Abstract
The reaction-diffusion equation is widely used in mathematical models of cancer. The calibration of model parameters based on limited clinical data is critical to using reaction-diffusion equation simulations for reliable predictions on a per-patient basis. Here, we focus on cell-level data as routinely available from tissue biopsies used for clinical cancer diagnosis. We analyze the spatial architecture in biopsy tissues stained with multiplex immunofluorescence. We derive a two-point correlation function and the corresponding spatial power spectral distribution. We show that this data-deduced power spectral distribution can fit the power spectrum of the solution of reaction-diffusion equations that can then identify patient-specific tumor growth and invasion rates. This approach allows the measurement of patient-specific critical tumor dynamical properties from routinely available biopsy material at a single snapshot in time.
Similar content being viewed by others
Introduction
Personalized treatment requires the right treatment at the right time for the right patient. However, there are multiple barriers to achieving these goals. Most standard-of-care clinical therapies are derived from clinical trials that aim to identify the treatment that improves the average desired outcome (local control, recurrence-free survival, disease-specific survival, overall survival, etc.) for the total population. Mean outcome, however, means that about half of the trial patient population has a better-than-average outcome, while the other half’s outcome is worse. There is a dire need to improve therapy for patients who are not average1. We must know critical tumor properties to understand how an individual tumor responds to treatment. There are multiple sources of patient-specific data—albeit with different temporal resolutions—including, but not limited to, patient demographics (including age, sex, race, and socio-economic status), medical history, patient-reported outcomes from medical surveys, blood samples (or liquid biopsies), radiology, and tissue biopsies typically for pathologic evaluation. Tumor dynamics, however, are currently not measurable with static pre-treatment routine clinical tests. This is the focus of the mathematical modeling technique developed herein.
Mathematical oncology, a burgeoning field of cancer research, seeks to develop quantitative methods to help understand, predict, and advance cancer biology and clinical oncology2,3,4,5. The explicit mathematical formulation of assumptions, analytical or numerical solution of the derived equation or systems of equations, and interpretation of the results within specified premises allow different hypotheses to be generated and tested6. For a mathematical model to make predictions, it is crucial to calibrate model parameters, validate model calibration on an independent data set, and test its predictive power on known outcomes7. Historical population data can train model parameter distributions, but the model and its parameters must be tailored to individual patients to help individualize cancer care. Thus, a significant barrier to model development, calibration, and validation is limited longitudinal clinical data to estimate patient-specific tumor dynamics before, during, and after treatment.
Among the most basic metrics of tumor dynamics are cell proliferation and the invasion rate into adjacent physiological tissue8. These are also critical parameters for mathematical models. One of the earliest and simplest models to simulate tumor growth and spread is the reaction-diffusion (R-D) equation, Fisher’s9, or Kolmogorov, Petrovsky, and Piskunov equation10. Gatenby and Gawlinski used the R-D equation to describe the tumor’s spatial spread and temporal development within the normal tissue. The model predicted a hypocellular interstitial gap between the growing tumor and the receding normal tissue, which was later confirmed in experimental studies and clinical tissue samples11.
The R-D equation has become omnipresent in mathematical oncology, often in multi-population model systems, during the past 25 years12,13,14,15,16,17,18,19. Swanson and colleagues demonstrated that different diffusion speeds in gray and white matter in the brain can simulate realistic glioma growth dynamics to help identify the locoregional spread of the tumor that may be below radiology detection thresholds20. In later work, Swanson demonstrated that the glioma R-D growth and invasion rates could be estimated from pre-treatment MRI features to predict survival21 and the efficacy of radiotherapy in individual glioblastoma patients22,23, as well as the effects of hypoxia on growth rates and recurrence location24. Enderling et al. have demonstrated that the R-D equation can be used to simulate the extent of cancer cells’ diffusion into adjacent tissue, which could help evaluate the therapeutic benefits of whole breast irradiation or targeted intraoperative radiotherapy25.
In addition to clinical data, cell diffusivity and cell proliferation rates can be inferred from cell scratch assay data26 and in vivo tumor growth dynamics27. With spatially resolved imaging data, extended R-D models can include anisotropy in different tissue regions28 and explicit consideration for molecularly distinct subpopulations29. Machine learning methods have been demonstrated to increase computational efficiency to enable numerical simulations of the R-D equation on richer data sets30. Integration of machine learning with the R-D equation also enables accurate predictions of variation in cell density of glioblastoma patients using multiparametric MRI31.
While multiple MRI features at multiple time points and rich pre-clinical data have been demonstrated to calibrate the R-D parameters in glioma patients, this approach cannot be used in many other clinical cancers without routine MRI imaging. Here, we hypothesized that routinely collected tissue biopsy data could be used to calibrate the R-D equation’s tumor growth and invasion parameters for individual patients. We demonstrate a method using spatial statistics and spectral analysis that permits an estimate of tumor growth and invasion rates when applied to histological images from breast cancer tissue biopsies.
Results
Different R-D equation parameters yield different power spectral densities (PSD)
We solve the R-D equation with arbitrary dimensionless parameters \(\hat{\gamma }=\{\mathrm{1,2,3}\}\) and \({\widehat{\mathcal{D}}}=\{{1,2,3}\}\) for an arbitrary time and space dimension. The time unit may be one day for tumor growth dynamics, and the corresponding parameter units for the R-D Eq. 1/day and μm2/day, respectively. Intuitively, a larger growth rate parameter, \(\gamma\), yields higher tumor densities, and a more significant diffusion rate, \({\mathcal{D}}\), leads to more spatially spread, invasive tumors. Thus, the most considerable localized tumor burden is simulated with a high \(\gamma\) and low \({\mathcal{D}}\) values (Fig. 1a). Analysis of the corresponding power spectral densities (PSD) at any given time (see Methods) reveals that variation in proliferation rates have a characteristic signature at low wavenumbers (intercept with the y-axis, Fig. 1b). In comparison, the influence at high wavenumbers is determined by variation in the diffusion coefficient (intercept with the x-axis) (Fig. 1c).
a Numerical solution to the R-D equation of cancer cell density \(\rho (\hat{x},\hat{y},\hat{t})\) at an arbitrary time for different unitless proliferation and diffusion rates. b The power spectral density (PSD) for different growth rates, \(\hat{\gamma }\), for the same diffusion rate \({\widehat{\mathcal{D}}}\) = 2. c The power spectral density (PSD) for different diffusion rates, \({\widehat{\mathcal{D}}}\), for the same growth rate \(\hat{\gamma }\) = 2. The mathematical hat symbol above each variable and parameter indicates arbitrary units without specific biological reference.
