Abstract
Production stability remains a major challenge in Chinese hamster ovary (CHO) cell-based therapeutic protein manufacturing, particularly during extended passaging where the underlying mechanisms of instability are not fully understood. Thus, in this study, we leveraged multivariate data analysis (MVDA) and flux balance analysis (FBA) with explainable AI (xAI) to mechanistically characterize the phenotypic differentiation between early (EP) and late passage (LP) of CHO cultures. Although EP and LP reached comparable peak viable cell densities, LP cultures exhibited a ~35% reduction in peak IgG titers and increased lactate and ammonia accumulation. Subsequent MVDA of temporal exometabolite profiles identified the exponential growth phase as the primary window of divergence, allowing us to interrogate metabolic rewiring via an FBA-xAI approach. This revealed that EP cells preferentially directed acetyl-CoA towards fatty acid biosynthesis to support proliferation. In contrast, LP prioritized oxidative stress mitigation by upregulating the trans-sulfuration pathway for de novo cysteine and glutathione synthesis while exhibiting heightened TCA cycle activity to maintain energy homeostasis. Overall, these mechanistic insights uncover a passage-associated shift from biosynthetic activity toward redox maintenance and identify the cysteine-glutathione axis as a critical metabolic lever for enhancing long-term stability and productivity in CHO cell culture.
Similar content being viewed by others
Introduction
Chinese hamster ovary (CHO) cells are the predominant mammalian host for biopharmaceutical manufacturing, producing more than 70% of therapeutic proteins1 due to their ability to perform human-like post-translational modifications2. Decades of advances in cell line and bioprocess developments have substantially improved key performance metrics such as cell growth, productivity and titer, and have also enhanced product quality and process robustness3,4,5. As industrial demand for high-producing cell lines has increased, cell line development (CLD) has relied on strong selection system combined with single-cell cloning, and more recently, with targeted integration strategies, to isolate CHO clones with robust and desirable productivity6,7. However, the intrinsic plasticity of CHO cells can cause clonal instability during CLD, leading to losses in productivity and product quality8,9,10. Thus, considerable effort is devoted to ensuring long-term stability of selected clones by performing extended passaging before being used in large-scale manufacturing11,12.
Prolonged passaging and long-term culture (LTC), however, introduce genetic and epigenetic alterations, including gene mutations and chromosomal rearrangements, which lead to metabolic drift and phenotypic change13,14,15. Previous studies have characterized these passage-associated changes primarily using transcriptomic and metabolomic analyses, identifying mechanistic contributors such as recombinant gene loss, genome-wide transcriptional imbalance, and metabolic rewiring events that can diminish productivity in certain CHO cell lines16,17,18,19. More recently, LTC has also been shown to affect product quality attributes, including charge heterogeneity, glycosylation patterns, and host-cell-protein composition, alongside morphological alterations and mitochondrial dysfunction that signify broader metabolic and redox imbalance20. Although these studies collectively reveal extensive reprogramming of CHO cell behavior during LTC, the underlying mechanisms remain elusive and do not resolve how specific metabolic pathways are rewired to generate phenotypic and production instability.
To address this gap, flux balance analysis (FBA) of genome-scale models (GEMs) has been widely utilized as a systems-level framework for mechanistically linking genotype to phenotype by quantitatively characterizing cellular behavior21,22. Since the first CHO GEM reconstruction23, various FBA-based approaches have guided rational cell line engineering4, optimized culture processes24, and also supported systematic media reformulation by integrating multivariate data analysis (MVDA)25,26. Most practical applications have focused on upstream process development27, whereas limited studies have explored passage-dependent effects on CHO cell stability during LTC22. While transcriptomic studies have highlighted shifts in lysosomal activity, redox balance, and lipid metabolism with increased passage number17, comprehensive mechanistic analyses linking these genetic changes to metabolic flux alterations remain lacking. This gap constrains targeted strategies such as tailored media supplements or cell engineering interventions to mitigate production instability.
Interestingly, recent integration of machine learning (ML) and artificial intelligence (AI) with the FBA framework has enhanced the exploration of complex metabolic networks in CHO cell cultures28,29. However, their adoption has been limited by the “black-box” nature of ML-based models30,31, which obscures how specific metabolites and reactions drive phenotypic outcomes. In this regard, explainable AI (xAI) could address this limitation by revealing which input features, such as fluxes of specific reactions, most strongly influence outcome variables (e.g., passage number, growth rate and productivity). For example, Wu et al. combined xAI with FBA to identify high-impact metabolic pathways and gene targets, such as succinate dehydrogenase, for enhancing ethanol production in S. cerevisiae32. Nonetheless, systematic frameworks for translating such macroscopic and metabolic insights into actionable, reaction-level targets in complex mammalian systems like CHO cells remain underdeveloped.
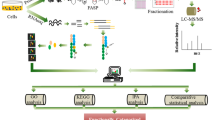
In this study, we present a comprehensive approach that integrates MVDA, xAI and flux-sum analysis within an enzyme capacity-constrained FBA (ecFBA) framework33 to investigate the metabolic underpinnings of phenotypic drift in CHO cells during LTC (Fig. 1). By continuously subculturing CHO cells for over 30 passages, we established early- and late-passage cultures and conducted four layers of analysis: (1) phenotypic profiling revealed decreased IgG production and elevated accumulation of toxic metabolites at higher passages; (2) MVDA identified key amino acids with altered uptake and secretion patterns distinguishing early and late passage cultures; (3) ecFBA coupled with SHapley Additive exPlanations (SHAP) highlighted critical metabolic reactions contributing to observed phenotypic changes; (4) in-depth flux analysis uncovered rewiring in central carbon and amino acid metabolism underlying productivity loss. Collectively, this framework demonstrates how xAI-enhanced metabolic modeling can pinpoint passage-induced reprogramming in CHO cells and guide rational interventions to improve production stability.
This framework is composed of (1) experimental cell culture characterization, (2) comparative phenotypic and metabolic profiling, (3) multivariate data analysis (MVDA), (4) enzyme-constrained flux balance analysis (ecFBA), (5) explainable AI (xAI)-driven interpretation, and (6) in-depth flux analysis to elucidate passage-dependent metabolic shifts. (Partial elements created with BioRender.com).
