolitis severity (test-set AUC, 0.828). This omics-cluster derived a molecular network, where innate immunity-related metabolites (e.g., ceramides) centralized and were characterized by toll-like receptor (TLR) and NF-κB signaling pathways (both FDR < 0.001). The network analyses identified eight modules and 50 existing drug candidates for repurposing, including prostaglandin I2 analogs (e.g., iloprost), which promote anti-inflammatory effects through TLR signaling. Our approach facilitates not only the identification of molecular networks underlying infant bronchiolitis but the development of pioneering treatment strategies.
Similar content being viewed by others
Introduction
Bronchiolitis is the most common lower respiratory infection in infants1. It is the leading cause of infant hospitalization in the U.S., accounting for 110,000 hospitalizations each year2. The intensive care unit use and the hospitalization costs for bronchiolitis increased substantially over the last decade3,4. However, the management of bronchiolitis is predominantly supportive, with no specific effective therapies available5. To develop treatment strategies for bronchiolitis, it is crucial to identify the molecular network driving bronchiolitis pathobiology.
Emerging research has partially elucidated the molecular mechanisms underlying infant bronchiolitis by leveraging omics data. For example, transcriptomics6,7,8,9,10,11,12 (profiling RNA expressions to uncover cellular gene regulation pathways) and metabolomics12,13,14,15,16,17,18,19,20 (characterizing metabolite abundances to reveal biochemical pathways) along with their integrated approaches12,17,18, have offered an elemental view of the molecular landscape in infant bronchiolitis. Yet, these findings have not accounted for intra-omics (i.e., within the transcriptome or metabolome) or inter-omics (i.e., between the transcriptome and metabolome) interactions, which are instrumental to accurately representing human pathobiology, inherently shaped by multi-omics network interactions. However, existing analytic methods have been unable to break through this barrier due to the limitation in representing the omics network interactions, and this has been considered a significant problem in omics analyses21, let alone in the study of infant bronchiolitis.
To address this knowledge gap, we employed a new analytical approach using explainable deep networks with image-converted integrative omics components. This approach enables us to consider intra- and inter-omics interactions and to identify the molecular networks driving the disease pathobiology. We applied this new approach to data from a multicenter prospective cohort study of infants with bronchiolitis using the nasopharyngeal transcriptome and metabolome profiles, and sought to identify the molecular networks driving bronchiolitis pathobiology, as well as the existing drug candidates for repurposing in infant bronchiolitis.
Results
Study design
Of the 1016 infants hospitalized with bronchiolitis and enrolled in the MARC-35 cohort, this study specifically focused on 397 infants (i.e., the analytic cohort) who underwent both nasopharyngeal transcriptome and metabolome profiling. There were no significant differences in the patient characteristics between the analytic and non-analytic cohorts (p ≥ 0.05; Supplementary Table 1). Among the analytic cohort, the median age was 3 months, 42% were female, 40% were non-Hispanic white, 23% were non-Hispanic black, and 33% were Hispanics. Overall, 20% (n = 79) were more-severe cases of bronchiolitis with respiratory support use during the index hospitalization (Table 1).
The explainable deep network using image-converted integrative omics components identifies severity-discriminatory mRNAs and metabolites
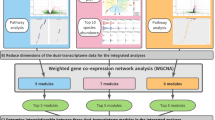
The overall analytic workflow is presented in Fig. 1. We first selected 588 mRNAs and 128 metabolites that were related (p < 0.05) to acute severity (i.e., respiratory support use) of infant bronchiolitis. We also presented the p-value distributions for the mRNAs and metabolites in Supplementary Fig. 1. Based on each distribution, we adopted a p < 0.05 threshold for mRNAs and metabolites to secure sufficient components for the following image conversion process. Next, we created variables for interaction by multiplying mRNA expression with metabolite abundance and retained 4835 variables for interaction related (FDR < 0.01) to the acute severity. Lastly, we prepared the severity-related components of each individual infant, comprising 588 mRNAs, 128 metabolites, and 4835 variables for interactions (Fig. 1A).
A The current analysis investigated 397 infants hospitalized with bronchiolitis who underwent both nasopharyngeal transcriptome and metabolome profiling. From all measured mRNAs and metabolites, we first selected 588 mRNAs and 128 metabolites that were related (p < 0.05) to acute severity (i.e., respiratory support use) of infant bronchiolitis. Next, we created variables for interaction by multiplying mRNA expression with metabolite abundance and retained 4835 variables for interaction related (FDR < 0.01) to the acute severity. Lastly, we prepared the severity-related components of each individual infant, comprising 588 mRNAs, 128 metabolites, and 4835 variables for interactions. B We used the DeepInsight method to convert the severity-related components of each infant into a single image. Within the image, the components represented by green pixels are mRNAs, those by red pixels are metabolites, and those by blue pixels are variables for interactions. The intensity of each pixel corresponds to the expression or abundance of each component. C We developed a convolutional neural network (CNN) that classifies the acute severity of bronchiolitis from individual images. The CNN was developed using 397 integrative omics images. We employed class activation mapping to identify the specific regions within the images representing the omics-cluster that discriminates the acute severity. We further demonstrated that the identified region comprises 401 mRNAs and 38 metabolites. D We performed a pathway analysis and network analysis using the severity-discriminatory 401 mRNAs and 38 metabolites. Additionally, we conducted a drug repurposing analysis using the 401 mRNAs to determine bronchiolitis drug candidates for repurposing. RNA ribonucleic acid, CNN convolutional neural network.
To represent the proximities and quantities of integrative omics components (i.e., mRNA, metabolites, and variables for interactions) in a single infant, we generated an integrative omics image for each infant (Fig. 1B). Within each image, the components represented by green pixels are mRNAs, those by red pixels are metabolites, and those by blue pixels are variables for interactions. The intensity of each pixel corresponds to the expression or abundance of each component (Fig. 2A, B). For the sensitivity analysis, we also prepared integrative omics images using 5983 variables for interactions—selected using a model that includes the main effects—instead of 4835 variables for interactions (Supplementary Fig. 2).
A To represent the proximities and quantities of integrative omics components (i.e., mRNA, metabolites, and their interactions) in a single infant, we generated an integrative omics image for each infant using the DeepInsight method. The intensity of each pixel corresponds to the expression or abundance of each omics component. The presented image illustrates the reference image wherein the intensities of all pixels are maximized to enhance the clarity of visual representation. B The upper panels present four integrative omics images extracted from infants with respiratory support use (i.e., severe cases; n = 79). The lower panels demonstrate four images extracted from infants without respiratory support use (i.e., less severe cases; n = 318). C To identify the omics-cluster that discriminates the acute severity of infant bronchiolitis, we applied class activation mapping to the convolutional neural network (CNN) model. The CNN model was trained with all integrative omics images (n = 397) to predict the acute severity. The resulting class activation mapping visualizes the importance scores, indicating the importance of each pixel to the prediction. The importance scores progressively increase from blue to yellow and then to red. RNA ribonucleic acid.