Biopsy tissue spatial statistics and power spectrum analysis calibrate reaction-diffusion equation parameters
We analyzed the spatial position of each individual cancer cell in three different tumor ROI, \({\mathcal{T}}\), (Fig. 2) for each patient using spatial statistics (Fig. 3). The 2-point correlation function (see Methods) reveals high correlations at low radii \(r=\sqrt{{x}^{2}+{y}^{2}+{z}^{2}}\) around each cancer cell (r < 50 μm), which decays into randomness with increasing distances from each cell (r\(\ge\)50 μm) (Fig. 3b). The corresponding power spectrum densities (see Methods) exhibit the highly characteristic “shoulder profile” as the PSD of the continuous R-D equation above (Fig. 3c). The PSD of the continuous R-D equation can be fit to the average PSD of the different \({\mathcal{T}}\) for each patient (Fig. 4). The comparison of the different tumor growth rates averaged per patient, \(\hat{\gamma }\), and tumor invasion rates averaged per patient, \({\widehat{\mathcal{D}}}\), shows a highly heterogeneous distribution of mean growth rates skewed towards slower growth and a narrow distribution of mean invasion rates (Fig. 5a). The corresponding probability distribution of \({\widehat{\mathcal{D}}}\)/\(\hat{\gamma }\) for all \({\mathcal{T}}\) ROI in the analysis is shown in Fig. 5b. The high patient-specificity of growth rates has been suggested to have significant implications for radiotherapy response, with the growth rate being highly correlated with radiosensitivity parameters22.
A board-certified pathologist selects 3 × 3 regions of interest (ROI), including tumor (\({\mathcal{T}}\)), stroma distant to the tumor (\({\mathcal{S}}\)), and tumor-stroma interface (\({\mathcal{T}}{/}{\mathcal{S}}\)) from breast cancer biopsy tissues for analysis.
a Three different tumor \({\mathcal{T}}\)-ROI for an individual patient (patient 20). b Corresponding 2pCF for each tissue. c Corresponding PSD for each 2pCF.
RD-equation provides excellent fit to the average data PSD.
a Distribution of average diffusion rates, \({\widehat{\mathcal{D}}}\), and growth rates, \(\hat{\gamma }\). b Histogram of all ratios of patient averaged invasion and growth rates, \({\widehat{\mathcal{D}}}\)/ \(\hat{\gamma }\).
Discussion
Understanding tumor growth and invasion dynamics before clinical intervention is paramount to personalizing patient care and improving outcomes. We developed a novel quantitative approach to infer such dynamic characteristics from a single biopsy at patient diagnosis. Patient tissues provide a single-time snapshot of the complex evolution of the cellular density in different spatial locations inside the resection area. The presented approach relies on the “fair representation” hypothesis that histopathology slides are reasonably representative of the clustering properties of the total tumor population. While locoregional properties may be well captured in individual biopsy tissue samples, the overall tumor may be highly heterogeneous. Therefore, the herein-derived methodology must undergo rigorous prospective testing to quantify confidence and uncertainty in the derived parameter values. Of note, however, these are the only collected clinical tissues, and current clinical staging and treatment planning is done on these samples assuming fair representation. Therefore, applying the derived methodology to these data remains a valid approach.
It is conceivable that the exact location of each cancer cell and cancer cell conglomerates in tissue samples is a manifestation of the tumor-intrinsic proliferation and invasion dynamics. We demonstrated that spatial statistical analyses of biopsy tissues could calibrate tumor growth and invasion parameters in the reaction-diffusion equation for individual patients. While the 2-point correlation function and the corresponding power spectrum distribution from clinical tissues are virtually free of assumption, the R-D equation was not derived from the data. It was chosen due to its simplicity and prevalence in mathematical oncology, and because of the interpretability of equation parameters with biological hallmarks of cancer processes. There is no a priori justification for the choice and form of the equation if not the practical analytical tractability, especially in Fourier space. While more physically involved approaches exist, such as Navier Stokes approaches as well as more complex non-linear mathematical models with numerous variables, numerical investigation of the power spectral density (which easily captures several orders of magnitude in wavelength) would involve multiscale physical parametrization that is not justified by the resolution and quantity of routinely collected data at cancer diagnosis. Thus, the focus on the R-D equation aligns with available clinical data and is poised to advance predictive quantitative oncology frameworks and help guide clinical decision-making. If additional data become available to help calibrate and validate additional model parameters, more complex models may be considered to inform radiosensitivity for individual patients before further therapy.
While prospective validation against clinical outcome data is the gold standard to confirm the herein presented methodology, validation against other data would provide further support of this novel approach. Relative abundance of tumor cell proliferation markers, such as KI6732,33, could validate lower versus higher proliferation rates in different patient tissue samples. Similarly, relative abundance of cell adhesion molecules34 and epithelial to mesenchymal transition (EMT) markers35,36 could validate the invasion rates abstracted in the presented methodology. As the major motivation for the presented work is to abstract model parameters to inform treatment sensitivity and treatment response, in particular radiation therapy, the abstracted model parameters could be compared against molecular signatures of radiosensitivity, such as RSI37,38,39 or GARD40,41. Additionally, the theoretical concept that we propose herein, could be validated in more hypothetical scenarios. The R-D equation could be solved numerically, and derived population densities would inform discrete cell counts at specific spatial locations. Such in silico spatial cell maps can then serve as input for the proposed methodology, and the abstracted parameters could be validated against the numerical solution ground truth parameters. This will also allow for testing of model robustness against increasing noise levels in the discretization of the continuous R-D equation solution. For further stochasticity, one can simulate tumor growth and invasion with agent-based models42,43,44,45 and input such single cell simulation results into the proposed methodology.