Results
Culture profiling for phenotypic characterization of CHO cells
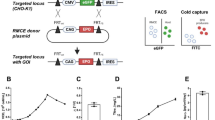
Since passaging up to 30 passages indicated the emergence of passage-dependent phenotypic drift specifically in growth rate and IgG production, two independent working cell banks (WCB) were derived from a single master cell bank (MCB) at passages 7 and 23. To evaluate the phenotypic effect of passaging, cells from each WCB were subcultured for an identical number of passages (i.e., +3 and +5 passages), resulting in four cultures comprising two early passage (EP) cultures (P10 and P12) and two late passage (LP) cultures (P26 and P28) for comparison (Fig. 2a). Notable differences between EP and LP cultures were observed, particularly in growth, productivity, and byproduct accumulation, whereas glucose consumption and viability remained comparable (Fig. 2b–i). Specifically, as the passage number increased, LP cultures exhibited higher viable cell densities with ~8.5% higher peak VCD than that of EP cultures throughout the exponential phase (Fig. 2b, c). LP showed a higher specific growth rate (SGR) during days 2–4 (0.6009 d−1) compared with EP (0.4667 d−1), indicating increased proliferative capacity in the LP group (Supplementary Fig. 1a). Despite their enhanced growth profiles, LP cultures showed reduced IgG production capacity with lower maximum IgG titer, whereas EP cultures accumulated IgG more steadily (Fig. 2d, e) and maintained elevated productivity for a longer duration (Supplementary Fig. 1a). In contrast, the differences of glucose and viability profiles were marginal (Fig. 2f, g), while LP cultures accumulated higher levels of lactate and ammonia (Fig. 2h, i), which are predominantly known as inhibitory byproducts that can perturb intracellular redox balance and inhibit the capacity of production.
a Schematic of the long-term passaging and banking schedule with asterisks (*) on sampling points. b–i Comparison between early passage (EP; P10 and P12, red symbols/lines/bars) and late passage (LP; P26 and P28, blue symbols/lines/bars) cultures showing: b, c Viable cell density (VCD) and maximum VCD; d, e IgG titer and maximum IgG titer; f Glucose concentration; g Cell viability; h, i Lactate and ammonia concentration profiles. (Partial elements created with BioRender.com).
To investigate whether passage-dependent metabolic reprogramming underlies the observed phenotypic shifts, we next closely examined the extracellular amino acid consumption and secretion profiles (Supplementary Fig. 2). During the exponential phase, EP showed a switch from net aspartate secretion to consumption specific to EP cultures while constant consumption was observed in LP. In addition, EP significantly increased secretion of glycine and proline during exponential growth, suggesting potential alterations within the interconnected aspartate-arginine-glutamine-proline metabolic axis, pathways closely linked to ammonia metabolism (i.e., asparaginase, arginine deiminase, and glutaminase)34. The differentiated engagement of this axis between EP and LP cultures is consistent with the observed disparities in lactate and ammonia accumulation, implying broader rewiring of nitrogen and redox-associated metabolism. Accordingly, these findings underscore the need to investigate how the interplay between amino acid metabolism and byproduct accumulation contributes to the observed phenotypic drift and production instability.
Multivariate data analysis for identification of highlighted metabolites
To systematically explore the metabolic drivers associated with passage-dependent characteristics, we conducted MVDA using the calculated specific consumption and secretion rates (pmol/cell-day) of glucose, lactate, ammonia, and 20 amino acids, over the culture intervals of days 0–2, 2–4, 4–6, 6–8, 8–10, 10–12, 12–14. Firstly, we generated a PLS model with the specific rates of metabolites as X variables and integrated viable cell density (IVCD) as the Y variable. The first PLS component (PLS1) was strongly correlated with sampling time (Spearman correlation = 0.921, p < 1 × 10−20), revealing a clear temporal trajectory that separated the exponential phase (days 0–6) from the stationary/death phase (days 6–14) (Fig. 3a, b). Applying PLS-tree analysis to this model further resolved the samples into six subgroups (G1–G6), with G1–G3 corresponding predominantly to the stationary/death phase and G4–G6 to the exponential phase. Interval-wise comparison of the PLS1 medians revealed how EP and LP diverged across culture intervals (Fig. 3c). Specifically, a reduced separation was observed during days 4–6 (+0.118), indicating a metabolic turning point before widening again in later intervals. Guided by the combined phenotypic and statistical evidence of the SGR peak (days 2–4) and the metabolic turning point (days 4–6), we targeted these specific intervals to systematically investigate the passage-dependent metabolic divergence and its subsequent stabilization.
a, b Score plot and corresponding loading plot from Partial Least Squares (PLS) analysis separating the overall metabolic profiles into distinct subgroups (G1–G6). c Comparison of interval-wise PLS1 medians between EP (red) and LP (blue) cultures. d Orthogonal PLS Discriminant Analysis (OPLS-DA) loading plot visualizing the maximal separation achieved between EP and LP metabolic profiles. e Variable Importance in Projection (VIP) scores derived from the OPLS model, identifying metabolites significantly contributing to the distinction between EP and LP conditions (dashed line indicates VIP = 1.0 threshold). Asterisks (*) indicate major metabolic drivers (Tyr, Met, and Trp) consistently identified as influential predictors.
To identify key metabolites distinguishing EP and LP cells during selected culture intervals, orthogonal partial least square discriminant analysis (OPLS-DA) was employed (Fig. 3d). The resultant loading plot revealed that EP cells were characterized by higher specific rates of glycine, arginine, aspartate, proline, and methionine, whereas LP cells showed reduced consumption/accumulation rates of serine, histidine, and leucine. Moreover, to further investigate metabolites that are highly associated with the IVCD in early and late passage conditions, OPLS-based VIP scores were obtained from the OPLS models and evaluated, with a cutoff of VIP > 1.0 (Fig. 3e). While ten metabolites (alanine, ammonium, asparagine, cysteine, glutamate, glutamine, glucose, lactate, phenylalanine, and valine) overlapped between the two conditions, tyrosine, methionine, and tryptophan were uniquely identified in EP cultures, suggesting potential shifts in metabolic dependencies or contributions to IVCD during prolonged cultivation. Taken together, these analyses highlighted subsets of amino acids as key drivers distinguishing EP and LP cells and significantly contributing to IVCD. These amino acids are known to participate in various metabolic pathways, including one-carbon metabolism and TCA cycle anaplerosis, which are central to cellular energy production, redox balance, and biomass synthesis2,35,36.