To identify the omics-cluster that discriminates the acute severity of infant bronchiolitis, we applied class activation mapping to the convolutional neural network (CNN) model. The CNN model was developed with all integrative omics images (n = 397) to predict the acute severity. The area under the receiver operating characteristic curve (AUROC) of the CNN model for the testing images was 0.828. We also confirmed the lower AUROC of the CNN model for the testing images in the sensitivity analysis using 5983 variables for interactions—selected using a model that includes the main effects—with the AUROC being 0.695. We employed class activation mapping to identify the specific regions within the images representing the omics-cluster that discriminates the acute severity (Fig. 2C). We further demonstrated that the omics-cluster in the identified region comprises 401 mRNAs and 38 metabolites, which we used as the severity-discriminatory components in the subsequent analyses (Fig. 1C, D)
The joint-pathway analysis reveals that the molecular networks of the severity-discriminatory components are characterized by inflammatory response pathways
In the functional enrichment joint-pathway analysis with the severity-discriminatory components (i.e., 401mRNAs and 38 metabolites), we identified 62 enriched Kyoto encyclopedia of genes and genomes (KEGG) pathways characterized by inflammatory response pathways, such as toll-like receptor (TLR) signaling pathway, nucleotide-binding oligomerization domain (NOD)-like receptor signaling pathway, nuclear factor kappa-light-chain-enhancer of activated B cells (NF-κB) signaling pathway, interleukin (IL)-17 signaling pathway, and phosphatidylinositol 3-kinase (PI3K)-protein kinase B (Akt) signaling pathway (all FDR < 0.01; Fig. 3A). In the subsequent joint-pathway analysis using RSV- or RV-related severity-discriminatory components—comprising 138 mRNAs and 15 metabolites for RSV and 179 mRNAs and 5 metabolites for RV—we observed both concordance (e.g., NOD-like receptor signaling pathway) and heterogeneity in the enriched KEGG pathways for each virus (Supplementary Fig. 3).
A The top 25 biological pathways according to the false discovery rate (FDR) value of each pathway in the functional enrichment joint-pathway analysis with Kyoto encyclopedia of genes and genomes (KEGG) terms using the severity-discriminatory 401 mRNAs and 38 metabolites. The horizontal axis represents the negative log10-transformed FDR. Plot color represents the percentage of hit components (mRNAs and metabolites) in each pathway. Plot size represents the impact of each pathway (i.e., pathway impact). This impact is computed by first determining the centrality of each component within a given pathway and then dividing the sum of these centrality measures by the total centrality measures for all components within the corresponding pathway. B The visualization of a pathway-based network generated from the severity-discriminatory 401 mRNAs and 38 metabolites. The dark green squares and red hexagons correspond to the severity-discriminatory mRNAs and metabolites identified in the analysis. The light green squares and pink hexagons represent the mRNAs and metabolites, derived from the KEGG and Edinburgh human metabolic network (EHMN) databases, that are adjacent to (i.e., surrounding) the severity-discriminatory mRNAs and metabolites. The size of the squares and hexagons represents the magnitude of the betweenness centrality of each node. The color of each edge represents the type of pathway related to the connected nodes, while the thickness of each edge indicates the magnitude of the edge betweenness of each edge. HIF-1 hypoxia-inducible factor 1, IL-17 interleukin 17, KEGG Kyoto encyclopedia of genes and genomes, NF-kappa B nuclear factor kappa B, NOD nucleotide-binding oligomerization domain, PI3K-Akt phosphatidylinositol 3-kinase / protein kinase B, RNA ribonucleic acid, TNF tumor necrosis factor.
The pathway-based network analysis demonstrates innate immunity-related metabolites centralized in a molecular network of the severity-discriminatory components
Through the pathway-based network analysis with the severity-discriminatory components, we identified 21 molecular networks. These networks comprised the severity-discriminatory (i.e., identified) mRNAs and metabolites, as well as the surrounding mRNAs and metabolites that are adjacent to the identified mRNAs and metabolites within pre-existing molecular networks derived from KEGG and Edinburgh human metabolic network (EHMN) databases. The largest molecular network identified in the analysis was centralized with the innate immunity-related metabolites (e.g., N-acylsphingosine, phosphatidylethanolamine) and involved in inflammatory and immune lipid metabolism pathways (e.g., glycosphingolipid metabolism, phosphatidylinositol phosphate metabolism; Fig. 3B). The remaining 20 molecular networks are presented in the supplementary material (Supplementary Fig. 4)
The drug-based network analysis identified eight sub-networks including 50 existing drug candidates for repurposing
Through the drug-based network analysis using the 401 severity-discriminatory mRNAs, we identified eight gene- and drug-based networks. In the network involving prostaglandin E receptor 2 (PTGER2), we confirmed six prostaglandin analogs or agonists (e.g., iloprost), three prostaglandin inhibitors, and one antibiotic drug (Fig. 4A). In another network involving Lyn tyrosine kinase, we confirmed 18 tyrosine kinase inhibitors (e.g., dasatinib, imatinib) and seven other types of drugs (Fig. 4B). The remaining five gene- and drug-based networks are presented in the supplementary material (Supplementary Fig. 5). In the subsequent network analysis using RSV- or RV-related severity-discriminatory mRNAs, tyrosine kinase inhibitors were included in the network derived from either the RSV- or RV-related mRNAs, although prostaglandin analogs or agonists were included only in the network derived from the RV-related mRNAs (Supplementary Fig. 6).
The visualization of gene- and drug-based networks generated from the severity-discriminatory 401 mRNAs and rankings of drug candidates for repurposing. In the network, the size of the squares represents the magnitude of the closeness centrality of each node. The orange, light blue, or purple squares represent drug candidates, while the green squares represent mRNAs. The thickness of each edge indicates the magnitude of the edge betweenness of each edge. Drug rankings were derived by employing the TrustRank algorithm on each network. In the rankings, the horizontal axis represents the log10-transformed Trustworthiness score calculated with the TrustRank algorithm. Plot size represents the magnitude of the closeness centrality of each node, while plot color represents the magnitude of the betweenness centrality of each node. A Drug network including prostaglandin E receptor 2. The orange squares represent prostaglandin analogues or agonists, the light blue squares represent prostaglandin inhibitors, and the purple squares represent other drug candidates. B Drug network including Lyn tyrosine kinase. The orange squares represent tyrosine kinase inhibitors, while the purple squares represent other drug candidates. RNA ribonucleic acid.
Discussion
By applying explainable deep networks using image-converted integrative omics components derived from a multi-center prospective cohort study of 397 infants with bronchiolitis, we identified an omics-cluster comprising 401 mRNAs and 38 metabolites that distinguishes bronchiolitis severity. These severity-discriminatory components derive molecular networks driving bronchiolitis pathobiology, which were characterized by inflammatory response pathways (e.g., TLR signaling pathway, NF-κB signaling pathway) and were centralized with innate immunity-related metabolites (e.g., N-acylsphingosine, phosphatidylethanolamine). We also identified eight sub-networks including existing 50 drug candidates for repurposing (e.g., iloprost, sunitinib) through the drug-based network analysis using the 401 severity-discriminatory mRNAs. To the best of our knowledge, this is the earliest investigation that has identified the molecular networks driving bronchiolitis severity by addressing intra- and inter-omics interactions of multi-omics datasets.
