Abstract
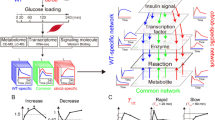
Starvation induces complex metabolic adaptations in skeletal muscle, a key tissue for maintaining energy homeostasis; however, these adaptations are largely impaired in obesity. How obesity alters global metabolic adaptations to starvation in skeletal muscle remains unclear. Here, we analyzed the metabolic adaptations on a trans-omics scale during starvation in skeletal muscle from wild-type (WT) and leptin-deficient obese (ob/ob) mice. We measured multi-omics data during starvation and constructed global trans-omics networks in WT and ob/ob mice. We found that starvation induces “responsiveness” in WT mice, characterized by increases or decreases in key regulator metabolites, including ATP and AMP, as well as enzyme proteins, leading to global regulation of metabolic pathways, which was lost in ob/ob mice. In contrast, during starvation, ob/ob mice exhibit “difference” in comparison to WT mice, manifested by the persistently elevated expression of metabolic enzymes. These features were similarly found in liver, another key metabolic organ. Thus, global loss of responsiveness and elevated enzyme proteins are systemic features of metabolic dysregulation in ob/ob mice.
Similar content being viewed by others
Data availability
The RNA-seq data generated during the current study are available in the DDBJ130 BioProject repository with links to BioProject accession number PRJDB19859. The mass spectrometry proteomics data generated during the current study are available in the jPOST repository131 with links to accession number JPST003499.
Code availability
The code used for the analysis in this paper is available at https://github.com/AmyLibzena/Starvation-Muscle-Analysis.
References
Tang, D. et al. Fasting: From Physiology to Pathology. Adv. Sci. (Weinh.) 10, e2204487 (2023).
Wang, Y. & Wu, R. The Effect of Fasting on Human Metabolism and Psychological Health. Dis. Markers 2022, 5653739 (2022).
Wei, M. et al. Life span extension by calorie restriction depends on Rim15 and transcription factors downstream of Ras/PKA, Tor, and Sch9. PLoS Genet 4, e13 (2008).
Kaeberlein, T. L. et al. Lifespan extension in Caenorhabditis elegans by complete removal of food. Aging Cell 5, 487–94 (2006).
Lee, G. D. et al. Dietary deprivation extends lifespan in Caenorhabditis elegans. Aging Cell 5, 515–24 (2006).
Williams, D. S., Cash, A., Hamadani, L. & Diemer, T. Oxaloacetate supplementation increases lifespan in Caenorhabditis elegans through an AMPK/FOXO-dependent pathway. Aging Cell 8, 765–8 (2009).
Greer, E. L. et al. An AMPK-FOXO pathway mediates longevity induced by a novel method of dietary restriction in C. elegans. Curr. Biol. 17, 1646–56 (2007).
Yang, Y., Zhou, H., Hou, L., Xing, K. & Shu, H. Transcriptional profiling of skeletal muscle reveals starvation response and compensatory growth in Spinibarbus hollandi. BMC Genomics 20, 938 (2019).
Sugasawa, T. et al. Gene Expression Profile Provides Novel Insights of Fasting-Refeeding Response in Zebrafish Skeletal Muscle. Nutrients. 14, 2239 (2022).
Chen, C. et al. Liver Transcriptome Analysis Reveals Energy Regulation and Functional Impairment of Onychostoma sima During Starvation. Mar. Biotechnol. (NY) 25, 247–58 (2023).
Gudiksen, A. et al. Muscle interleukin-6 and fasting-induced PDH regulation in mouse skeletal muscle. Am. J. Physiol. Endocrinol. Metab. 312, E204–E14 (2017).
Collins-Hooper, H. et al. Symmorphosis through dietary regulation: a combinatorial role for proteolysis, autophagy and protein synthesis in normalising muscle metabolism and function of hypertrophic mice after acute starvation. PLoS One 10, e0120524 (2015).
Kvedaras, M., Minderis, P., Cesanelli, L., Cekanauskaite, A. & Ratkevicius, A. Effects of fasting on skeletal muscles and body fat of adult and old C57BL/6J mice. Exp. Gerontol. 152, 111474 (2021).
Zhang, F., Xu, X., Zhou, B., He, Z. & Zhai, Q. Gene expression profile change and associated physiological and pathological effects in mouse liver induced by fasting and refeeding. PLoS One 6, e27553 (2011).