Combining ecFBA with xAI for targeting metabolic reactions
In order to further elucidate metabolic behaviors during the observed phenotypic changes, we have exploited FBA framework with xAI (see Methods). Although previous MVDA identified statistically significant metabolites, such a broad spectrum of metabolites and the inherent complexity of metabolic networks render it difficult to pinpoint the key drivers of metabolic shifts. Therefore, to mechanistically examine how metabolism changes with increasing passage number, we performed ecFBA33 across 14 experimental conditions, derived from seven biological samples (duplicates of P10, P26, P28, and a single P12) measured over two specific culture intervals (days 2–4 and 4–6). For each condition, the five flux distributions with the highest growth prediction accuracy were selected (Supplementary Fig. 3), yielding a total of 70 flux distributions (see Methods). Using these flux distribution sets, we exploited SHAP analysis to quantify the contribution of individual reactions to predicting passage number (Fig. 4), thereby allowing us to systematically filter the extensive reaction network of CHO-GEM and subsequently identify the key metabolic drivers distinguishing EP and LP cultures. They include the reactions in central carbon metabolism (i.e., pyruvate metabolism and TCA cycle) as well as amino acid pathways such as cysteine, methionine, arginine, and proline metabolism. To corroborate these reaction-level insights from a metabolite-centric perspective, we additionally performed flux-sum analysis on the selected flux distributions37. Notably, the metabolites identified as major drivers based on VIP scores (tyrosine, methionine, and tryptophan) were consistently highlighted as influential predictors of passage number in the flux-sum SHAP model (Supplementary Fig. 4).
The 15 top-ranked reactions based on overall predictive importance (left), quantified as the mean absolute SHAP value (mean(|SHAP value|), gray bars). The vertical yellow dotted line indicates the threshold value applied for prioritizing reactions, determined as the average value of absolute summed non-zero SHAP value of each reaction. The positive (+) and negative (−) indicators to the left of the reaction names denote the directionality of the mean contribution, distinguishing reactions primarily associated with LP (class 1) and EP (class 0) phenotypes, respectively. The corresponding SHAP summary plot (right) illustrates the distribution of SHAP values for individual flux simulation samples. The x-axis indicates the SHAP value (impact on model prediction), while the color scale represents the relative flux value for that reaction in each sample.
Given the overall contributions of metabolic reactions assessed via SHAP analysis (Fig. 4, gray bars in the left panel), we sought to identify reactions that serve as dependable and robust indicators of the passage phenotype. To this end, we developed a composite scoring system that integrates feature importance with a linearity metric (R²), enabling the prioritization of reactions whose flux magnitudes exhibit a predictable relationship with the phenotypic outcome (see Methods). Applying this prioritization showed that the reactions such as mitochondrial L-lactate dehydrogenase (LDH_Lm), tyrosine transaminase (TYRTA) and phosphatidylethanolamine N-methyltransferase (PETOHMm_cho), despite ranking among the top 15 in overall impact, displayed broad and complex distributions of feature values (fluxes) relative to their SHAP scores (Fig. 4, right panel). Although such patterns likely reflect underlying biological complexity, such as multi-factor interactions or regulatory effects, their non-linear dependencies make them less suitable as straightforward candidates for subsequent in-depth flux analyses. Consequently, these reactions were deprioritized or excluded as primary targets for the subsequent in-depth flux analysis. On the other hand, reactions including cystathionine γ-lyase (CYSTGL), cystathionine β-synthase (CYSTS) and mitochondrial alanine transaminase (ALATA_Lm), were identified as robust candidates. Their high flux values consistently contribute to the classification of late passage cultures, whereas PHCDym was characterized by low flux values in most LP-classified samples.
In-depth flux distribution analysis of the targeted pathways
Leveraging the key reactions identified through xAI-ecFBA analysis, we conducted an in-depth flux analysis. Here, we selected the culture intervals of days 4–6 as this transition period captures the critical moment where cells actively reconfigure their metabolic networks after peak growth divergence during days 2–4. In addition, the prioritized reactions clustered in central carbon metabolism and amino acid pathways linked to cysteine-methionine and arginine-proline metabolism, we implemented in-depth analysis of flux distribution focused on those pathways. In glycolysis (Fig. 5a), glucose consumption varied across samples, with P10 exhibiting the highest uptake and P28 the lowest, reflecting their trends in downstream glycolytic fluxes. In line with the SHAP results, LDH_Lm showed elevated flux in EP cultures, resulting in elevated pyruvate synthesis and increased acetyl-CoA production. Moreover, fluxes through 3-hydroxyisobutyryl-CoA hydrolase (3HBCOAHLm) and L-threonine 3-dehydrogenase (THDm) were increased, further supplying more acetyl-CoA via valine and threonine catabolism. While EP cultures utilized acetyl-CoA primarily for fatty acid metabolism to synthesize structural precursors such as malonyl-CoA, HMG-CoA, and acyl-ACP to support rapid membrane biogenesis, LP cultures exhibited greater TCA cycle activity. This suggests that EP cultures prioritize biosynthetic processes, while LP cultures shift toward oxidative phosphorylation to sustain energy production under prolonged culture conditions, reflecting a trade-off between growth and maintenance.
Key metabolites and reactions of ‘central carbon metabolism’ and ‘methionine and cysteine metabolism’ are depicted in (a) and (b), respectively. Adjacent plots show flux distributions for indicated reactions at P10 (red), P12 (dark red), P26 (blue) and P28 (dark blue), and mean flux values for P10, P12 (red and dark red), P26, and P28 (blue and dark blue).
