Abstract
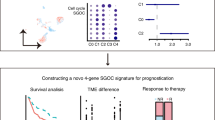
Head and neck squamous cell carcinoma (HNSCC) is a complex multivariable disease posing a significant challenge in therapeutics. OXPHOS and PPP are centrally implicated in metabolic heterogeneity influencing tumor behavior, treatment response, and patient outcomes. This study aims to stratify HNSCC based on OXPHOS and PPP gene expression to construct a clinical outcome risk predictive model. HNSCC patients were stratified into four metabolic subtypes including mixed, OXPHOS-leaning, PPP-leaning, and quiescent. The quiescent metabolic subtype showed longest survival with lower proliferation scores, whereas mixed subtype showed the worst survival with higher proliferation scores. Metabolic proteins of the CPTAC and ICPC cohorts affirmed the existence of metabolic heterogeneity among tumor samples. A prognostic risk predictive model was developed based on thirteen genes, which performed better than the OXPHOS-PPP-glycolysis model and other predictive forty-seven metabolic gene’s model. Existence of metabolic heterogeneity was successfully determined in HNSCC. OXPHOS and PPP genes enriched in mixed metabolic subtype having the worst clinical outcome are suggestive of higher metastatic potential, which might offer advisement in personalized therapeutics.

Similar content being viewed by others
Data availability
The proteomics dataset generated in this study has been deposited at the ProteomeXchange Consortium via the PRIDE83 partner repository with the dataset identifier PXD073947. The data analysis workflow and code is available at https://github.com/AK2-lab/Metabolic-Heterogeneity-Code/tree/main.
References
Rumgay, H. et al. Global burden of oral cancer in 2022 attributable to smokeless tobacco and areca nut consumption: a population attributable fraction analysis. Lancet Oncol. 25, 1413–1423 (2024).
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 74, 229–263 (2024).
Abdulla, R. et al. Clinicopathological analysis of oral squamous cell carcinoma among the younger age group in coastal Karnataka, India: a retrospective study. J. Oral. Maxillofac. Pathol. 22, 180–187 (2018).
Johnson, D. E. et al. Head and neck squamous cell carcinoma. Nat. Rev. Dis. Prim. 6, 92 (2020).
Ekanayaka, R. P. & Tilakaratne, W. M. Impact of histopathological parameters in prognosis of oral squamous cell carcinoma. Oral. Dis. 31, 1420–1438 (2025).
Hoesseini, A. et al. Head and neck cancer patients’ preferences for individualized prognostic information: a focus group study. BMC Cancer 20, 399 (2020).
Hoesseini, A. et al. Prognostic model for overall survival of head and neck cancer patients in the palliative phase. BMC Palliat. Care. 23, 54 (2024).
Kim, J. & DeBerardinis, R. J. Mechanisms and implications of metabolic heterogeneity in cancer. Cell Metab. 30, 434–446 (2019).
Demicco, M., Liu, X. Z., Leithner, K. & Fendt, S. M. Metabolic heterogeneity in cancer. Nat. Metab. 6, 18–38 (2024).
Pavlova, N. N. & Thompson, C. B. The emerging hallmarks of cancer metabolism. Cell Metab. 23, 27–47 (2016).
Warburg, O. On respiratory impairment in cancer cells. Science 124, 269–270 (1956).
Ohshima, K. & Morii, E. Metabolic reprogramming of cancer cells during tumor progression and metastasis. Metabolites 11, 28 (2021).
Nguyen, D. X. & Massague, J. Genetic determinants of cancer metastasis. Nat. Rev. Genet. 8, 341–352 (2007).
Evers, T. M. J. et al. Deciphering metabolic heterogeneity by single-cell analysis. Anal. Chem. 91, 13314–13323 (2019).
Gong, Y. et al. Metabolic-pathway-based subtyping of triple-negative breast cancer reveals potential therapeutic targets. Cell Metab. 33, 51–64.e9 (2021).
Flint, L. E. et al. Characterization of an aggregated three-dimensional cell culture model by multimodal mass spectrometry imaging. Anal. Chem. 92, 12538–12547 (2020).
Xiao, Z., Dai, Z. & Locasale, J. W. Metabolic landscape of the tumor microenvironment at single cell resolution. Nat. Commun. 10, 3763 (2019).
DeBerardinis, R. J. & Chandel, N. S. Fundamentals of cancer metabolism. Sci. Adv. 2, e1600200 (2016).