Jia, W. H. et al. Effect of skeletal muscle phenotype and gender on fasting-induced myokine expression in mice. Biochem Biophys. Res Commun. 514, 407–14 (2019).
Ieko, T. et al. Mechanism of skeletal muscle atrophy by muscle fiber types in male rats under long-term fasting stress. Steroids 200, 109328 (2023).
Gartner, U., Goeser, T. & Wolkoff, A. W. Effect of fasting on the uptake of bilirubin and sulfobromophthalein by the isolated perfused rat liver. Gastroenterology 113, 1707–13 (1997).
Bosron, W. F., Crabb, D. W., Housinger, T. A. & Li, T. K. Effect of fasting on the activity and turnover of rat liver alcohol dehydrogenase. Alcohol Clin. Exp. Res. 8, 196–200 (1984).
Bak, A. M. et al. Prolonged fasting-induced metabolic signatures in human skeletal muscle of lean and obese men. PLoS One 13, e0200817 (2018).
Vendelbo, M. H. et al. Fasting increases human skeletal muscle net phenylalanine release and this is associated with decreased mTOR signaling. PLoS One 9, e102031 (2014).
Browning, J. D., Baxter, J., Satapati, S. & Burgess, S. C. The effect of short-term fasting on liver and skeletal muscle lipid, glucose, and energy metabolism in healthy women and men. J. Lipid Res 53, 577–86 (2012).
Kummitha, C.M., Kalhan, S.C., Saidel, G.M. & Lai, N. Relating tissue/organ energy expenditure to metabolic fluxes in mouse and human: experimental data integrated with mathematical modeling. Physiol. Rep. 2, e12159 (2014).
Argiles, J. M., Campos, N., Lopez-Pedrosa, J. M., Rueda, R. & Rodriguez-Manas, L. Skeletal Muscle Regulates Metabolism via Interorgan Crosstalk: Roles in Health and Disease. J. Am. Med Dir. Assoc. 17, 789–96 (2016).
Henriksson, J. The possible role of skeletal muscle in the adaptation to periods of energy deficiency. Eur. J. Clin. Nutr. 44 Suppl 1, 55–64 (1990).
Krois, C. R. et al. RDH1 suppresses adiposity by promoting brown adipose adaptation to fasting and re-feeding. Cell Mol. Life Sci. 76, 2425–47 (2019).
Bredella, M. A. et al. Bone marrow adipose tissue composition following high-caloric feeding and fasting. Bone 152, 116093 (2021).
Zhang, X., Yang, S., Chen, J. & Su, Z. Unraveling the Regulation of Hepatic Gluconeogenesis. Front Endocrinol. (Lausanne) 9, 802 (2018).
Wolfe, R. R. The underappreciated role of muscle in health and disease. Am. J. Clin. Nutr. 84, 475–82 (2006).
Morita, K. et al. Structural robustness and temporal vulnerability of the starvation-responsive metabolic network in liver of healthy and obese mice. bioRxiv. 2024:2024.06.17.599249.
Adeva-Andany, M. M., Gonzalez-Lucan, M., Donapetry-Garcia, C., Fernandez-Fernandez, C. & Ameneiros-Rodriguez, E. Glycogen metabolism in humans. BBA Clin. 5, 85–100 (2016).
Mong, F. S., Poland, J. L. & Poland, J. W. Glycogen and histological changes in muscle grafts of rats during fasting or exercise. Can. J. Physiol. Pharm. 60, 387–91 (1982).
Conlee, R. K., Rennie, M. J. & Winder, W. W. Skeletal muscle glycogen content: diurnal variation and effects of fasting. Am. J. Physiol. 231, 614–18 (1976).
Conlee, R. K., Lawler, R. M. & Ross, P. E. Effects of glucose or fructose feeding on glycogen repletion in muscle and liver after exercise or fasting. Ann. Nutr. Metab. 31, 126–32 (1987).
Kokubun, E., Hirabara, S. M., Fiamoncini, J., Curi, R. & Haebisch, H. Changes of glycogen content in liver, skeletal muscle, and heart from fasted rats. Cell Biochem Funct. 27, 488–95 (2009).