Additionally, elevated fluxes through CYSTS and CYSTGL in LP cultures were observed (Fig. 5b), indicating that the cells were reprogrammed from extracellular cysteine uptake to de novo cysteine synthesis via the trans-sulfuration pathway to support glutathione production. This metabolic reprogramming is driven by an adaptive response to prioritize glutathione production for survival against ROS38,39,40. However, as cysteine serves as a rate-limiting precursor for both glutathione biosynthesis and disulfide bond formation in the endoplasmic reticulum (ER), this redirection of cysteine, despite minimal changes in extracellular cysteine levels, can reduce the pool available for efficient IgG folding41. In addition, the increased flux through CYSTGL not only channels the metabolites toward glutathione synthesis but also produces ammonia as an obligatory byproduct, representing a metabolic trade-off between redox homeostasis maintenance and toxic byproduct generation. This may, in turn, contribute to differences in productivity. Concurrently, increased ALATA_Lm flux utilizes alanine to synthesize TCA cycle intermediates to sustain ATP/NADH production, while suppressed HISDC/FTHFDH activity limits folate-mediated NADPH regeneration. Taken together, these flux adjustments propose a reprogrammed metabolism in which central-carbon and amino-acid reactions are repurposed to prioritize antioxidant defense rather than biosynthetic throughput required for high-level IgG production. This complex interplay provides a mechanistic explanation for the productivity loss observed in long-term cultures, importantly highlighting rational metabolic levers for media reformulation, such as trade-offs in cysteine utilization and alterations in one-carbon (C1) metabolism.
Discussion
In this study, we observed key phenotypic differences of prolonged passaging, wherein later passage cultures revealed a 35% drop in peak IgG production, despite exhibiting enhanced peak VCD and average SGR. To mechanistically understand these phenotypic shifts, we employed explainable AI-driven flux balance analysis to rationally identify key metabolic reactions that are highly potent contributors to the observed phenotypic drift. The initial search space with thousands of reactions was narrowed down to a concise set of predictive features, thereby facilitating the identification and interpretation of metabolic levers attributed to the distinct profiles of EP and LP cultures. Subsequent in-depth analysis revealed that LP cultures undergo metabolic reprogramming, shifting from extracellular cysteine uptake toward cysteine synthesis via trans-sulfuration pathways (CYSTS, CYSTGL) and reallocating acetyl-CoA from biosynthesis to oxidative phosphorylation. This observation not only provides mechanistic explanations for productivity loss under LP culture, but also lays a rational foundation for media reformulation strategies, pointing specifically toward supplementing cysteine-sparing compounds, alternative antioxidants, or one-carbon donors to mitigate oxidative stress and sustain productivity42,43.
Oxidative stress, which occurs when cellular defense systems fail to counterbalance the generation of free radicals, has been recognized as a major contributor to impaired cell growth, reduced productivity and increased protein microheterogeneity38,44,45. Especially, during prolonged passage and culture, cells continuously modulate genes involved in lysosomal function and lipid metabolism, which can lead to an accumulation of reactive oxygen species46,47. To survive this detrimental environment, cells upregulate their antioxidant defenses17,19, which shows good agreement with our finding that late-passage cells specifically reprogram cysteine metabolism toward antioxidant synthesis as a primary mechanism to mitigate oxidative stress. Additionally, production instability in long-term culture is a multifactorial issue which can also be contributed by other mechanisms, such as the direct loss or silencing of the recombinant gene16,48 and epigenetic and transcriptional alterations49. Therefore, future investigations employing transcriptomic and genomic analyses are thus required to assess holistic understanding of the interplay between metabolic, genetic, and epigenetic factors that drives production instability. By integrating our metabolic model with the expression dynamics of genes involved in cell cycle progression and DNA repair, thus, our systematic framework can be extended to provide more comprehensive and robust solutions for both phenotype and production instability in CHO cells during the LTC.
While our ecFBA–SHAP framework successfully identified robust metabolic levers, a potential limitation of the current scoring framework arises from its prioritization of linearity. This approach inherently deprioritizes reactions such as LDH_Lm and TYRTA, which exhibit high feature importance but complex, non-linear distributions. However, in specific metabolic contexts, these reactions exert substantial influence on cell culture performance by serving as pivotal metabolic switches on glycolytic and amino acid pathways. For instance, LDH_Lm which our analysis captured as exhibiting clear flux divergence between EP and LP cultures but was deprioritized due to its non-linear behavior, modulates glycolytic overflow and mitochondrial redox balance, thereby mediating lactate shift in mammalian cell culture50,51. Similarly, TYRTA facilitates the rewiring of tyrosine metabolism, thereby regulating amino acid homeostasis known to affect both cell growth kinetics and specific productivity52,53,54. Despite their biological significance, the incorporation of a linearity metric (R2) in our scoring framework was a deliberate strategy to highlight metabolic drivers that can be more readily translated into actionable engineering targets following in-depth flux analysis. To maintain flexibility of the scoring system, a tunable coefficient (k) is also introduced to balance between impact magnitude and linearity. This ensures that future applications of the framework can capture non-linear regulatory behaviors, particularly when analyzing larger or more diverse datasets.
Recently, systematic frameworks integrating data-driven approach of omics data with mechanistic metabolic models represent a state-of-the-art strategy for investigating complex cellular metabolism and developing upstream bioprocess22, such as media development25,26,55 and metabolic reprogramming observed during CHO cell long-term culture9. This synergy allows researchers to correlate patterns identified in experimental data with plausible underlying flux distributions simulated mechanistically. Nonetheless, a significant challenge remains because the phenotypic shifts in long-term culture are driven by the intricate, non-linear interplay of the entire metabolic network—rather than by single, independent reactions—making it difficult to pinpoint the most influential flux changes from high-dimensional simulation data. Accordingly, xAI approach, particularly SHAP based on cooperative game theory56, offers a robust approach for providing quantitative, interpretable metrics for the contribution of each individual input feature. Moreover, the application of AI/ML as well as xAI to move beyond the limitations of “black-box” data-driven models is an accelerating trend, not only in biomedical fields57,58 such as drug and cancer diagnostics but also emerging in bioprocess modeling59. For example, a recent study successfully utilized SHAP to deconstruct the complex interplay between media components and critical quality attributes in CHO cell cultures, enabling rational media design60. In this regard, our integrated ecFBA-SHAP approach provides a systematic methodology for translating complex flux data into rational, actionable strategies for cell engineering and media reformulation. Thus, the proposed framework could serve as a foundational component in the shift from data-driven analysis toward AI-powered, mechanistic, and explainable decision-making in bioprocessing and biomanufacturing.