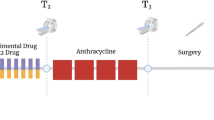
This study’s analyzed tissues and model-calibrated parameters cannot be correlated with clinical outcome. Patients were either treated with neoadjuvant hypofractionated radiation (followed by surgery) or intraoperative radiotherapy. Inclusion of surgery in the treatment plan prevents correlation of R-D growth and invasion parameters with locoregional control or disease-free survival. It is conceivable that targeted local aggressive therapies (e.g., surgery or radiation therapy) would be most successful for a patient with a high proliferative index and a low diffusivity, as previously demonstrated for glioblastoma46. Seminal work by Rockne and Swanson demonstrated that R-D equation-derived tumor growth rates correlate with patient-specific radiosensitivity parameters and clinical radiation response dynamics22. Vice versa, systemic treatments (e.g., immunotherapy, chemotherapy) would be warranted for patients with higher diffusion rates, indicating long-range tissue invasion and an increased risk of metastasis.
The presented approach represents a fundamental step toward integrating quantitative methods into clinical decision-making to improve treatment responses and outcomes47. A possible prospective clinical validation study of the developed approach is to classify patients as high or low risk for locoregional radiation failure based on the R-D equation parameter ratio, \(\rho /{\mathcal{D}}\), as demonstrated for glioblastoma radiation response23. Such predictive biomarker can be synergized with other radiation response signatures37,39,40,48 to arrive at multimodal biomarkers to safely escalate or de-escalate radiation therapy for each individual patient. In addition to risk classification, the power of mechanistic modeling includes simulation of radiation response to different radiation doses and dose fractionations13,25,49,50,51,52. With a calibrated R-D equation, it is conceivable to simulate N = 1 clinical trials in a digital twin, and predict the most successful radiation protocol on a per patient basis1,53,54,55.
Our analysis suggests a narrow distribution of diffusion coefficients, \({\mathcal{D}}\), compared to the wider spread of proliferation rates, \(\gamma\). The nearly homogenous diffusion coefficient across all patients may reflect the selection of early-stage breast cancer patients with localized, non-invasive disease. For more invasive and diffusive diseases, such as glioblastoma, a wider distribution in diffusion coefficients compared to proliferation rates has been demonstrated56. Following the correlation of proliferation rates with radiosensitivity, a wide inter-patient heterogeneity of radiosensitivity has been demonstrated for many cancers, including breast cancer48.
While previous mathematical models and parameter abstraction methodologies require clinical measurements at multiple time points to calibrate tumor dynamics parameters21,57, this is the first solution that requires clinical data from only a single time point. Future steps will need the correlation of R-D equation-derived parameters with clinical outcomes, for which datasets with significant differences in clinical outcomes will be necessary. Herein, we developed the concept based on breast cancer tissue analysis. Future work will include pan-cancer analysis of calibrating tumor growth and invasion parameters with spectral-spatial analysis of cancer biopsy tissues to understand the methodology’s range and limits of applicability.
Methods
Patient cohort
\(44\) early-stage breast cancer patients were included in this analysis (NCT03137693: Preoperative Stereotactic Ablative Body Radiotherapy (SABR) for Early-Stage Breast Cancer; Pro00044616, Advarra: Immunobiology of ER+ and ER- Breast Cancer Tumors). Tissue collection and analysis was approved by the Moffitt Cancer Center Institutional Review Board. The IRB granted a Waiver of Consent per 45 CFR 46.116(d) and a Waiver of HIPAA Authorization per 45 CFR 164.512(i)(2) because the study is retrospective in nature and did not affect the way patients were treated. All data analyzed were collected as part of routine clinical care, with no additional interventions or procedures conducted for the research. A 5 μm thick unstained tumor slide from a formalin-fixed paraffin-embedded tumor block was acquired from patients’ untreated tissue biopsies and analyzed by the Moffitt Cancer Center tissue core and digital imaging laboratory. Formalin-fixed and paraffin-embedded (FFPE) tissue samples were immunostained using the AKOAYA Biosciences OPAL TM 7-Color Automation IHC kit (Waltham, MA) on the BOND RX autostainer (Leica Biosystems, Vista, CA). The OPAL 7-color kit uses tyramide signal amplification (TSA)-conjugated to individual fluorophores to detect various targets within the multiplex assay. Sections were baked at 65 C for one hour, and then transferred to the BOND RX (Leica Biosystems). All subsequent steps (ex., deparaffinization, antigen retrieval) were performed using an automated OPAL IHC procedure (AKOYA). OPAL staining of each antigen occurred as follows: heat-induced epitope retrieval (HIER) was achieved with Citrate pH 6.0 buffer for 20 min at 95 °C before the slides were blocked with AKOYA blocking buffer for 10 min. Then slides were incubated with primary antibody, CD68 (CST, D4BAC, 1:300, dye 520) at RT for 60 min followed by OPAL HRP polymer and one of the OPAL fluorophores during the final TSA step. Individual antibody complexes are stripped after each round of antigen detection. This was repeated five more times using the following antibodies; CD8 (DAKO, C8/144B, HIER-EDTA pH 9.0, 1:100, dye540), CD4 (CM, EP204, HIER- EDTA pH 9.0, 1:100, dye570), CD3 (Thermofisher, SP7, HIER-EDTA pH 9.0, 1:500, dye 570), FOXP3 (ABCAM, 236 A/E7, HIER- EDTA pH 9.0, 1:500, dye650), and PCK (DAKO, AE1/AE3, HIER- Citrate pH 6.0, 1:200, dye690). After the final stripping step, DAPI counterstain is applied to the multiplexed slide and is removed from BOND RX for coverslipping with ProLong Diamond Antifade Mountant (ThermoFisher Scientific). All slides were imaged with the Vectra®3 Automated Quantitative Pathology Imaging System, and the exact (x,y) position for each individual cell was abstracted. Three Regions of Interest (ROI) were selected by a pathologist, including tumor (\({\mathcal{T}}\)), stroma distant to the tumor (\({\mathcal{S}}\)), and tumor and stroma interface (\({\mathcal{T}}{/}{\mathcal{S}}\)) (Fig. 2). Here, we focus only on the tumor ROI, \({\mathcal{T}}\), and the cancer cell staining DAPI + PCK.