Recent studies demonstrated the molecular landscape underlying infant bronchiolitis by incorporating omics data, including the upper airway transcriptome and metabolome profiles. For example, a single-center study of 55 infants hospitalized for RSV bronchiolitis using nasopharyngeal transcriptome profiling reported that lower type I interferon responses were associated with higher severity22. Similarly, studies of infants with bronchiolitis using nasopharyngeal transcriptome profiling suggested inflammatory responses (e.g., type 1 interferon [IFN], neutrophil, and regulatory T cells) were associated with disease severity9. In another study of 144 infants hospitalized for bronchiolitis using nasopharyngeal metabolome profiling showed that sphingolipid metabolism pathway was significantly enriched in infants with positive pressure ventilation use23. A previous integrated-omics analysis of infants with RSV bronchiolitis—which includes transcriptome and metabolome profiling—also found that the most-severe endotype had unique inflammatory response profiles (e.g., low type I IFN response) and a higher abundance of sphingolipids17. The present study—applying explainable deep networks using image-converted integrative omics components—corroborates these prior reports and extends them by demonstrating the molecular networks driving bronchiolitis pathobiology in a large prospective study.
The mechanisms underlying the observed molecular networks—characterized by inflammatory response pathways (e.g., TLR signaling pathway, NF-κB signaling pathway) and centralized with immunity-related metabolites (e.g., N-acylsphingosine, phosphatidylethanolamine)—warrant clarification. In concordance with our data, studies suggested the role of these inflammatory response pathways and innate immunity-related metabolites in bronchiolitis pathobiology11,24,25,26. First, research showed that neutrophil TLR4 expression was deficient in both the blood and the airways among infants with severe bronchiolitis24 and that airway neutrophils of life-threatening RSV bronchiolitis infants were characterized by NF-κB signaling pathway and upregulated inflammatory responses (i.e., interleukin [IL]-6 and IFN-based responses)11. Similarly, TLR engagements activate the NF-κB pathway through combinations of the adaptor molecules (e.g., MyD88, TRIF), thereby mediating the production of both pro- and anti-inflammatory cytokines in innate immune cells25. Second, a recent study showed the most-severe endotype of infants with bronchiolitis had a higher abundance of ceramide (i.e., N-acylsphingosine) and phosphatidylethanolamine14. Ceramides are the central core of sphingolipid metabolism27,28 and their chronic accumulation in the lung can induce pulmonary inflammation29. Another study also showed that phosphatidylethanolamine was correlated with excessive proinflammatory responses and was over-represented in patients with severe respiratory illness30. Lastly, recent experimental studies demonstrated that ceramide and phosphatidylethanolamine play a key role in apoptosis, inflammation, and stress responses involving TLR and/or NF-κB signaling pathways31,32,33,34,35,36. For example, ceramide metabolism is regulated by TLR stimulation and modulates TLR-induced IL-6 release31. The phosphatidylethanolamine binding protein 4 knockout mice showed upregulated levels of TLR4 and NF-κB with increased neutrophil infiltration and cytokine secretions (e.g., IL-1β, TNFα, and cyclooxygenase-2)32.
The drug repurposing candidates identified from these molecular networks—such as prostaglandin analogs or agonists (e.g., iloprost) and tyrosine kinase inhibitors (e.g., dasatinib, imatinib)—also warrant clarification. Prostaglandin E2 (PGE2), prostaglandin I2 (PGI2), and prostaglandin D2 (PGD2) have been shown to modulate inflammation and the immune system by regulating the expression and concentration of cytokines, such as IL-1β and IL-637,38. Consequently, these are being considered potential therapeutic targets for the treatment of viral infections39,40,41,42,43,44. For example, recent studies showed the EP2 and EP4 receptors of PGE2 mediate inflammation in innate and adaptive immune cells through NF-κB signaling and PI3K signaling pathways39,40, and could be effective anti-inflammatory targets in human lung disease41. Additionally, PGI2 analogs including iloprost regulate the innate immune systems and promote anti-inflammatory effects by modifying TLR4 expression42,43. Another study also supported a protective role for PGI2 analogs in RSV-induced illness and suggested a possible therapy for acute RSV infection44. In contrast, kinase inhibitors—one of the main therapeutic approaches to alleviate dysregulated inflammation—target the activity of kinases that regulate the production of inflammatory mediators45. Previous studies suggested that Lyn tyrosine kinase and receptor tyrosine kinase KIT have a pivotal role in the initiation of inflammatory reactions through mast cell activation, and their inhibitors could potentially serve as therapeutic targets for inflammatory diseases46,47,48,49. For example, a mouse study showed that Lyn kinase inhibition by dasatinib suppressed the secretion of inflammatory cytokines (i.e., IL-4 and TNFα)48. A randomized trial of patients with severe refractory asthma showed that KIT inhibition by imatinib decreased airway hyperresponsiveness, mast cell counts and tryptase release49. Notwithstanding the complexity, the present study consolidates the preceding insights into the integrative omics molecular networks—which incorporate the intra- and inter-omics interactions representing human pathobiology—along with potential drug candidates for repurposing.
In summary, by applying explainable deep networks using image-converted integrative omics components derived from a multicenter prospective cohort study of infants with bronchiolitis, we identified an omics-cluster comprising 401 mRNAs and 38 metabolites that distinguishes bronchiolitis severity. These severity-discriminatory components derive molecular networks driving bronchiolitis pathobiology, which were characterized by inflammatory response pathways and were centralized with innate immunity-related metabolites. We also identified eight sub-networks including existing 50 drug candidates for repurposing through the drug-based network analysis. While external validation is warranted, our data suggest the integrative omics molecular networks and existing drug candidates for repurposing in infant bronchiolitis. Our approach facilitates not only the identification of molecular networks underlying infant bronchiolitis but also the development of pioneering treatment strategies. Furthermore, our new analytic approach may provide a solution for the holistic understanding of disease pathobiology, extending beyond infant bronchiolitis.
Limitation of the study
The current study has several potential limitations. First, our study did not have “healthy controls”. However, the study objective was not to evaluate the role of the transcriptome and metabolome in the development of bronchiolitis but to investigate their relationship with the disease severity within infants with bronchiolitis. Second, bronchiolitis involves inflammation of both upper and lower airways, while this study is based on nasopharyngeal specimens. Regardless, the use of upper airway samples is preferable because lower airway sampling (e.g., bronchoscopy) would be invasive in these young infants. Additionally, studies have suggested that upper airway sampling represents the lung transcriptome50 and metabolome51 profiles in children. Third, the present study did not have mechanistic experiments to validate the identified molecular networks and their functions. Fourth, our inferences might be biased due to the relationship between the timing of treatments, specimen collections, and respiratory support use despite that the specimens were collected within a short time frame. Fifth, although this study offers well-calibrated hypotheses that facilitate future experiments, our inferences warrant external validation, particularly in infant populations. In addition, the primary data source for our network analyses predominantly represents adult populations; thus, we need further studies and clinical trials to confirm the applicability and safety of our findings in infants. Sixth, we did not have a testing dataset that is completely kept hidden during the whole model training and tuning processes, which could potentially increase the risk of overfitting problems. Our study aims not primarily at constructing a highly predictive CNN model, but rather at developing a model that better represents the pathobiology by using our data to its fullest extent. Therefore, we chose to maximize our training dataset and minimized overfitting problems by optimizing our model with a minimal set of parameters using our testing (i.e., validation) dataset. Yet, we need to assess the validity of our model using an additional dataset in future studies. Lastly, while the study sample included a racially/ethnically and geographically diverse multicenter cohort, our inferences should be generalized cautiously beyond infants hospitalized for bronchiolitis. Nonetheless, our findings remain relevant for 110,000 U.S. children hospitalized annually—a demographic carrying a substantial health burden2.