Pilegaard, H., Saltin, B. & Neufer, P. D. Effect of short-term fasting and refeeding on transcriptional regulation of metabolic genes in human skeletal muscle. Diabetes 52, 657–62 (2003).
Frier, B. C., Jacobs, R. L. & Wright, D. C. Interactions between the consumption of a high-fat diet and fasting in the regulation of fatty acid oxidation enzyme gene expression: an evaluation of potential mechanisms. Am. J. Physiol. Regul. Integr. Comp. Physiol. 300, R212–21 (2011).
Tunstall, R. J., Mehan, K. A., Hargreaves, M., Spriet, L. L. & Cameron-Smith, D. Fasting activates the gene expression of UCP3 independent of genes necessary for lipid transport and oxidation in skeletal muscle. Biochem Biophys. Res Commun. 294, 301–8 (2002).
Yuan, C. L. et al. Preserved protein synthesis in the heart in response to acute fasting and chronic food restriction despite reductions in liver and skeletal muscle. Am. J. Physiol. Endocrinol. Metab. 295, E216–22 (2008).
Oost, L. J., Kustermann, M., Armani, A., Blaauw, B. & Romanello, V. Fibroblast growth factor 21 controls mitophagy and muscle mass. J. Cachexia Sarcopenia Muscle 10, 630–42 (2019).
Zhao, Y. et al. Histone phosphorylation integrates the hepatic glucagon-PKA-CREB gluconeogenesis program in response to fasting. Mol. Cell 83, 1093–108.e8 (2023).
Rah, S. Y. et al. CD38/ADP-ribose/TRPM2-mediated nuclear Ca(2+) signaling is essential for hepatic gluconeogenesis in fasting and diabetes. Exp. Mol. Med. 55, 1492–505 (2023).
Rodgers, J. T. et al. Nutrient control of glucose homeostasis through a complex of PGC-1alpha and SIRT1. Nature 434, 113–8 (2005).
Ruppert, P. M. M. & Kersten, S. Mechanisms of hepatic fatty acid oxidation and ketogenesis during fasting. Trends Endocrinol. Metab. 35, 107–24 (2024).
Koronowski, K. B. et al. Ketogenesis impact on liver metabolism revealed by proteomics of lysine beta-hydroxybutyrylation. Cell Rep. 36, 109487 (2021).
Ghimire, P., Dhamoon, A.S. Ketoacidosis. StatPearls. Treasure Island (FL) (2024).
Kelley, D. E., Goodpaster, B., Wing, R. R. & Simoneau, J. A. Skeletal muscle fatty acid metabolism in association with insulin resistance, obesity, and weight loss. Am. J. Physiol. 277, E1130–41 (1999).
Wijngaarden, M. A., van der Zon, G. C., van Dijk, K. W., Pijl, H. & Guigas, B. Effects of prolonged fasting on AMPK signaling, gene expression, and mitochondrial respiratory chain content in skeletal muscle from lean and obese individuals. Am. J. Physiol. Endocrinol. Metab. 304, E1012–21 (2013).
Khamzina, L., Veilleux, A., Bergeron, S. & Marette, A. Increased activation of the mammalian target of rapamycin pathway in liver and skeletal muscle of obese rats: possible involvement in obesity-linked insulin resistance. Endocrinology 146, 1473–81 (2005).
Lewis, T. & Stone, W.L. Biochemistry, Proteins Enzymes. StatPearls. Treasure Island (FL)2024.
Fendt, S. M. et al. Tradeoff between enzyme and metabolite efficiency maintains metabolic homeostasis upon perturbations in enzyme capacity. Mol. Syst. Biol. 6, 356 (2010).
Hicks, K. G. et al. Protein-metabolite interactomics of carbohydrate metabolism reveal regulation of lactate dehydrogenase. Science 379, 996–1003 (2023).
Ardito, F., Giuliani, M., Perrone, D., Troiano, G. & Lo Muzio, L. The crucial role of protein phosphorylation in cell signaling and its use as targeted therapy (Review). Int J. Mol. Med. 40, 271–80 (2017).
Lorendeau, D., Christen, S., Rinaldi, G. & Fendt, S. M. Metabolic control of signalling pathways and metabolic auto-regulation. Biol. Cell 107, 251–72 (2015).