Reaction-diffusion equation and 2-point correlation function
The diffusion of a cluster of cancer cells with density \(\rho =\rho \left({\boldsymbol{r}},t\right)\) at position \({\boldsymbol{r}}\) and at a time \(t\) is described by the combination of the continuity equation without source and sink terms as \({\partial }_{t}\rho +{\boldsymbol{\nabla }}.{\boldsymbol{\varphi }}=0\) (with \(\partial\) being the partial derivative and \({\boldsymbol{\nabla }}\) the gradient operator) and Fick’s first law of proportionality between flux \({\boldsymbol{\varphi }}\) and density gradient \({\boldsymbol{\nabla }}\rho\), \({\boldsymbol{\varphi }}={\boldsymbol{-}}{\mathcal{D}}{\mathscr{.}}{\boldsymbol{\nabla }}\rho\), to get (for a generic anisotropic flow) \({\partial }_{t}\rho =-{\boldsymbol{\nabla }}.\left({\mathcal{D}}{{.}}{\boldsymbol{\nabla }}\rho \right)\) where \({\mathcal{D}}\) is the diffusion coefficient matrix. In a 3D system of reference \(S\left({\boldsymbol{O}},{\boldsymbol{r}}=\left\{x,y,z\right\}\right)\), with \({r}^{2}={x}^{2}+{y}^{2}+{z}^{2}\) being the squared radial direction from an arbitrary origin \({\boldsymbol{O}}\), we can write the diffusion equation for the locally homogeneous and isotropic case as \({\partial }_{t}\rho ={\mathcal{D}}{{\boldsymbol{\nabla }}}^{2}\rho ,\) where \({{\boldsymbol{\nabla }}}^{2}\) is the Laplacian operator. Green functions offer a natural solution for cell-like source points, even in the presence of an explicit exponential growth factor \(\gamma\), i.e., \({\partial }_{t}\rho ={\mathcal{D}}{\nabla }^{2}\rho +\gamma \rho\) of interest here, in the form:
This approach follows exponential tumor growth without carrying capacity constraints. We assume that the cells are spread over a small 3D volume \({d}^{3}{\boldsymbol{r}}\) so that, as long as this volume is asymptotically smaller than the cell migration movement per unit time, i.e., to the first order in time, it holds on a linear scale \(d{\boldsymbol{r}}\prec O\left(\sqrt{{\mathcal{D}}{dt}}\right)\), we can generalize Eq. (1) as \({d}^{3}\rho \left({\boldsymbol{r}},t\right)={\rho }_{0}{d}^{3}{\boldsymbol{r}}{\left(4\pi {\mathcal{D}}t\right)}^{-\frac{3}{2}}{e}^{-\frac{{\|{\boldsymbol{r}}\|}^{2}}{4{\mathcal{D}}t}+\gamma t}.\)
In the case of \(n\) different cells \({d}^{3}\rho \left({\boldsymbol{r}},t\right)=\sum _{i}{\left(4\pi {\mathcal{D}}t\right)}^{-\frac{3}{2}}{\rho }_{0i}{e}^{-\frac{{\|{{\boldsymbol{r}}}_{i}\|}^{2}}{4{\mathcal{D}}t}+\gamma t}{d}^{3}{\boldsymbol{r}}\) or, in the passage to the continuous limit, we get the classical heat-equation result that we are going to exploit here for the R-D equation:
The last term in the equation is of interest here. The cell concentration at the generic time \(t\) is the convolution, denoted here by “*” of the initial concentration with a Gaussian, that by borrowing the terminology from Astronomy58 we call R-D point-spread-function \({f}_{{\rm{RDPS}}}\). Here, cell concentration at \({\mathbf{+}}{\mathbf{\infty}}\) is assumed to be null. Furthermore, taking constant temperature and pressure in the tissue sample, we justify \({\mathcal{D}}\) to be constant and elaborate on the \({f}_{{\rm{RDPS}}}\) as the function that carries the temporal dependence.
To proceed further, we adopt the “fair representation” hypothesis59, that the sampled tissues are reasonably representative of the total tumor population, to proceed with a mean-field solution for the R-D equation. By performing this Gibbs average60, we assume that the average volume is large enough to contain sufficient cells to perform statistical analyses. Still, it is small enough to neglect large-scale gradients. In this case, we obtain from Eq. (1):
where we define the average over the time of the R-D spread function as:
with \(\Delta t\) sufficiently small that accounts for the non-equilibrium dynamics of diffusion and proliferation required by the average in \({\rho }_{0}\). Finally, by considering the non-normalized autocorrelation of the average cell concentration \(\left\langle \rho \right\rangle\), we obtain
where the initial density is non-correlated, and the correlation function reduces to its variance \({\sigma }_{\rho }\). The biological implication is that diffusing cells enhances cell proliferation—a visualization of loss of space and contact inhibition. Therefore, these reactions are stochastically correlated, and Eq. (4) assumes cell diffusion is the only mechanism by which cells become spatially correlated. The autocorrelation of \({f}_{{\rm{RD}}}\) captures this spatial correlation thus not contradicting Eq. (2) as ergodicity does not imply mixing considering our definition of averages.
Power spectrum of the reaction-diffusion equation
The R-D equation’s analytical solution is unavailable for arbitrary reaction terms; therefore, working with power spectral densities is more accessible than with 2-point correlation functions to connect cancer diffusion with its spatial distribution. Performing Eq. (3) on Eq. (4), we obtain \({f}_{{\rm{RD}}}\) as
where \({\rm{erfc}}()\) is the complementary error function and \(\iota\) the complex unit. Finally, considering Eq. (4) and Eq. (5) together with the definition of power spectral density, we obtain
where for easier comparison with Eq. (13) we normalized with \({P}_{0}\) to the zero-wavelength by computing \({\int_{{{\mathbb{R}}}^{3}}}P\left({\boldsymbol{k}}\right){d}^{3}{\boldsymbol{k}}\).