Methods
Cell culture and media
A recombinant Immunoglobulin G (IgG) antibody producing CHO-K1 cell line was used in this study. Master cell bank (MCB) vials at passage numbers P7 and P23 were aliquoted and stored as independent WCBs. Selected vials from each WCB were thawed and expanded under identical culture conditions. For long-term stability monitoring, one expansion series was continued up to passage P30, while a decline in productivity was first observed during prolonged expansion at passages P23-P24. For the comparative phenotypic and metabolic analyses reported here, cultures were harvested at defined endpoints: P10 and P12 (early-passage samples originating from the P7 bank) and P26 and P28 (late-passage samples originating from the P23 bank), to ensure that EP and LP groups experienced the same subculture steps, thereby enabling an unbiased comparison of passage-dependent phenotypes. The cells were initially cultured in Lonza powerCHO™2 basal medium supplemented with L-glutamine 200 mM (100×) solution (Cytiva) to a final concentration of 8 mmol. After thawing, cells were passaged to expand the cell population up to passage number 30. For cell passaging and expansion, cells were centrifuged at 400 × g for 5 min using a Thermo Fisher centrifuge during each passage transfer to remove accumulated byproducts. For passage numbers 10 and 26 (P10, P26), cells were seeded at a viable cell density of 5 ×105 cells/mL into a total volume of 100 mL of the supplemented Lonza powerCHO™2 medium in 250 mL Erlenmeyer shake flasks (Corning Life Sciences). For passage numbers 12 and 28 (P12, P28), cells were seeded at a viable cell density of 5 ×105 cells/mL into a total volume of 200 mL of the supplemented with the same medium in 500 mL Erlenmeyer shake flasks. The flasks were placed on an orbital shaker at 120 rpm in a humidified incubator at 5% CO2 in air at 37 °C. For the selected early (P10, P12) and late (P26, P28) passages, dedicated sampling flasks were seeded at a viable cell density of 5 ×105 cells/mL into a total volume of 30 mL of the supplemented with the medium in 150 mL Erlenmeyer shake flasks. All experiments for P10, P12, P26, and P28 were performed in duplicates, however, one replicate of P12 was identified as an outlier, so only a single data set is presented for P12. To ensure that observed metabolic profile differences were attributable to LTC and not to variations in media conditions, identical media were used across all cultures throughout the study. These flasks were established, from which bi-daily samples were collected from day 0 to day 14 to monitor cell growth and other parameters.
Metabolite analyzer and UPLC sample preparation and data acquisition
Cell culture samples were collected for main flasks on day 0, 3, and 4, and sampling flasks with bi-daily manner for cell count and viability determination, which were measured with LUNA-II™ (Logos Biosystems) and every two days for metabolite and amino acid concentrations. Viable cell density (VCD) and viability were measured with Cedex Bio (Roche, CustomBiotech). Glucose, lactate, ammonia, and IgG concentration were measured with Cedex Bio analyzer (Roche CustomBiotech).
For amino acids measurements, supernatant was collected by centrifuging flask culture samples for 3 min at 5000 rpm followed by filtration using 0.2 µm syringe filters to eliminate residual cell debris. The AccQ-Tag Ultra Derivatization kit (Waters, Milford, MA, USA) was used in accordance with the manufacturer’s instructions. A C18 column (AccQ-Tag ultra, 1.7 μm, 2.1 × 100 mm, Waters, Milford, MA, USA) equipped with the ultra-high-performance liquid chromatography system (ACQUITY I-class UPLC, Waters, Milford, USA) was employed for separation of derivatized amino acids. The column was pre-heated and sustained at 55°C during separation and mobile phase A consisted of 5% (v/v) eluent A (AccQ-Tag Ultra, Waters, Milford, MA, USA) in UPLC grade water, while mobile phase B comprise 100% eluent B (AccQ-Tag Ultra, Waters, Milford, MA, USA). A gradient profile of mobile phase was used under a constant flow rate of 0.7 mL/min. A photodiode array detector (Acquity UPLC®, Waters, Milford, USA) was set to 260 nm in 2D mode.
Multivariate data analysis
To determine the media components that contribute to the difference in culture performance over time, we calculated the SGR (day−1) and specific productivity (Qp, pg cell−1 day−1) using the method described in the previous study26. The concentrations of metabolites such as glucose, lactate, ammonia, and 20 amino acids were also converted into cell-specific consumption or secretion rates between each time point. Principal component analysis (PCA) was performed as it represents the stage where early and late passage (EP and LP) conditions begin to cluster closely within conditions in the PCA analysis of the specific rates of measured metabolites. For this purpose, the specific rates of 20 amino acids, glucose, lactate, and ammonia were used as X variables for PCA. Specific rate calculations were performed for the bidaily sampling intervals of days 0–2, 2–4, 4–6, 6–8, 8–10, 10–12, and 12–14, and PCA was conducted on these intervals, as well as on the initial intervals of 0–2, 2–4, and 4–6. Prior to this, OPLS-DA was conducted to identify the differentiating metabolites between EP and LP using the data from days 2–4 and 4–6. Subsequently, to examine the influence of specific metabolite rates (input variables, X) on IVCD (output variable, Y), variable influence on projection (VIP) scores were obtained from batch evolution model (BEM)-based orthogonal partial least square (OPLS) analysis for each passage group with intervals of days 2–4 and 4–6. To identify the key metabolites among the feed media components and narrow down the number of metabolites for further pathway analysis, a VIP score greater than 1.0 was applied for these selected intervals (days 2–4 and 4–6) and exhibited a two-fold difference between EP and LP conditions. These metabolites that met these criteria were considered significant and were analyzed in-depth using a genome-level mechanistic model.