Methods
Study design, setting, and participants
We analyzed data from the 35th Multicenter Airway Research Collaboration (MARC-35)—a multicenter prospective cohort study of infants hospitalized with bronchiolitis52. MARC-35 is coordinated by the Emergency Medicine Network (EMNet), a collaboration of 247 participating hospitals53. Investigators enrolled infants (age <1 year) hospitalized with bronchiolitis at 17 sites across 14 U.S. states (Supplementary Table 2) using a standardized protocol during three consecutive bronchiolitis seasons (from November 1 through April 30) during 2011–2014. Bronchiolitis was defined by the American Academy of Pediatrics guidelines54 as an acute respiratory illness with some combination of rhinitis, cough, tachypnoea, wheezing, crackles, and retractions54 and was diagnosed by an attending physician. We excluded infants who were transferred to a participating hospital >24 h after the original hospitalization, those who were consented >24 h after hospitalization, or those with known heart-lung disease, immunodeficiency, immunosuppression, or gestational age of <32 weeks. All patients were treated at the discretion of the treating physicians.
Of 1016 infants enrolled in the MARC-35 cohort, the current analysis investigated 397 infants who underwent both nasopharyngeal transcriptome and metabolome profiling. The institutional review board at each of the participating hospitals approved the study. Written informed consent was obtained from the parent or guardian.
Data collection
Clinical data (demographic characteristics; medical, environmental, and family history; and details of the acute illness and hospital course) were collected via structured interview and chart reviews using a standardized protocol55. All data were reviewed at the EMNet Coordinating Center (Boston, Massachusetts, USA), and site investigators were queried about missing data and discrepancies identified by manual data checks.
In addition to the clinical data, investigators also collected nasopharyngeal airway specimens within 24 h of hospitalization by using standardized protocols55,56. All sites used the same collection equipment and collected the samples within 24 h of a child’s arrival in the medical ward or intensive care unit. Frozen nasopharyngeal specimens were shipped in batches to Baylor College of Medicine (Houston, Texas, USA) where they performed respiratory virus testing using real-time polymerase chain reaction (RT-PCR), transcriptome profiling by total RNA-sequencing, and metabolome profiling using liquid chromatography mass spectrometry (LC-MS)55,56,57,58.
Respiratory virus testing
Nasopharyngeal samples were tested for 17 respiratory viruses (including respiratory syncytial virus [RSV] and rhinovirus [RV]) by using RT-PCR assays58. For RV detection, complementary DNA was generated using virus-specific primers for RV and singleplex RT-PCR was used. The details of the RV primers and probes have been described elsewhere59.
Nasopharyngeal total RNA extraction and RNA-seq
Total RNA was isolated from 398 randomly-selected nasopharyngeal specimens using Trizol LS reagent (ThermoFisher Scientific, Waltham, MA, U.S.A.) in combination with the Direct-zol RNA Miniprep Kit (Zymo Research, Irvine, CA, U.S.A.). RNA quantity was measured with the Qubit 2.0 fluorometer (ThermoFisher Scientific). Its quality was assessed with the Agilent Bioanalyzer 2100 (Agilent, Palo Alto, CA, U.S.A.) using the RNA 6000 Nano kit. Total RNA underwent DNase treatment using the TURBO DNA-free™ Kit (ThermoFisher Scientific) and rRNA reduction for both human and bacterial rRNA using NEBNext rRNA Depletion Kits (New England Biolabs, Ipswich, MA, U.S.). RNA was prepared for sequencing using the NEBNext Ultra II Directional RNA Library Prep Kit (New England Biolabs) and sequenced on a NovaSeq6000 (Illumina, San Diego, CA, U.S.A.) using an S4 100 bp PE Flowcell (Illumina). All RNA-seq samples had sufficient sequence depth (mean, 48,014,679 pair-end reads/sample) to obtain a high degree of sequence coverage.
Nasopharyngeal transcriptome profiling
Raw sequence reads were trimmed and filtered for adapters and contaminants using the k-mers strategy in bbduk60 and default settings (i.e., minlength = 14, minavgquality = 20, maxns = 0). We estimated transcript abundances from the filtered and trimmed reads by Salmon 1.9.061 using the human transcriptome (hg38) and the mapping-based mode. We generated a decoy-aware transcriptome and then quantified the reads using Salmon’s default settings and the following flags: –validateMappings, – recoverOrphans, –seqBias, and –gcBias. Salmon is fast and accurate, corrects for potential changes in gene length across samples (e.g., from differential isoform usage), and has higher sensitivity at the same false discovery rate (FDR) in differential expression gene analysis.
Nasopharyngeal airway metabolome profiling
Metabolome profiling used 125 μl of nasopharyngeal airway sample. All samples were blinded to Metabolon and processed in random order. The metabolic profiling used ACQUITY ultra-high performance liquid chromatography (UPLC) (Waters, Milford, MA, USA) and Q-Exactive high resolution/accurate mass spectrometry (MS) interfaced with a heated electrospray ionization (HESI-II) source and Orbitrap mass analyzer operated at 35,000 mass resolution (Thermo Fisher Scientific, Waltham, MA, USA).
Sample preparation was carried out as described previously62. In brief, recovery standards were added prior to the first step in the extraction process for quality control purposes. Proteins were precipitated with methanol under vigorous shaking for two minutes (Glen Mills Genogrinder 2000; Clifton, NJ, USA) followed by centrifugation. The resulting extract was divided into four fractions. The first aliquot was analyzed using acidic positive ion conditions, chromatographically optimized for more hydrophilic compounds. The second aliquot was also analyzed using acidic positive ion conditions and was chromatographically optimized for more hydrophobic compounds. The third aliquot was analyzed with basic negative ion optimized conditions using a separate dedicated C18 column. The fourth aliquot was analyzed via negative ionization following elution from a HILIC column (Waters UPLC BEH Amide). The MS analysis alternated between MS and data-dependent MSn scans using dynamic exclusion.