The 2-point correlation function of spatial cell data
We implement the 2-point correlation function (2pCF) as a spatial statistic estimator for discretized cell data61,62,63,64,65. The spatial n-point autocorrelation function \({c}_{\textit{np}}\) is defined as the probability of finding n cells at coordinates \({{\boldsymbol{x}}}_{1}\), \({{\boldsymbol{x}}}_{2}\), …, \({{\boldsymbol{x}}}_{n}\) in a suitably defined reference frame \(S\left({\boldsymbol{O}},{\boldsymbol{x}}\right)\) centered in \({\boldsymbol{O}}\) with \({{\boldsymbol{x}}}_{i}={\left\{{x}_{1},{x}_{2},{x}_{3}\right\}}_{i}\) as spatial coordinates for the \(i=1,\,2,\,3,\ldots ,n\). We formalize \({c}_{{np}}\) as \({c}_{{np}}=\left\langle \delta \left({{\boldsymbol{x}}}_{1}\right),\delta \left({{\boldsymbol{x}}}_{2}\right),\ldots ,\delta \left({{\boldsymbol{x}}}_{n}\right)\right\rangle\). We note the significant differences between npCf and cooccurrence45, aside from normalization factors, the pixeled/grid-free definition of the npCf and the possibility to capture both geometry and statistical nature of the images analyzed without gray-levels definition. Because of the connection with the continuous R-D equation, we implement the following definition for the 2pCF:
with \(\left\langle \rho \right\rangle =n\) the number of cells for volume unit, and \(\Delta t\) is a time interval. Because the same tissue section cannot be sampled twice, we consider the time dependence of the autocorrelation. Therefore, Eq. (7) simplifies to:
where \(r=\|{\boldsymbol{r}}\|{\boldsymbol{\equiv }}\|{{\boldsymbol{x}}}_{{\boldsymbol{1}}}{\boldsymbol{-}}{{\boldsymbol{x}}}_{{\boldsymbol{2}}}\|\). Then, if the probability of finding a cell in the volume \({dV}\) is \({dP}={ndV}\), the average number of cells in the finite volume spanned by the tissue ROI is \(\left\langle N\right\rangle ={nV}\), and the joined probability of finding two cells (say cell 1 and cell 2) at a given distance \(r\), is
If the cell distribution is a 3D random Poisson point process, then the probability of finding cells in \(d{V}_{1}\) and \(d{V}_{2}\) are independent, i.e., \({c}_{2{\rm{p}}}=0\). If the cell positions are correlated, \({c}_{2{\rm{p}}}\, > \,0\), and if the positions are anticorrelated \({c}_{2{\rm{p}}}\in \left[-{1,0}\right]\).
To account for the 3D overlapping of cells in the spatial analysis of the 2pCF, we tested Eq. (8) with several kernels, boundary conditions, and bandwidth, always finding convergence (but not a coincidence) of results.
Power spectrum of spatial cell data
The Fourier transform for the distribution of cells with a density \(\rho \left({\boldsymbol{x}},t\right)={\left(\left\langle \rho \right\rangle V\right)}^{-1}\sum _{i}\delta \left({\boldsymbol{x}}-{{\boldsymbol{x}}}_{i}\left(t\right)\right)\) is
with wavenumber \(k=\|{\boldsymbol{k}}\|\), with \({\|\,\|=\|\,\|}_{2}\) Euclidean norm, \({\boldsymbol{k}}=\{{{\boldsymbol{k}}}_{{\boldsymbol{x}}},{{\boldsymbol{k}}}_{{\boldsymbol{y}}},{{\boldsymbol{k}}}_{{\boldsymbol{z}}}\}\), where the barycenter of the ith cell is located at \({{\boldsymbol{x}}}_{i}\) and \({e}^{\iota {\boldsymbol{k}}{{\cdot }}{{\boldsymbol{x}}}_{{\boldsymbol{i}}}}\) is periodic in \(V\) (again here \(\iota\) is the complex imaginary unit, “⋅” the inner product). We slice the volume \(V\) (we refer to the slice and its volume with the same symbol \(V\) without loss of generality) into infinitesimal cells with unitary function \({{\boldsymbol{1}}}_{{dV}}\) (\({{\boldsymbol{1}}}_{{dV}}=1\) if the cell is in the volume \(V\), or \({{\boldsymbol{1}}}_{{dV}}=0\) if it is not). Therefore Eq. (10) is equivalent to
The averaged two-cell contribution to the spectrum of wavelengths, say cell \(1\) in \(d{V}_{1}\) and cell 2 in \(d{V}_{2}\), reads (note how because \(\delta \left({\bf{x}}\right)\) is real, the complex-conjugate \({\delta }_{{\bf{k}}}^{* }={\delta }_{-{\bf{k}}}\)):
because \(\langle {{\boldsymbol{1}}}_{1}{{\boldsymbol{1}}}_{2}\rangle ={n}^{2}d{V}_{1}d{V}_{2}(1+{c}_{2{\rm{p}}})\) as results of considering Eq. (9) (with \({{\boldsymbol{x}}}_{12}\) Euclidean distance between cell 1 and 2). Because Fourier components belong to different \({\boldsymbol{k}}\) are statistically independent, the integrals in the previous vanish if \({\boldsymbol{k}}{\;{\ne}\;}{{\boldsymbol{k}}}^{{{{\prime} }}}\), while \({\boldsymbol{k}}={{\boldsymbol{k}}}^{{{{\prime} }}}\) yields the spectrum, or power spectral density, PSD, of the cell distribution:
for \({\boldsymbol{k}}\,{{\ne }}\,{\boldsymbol{0}}\), and \(\langle {|{\delta }_{0}|}^{2}\rangle ={\int_{{{\mathbb{R}}}^{3}}}\frac{{d}^{3}{\boldsymbol{x}}}{V}{c}_{2{\rm{p}}}+\frac{1}{{nV}}\) otherwise. The power spectrum measures the mean number of neighbors over a random Poisson distribution within a distance of \({{ \sim }}{k}^{-1}\) from a randomly chosen cell. The extra term \({\left({nV}\right)}^{-1}\) is due to shot noise (a contribution visible as white noise \({P}\left(k\right)\propto {k}^{0}\)). If the cell distribution is Poisson, then \({\int }_{{{\mathbb{R}}}^{3}}\frac{{d}^{3}{\textbf{x}}}{V}{c}_{2{\rm{p}}}\left(r\right){e}^{\iota {\textbf{k}} \cdot {\textbf{x}}}=0\), but the Fourier modes have a not null variance \(\langle {|{\delta }_{k}|}^{2}\rangle ={({nV})}^{-1}\) due to the discreteness. This classical result66,67 is often coupled with filter-design windows functions commonly implemented in power spectrum determination to optimize its analytical treatment, which we also tested with consistent results (next section). Nuttall-softening length33 will be preferred if necessary.