Kokaji, T. et al. In vivo transomic analyses of glucose-responsive metabolism in skeletal muscle reveal core differences between the healthy and obese states. Sci. Rep. 12, 13719 (2022).
Kokaji, T. et al. Transomics analysis reveals allosteric and gene regulation axes for altered hepatic glucose-responsive metabolism in obesity. Sci. Signal. 13, eaaz1236 (2020).
Steinberg, G. R. & Hardie, D. G. New insights into activation and function of the AMPK. Nat. Rev. Mol. Cell Biol. 24, 255–72 (2023).
Salt, I. P., Johnson, G., Ashcroft, S. J. & Hardie, D. G. AMP-activated protein kinase is activated by low glucose in cell lines derived from pancreatic beta cells, and may regulate insulin release. Biochem J. 335, 533–9 (1998).
Oakhill, J. S. et al. beta-Subunit myristoylation is the gatekeeper for initiating metabolic stress sensing by AMP-activated protein kinase (AMPK). Proc. Natl. Acad. Sci. USA 107, 19237–41 (2010).
Oakhill, J. S. et al. AMPK is a direct adenylate charge-regulated protein kinase. Science 332, 1433–5 (2011).
Mihaylova, M. M. & Shaw, R. J. The AMPK signalling pathway coordinates cell growth, autophagy and metabolism. Nat. Cell Biol. 13, 1016–23 (2011).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–40 (2010).
McCarthy, D. J., Chen, Y. & Smyth, G. K. Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucleic Acids Res. 40, 4288–97 (2012).
Chen, Y., Lun, A. T. & Smyth, G. K. From reads to genes to pathways: differential expression analysis of RNA-Seq experiments using Rsubread and the edgeR quasi-likelihood pipeline. F1000Res 5, 1438 (2016).
Kanehisa, M. Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Velten, B. et al. Identifying temporal and spatial patterns of variation from multimodal data using MEFISTO. Nat. Methods 19, 179–86 (2022).
Yang, J., Nishihara, R., Zhang, X., Ogino, S. & Qian, Z. R. Energy sensing pathways: Bridging type 2 diabetes and colorectal cancer?J. Diab Complications 31, 1228–36 (2017).
Cespedes, A. et al. Energy-Sensing Pathways in Ischemia: The Counterbalance Between AMPK and mTORC. Curr. Pharm. Des. 25, 4763–70 (2019).
Hardie, D. G. Energy sensing by the AMP-activated protein kinase and its effects on muscle metabolism. Proc. Nutr. Soc. 70, 92–9 (2011).
Hardie, D. G. Sensing of energy and nutrients by AMP-activated protein kinase. Am. J. Clin. Nutr. 93, 891S–6 (2011).
Kelley, D. E., Mokan, M., Simoneau, J. A. & Mandarino, L. J. Interaction between glucose and free fatty acid metabolism in human skeletal muscle. J. Clin. Invest 92, 91–8 (1993).
Kelley, D. E. & Mandarino, L. J. Hyperglycemia normalizes insulin-stimulated skeletal muscle glucose oxidation and storage in noninsulin-dependent diabetes mellitus. J. Clin. Invest 86, 1999–2007 (1990).
Kelley, D. E. & Simoneau, J. A. Impaired free fatty acid utilization by skeletal muscle in non-insulin-dependent diabetes mellitus. J. Clin. Invest 94, 2349–56 (1994).
Lu, B. et al. Metabolic crosstalk: molecular links between glycogen and lipid metabolism in obesity. Diabetes 63, 2935–48 (2014).
Waskiewicz, A. J. et al. Phosphorylation of the cap-binding protein eukaryotic translation initiation factor 4E by protein kinase Mnk1 in vivo. Mol. Cell Biol. 19, 1871–80 (1999).
Pyronnet, S. et al. Human eukaryotic translation initiation factor 4G (eIF4G) recruits mnk1 to phosphorylate eIF4E. EMBO J. 18, 270–9 (1999).
Ferrari, S., Bandi, H. R., Hofsteenge, J., Bussian, B. M. & Thomas, G. Mitogen-activated 70K S6 kinase. Identification of in vitro 40 S ribosomal S6 phosphorylation sites. J. Biol. Chem. 266, 22770–5 (1991).