Metabolites were identified by automated comparison of the ion features in the experimental samples to a reference library of chemical standard entries that include retention time, molecular weight (m/z), preferred adducts, and in-source fragments as well as associated MS spectra, and curated by visual inspection for quality control using QUICS software63. Identification of known chemical entities was based on comparisons to metabolomic library entries of >3000 purified standards. Peaks were quantified using area-under-the-curve. The raw area counts for each metabolite in each sample were normalized to correct for variation due to instrument inter-day tuning differences by the median value for each run-day, setting the median to 1.0 for each run. Missing values were imputed with the observed minimum for that particular compound.
Four types of quality controls were analyzed in concert with the specimens: 1) samples generated from a pool from a small portion of each experimental specimen that served as technical replicate; 2) extracted water samples that served as process blanks; 3) samples of solvent used in extraction; and 4) a cocktail of standards spiked into every analyzed specimen that allowed instrument performance monitoring. The median relative standard deviation (RSD) for the standards that are added to each sample—a measure of instrument variability—was <5%.
Clinical outcome measure
The primary outcome was respiratory support use, defined as the use of continuous positive airway ventilation and/or mechanical ventilation and/or high-flow oxygen at any time during the index hospitalization64.
Data normalization
Before the main analyses, we executed a series of data processing steps on the mRNA data. First, we filtered the mRNAs with a total read count of <10 across all subjects, reducing the number of mRNAs from 21,136 to 20,596. Second, we normalized the read count by the R DESeq2 package65 using default settings. Third, we applied a log(x + 1) transformation and robust scaling to the mRNA data. In robust scaling, each value of a feature (i.e., mRNA) is scaled by removing the median and dividing by its interquartile range (IQR: the range between the 1st quartile [Q1] and 3rd quartile [Q3]). Fourth, we modified outliers of the mRNA data that fall below \(Q1-1.5* {IQR}\) or above \(Q3+1.5* {IQR}\) into \(Q1-1.5* {IQR}\) or \(Q3+1.5* {IQR}\), respectively. Finally, we applied a min-max transformation to the mRNA data to ensure all values fell within the range of 0 to 1 as a preparation for the subsequent image transformation.
We also executed a series of data processing steps on the metabolome data. We firstly adjusted the metabolome data for potential batch effect by using empirical Bayes models (ComBat method66). Next, we adopted probabilistic quotient normalization (PQN)67 to the metabolome data in accordance with the previous study13. Lastly, the metabolome data underwent the same sequence of processing as the mRNA data: log(x + 1) transformation, robust scaling, outlier modifications, and min-max transformation.
Component selection
From all measured mRNAs and metabolites, we first selected mRNAs and metabolites that were related to acute severity (i.e., respiratory support use) of infant bronchiolitis to filter candidates for the following image conversion process. To select the mRNAs and metabolites, we used logistic regression models with a threshold of p < 0.05, adjusting for the sex, age, and race/ethnicity of the infants. Next, we created variables for interaction by multiplying mRNA expression with metabolite abundance and selected variables for interaction related to the acute severity. To address the issue of multiple testing, we used a regression model excluding the main effects to select variables for interaction, as this approach allowed us to secure a sufficient number of variables for interaction meeting the FDR <0.01 threshold. However, excluding the main effects of mRNA and metabolite may confound the estimate of the interaction effect between mRNA and metabolite. Therefore, as a sensitivity analysis, we also selected variables for interaction using a regression model including the main effects with the p < 0.05 threshold, then compared the accuracy of the constructed model to the main analysis. Each variable for interaction underwent the same sequence of processing as the mRNA and metabolome data: log(x + 1) transformation, robust scaling, outlier modifications, and Min-max transformation. Lastly, we prepared the severity-related components of each individual infant, comprising 588 mRNAs, 128 metabolites, and 4835 variables for interactions.
Image conversion of integrative omics components
We transformed the severity-related components of each infant into a single image using the DeepInsight method68, which converts non-image components into an organized image form. In the conversion process, we used t-distributed stochastic neighbor embedding (t-SNE)69 to create a 224-pixel square image for each infant. Specifically, we firstly reduced the dimensions of feature set of the severity-related components to a 2D plane using t-SNE, which determined the feature’s locations. t-SNE calculates feature proximities in high-dimensional data by first computing pairwise similarities using a Gaussian distribution, and then mapping these features to a 2D plane using a t-distribution. Second, we identified the smallest rectangle enclosing all points on the 2D plane. Second, we identified the smallest rectangle enclosing all points on the 2D plane. The points in this Cartesian plane (i.e., 2D plane) represent the mRNAs, metabolites, or variables for interactions. We performed rotation on the identified rectangle since the image should be framed in a horizontal or vertical form for the CNN architecture. For rotation, a gradient of two corner coordinates of the rectangle is considered. Following this procedure, the Cartesian coordinates of all types of features (i.e., mRNAs, metabolites, and variables for interaction) are converted to pixels. Therefore, the pixel frame consists of the positions of all features for each sample. Third, we mapped values of the components to pixel locations. The intensity of each pixel corresponds to the expression or abundance of each component. Additionally, in each image, we assigned different colors to the pixels corresponding to metabolites (red), mRNAs (green), and variables for interaction (blue) to highlight the differences among the different types of components. When a single pixel corresponds to multiple components, we overlaid all the corresponding colors.
Severity-discriminatory omics-cluster identification by explainable deep neural networks
We developed a convolutional neural network (CNN) model70 that classifies the acute severity of bronchiolitis from individual images. Specifically, we first randomly partitioned the 397 images into an 8:2 ratio, yielding 317 for training and 80 for testing. To address the class imbalance in the training images, we used the synthetic minority over-sampling technique71 to equalize positive and negative outcomes (i.e., respiratory support use), augmenting the number of training images from 317 to 508. We secondly created the CNN model with the augmented training images using the SqueezeNet architecture72, adjusting the final convolution layer for binary classification tasks. To assess the potential overfitting problem, the model performance was evaluated with the testing images using the AUROC after completing the 1500 training epochs. The learning rate was determined manually through comparisons of the model performance. The other parameters of the model are presented on the shared code (See Data and code availability). We then employed class activation mapping (CAM)73 with the CNN model using the DeepFeature method74 to identify the specific regions within the images representing the omics-cluster that discriminates the acute severity. CAM used the final convolutional layer of a CNN model to identify important regions in an image for classification. The importance scores were assigned to each pixel in every individual sample’s image, based on the activation of units in this layer and their weights associated with a specific class.
To integrate important pixels from images of all samples, we employed a region accumulation step, which aggregated the important scores in each pixel across all images, thereby identifying pixels (i.e., omics components) essential for classifying severity. The region accumulation step integrates pixels of importance by performing a union operation on individual activation pixels corresponding to each sample. In this process, we consider a set of samples, each belonging to a specific category. The active pixels from each sample in the specific category are integrated to form a single comprehensive pixel set. This integrated pixel set includes all significant features necessary for the classification of each category. We extracted the mRNAs, metabolites, and variables for interactions from the integrated pixel set of samples with severe bronchiolitis using the max flatten method with a threshold of 0.001; this method selected the maximum importance scores across all images for each pixel, thus capturing any pixel that exceeds the threshold in each class, resulting in a comprehensive feature set relevant to the class prediction. We finally selected the severity-discriminatory mRNAs and metabolites—either originally extracted or included within the extracted variables for interaction—for the subsequent biological interpretation analyses.