Enzyme-capacity constraint flux balance analysis (ecFBA) and flux-sum analysis
To investigate metabolic states under various conditions, we utilized the iCHO2291 model, a comprehensive reconstruction of the CHO cell metabolic network33, adapted by incorporating experimentally measured uptake and secretion rates as boundary constraints for each condition, with exchange flux bounds for unmeasured or unexpected metabolites set to zero to prevent artificial nutrient uptake, while adjusting the bounds for glucose uptake, lactate secretion, and the biomass reaction to ±10% of their measured rates to accommodate potential experimental errors, a range determined to effectively simulate physiological behavior. In cases where initial bounds for threonine and arginine led to infeasible solutions, threonine was assigned to a minimal uptake rate of −0.001 mmol/gDCW/hr. For P12 culture, arginine bounds were set to the average of duplicate conditions (P12A, P12B); similarly, average specific rates from duplicate experiments served as boundary conditions for passages P10, P26, and P28. Growth rates were predicted by setting the biomass reaction in the iCHO2291 model as the objective function and applying enzymatic capacity constraint flux balance analysis (ecFBA), which, unlike conventional FBA (e.g., FBA, FVA, pFBA), integrates enzyme kinetic constraints like turnover numbers (kcat) alongside standard FBA constraints, enhancing the accuracy of intracellular flux distributions by accounting for enzyme efficiency and enabling exploration of metabolic resource allocation33. For each experimental condition, we performed ecFBA with various maintenance energy constraints on data from days 2–4 and 4–6. To reflect varying maintenance energy needs, the DM_atp reaction was varied from 0.1 to 10.0 mmol/gDCW/hr with 0.1 intervals. Measured metabolites such as 20 amino acids, glucose, lactate and ammonia were constrained as uptake or secretion bounds with experimentally determined flux values with a ± 10% error relaxation (Supplementary Table 1). From each condition, we selected the five sets of flux distributions that best predicted the experimentally measured growth rate (Supplementary Fig. 3). Thus, resulting 70 sets served as the primary dataset for subsequent SHAP analysis. Moreover, flux-sum analysis was implemented to assess compartmentalized metabolite turnover rates for a metabolite-focused perspective37,61 based on the selected ecFBA solutions, resulting in the quantification of the net metabolic contribution towards these experimentally measured exchange fluxes. This allowed for an estimation of the overall intracellular metabolic activity, or ‘indirect metabolic pool’, associated with the uptake and secretion balance of the 20 amino acids, glucose, lactate, and ammonia under each simulated condition. All simulations were executed using the cobrapy package in a Jupyter notebook and COBRA Toolbox version 3.062 in MATLAB 2020a, employing the Gurobi solver for linear programming optimizations.
SHAP analysis
Given the high-dimensional nature of the flux data relative to the sample size (n = 70), we assessed the model’s predictive performance and potential overfitting using a stratified 5-fold cross-validation approach. The RF model consistently achieved an average test Accuracy of 1.0 and an F1-score of 1.0 across the folds (Supplementary Table 2). While this perfect score suggests a high risk of overfitting, it also provides strong evidence that the metabolic flux profiles of EP and LP cultures are highly separable. Thus, the model was deemed suitable for mechanistic interpretation via SHAP analysis rather than generalized prediction. Accordingly, we employed the random forest (RF) algorithm from the open-source Python library Scikit-learn (https://scikit-learn.org).
The interpretability of the developed RF model was further enhanced by employing the SHAP (SHapley Additive exPlanations) framework, a game-theoretic approach that elucidates the output of machine learning models63. SHAP values provide insights into the contribution of each feature (average flux value of each reaction) to the model’s prediction for each sample, acting as equivalent positive and negative weight coefficients. Here, passage 10 and 12 are classified as 0 for y-variable, and 26 and 28 as 1. In a separate analysis utilizing flux-sum analysis, the results for measured metabolites were derived from each of the 70 flux distribution sets to further investigate the relationship between metabolite levels and class of passages. This allowed for highlighting metabolic reactions and metabolites derived from individual fluxes and metabolite-centered flux-sum results.
Given that the resultant flux distributions can exhibit sample-dependent variability, further criteria were applied to prioritize metabolic reactions that exhibit a predictable and consistent relationship between flux magnitude and phenotypic based on the following formula:
Here, SHAPi represents the SHAP values across all samples (i) for the reaction with non-zero SHAP value and R² is the squared Pearson correlation coefficient between the reaction’s flux values and its corresponding SHAP values across all samples, measuring the linearity and consistency of the reaction’s impact on the prediction. Reactions were then prioritized if their Score exceeded a defined threshold. While reactions with non-linear dependencies may be biologically impactful, they were deprioritized in this study to streamline the selection of candidate reactions. The threshold was determined using a factor, denoted as k, which scales the average impact magnitude across reactions. In this study, k was set to 1. Employing a higher value for the threshold factor k would impose a more stringent selection criterion, preferentially identifying reactions with the strongest linear correlation (R²) between flux and predictive contribution.
Data availability
The metabolic model used in this study is publicly available in the BioModels Database (ID: MODEL1912180001). All experimental data and model constraints analyzed during this study are included in this published article and its supplementary information file. The internal scripts used for analysis are not publicly available as they are not designed as a deployable software tool but are available from the corresponding author on reasonable request.
References
Walsh, G. & Walsh, E. Biopharmaceutical benchmarks 2022. Nat. Biotechnol. 40, 1722–1760 (2022).
Pereira, S., Kildegaard, H. F. & Andersen, M. R. Impact of CHO metabolism on cell growth and protein production: an overview of toxic and inhibiting metabolites and nutrients. Biotechnol. J. 13, 1–13 (2018).
Shin, S. W., Kyeong, M. & Lee, J. S. Next-generation cell engineering platform for improving recombinant protein production in mammalian cells. In Cell Culture Engineering and Technology. Cell Engineering, vol 10 (ed. Pörtner, R.) 189–224 https://doi.org/10.1007/978-3-030-79871-0_7 (Springer, 2021).
Hong, J. K., Lakshmanan, M., Goudar, C. & Lee, D.-Y. Towards next generation CHO cell line development and engineering by systems approaches. Curr. Opin. Chem. Eng. 22, 1–10 (2018).
Fouladiha, H. et al. A metabolic network-based approach for developing feeding strategies for CHO cells to increase monoclonal antibody production. Bioprocess Biosyst. Eng. 43, 1381–1389 (2020).
Browne, S. M. & Al-rubeai, M. Selection methods for high-producing mammalian cell lines. Trends Biotechnol. 25, 425–432 (2010).
Lee, J. S., Kildegaard, H. F., Lewis, N. E. & Lee, G. M. Mitigating clonal variation in recombinant mammalian cell lines. Trends Biotechnol 37, 931–942 (2019).
Tharmalingam, T. et al. Characterization of phenotypic and genotypic diversity in subclones derived from a clonal cell line. Biotechnol. Prog. 34, 613–623 (2018).
Dahodwala, H. & Lee, K. H. The fickle CHO: a review of the causes, implications, and potential alleviation of the CHO cell line instability problem. Curr. Opin. Biotechnol. 60, 128–137 (2019).