Flotow, H. & Thomas, G. Substrate recognition determinants of the mitogen-activated 70K S6 kinase from rat liver. J. Biol. Chem. 267, 3074–8 (1992).
de Haro, C., Mendez, R. & Santoyo, J. The eIF-2alpha kinases and the control of protein synthesis. FASEB J. 10, 1378–87 (1996).
Hinnebusch, A. G. The eIF-2 alpha kinases: regulators of protein synthesis in starvation and stress. Semin Cell Biol. 5, 417–26 (1994).
Namkoong, S., Cho, C. S., Semple, I. & Lee, J. H. Autophagy Dysregulation and Obesity-Associated Pathologies. Mol. Cells 41, 3–10 (2018).
Welle, S., Barnard, R. R., Statt, M. & Amatruda, J. M. Increased protein turnover in obese women. Metabolism 41, 1028–34 (1992).
Glynn, E. L. et al. Impact of combined resistance and aerobic exercise training on branched-chain amino acid turnover, glycine metabolism and insulin sensitivity in overweight humans. Diabetologia 58, 2324–35 (2015).
Jensen, M. D. & Haymond, M. W. Protein metabolism in obesity: effects of body fat distribution and hyperinsulinemia on leucine turnover. Am. J. Clin. Nutr. 53, 172–6 (1991).
Solini, A., Bonora, E., Bonadonna, R., Castellino, P. & DeFronzo, R. A. Protein metabolism in human obesity: relationship with glucose and lipid metabolism and with visceral adipose tissue. J. Clin. Endocrinol. Metab. 82, 2552–8 (1997).
Patterson, B. W. et al. Regional muscle and adipose tissue amino acid metabolism in lean and obese women. Am. J. Physiol. Endocrinol. Metab. 282, E931–6 (2002).
Guillet, C. et al. Changes in basal and insulin and amino acid response of whole body and skeletal muscle proteins in obese men. J. Clin. Endocrinol. Metab. 94, 3044–50 (2009).
Lopez-Soldado, I., Guinovart, J. J. & Duran, J. Increased liver glycogen levels enhance exercise capacity in mice. J. Biol. Chem. 297, 100976 (2021).
Mengeste, A. M., Rustan, A. C. & Lund, J. Skeletal muscle energy metabolism in obesity. Obes. (Silver Spring) 29, 1582–95 (2021).
Kelley, D. E. & Mandarino, L. J. Fuel selection in human skeletal muscle in insulin resistance: a reexamination. Diabetes 49, 677–83 (2000).
Galgani, J. E., Moro, C. & Ravussin, E. Metabolic flexibility and insulin resistance. Am. J. Physiol. Endocrinol. Metab. 295, E1009–17 (2008).
Kelley, D. E. Skeletal muscle fat oxidation: timing and flexibility are everything. J. Clin. Invest 115, 1699–702 (2005).
Goodpaster, B. H. & Sparks, L. M. Metabolic Flexibility in Health and Disease. Cell Metab. 25, 1027–36 (2017).
Shulman, G. I. Ectopic fat in insulin resistance, dyslipidemia, and cardiometabolic disease. N. Engl. J. Med 371, 2237–8 (2014).
Samuel, V. T. & Shulman, G. I. The pathogenesis of insulin resistance: integrating signaling pathways and substrate flux. J. Clin. Invest 126, 12–22 (2016).
Healy, J. E., Gearhart, C. N., Bateman, J. L., Handa, R. J. & Florant, G. L. AMPK and ACCchange with fasting and physiological condition in euthermic and hibernating golden-mantled ground squirrels (Callospermophilus lateralis). Comp. Biochem Physiol. A Mol. Integr. Physiol. 159, 322–31 (2011).
Lee-Young, R. S. et al. Obesity impairs skeletal muscle AMPK signaling during exercise: role of AMPKalpha2 in the regulation of exercise capacity in vivo. Int J. Obes. (Lond.) 35, 982–9 (2011).
Steinberg, G. R. & Carling, D. AMP-activated protein kinase: the current landscape for drug development. Nat. Rev. Drug Discov. 18, 527–51 (2019).
Foretz, M., Guigas, B., Bertrand, L., Pollak, M. & Viollet, B. Metformin: from mechanisms of action to therapies. Cell Metab. 20, 953–66 (2014).