Modulation analysis
We can normalize the 2pCF to the unitary sphere in \({{\mathbb{R}}}^{3}\) to test the impact of the most common non-parametric kernel function \(\kappa\), wherefrom signal-analysis literature, we tested the uniform (default in this work) \(\kappa \left(u\right)=\frac{1}{2}\), triangular \(\kappa \left(u\right)=1-\left|u\right|\), parabolic \(\kappa \left(u\right)=\frac{3}{4}\left(1-{u}^{2}\right)\) with support \(\left|u\right|\le 1\), quadratic (biweight) \(\kappa \left(u\right)=\frac{15}{16}{\left(1-{u}^{2}\right)}^{2}\) over \(\left|u\right|\le 1\), cosine \(\kappa \left(u\right)=\frac{\pi }{4}\cos \left(\frac{\pi }{2}u\right)\) with support \(\left|u\right|\le 1\), Gaussian, SemiCircle, and Triweight \(\kappa \left(u\right)=\frac{35}{32}{(1-{u}^{2})}^{3}\) over \({|u|} \le 1\). Although many powerful parametric kernels might help the fitting process, we avoid arbitrary parameter introductions. By following the scheme in65, we can detect a range of interest in the bandwidth, and the impact of the window function on the final result is generally minor (Fig. 6). Finally, the radial distribution function \({c}_{2p}\) is generally computed without uncertainty bars attached. \({c}_{2p}\) is indeed an average over many different measurements of the same tissue tumor ROI, \({\mathscr{T}}\). Therefore, the relatively small standard error in the mean (because of analyses across many cells in the same ROI) could be worked out. Advanced bootstrap simulations found some attempts to account for this error in an astronomical context68. Another approach, different from what we have done, works on binned data, thus obtaining error bars as standard errors65.
Analysis demonstrates no significant difference between different window functions.
Isotropy analysis
Not every image satisfies the hypothesis of slight deviation from isotropy for several reasons (sampling size, tumor clustering grade, etc.), as in the single thick dots ROI in the bottom left of panel 1 of Fig. 7. The corresponding rose-wind plot is in the central panel. Where necessary, we analyze the original ROI copied several times (9 copies in the left panel) until isotropy distribution is achieved (corresponding rose-wind plot in the right panel). Note how this poor approach assumes that the 2D isotropic distribution correctly represents the unknown 3D isotropic distribution. Unfortunately, no 3D information is available to inform a 3D diffusion tensor.
The original (bottom left in the left plot) is with darker dots. The corresponding wind plot in the central panel. The distribution is replicated (3 lighter gray copies, 9 lighter gray copies etc.) until isotropy is reached at a tolerable level (20% of the overall distribution).
Data availability
All data generated and analyzed during this study are available from the corresponding author upon reasonable request.
Code availability
The Mathematica codes are available from the corresponding author upon reasonable request.
References
Yankeelov, T. E. et al. Designing clinical trials for patients who are not average. iScience 27, 108589 (2024).
Araujo, R. A history of the study of solid tumour growth: the contribution of mathematical modelling. Bull. Math. Biol. 66, 1039–1091 (2004).
Anderson, A. R. A. & Quaranta, V. Integrative mathematical oncology. Nat. Rev. Cancer 8, 227–234 (2008).
Altrock, P. M., Liu, L. L. & Michor, F. The mathematics of cancer: integrating quantitative models. Nat. Rev. Cancer 15, 730–745 (2015).
Enderling, H., Alfonso, J. C. L., Moros, E., Caudell, J. J. & Harrison, L. B. Integrating Mathematical Modeling into the Roadmap for Personalized Adaptive Radiation Therapy. Trends Cancer 5, 467–474 (2019).
Enderling H. & Wolkenhauer O. Are all models wrong? Computational and Systems Oncology. 2021;1. https://doi.org/10.1002/cso2.1008.
Brady, R. & Enderling, H. Mathematical Models of Cancer: When to Predict Novel Therapies, and When Not to. Bull. Math. Biol. 81, 3722–3731 (2019).
Hanahan, D. & Weinberg, R. A. The Hallmarks of Cancer. Cell 100, 57–70 (2000).
Fisher, R. A. The Wave of Advance of Advantageous Genes. Ann. Eugen. 7, 355–369 (1937).
Kolmogorov AP, I. & Piskunov, N. Studies of the Diffusion with the increasing quantity of the substance; its application to a biological problem. Mosc. Univ. Math. Bull. 1, 1–26 (1937).
Gatenby, R. A. & Gawlinski, E. T. A reaction-diffusion model of cancer invasion. Cancer Res 56, 5745–5753 (1996).
Anderson ARAC, M. A. J., Newman, E. L., Steele, R. J. C. & Thompson, A. M. Mathematical modelling of tumour invasion and metastasis. J. Theor. Med. 2, 129–154 (2000).
Enderling, H., Chaplain, M. A. J., Anderson, A. R. A. & Vaidya, J. S. A mathematical model of breast cancer development, local treatment and recurrence. J. Theor. Biol. 246, 245–259 (2007).
Ramis-Conde, I., Chaplain, M. A. J. & Anderson, A. R. A. Mathematical modelling of cancer cell invasion of tissue. Math. Computer Model. 47, 533–545 (2008).
Stamper, I. J., Owen, M. R., Maini, P. K. & Byrne, H. M. Oscillatory dynamics in a model of vascular tumour growth - implications for chemotherapy. Biol. Direct 5, 27 (2010).
Andasari, V., Gerisch, A., Lolas, G., South, A. P. & Chaplain, M. A. J. Mathematical modeling of cancer cell invasion of tissue: biological insight from mathematical analysis and computational simulation. J. Math. Biol. 63, 141–171 (2010).
Massey, S. C., Assanah, M. C., Lopez, K. A., Canoll, P. & Swanson, K. R. Glial progenitor cell recruitment drives aggressive glioma growth: mathematical and experimental modelling. J. R. Soc. Interface 9, 1757–1766 (2012).
Bertolusso, R. & Kimmel, M. Modeling Spatial Effects in Early Carcinogenesis : Stochastic Versus Deterministic Reaction-Diffusion Systems. Math. Model. Nat. Phenom. 7, 245–260 (2012).
Hormuth, D. A. et al. A mechanically coupled reaction–diffusion model that incorporates intra-tumoural heterogeneity to predict in vivo glioma growth. J. R. Soc. Interface 14, 20161010 (2017).