Pathway analysis of severity-discriminatory mRNAs and metabolites
We conducted the functional enrichment joint-pathway analysis with the severity-discriminatory components (i.e., mRNAs and metabolites) using MetaboAnalyst (v.5.0)75. For the joint-pathway analysis, we converted the Ensemble IDs of the identified mRNAs to official gene symbols and the names of identified metabolites to the human metabolome database (HMDB)76 IDs. We used the Kyoto Encyclopedia of genes and genomes (KEGG) database77 to select enriched pathways for both mRNAs and metabolites. For the joint-pathway selection algorithm, we used the hypergeometric test for enrichment analysis, degree centrality as the topology measure, and combined queries as the integration method. As previous research1,2,3,4,5,20 showed that the infecting virus leads to heterogeneity in the pathobiology and severity of bronchiolitis, we also conducted the functional enrichment joint-pathway analysis using the RSV- or RV-related severity-discriminatory components to examine the heterogeneity and demonstrate the validity of our methods. We selected RSV- or RV-related components by using linear regression models, adjusting for the age, sex, and race/ethnicity of the infants.
Network analysis of severity-discriminatory mRNAs and metabolites
We identified pathway-based molecular networks with the severity-discriminatory mRNAs and metabolites using MetScape (v.3.1.3)78 and Cytoscape (v.3.9.1)79. After converting the Ensemble IDs of the identified mRNAs to Entrez gene IDs and the names of identified metabolites to KEGG IDs77, we imported these components into the MetScape application to identify networks. In the visualization of the largest network identified from the analysis, we set thresholds of the nodes at a closeness centrality of ≥0.125 and a betweenness centrality of ≥0.01 to enhance the interpretability of the result. We also identified molecular networks with the RSV- or RV-related severity-discriminatory components selected in the pathway analysis section.
Identification of potential drug candidates for repurposing
We identified potential drug candidates for repurposing through gene- and drug-based network analyses with the severity-discriminatory mRNAs using NeDRexApp (v.1.1.2)80. First, we imported a network query from NeDRexDB80—which integrates eight publicly available biomedical databases—with Gene-Disorder and Disorder-Disorder association options, which allowed us to construct a gene- and drug-based network based on the relationships between genes and disorders in NeDRexDB. Second, we identified the disease module with DIAMOnD algorithm81 using the input seeds of gene symbols of the severity-discriminatory mRNAs. Specifically, the DIAMOnD algorithm starts with the input seeds as an initial disease module and iteratively expands the module by adding genes (i.e., nodes) that show the highest connectivity significance to the already selected genes in the module. In each iteration, the algorithm assesses the connectivity significance of all direct neighbors of the existing module and integrates the most significantly connected node. We set the number of iterations to 200, which implies that the module expands to include 200 additional genes, based on their connectivity to the initial seeds. We assigned each initial seed weight of 1, signifying equal importance of each seed in the initial module. Third, to visualize gene- and drug-based networks and to identify drug candidates for repurposing, we performed drug prioritization with TrustRank algorithm82 using the identified disease module. TrustRank algorithm calculates a trust score for each drug based on how well-connected it is to the genes within the disease module. The trust scores are spread from the seed nodes to other nodes (i.e., other genes and potential drug targets) in the network. The damping factor controls how much of this score is passed from one node to the next. We set the damping factor as 0.85 and selected 50 top-ranked drugs with the highest trust scores. Finally, we ranked the identified drug candidates based on the log10 Trustworthiness score82 using the largest or second largest gene- and drug-based network.
Data availability
The RNA-seq and metabolome profiling data that support the findings of this study are available on the NIH/NIAID ImmPort (https://www.immport.org/shared/study/SDY1883), with the accession number SDY1883, upon a request from the researchers whose research investigates severe bronchiolitis, recurrent wheezing, asthma, and related concepts. The data are not available without restriction to be compliant with the informed consent forms of MARC-35 study and the genomic data sharing plan.
Code availability
All original code has been deposited on the GitHub page (https://github.com/HasegawaLab/omics_deep_networks) and is publicly available as of the date of publication.
References
Dalziel, S. R. et al. Bronchiolitis. Lancet 400, 392–406 (2022).
Fujiogi, M. et al. Trends in bronchiolitis hospitalizations in the United States: 2000-2016. Pediatrics 144, e20192614 (2019).
Pelletier, J. H., Au, A. K., Fuhrman, D., Clark, R. S. B. & Horvat, C. Trends in Bronchiolitis ICU Admissions and Ventilation Practices: 2010-2019. Pediatrics 147. https://doi.org/10.1542/peds.2020-039115 (2021).
Mahant, S. et al. Rates in bronchiolitis hospitalization, intensive care unit use, mortality, and costs from 2004 to 2018. JAMA Pediatr. 176, 270–279 (2022).
Kirolos, A. et al. A Systematic Review of Clinical Practice Guidelines for the Diagnosis and Management of Bronchiolitis. J. Infect. Dis. 222, S672–S679 (2020).
Jong, V. L. et al. Transcriptome assists prognosis of disease severity in respiratory syncytial virus infected infants. Sci. Rep. 6, https://doi.org/10.1038/srep36603 (2016).
Jones, A. C. et al. Personalized transcriptomics reveals heterogeneous immunophenotypes in children with viral bronchiolitis. Am. J. Respir. Crit. Care Med. 199, 1537–1549 (2019).
Zhu, Z. et al. Nasopharyngeal airway dual-transcriptome of infants with severe bronchiolitis and risk of childhood asthma: A multicenter prospective study. J. Allergy Clin. Immunol. 150, 806–816 (2022).
Fujiogi, M. et al. Integrated relationship of nasopharyngeal airway host response and microbiome associates with bronchiolitis severity. Nat. Commun. 13, 4970 (2022).
Fujiogi, M. et al. In infants with severe bronchiolitis: dual-transcriptomic profiling of nasopharyngeal microbiome and host response. Pediatr. Res. 88, 144–146 (2020).
Besteman, S. B. et al. Transcriptome of airway neutrophils reveals an interferon response in life-threatening respiratory syncytial virus infection. Clin. Immunol. 220, 108593 (2020).
Raita, Y. et al. Integrated-omics endotyping of infants with rhinovirus bronchiolitis and risk of childhood asthma. J. Allergy Clin. Immunol. 147, 2108–2117 (2021).
Ooka, T. et al. Integrative genetics-metabolomics analysis of infant bronchiolitis-childhood asthma link: A multicenter prospective study. Front. Immunol. 13, 1111723 (2022).
Fujiogi, M. et al. Nasopharyngeal lipidomic endotypes of infants with bronchiolitis and risk of childhood asthma: a multicentre prospective study. Thorax 77, 1059–1069 (2022).
Kyo, M. et al. Association of Nasopharyngeal and Serum Glutathione Metabolism with Bronchiolitis Severity and Asthma Risk: A Prospective Multicenter Cohort Study. Metabolites 12, https://doi.org/10.3390/metabo12080674 (2022).