Myers, R. W. et al. Systemic pan-AMPK activator MK-8722 improves glucose homeostasis but induces cardiac hypertrophy. Science 357, 507–11 (2017).
Attina, A. et al. Fasting: How to Guide. Nutrients 13, 1570 (2021).
He, C. Balancing nutrient and energy demand and supply via autophagy. Curr. Biol. 32, R684–R96 (2022).
Wang, T., Wiater, E., Zhang, X., Thomas, J.B. & Montminy, M. Crtc modulates fasting programs associated with 1-C metabolism and inhibition of insulin signaling. Proc. Natl. Acad. Sci. USA. 118, e2024865118 (2021).
Phillips, P. J. Oral glucose tolerance testing. Aust. Fam. Physician 41, 391–3 (2012).
Steele, R., Bjerknes, C., Rathgeb, I. & Altszuler, N. Glucose uptake and production during the oral glucose tolerance test. Diabetes 17, 415–21 (1968).
Jackson, R. A. et al. Forearm glucose uptake during the oral glucose tolerance test in normal subjects. Diabetes 22, 442–58 (1973).
Whichelow, M. J. & Butterfield, W. J. Peripheral glucose uptake during the oral glucose-tolerance test in normal and obese subjects and borderline and frank diabetics. Q J. Med. 40, 261–73 (1971).
Kajita, K. et al. Effect of fasting on PPARgamma and AMPK activity in adipocytes. Diab Res Clin. Pr. 81, 144–9 (2008).
Zou, Z., Ohta, T. & Oki, S. ChIP-Atlas 3.0: a data-mining suite to explore chromosome architecture together with large-scale regulome data. Nucleic Acids Res. 52, W45–W53 (2024).
Noh, S. G. et al. Regulation of Circadian Genes Nr1d1 and Nr1d2 in Sex-Different Manners during Liver Aging. Int. J. Mol. Sci. 23, 10032 (2022).
Minokoshi, Y. et al. Leptin stimulates fatty-acid oxidation by activating AMP-activated protein kinase. Nature 415, 339–43 (2002).
Janovska, A. et al. AMPK and ACC phosphorylation: effect of leptin, muscle fibre type and obesity. Mol. Cell Endocrinol. 284, 1–10 (2008).
Hong, J. et al. Butyrate alleviates high fat diet-induced obesity through activation of adiponectin-mediated pathway and stimulation of mitochondrial function in the skeletal muscle of mice. Oncotarget 7, 56071–82 (2016).
Song, J. et al. Normobaric hypoxia shows enhanced FOXO1 signaling in obese mouse gastrocnemius muscle linked to metabolism and muscle structure and neuromuscular innervation. Pflug. Arch. 475, 1265–81 (2023).
Soga, T. & Heiger, D. N. Amino acid analysis by capillary electrophoresis electrospray ionization mass spectrometry. Anal. Chem. 72, 1236–41 (2000).
Soga, T. et al. Differential metabolomics reveals ophthalmic acid as an oxidative stress biomarker indicating hepatic glutathione consumption. J. Biol. Chem. 281, 16768–76 (2006).
Soga, T. et al. Metabolomic profiling of anionic metabolites by capillary electrophoresis mass spectrometry. Anal. Chem. 81, 6165–74 (2009).
Hirayama, A. et al. The use of a double coaxial electrospray ionization sprayer improves the peak resolutions of anionic metabolites in capillary ion chromatography-mass spectrometry. J. Chromatogr. A 1619, 460914 (2020).
Cunningham, F. et al. Ensembl 2015. Nucleic Acids Res. 43, D662–9 (2015).
Flicek, P. et al. Ensembl 2014. Nucleic Acids Res. 42, D749–55 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinforma. 12, 323 (2011).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–82 (2012).
Gene, et al. The Gene Ontology knowledgebase in 2023. Genetics. 224, iyad031 (2023).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet 25, 25–9 (2000).
UniProt, C. UniProt: the Universal Protein Knowledgebase in 2025. Nucleic Acids Res. 53, D609–D17 (2025).
Schomburg, I. et al. BRENDA: a resource for enzyme data and metabolic information. Trends Biochem Sci. 27, 54–6 (2002).