Swanson, K. R., Alvord, E. C. & Murray, J. D. A quantitative model for differential motility of gliomas in grey and white matter. Cell Prolif. 33, 317–329 (2000).
Swanson, K. R., Rostomily, R. C. & Alvord, E. C. A mathematical modelling tool for predicting survival of individual patients following resection of glioblastoma: a proof of principle. Br. J. Cancer 98, 113–119 (2008).
Rockne, R. et al. Predicting the efficacy of radiotherapy in individual glioblastoma patientsin vivo:a mathematical modeling approach. Phys. Med. Biol. 55, 3271–3285 (2010).
Massey, S. C. et al. Image-based metric of invasiveness predicts response to adjuvant temozolomide for primary glioblastoma. PLoS ONE 15, e0230492 (2020).
Curtin, L., Hawkins-Daarud, A., van der Zee, K. G., Swanson, K. R. & Owen, M. R. Speed Switch in Glioblastoma Growth Rate due to Enhanced Hypoxia-Induced Migration. Bull. Math. Biol. 82, 43 (2020).
Enderling, H., Anderson, A. R. A., Chaplain, M. A. J., Munro, A. J. & Vaidya, J. S. Mathematical modelling of radiotherapy strategies for early breast cancer. J. Theor. Biol. 241, 158–171 (2006).
Johnston, S. T., Shah, E. T., Chopin, L. K., Sean McElwain, D. L. & Simpson, M. J. Estimating cell diffusivity and cell proliferation rate by interpreting IncuCyte ZOOM™ assay data using the Fisher-Kolmogorov model. BMC Syst. Biol. 9, 38 (2015).
Rutter, E. M. et al. Mathematical Analysis of Glioma Growth in a Murine Model. Sci. Rep. 7, 2508 (2017).
Swan, A., Hillen, T., Bowman, J. C. & Murtha, A. D. A Patient-Specific Anisotropic Diffusion Model for Brain Tumour Spread. Bull. Math. Biol. 80, 1259–1291 (2018).
Morris, B. et al. Identifying the spatial and temporal dynamics of molecularly-distinct glioblastoma sub-populations. Math. Biosci. Eng. 17, 4905–4941 (2020).
Viguerie, A., Grave, M., Barros, G. F., Lorenzo, G., Reali, A. & Coutinho, A. L. G. A. Data-Driven Simulation of Fisher–Kolmogorov Tumor Growth Models Using Dynamic Mode Decomposition. J. Biomech. Eng. 144, 121001 (2022).
Gaw, N. et al. Integration of machine learning and mechanistic models accurately predicts variation in cell density of glioblastoma using multiparametric MRI. Sci. Rep. 9, 10063 (2019).
Inwald, E. C. et al. Ki-67 is a prognostic parameter in breast cancer patients: results of a large population-based cohort of a cancer registry. Breast Cancer Res Treat. 139, 539–552 (2013).
Ellis, M. J. et al. Ki67 Proliferation Index as a Tool for Chemotherapy Decisions During and After Neoadjuvant Aromatase Inhibitor Treatment of Breast Cancer: Results From the American College of Surgeons Oncology Group Z1031 Trial (Alliance). J. Clin. Oncol. 35, 1061–1069 (2017).
Ramis-Conde, I., Drasdo, D., Anderson, A. R. & Chaplain, M. A. Modeling the influence of the E-cadherin-beta-catenin pathway in cancer cell invasion: a multiscale approach. Biophys. J. 95, 155–165 (2008).
Pearson, G. W. Control of Invasion by Epithelial-to-Mesenchymal Transition Programs during Metastasis. J. Clin. Med 8, 646 (2019).
Jolly, M. K., Ware, K. E., Gilja, S., Somarelli, J. A. & Levine, H. EMT and MET: necessary or permissive for metastasis? Mol. Oncol. 11, 755–769 (2017).
Eschrich, S. et al. Systems biology modeling of the radiation sensitivity network: a biomarker discovery platform. Int J. Radiat. Oncol. Biol. Phys. 75, 497–505 (2009).
Torres-Roca, J. F. et al. Prediction of radiation sensitivity using a gene expression classifier. Cancer Res. 65, 7169–7176 (2005).
Alfonso, J. C. L. et al. Tumor-immune ecosystem dynamics define an individual Radiation Immune Score to predict pan-cancer radiocurability. Neoplasia 23, 1110–1122 (2021).
Scott, J. G. et al. A genome-based model for adjusting radiotherapy dose (GARD): a retrospective, cohort-based study. Lancet Oncol. 18, 202–211 (2017).
Scott, J. G. et al. Pan-cancer prediction of radiotherapy benefit using genomic-adjusted radiation dose (GARD): a cohort-based pooled analysis. Lancet Oncol. 22, 1221–1229 (2021).
Enderling, H., Anderson, A. R., Chaplain, M. A., Beheshti, A., Hlatky, L. & Hahnfeldt, P. Paradoxical dependencies of tumor dormancy and progression on basic cell kinetics. Cancer Res 69, 8814–8821 (2009).
Enderling, H., Hlatky, L. & Hahnfeldt, P. Migration rules: tumours are conglomerates of self-metastases. Br. J. Cancer 100, 1917–1925 (2009).
Poleszczuk, J. & Enderling, H. A High-Performance Cellular Automaton Model of Tumor Growth with Dynamically Growing Domains. Appl Math. (Irvine) 5, 144–152 (2014).
Poleszczuk, J., Macklin, P. & Enderling, H. Agent-Based Modeling of Cancer Stem Cell Driven Solid Tumor Growth. Methods Mol. Biol. 1516, 335–346 (2016).
Baldock, A. L. et al. Patient-Specific Metrics of Invasiveness Reveal Significant Prognostic Benefit of Resection in a Predictable Subset of Gliomas. PLoS ONE 9, e99057 (2014).
Pasetto S., Gatenby R. A. & Enderling H. Bayesian Framework to Augment Tumor Board Decision Making. JCO Clinical Cancer Informatics. 2021:508-517. https://doi.org/10.1200/cci.20.00085.
Grass, G. D. et al. The Radiosensitivity Index Gene Signature Identifies Distinct Tumor Immune Microenvironment Characteristics Associated With Susceptibility to Radiation Therapy. Int J Radiat Oncol Biol Phys 113, 635–647 (2022).