Teoh, S. T. et al. Combined plasma and urinary metabolomics uncover metabolic perturbations associated with severe respiratory syncytial viral infection and future development of asthma in infant patients. Metabolites 12, 178 (2022).
Raita, Y. et al. Integrated omics endotyping of infants with respiratory syncytial virus bronchiolitis and risk of childhood asthma. Nat. Commun. 12, 3601 (2021).
Zhu, Z. et al. Metabolome subtyping of severe bronchiolitis in infancy and risk of childhood asthma. J. Allergy Clin. Immunol. 149, 102–112 (2022).
Fujiogi, M. et al. Integrated associations of nasopharyngeal and serum metabolome with bronchiolitis severity and asthma: A multicenter prospective cohort study. Pediatr. Allergy Immunol. 32, 905–916 (2021).
Fujiogi, M. et al. Association of rhinovirus species with nasopharyngeal metabolome in bronchiolitis infants: A multicenter study. Allergy 75, 2379–2383 (2020).
Maghsoudi, Z., Nguyen, H., Tavakkoli, A. & Nguyen, T. A comprehensive survey of the approaches for pathway analysis using multi-omics data integration. Brief. Bioinform. 23, https://doi.org/10.1093/bib/bbac435 (2022).
Thwaites, R. S. et al. Reduced nasal viral load and IFN responses in infants with respiratory syncytial virus bronchiolitis and respiratory failure. Am. J. Respir. Crit. Care Med. 198, 1074–1084 (2018).
Stewart, C. J. et al. Associations of nasopharyngeal metabolome and microbiome with severity among infants with bronchiolitis. A multiomic analysis. Am. J. Respir. Crit. Care Med. 196, 882–891 (2017).
Halfhide, C. P. et al. Neutrophil TLR4 expression is reduced in the airways of infants with severe bronchiolitis. Thorax 64, 798–805 (2009).
Wang, H. et al. The role of JAK-3 in regulating TLR-mediated inflammatory cytokine production in innate immune cells. J. Immunol. 191, 1164–1174 (2013).
Zhu, Z. et al. and MARC-35 Investigators. Nasal airway microRNA profiling of infants with severe bronchiolitis and risk of childhood asthma: a multicentre prospective study. Eur. Respir. J. 62, https://doi.org/10.1183/13993003.00502-2023 (2023).
Hannun, Y. A. & Obeid, L. M. Principles of bioactive lipid signalling: lessons from sphingolipids. Nat. Rev. Mol. Cell Biol. 9, 139–150 (2008).
Maceyka, M. & Spiegel, S. Sphingolipid metabolites in inflammatory disease. Nature 510, 58–67 (2014).
Ziobro, R., Henry, B., Edwards, M. J., Lentsch, A. B. & Gulbins, E. Ceramide mediates lung fibrosis in cystic fibrosis. Biochem. Biophys. Res. Commun. 434, 705–709 (2013).
Wu, J. et al. Lipidomic signatures align with inflammatory patterns and outcomes in critical illness. Nat. Commun. 13, 6789 (2022).
Köberlin, M. S. et al. A conserved circular network of coregulated lipids modulates innate immune responses. Cell 162, 170–183 (2015).
Qu, X.-Q. et al. Hepatocyte-conditional knockout of phosphatidylethanolamine binding protein 4 aggravated LPS/D-GalN-induced acute liver injury via the TLR4/NF-κB pathway. Front. Immunol. 13, 901566 (2022).
Kiefer, K., Casas, J., García-López, R. & Vicente, R. Ceramide Imbalance and Impaired TLR4-Mediated Autophagy in BMDM of an ORMDL3-Overexpressing Mouse Model. Int. J. Mol. Sci. 20, 1391 (2019).
Zhang, Y. et al. Short-Chain Fatty Acids Attenuate 5-Fluorouracil-Induced THP-1 Cell Inflammation through Inhibiting NF-κB/NLRP3 Signaling via Glycerolphospholipid and Sphingolipid Metabolism. Molecules 28, 494 (2023).
Bae, M. et al. Akkermansia muciniphila phospholipid induces homeostatic immune responses. Nature 608, 168–173 (2022).
Olona, A. et al. Sphingolipid metabolism during Toll-like receptor 4 (TLR4)-mediated macrophage activation. Br. J. Pharmacol. 178, 4575–4587 (2021).
Sander, W. J., O’Neill, H. G. & Pohl, C. H. Prostaglandin E2 as a modulator of viral infections. Front. Physiol. 8, 89 (2017).
Ricciotti, E. & FitzGerald, G. A. Prostaglandins and inflammation. Arterioscler. Thromb. Vasc. Biol. 31, 986–1000 (2011).
Thumkeo, D. et al. PGE2-EP2/EP4 signaling elicits immunosuppression by driving the mregDC-Treg axis in inflammatory tumor microenvironment. Cell Rep. 39, 110914 (2022).
Martínez-Colón, G. J. & Moore, B. B. Prostaglandin E2 as a regulator of immunity to pathogens. Pharmacol. Ther. 185, 135–146 (2018).
Gill, S. K. et al. The anti-inflammatory effects of PGE2 on human lung macrophages are mediated by the EP4 receptor. Br. J. Pharmacol. 173, 3099–3109 (2016).
Wu, X. et al. A transcriptomics-guided drug target discovery strategy identifies receptor ligands for lung regeneration. Sci. Adv. 8, eabj9949 (2022).
Toki, S., Zhou, W., Goleniewska, K., FitzGerald, G. A. & Peebles, R. S. Jr Prostaglandin I2 receptor (IP) signaling inhibits TLR4 cell surface expression and reduces LPS-induced lung inflammation. J. Allergy Clin. Immunol. 127, AB92–AB92 (2011).
Hashimoto, K. et al. Functional polymorphism of the promoter region of the prostacyclin synthase gene and severity of RSV infection in hospitalized children. J. Med. Virol. 80, 2015–2022 (2008).
Zarrin, A. A., Bao, K., Lupardus, P. & Vucic, D. Kinase inhibition in autoimmunity and inflammation. Nat. Rev. Drug Discov. 20, 39–63 (2021).
Brian, B. F. 4th & Freedman, T. S. The Src-family kinase Lyn in immunoreceptor signaling. Endocrinology 162, bqab152 (2021).
Gilfillan, A. M. & Rivera, J. The tyrosine kinase network regulating mast cell activation. Immunol. Rev. 228, 149–169 (2009).
Lee, D. et al. Dasatinib inhibits Lyn and Fyn Src-family kinases in mast cells to suppress type I hypersensitivity in mice. Biomol. Ther. 28, 456–464 (2020).
Cahill, K. N. et al. KIT Inhibition by Imatinib in Patients with Severe Refractory Asthma. N. Engl. J. Med. 376, 1911–1920 (2017).
Poole, A. et al. Dissecting childhood asthma with nasal transcriptomics distinguishes subphenotypes of disease. J. Allergy Clin. Immunol. 133, 670–8.e12 (2014).