Saier, M. H. et al. The Transporter Classification Database (TCDB): 2021 update. Nucleic Acids Res. 49, D461–D7 (2021).
Mudunuri, U., Che, A., Yi, M. & Stephens, R. M. bioDBnet: the biological database network. Bioinformatics 25, 555–6 (2009).
Hartmann, A. & Jozefowicz, A. M. VANTED: A Tool for Integrative Visualization and Analysis of -Omics Data. Methods Mol. Biol. 1696, 261–78 (2018).
Kodama, Y., Mashima, J., Kosuge, T. & Ogasawara, O. DDBJ update: the Genomic Expression Archive (GEA) for functional genomics data. Nucleic Acids Res. 47, D69–D73 (2019).
Okuda, S. et al. jPOSTrepo: an international standard data repository for proteomes. Nucleic Acids Res. 45, D1107–D11 (2017).
Acknowledgements
We thank Maki Ohishi and Ayano Ueno (Keio University) for their expertise and assistance with metabolome analysis using CE-MS. We also thank Kazusa DNA Research Institute for conducting the proteomic measurements. We also thank our laboratory members for critically reading this manuscript and technical assistance with the experiments. The computational analysis of this work was performed in part with support of the supercomputer system of the National Institute of Genetics (NIG), Research Organization of Information and Systems (ROIS). This study was supported by the Japan Society for the Promotion of Science (JSPS) KAKENHI grant numbers JP17H06299, JP17H06300, JP18H03979, JP21H04759, JP23H04939, and JP23H04946 to S.K., and by Japan Science and Technology Agency (JST) as part of CREST (JPMJCR2123 to S.K., Y. Inaba, and T.S.) and of Adopting Sustainable Partnerships for Innovative Research Ecosystem (ASPIRE), Grant Number JPMJAP24B1; The Uehara Memorial Foundation (to S.K.); K.M. receives funding from a Grant-in-Aid for Early-Career Scientists (JP21K15342). T.K. receives funding from a Grant-in-Aid for Early-Career Scientists (JP21K16349). A.Hatano receives funding from a Grant-in-Aid for Early-Career Scientists (JP22K15034). This work was also supported by the Japan Society for the Promotion of Science (JSPS) KAKENHI grant number JP22H04925 (PAGS); in part by the MEXT Cooperative Research Project Program, Medical Research Center Initiative for High Depth Omics, and CURE:JPMXP1323015486 for MIB, Kyushu University; and AMED Grant Number JP21zf0127001 (T.S.), JST, MEXT KAKENHI Grant Number JP23H04946 (T.S.) and World Premier International Research Center Initiative (WPI), Human Biology-Microbiome-Quantum Research Center (Bio2Q) (T.S.), MEXT, Japan.; and JST FOREST Program (Grant Number JPMJFR2052, to A. Hirayama). T.T. receives funding from a Grant-in-Aid for Early-Career Scientists (JP20K19915). H.O. receives funding from a Grant-in-Aid for Early-Career Scientists (JP22K17992). S.O. receives funding from a Grant-in-Aid for Early-Career Scientists (JP21K14467). Y. Inaba also receives AMED-PRIME (JP23gm6910002).
Author information
Authors and Affiliations
Contributions
D.L., A. Hatano, S.O., K.M., T.K., and S.K. designed the project. A. Hatano performed animal experiments. A. Hatano and K.M. prepared samples for omic measurement. T.S. and A. Hirayama performed metabolomic analysis using CE-MS and IC-QEMS. Y.S. performed transcriptomic analysis using RNA-seq. D.L., T.T., and H.O. analyzed the RNA-seq data. M.M. and A. Hatano performed proteomic analysis using MS. D.L. performed the western blot analysis. D.L. analyzed the omics data. D.L. performed the visualization of the results. H.I. and Y.I. helped with the biological interpretation. H.M. performed MEFISTO analysis. H.S., Y.P., and K.M. provided critical intellectual contributions to the omic analysis, including the optimization of bioinformatic pipelines and the biological interpretation of complex data sets. D.L. and S.K. wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, D., Morita, K., Kokaji, T. et al. Global loss of metabolic responsiveness and elevated enzyme in leptin deficient obese mice during starvation. npj Syst Biol Appl (2026). https://doi.org/10.1038/s41540-026-00678-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41540-026-00678-3