Bruningk, S. C. et al. Intermittent radiotherapy as alternative treatment for recurrent high grade glioma: a modeling study based on longitudinal tumor measurements. Sci. Rep. 11, 20219 (2021).
Poleszczuk, J., Walker, R., Moros, E. G., Latifi, K., Caudell, J. J. & Enderling, H. Predicting Patient-Specific Radiotherapy Protocols Based on Mathematical Model Choice for Proliferation Saturation Index. Bull. Math. Biol. 80, 1195–1206 (2018).
Prokopiou, S. et al. A proliferation saturation index to predict radiation response and personalize radiotherapy fractionation. Radiat. Oncol. 10, 159 (2015).
Zahid, M. U. et al. Dynamics-Adapted Radiotherapy Dose (DARD) for Head and Neck Cancer Radiotherapy Dose Personalization. J. Pers. Med 11, 1124 (2021).
Aherne, N. J., Dhawan, A., Scott, J. G. & Enderling, H. Mathematical oncology and it’s application in non melanoma skin cancer - A primer for radiation oncology professionals. Oral. Oncol. 103, 104473 (2020).
Laubenbacher, R. et al. Toward mechanistic medical digital twins: some use cases in immunology. Front Digit Health 6, 1349595 (2024).
Wu, C. et al. Integrating mechanism-based modeling with biomedical imaging to build practical digital twins for clinical oncology. Biophys. Rev. (Melville) 3, 021304 (2022).
Harpold, H. L. P., Alvord, E. C. & Swanson, K. R. The Evolution of Mathematical Modeling of Glioma Proliferation and Invasion. J. Neuropathol. Exp. Neurol. 66, 1–9, https://doi.org/10.1097/nen.0b013e31802d9000 (2007).
Harshe, I., Enderling, H. & Brady-Nicholls, R. Predicting Patient-Specific Tumor Dynamics: How Many Measurements Are Necessary? Cancers 15, 1368, https://doi.org/10.3390/cancers15051368 (2023).
Ahi, K. Mathematical Modeling of THz Point Spread Function and Simulation of THz Imaging Systems. IEEE Trans. Terahertz Sci. Technol. 7, 747–754, https://doi.org/10.1109/tthz.2017.2750690 (2017).
Grafström, A. & Schelin, L. How to Select Representative Samples. Scand. J. Stat. 41, 277–290, https://doi.org/10.1111/sjos.12016 (2013).
de Alfaro, V., Fubini, S. & Furlan, G. Gibbs average in the functional formulation of quantum field theory. Phys. Lett. B 105, 462–466, https://doi.org/10.1016/0370-2693(81)91205-3 (1981).
Ridgway RI, M. O., Machiraju, R., Huang, K., Wenzel, P., deBruin, A. & Leone, G. Image Segmentation with Tensor–Based Classification of N–Point Correlation Functions. MICCAI Workshop on Medical Image Analysis with Applications in. Biology 13, 156–166 (2006).
Mosaliganti, K. et al. Tensor classification of N-point correlation function features for histology tissue segmentation. Med. Image Anal. 13, 156–166 (2009).
Cooper, L. S., Machiraju, R. & Huang, K. Two-Point Correlation as a Feature for Histology Images: Feature Space Structure and Correlation Updating. 2010 IEEE Computer Society Conference on Computer Vision and Pattern Recognition - Workshops. 2010:79–86. https://doi.org/10.1109/CVPRW.2010.5543453.
Cooper, L. A. D., Saltz, J. H., Catalyurek, U. & Huang, K. Acceleration of Two Point Correlation Function Calculation for Pathology Image Segmentation. Proc IEEE Int Conf Healthc Inform Imaging Syst Biol 2011, 174–181 (2011).
Binder, B. J. & Simpson, M. J. Spectral analysis of pair-correlation bandwidth: application to cell biology images. Royal Society Open Science. 2015;2. 140494 https://doi.org/10.1098/rsos.140494.
Peebles, P. J. E. Statistical Analysis of Catalogs of Extragalactic Objects. I. Theory. The Astrophysical Journal. 1973;185. https://doi.org/10.1086/152431.
Epanechnikov, V. A. Non-Parametric Estimation of a Multivariate Probability Density. Theory Probab. Its Appl. 14, 153–158, https://doi.org/10.1137/1114019 (1969).
Mo, H. J., Jing, Y. P. & Boerner, G. On the error estimates of correlation functions. The Astrophysical Journal. 1992;392. https://doi.org/10.1086/171444.
Acknowledgements
This work was supported in part by NIH/NCI grant U01CA244100 and the Tissue Core Facility at the H. Lee Moffitt Cancer Center & Research Insitute, an NCI-designated Comprehensive Cancer Center (P30-CA076292). This work was supported by The JAYNE KOSKINAS TED GIOVANIS FOUNDATION FOR HEALTH AND POLICY, a Maryland private foundation dedicated to effecting change in health care for the public good. The opinions, findings, and conclusions or recommendations expressed in this material are those of the authors and do not necessarily represent the official views of the JAYNE KOSKINAS TED GIOVANIS FOUNDATION FOR HEALTH AND POLICY, its directors, officers, or staff. Special thanks to Jordan Pietz, MA, CMI from the MD Anderson Biomedical Visualization team for his assistance in with the visual artwork in Fig. 2.
Author information
Authors and Affiliations
Contributions
S.P., M.M., R.D., and H.E. conceptualized the study. M.M., M.R., R.D., S.P., H.E. assembled the data. S.P. and H.E. developed and designed the methodology, performed computational and statistical analysis, and visualized the results. S.P., M.M., M.Z., M.R., R.G., R.D., H.E. wrote the original draft of the manuscript. P.S. reviewed and edited the manuscript. All authors have read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pasetto, S., Montejo, M., Zahid, M.U. et al. Calibrating tumor growth and invasion parameters with spectral spatial analysis of cancer biopsy tissues. npj Syst Biol Appl 10, 112 (2024). https://doi.org/10.1038/s41540-024-00439-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41540-024-00439-0
This article is cited by
-
Calibrating tissue level PDE models of ligand dynamics using single cell and spatial transcriptomics data
npj Systems Biology and Applications (2026)
-
Pattern Formation as a Resilience Mechanism in Cancer Immunotherapy
Bulletin of Mathematical Biology (2025)