Dorai, H. et al. Early prediction of instability of chinese hamster ovary cell lines expressing recombinant antibodies and antibody-fusion proteins. Biotechnol. Bioeng. 109, 1016–1030 (2012).
Tihanyi, B. & Nyitray, L. Recent advances in CHO cell line development for recombinant protein production. Drug Discov. Today Technol 38, 25–34 (2020).
O’Flaherty, R. et al. Mammalian cell culture for production of recombinant proteins: a review of the critical steps in their biomanufacturing. Biotechnol. Adv. 43, 107552 (2020).
Wurm, F. & Wurm, M. Cloning of CHO. Cells, Productivity and Genetic Stability—A Discussion. Processes 5, 20 (2017).
Marx, N., Eisenhut, P., Weinguny, M., Klanert, G. & Borth, N. How to train your cell - Towards controlling phenotypes by harnessing the epigenome of Chinese hamster ovary production cell lines. Biotechnol. Adv. 56, 107924 (2022).
Feichtinger, J. et al. Comprehensive genome and epigenome characterization of CHO cells in response to evolutionary pressures and over time. Biotechnol. Bioeng. 113, 2241–2253 (2016).
Bailey, L. A., Hatton, D., Field, R. & Dickson, A. J. Determination of Chinese hamster ovary cell line stability and recombinant antibody expression during long-term culture. Biotechnol. Bioeng. 109, 2093–2103 (2012).
Qian, Y. et al. New insights into genetic instability of an industrial CHO cell line by orthogonal omics. Biochem. Eng. J. 164, 107799 (2020).
Sowa, S. W. et al. Metabolomic and quality data for early and late passages of an antibody-producing industrial CHO cell line. Data Br 33, 106591 (2020).
Torres, M. et al. Long term culture promotes changes to growth, gene expression, and metabolism in CHO cells that are independent of production stability. Biotechnol. Bioeng. 120, 2389–2402 (2023).
Kaur, R., Jain, R., Budholiya, N. & Rathore, A. S. Long term culturing of CHO cells: phenotypic drift and quality attributes of the expressed monoclonal antibody. Biotechnol. Lett. 45, 357–370 (2023).
Suthers, P. F., Foster, C. J., Sarkar, D., Wang, L. & Maranas, C. D. Recent advances in constraint and machine learning-based metabolic modeling by leveraging stoichiometric balances, thermodynamic feasibility and kinetic law formalisms. Metab. Eng. 63, 13–33 (2021).
Park, S.-Y. et al. Driving towards digital biomanufacturing by CHO genome-scale models. Trends Biotechnol. 42, 1192–1203 (2024).
Hefzi, H. et al. A consensus genome-scale reconstruction of Chinese hamster ovary cell metabolism. Cell Syst. 3, 434–443.e8 (2016).
Strain, B., Morrissey, J., Antonakoudis, A. & Kontoravdi, C. Genome-scale models as a vehicle for knowledge transfer from microbial to mammalian cell systems. Comput. Struct. Biotechnol. J. 21, 1543–1549 (2023).
Hong, J. K. et al. Data-driven and model-guided systematic framework for media development in CHO cell culture. Metab. Eng. 73, 114–123 (2022).
Park, S.-Y. et al. Debottlenecking and reformulating feed media for improved CHO cell growth and titer by data-driven and model-guided analyses. Biotechnol. J. 18, 1–14 (2023).
Kavoni, H., Savizi, I. S. P., Lewis, N. E. & Shojaosadati, S. A. Recent advances in culture medium design for enhanced production of monoclonal antibodies in CHO cells: a comparative study of machine learning and systems biology approaches. Biotechnol. Adv. 78, 108480 (2025).
Khaleghi, M. K., Savizi, I. S. P., Lewis, N. E. & Shojaosadati, S. A. Synergisms of machine learning and constraint-based modeling of metabolism for analysis and optimization of fermentation parameters. Biotechnol. J. 16, 2100212 (2021).
Sahu, A., Blätke, M. A., Szymański, J. J. & Töpfer, N. Advances in flux balance analysis by integrating machine learning and mechanism-based models. Comput. Struct. Biotechnol. J. 19, 4626–4640 (2021).
Hassija, V. et al. Interpreting black-box models: a review on explainable artificial intelligence. Cognit. Comput. 16, 45–74 (2024).
Hashizume, T. & Ying, B. W. Challenges in developing cell culture media using machine learning. Biotechnol. Adv. 70, 108293 (2024).
Wu, D. et al. Towards a hybrid model-driven platform based on flux balance analysis and a machine learning pipeline for biosystem design. Synth. Syst. Biotechnol. 9, 33–42 (2024).
Yeo, H. C., Hong, J., Lakshmanan, M. & Lee, D.-Y. Enzyme capacity-based genome scale modelling of CHO cells. Metab. Eng. 60, 138–147 (2020).
Kuo, M., Chen, H., Feun, L. & Savaraj, N. Targeting the proline–glutamine–asparagine–arginine metabolic axis in amino acid starvation cancer therapy. Pharmaceuticals 14, 72 (2021).
Owen, O. E., Kalhan, S. C. & Hanson, R. W. The key role of anaplerosis and cataplerosis for citric acid cycle function. J. Biol. Chem. 277, 30409–30412 (2002).
Yang, M. & Vousden, K. H. Serine and one-carbon metabolism in cancer. Nat. Rev. Cancer 16, 650–662 (2016).
Chung, B. K. S. & Lee, D.-Y. Flux-sum analysis: a metabolite-centric approach for understanding the metabolic network. BMC Syst. Biol. 3, 117 (2009).
Handlogten, M. W., Zhu, M. & Ahuja, S. Intracellular response of CHO cells to oxidative stress and its influence on metabolism and antibody production. Biochem. Eng. J. 133, 12–20 (2018).
Ha, T. K., Hansen, A. H., Kol, S., Kildegaard, H. F. & Lee, G. M. Baicalein reduces oxidative stress in CHO cell cultures and improves recombinant antibody productivity. Biotechnol. J. 13, 1–10 (2018).
Chevallier, V., Andersen, M. R. & Malphettes, L. Oxidative stress-alleviating strategies to improve recombinant protein production in CHO cells. Biotechnol. Bioeng. 117, 1172–1186 (2020).