Farne, H. et al. Comparative metabolomic sampling of upper and lower airways by four different methods to identify biochemicals that may support bacterial growth. Front. Cell. Infect. Microbiol. 8, 432 (2018).
Hasegawa, K. et al. Association of Rhinovirus C Bronchiolitis and Immunoglobulin E Sensitization During Infancy With Development of Recurrent Wheeze. JAMA Pediatr. 173, 544–552 (2019).
Emergency Medicine Network http://www.emnet-usa.org/.
Ralston, S. L. et al. Clinical practice guideline: the diagnosis, management, and prevention of bronchiolitis. Pediatrics 134, e1474–e1502 (2014).
Hasegawa, K. et al. Association of nasopharyngeal microbiota profiles with bronchiolitis severity in infants hospitalised for bronchiolitis. Eur. Respir. J. 48, 1329–1339 (2016).
Hasegawa, K. et al. Respiratory syncytial virus genomic load and disease severity among children hospitalized with bronchiolitis: multicenter cohort studies in the United States and Finland. J. Infect. Dis. 211, 1550–1559 (2015).
Mansbach, J. M. et al. Prospective multicenter study of viral etiology and hospital length of stay in children with severe bronchiolitis. Arch. Pediatr. Adolesc. Med. 166, 700–706 (2012).
Mansbach, J. M. et al. Prospective multicenter study of children with bronchiolitis requiring mechanical ventilation. Pediatrics 130, e492–e500 (2012).
Lu, X. et al. Real-time reverse transcription-PCR assay for comprehensive detection of human rhinoviruses. J. Clin. Microbiol. 46, 533–539 (2008).
Bushnell, B. BBMap: a fast, accurate, splice-aware aligner. (No. LBNL-7065E). Lawrence Berkeley National Lab. (LBNL), Berkeley, CA (United States). (2014).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Evans, A. M., DeHaven, C. D., Barrett, T., Mitchell, M. & Milgram, E. Integrated, nontargeted ultrahigh performance liquid chromatography/electrospray ionization tandem mass spectrometry platform for the identification and relative quantification of the small-molecule complement of biological systems. Anal. Chem. 81, 6656–6667 (2009).
Dehaven, C. D., Evans, A. M., Dai, H. & Lawton, K. A. Organization of GC/MS and LC/MS metabolomics data into chemical libraries. J. Cheminform. 2, 9 (2010).
Ooka, T. et al. Proteome signature difference between respiratory viruses is associated with severity of bronchiolitis. Pediatr. Allergy Immunol. 32, 1869–1872 (2021).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Dieterle, F., Ross, A., Schlotterbeck, G. & Senn, H. Probabilistic quotient normalization as robust method to account for dilution of complex biological mixtures. Application in 1H NMR metabonomics. Anal. Chem. 78, 4281–4290 (2006).
Sharma, A., Vans, E., Shigemizu, D., Boroevich, K. A. & Tsunoda, T. DeepInsight: A methodology to transform a non-image data to an image for convolution neural network architecture. Sci. Rep. 9, 11399 (2019).
Van der Maaten, L. & Hinton, G. Visualizing Data using t-SNE. J. Mach. Learn. Res. 9, (2008).
O’Shea, K. & Nash, R. An Introduction to Convolutional Neural Networks. arXiv. https://doi.org/10.48550/arXiv.1511.08458 (2015).
Chawla, N. V., Bowyer, K. W., Hall, L. O. & Kegelmeyer, W. P. SMOTE: synthetic minority over-sampling technique. J. Artif. Intell. Res. 16, 321–357 (2002).
Iandola, F. N. et al. SqueezeNet: AlexNet-level accuracy with 50x fewer parameters and <0.5MB model size. arXiv. https://doi.org/10.48550/arXiv.1602.07360 (2016).
Zhou, B., Khosla, A., Lapedriza, A., Oliva, A. & Torralba, A. Learning Deep Features for Discriminative Localization. arXiv. https://doi.org/10.48550/arXiv.1512.04150.
Sharma, A., Lysenko, A., Boroevich, K. A., Vans, E. & Tsunoda, T. DeepFeature: feature selection in nonimage data using convolutional neural network. Brief. Bioinform 22, bbab297 (2021).
Pang, Z. et al. MetaboAnalyst 5.0: narrowing the gap between raw spectra and functional insights. Nucleic Acids Res. 49, W388–W396 (2021).
Wishart, D. S. et al. HMDB 5.0: the Human Metabolome Database for 2022. Nucleic Acids Res. 50, D622–D631 (2022).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Karnovsky, A. et al. Metscape 2 bioinformatics tool for the analysis and visualization of metabolomics and gene expression data. Bioinformatics 28, 373–380 (2012).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Sadegh, S. et al. Network medicine for disease module identification and drug repurposing with the NeDRex platform. Nat. Commun. 12, 6848 (2021).
Ghiassian, S. D., Menche, J. & Barabási, A.-L. A DIseAse MOdule Detection (DIAMOnD) algorithm derived from a systematic analysis of connectivity patterns of disease proteins in the human interactome. PLoS Comput. Biol. 11, e1004120 (2015).
Gyöngyi, Z., Garcia-Molina, H. & Pedersen, J. Combating web spam with trustrank. In Proc. of the Thirtieth international conference on Very large data bases-Volume. 30, 576–587 (2004)
Acknowledgements
This study was supported by grants (UG3/UH3 OD-023253, R01 AI-127507, R01 AI-134940, R01 AI-137091, R01 AI-148338, and K01 AI-153558) from the National Institutes of Health (Bethesda, MD). The content of this manuscript is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. The funding organization was not involved in the collection, management, or analysis of the data; preparation or approval of the manuscript; or decision to submit the manuscript for publication.
Author information
Authors and Affiliations
Contributions
T.O. carried out the data processing, optimized the existing deep network methodology for the present study, conducted the main statistical analysis, drafted the initial manuscript, and approved the final manuscript as submitted. N.U. supervised the methodology development and code writing, interpreted data, reviewed and revised the initial manuscript, and approved the final manuscript as submitted. M.K., R.S., and Z.Z. verified the underlying data, reviewed and revised the initial manuscript, and approved the final manuscript as submitted. J.M. collected the study data, reviewed and revised the initial manuscript, and approved the final manuscript as submitted. C.C. and K.H. conceptualized the study, obtained funding, supervised the statistical analysis, reviewed and revised the initial manuscript, and approved the final manuscript as submitted. All authors contributed to the article and approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
N.U. is an employee of Microsoft, as denoted in the authorship. However, the scope of this work is strictly within research exploration and does not involve any Microsoft product or commercial sponsorship. The remaining authors have no conflicts of interest relevant to this article to disclose.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ooka, T., Usuyama, N., Shibata, R. et al. Integrated-omics analysis with explainable deep networks on pathobiology of infant bronchiolitis. npj Syst Biol Appl 10, 93 (2024). https://doi.org/10.1038/s41540-024-00420-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41540-024-00420-x
This article is cited by
-
Escaping the forest: a sparse, interpretable, and foundational neural network alternative for tabular data
npj Artificial Intelligence (2026)