Feige, M. J. & Hendershot, L. M. Disulfide bonds in ER protein folding and homeostasis. Curr. Opin. Cell Biol. 23, 167–175 (2011).
Mulukutla, B. C. et al. Metabolic engineering of Chinese hamster ovary cells towards reduced biosynthesis and accumulation of novel growth inhibitors in fed-batch cultures. Metab. Eng. 54, 54–68 (2019).
Kundu, P., Beura, S., Mondal, S., Das, A. K. & Ghosh, A. Machine learning for the advancement of genome-scale metabolic modeling. Biotechnol. Adv. 74, 108400 (2024).
Hecklau, C. et al. S-Sulfocysteine simplifies fed-batch processes and increases the CHO specific productivity via anti-oxidant activity. J. Biotechnol. 218, 53–63 (2016).
Kaur, R. & Rathore, A. S. Role of oxidative stress in modulating CHO cell culture performance: Impact on titer and quality attributes of a monoclonal antibody therapeutic. J. Chem. Technol. Biotechnol. 98, 651–660 (2023).
Jaishy, B. & Abel, E. D. Lipids, lysosomes, and autophagy. J. Lipid Res. 57, 1619–1635 (2016).
Su, L.-J. et al. Reactive oxygen species-induced lipid peroxidation in apoptosis, autophagy, and ferroptosis. Oxid. Med. Cell. Longev. 2019, 1–13 (2019).
Kim, M., O’Callaghan, P. M., Droms, K. A. & James, D. C. A mechanistic understanding of production instability in CHO cell lines expressing recombinant monoclonal antibodies. Biotechnol. Bioeng. 108, 2434–2446 (2011).
Veith, N., Ziehr, H., MacLeod, R. A. F. & Reamon-Buettner, S. M. Mechanisms underlying epigenetic and transcriptional heterogeneity in Chinese hamster ovary (CHO) cell lines. BMC Biotechnol. 16, 1–16 (2016).
Mulukutla, B. C., Gramer, M. & Hu, W.-S. On metabolic shift to lactate consumption in fed-batch culture of mammalian cells. Metab. Eng. 14, 138–149 (2012).
Hartley, F., Walker, T., Chung, V. & Morten, K. Mechanisms driving the lactate switch in Chinese hamster ovary cells. Biotechnol. Bioeng. 115, 1890–1903 (2018).
Song, J., Park, S.-Y. & Lee, D.-Y. Characterization and design of dipeptide media formulation for scalable therapeutic production. Appl. Microbiol. Biotechnol. 109, 7 (2025).
Tang, H. et al. Insight into the roles of tyrosine on rCHO cell performance in fed-batch cultures. Appl. Microbiol. Biotechnol. 103, 6483–6494 (2019).
Bröer, S. & Gauthier-coles, G. Amino acid homeostasis in mammalian cells with a focus on amino acid transport. J. Nutr 152, 16–28 (2021).
Hong, J. K. et al. In silico model-based characterization of metabolic response to harsh sparging stress in fed-batch CHO cell cultures. J. Biotechnol. 308, 10–20 (2020).
Lundberg, S. M. et al. From local explanations to global understanding with explainable AI for trees. Nat. Mach. Intell. 2, 56–67 (2020).
Bifarin, O. O. & Fernández, F. M. Automated machine learning and explainable AI (AutoML-XAI) for metabolomics: improving cancer diagnostics. J. Am. Soc. Mass Spectrom. 35, 1089–1100 (2024).
Ding, Q. et al. Explainable artificial intelligence in the field of drug research. Drug Des. Devel. Ther. 19, 4501–4516 (2025).
Rogers, A. W., Vega-Ramon, F., Lane, A., Martin, P. & Zhang, D. Interpretable-AI-based model structural transfer learning to accelerate bioprocess model construction. Biotechnol. Bioeng. 1–13, https://doi.org/10.1002/bit.70026 (2025).
Gangwar, N., Balraj, K. & Rathore, A. S. Explainable AI for CHO cell culture media optimization and prediction of critical quality attribute. Appl. Microbiol. Biotechnol. 108, 308 (2024).
Lakshmanan, M., Kim, T. Y., Chung, B. K. S., Lee, S. Y. & Lee, D. Y. Flux-sum analysis identifies metabolite targets for strain improvement. BMC Syst. Biol. 9, 1–11 (2015).
Heirendt, L. et al. Creation and analysis of biochemical constraint-based models using the COBRA Toolbox v.3.0. Nat. Protoc. 14, 639–702 (2019).
Lundberg, S. & Lee, S.-I. A unified approach to interpreting model predictions. In Advances in Neural Information Processing Systems 4766–4775 http://arxiv.org/abs/1705.07874 (2017).
Acknowledgements
This study was supported by the National Research Foundation of Korea (NRF) (grant number RS-2024-00341312) funded by the Korea government (MSIT), and the Industrial Technology Innovation Program (grant number RS-2025-02305152) funded by the Ministry of Trade, Industry & Energy of the Republic of Korea. The funders played no role in study design, data collection, analysis and interpretation of data, or the writing of this manuscript. Parts of Figs. 1 and 2 were created with BioRender.com.
Author information
Authors and Affiliations
Contributions
D-H.C. performed the formal analysis of the systematic framework, conducted the investigation, and wrote the original draft, including review and editing. S-J.K. conducted the investigation and performed the ML modeling and analysis. J.S.S. performed, and D-H.C. supported the generation of the UPLC measurement dataset. D-H.C., S-J.K., J.S.S., J.H.L., and C-H.P. performed experiments and generated the experimental dataset. S-Y.P. contributed to methodology, conceptualization, and investigation. D-Y.L. provided the conceptualization, supervised the project, contributed to the investigation, and was a major contributor in writing, reviewing, and editing the manuscript. All authors reviewed the findings and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests. Author D.-Y.L. is a Guest Editor for the Collection “Next-Generation Mammalian Cell Bioprocessing: Systems Biology, Synthetic Biology, and Beyond” in npj Systems Biology and Applications. D.-Y.L. was not involved in the journal’s review of, or decisions related to, this manuscript.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Choi, DH., Kim, SJ., Song, J. et al. Exploring CHO cell stability during prolonged passaging via eXplainable AI driven flux balance analysis. npj Syst Biol Appl 12, 36 (2026). https://doi.org/10.1038/s41540-026-00660-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41540-026-00660-z