Jin, J., Byun, J. K., Choi, Y. K. & Park, K. G. Targeting glutamine metabolism as a therapeutic strategy for cancer. Exp. Mol. Med. 55, 706–715 (2023).
Miki, K. et al. Glutaminolysis is associated with mitochondrial pathway activation and can be therapeutically targeted in glioblastoma. Cancer Metab. 12, 35 (2024).
Lee, K. M. et al. MYC and MCL1 cooperatively promote chemotherapy-resistant breast cancer stem cells via regulation of mitochondrial oxidative phosphorylation. Cell Metab. 26, 633–47.e7 (2017).
Zhang, L. et al. Metabolic reprogramming toward oxidative phosphorylation identifies a therapeutic target for mantle cell lymphoma. Sci. Transl. Med. 11, eaau1167 (2019).
Viale, A. et al. Oncogene ablation-resistant pancreatic cancer cells depend on mitochondrial function. Nature 514, 628–632 (2014).
Hu, J. et al. Heterogeneity of tumor-induced gene expression changes in the human metabolic network. Nat. Biotechnol. 31, 522–529 (2013).
TeSlaa, T., Ralser, M., Fan, J. & Rabinowitz, J. D. The pentose phosphate pathway in health and disease. Nat. Metab. 5, 1275–1289 (2023).
Patra, K. C. & Hay, N. The pentose phosphate pathway and cancer. Trends Biochem. Sci. 39, 347–354 (2014).
Giacomini, I., Ragazzi, E., Pasut, G., & Montopoli M. The pentose phosphate pathway and its involvement in cisplatin resistance. Int. J. Mol. Sci. 21, 937 (2020).
Gong, Y. et al. SRC enhanced cisplatin resistance in bladder cancer by reprogramming glycolysis and pentose phosphate pathway. Commun. Biol. 8, 36 (2025).
Wilkerson, M. D. & Hayes, D. N. ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics 26, 1572–1573 (2010).
Tranby, E. P. et al. Oral cancer prevalence, mortality, and costs in medicaid and commercial insurance claims data. Cancer Epidemiol. Biomark. Prev. 31, 1849–1857 (2022).
Zhou, X. et al. Genomic alterations in oral multiple primary cancers. Int J. Oral. Sci. 16, 13 (2024).
Ward, P. S. & Thompson, C. B. Metabolic reprogramming: a cancer hallmark even warburg did not anticipate. Cancer Cell. 21, 297–308 (2012).
Weinberg, F. et al. Mitochondrial metabolism and ROS generation are essential for Kras-mediated tumorigenicity. Proc. Natl. Acad. Sci. 107, 8788–8793 (2010).
LeBleu, V. S. et al. PGC-1alpha mediates mitochondrial biogenesis and oxidative phosphorylation in cancer cells to promote metastasis. Nat. Cell Biol. 16, 992–1003 (2014). 1-15.
Yao, C. H. et al Mitochondrial fusion supports increased oxidative phosphorylation during cell proliferation. Elife. 8, e41351 (2019).
Davis, R. T. et al. Transcriptional diversity and bioenergetic shift in human breast cancer metastasis revealed by single-cell RNA sequencing. Nat. Cell Biol. 22, 310–320 (2020).
Danko, B. et al. Metabolic pathway-based subtypes associate glycan biosynthesis and treatment response in head and neck cancer. NPJ Precis. Oncol. 8, 116 (2024).
Bansal, A. & Simon, M. C. Glutathione metabolism in cancer progression and treatment resistance. J. Cell Biol. 217, 2291–2298 (2018).
Harris, I. S. et al. Glutathione and thioredoxin antioxidant pathways synergize to drive cancer initiation and progression. Cancer Cell. 27, 211–222 (2015).
Uslu, C., Kapan, E. & Lyakhovich, A. Cancer resistance and metastasis are maintained through oxidative phosphorylation. Cancer Lett. 587, 216705 (2024).
Li, S. S. et al. FAO-fueled OXPHOS and NRF2-mediated stress resilience in MICs drive lymph node metastasis. Proc. Natl. Acad. Sci. 122, e2411241122 (2025).
Xu, Y., Xue, D., Bankhead, A. 3rd & Neamati, N. Why all the fuss about oxidative phosphorylation (OXPHOS)?. J. Med Chem. 63, 14276–14307 (2020).
Beerkens, A. P. M. et al. Mitochondria targeting of oxidative phosphorylation inhibitors to alleviate hypoxia and enhance anticancer treatment efficacy. Clin. Cancer Res. 31, 1186–1193 (2025).
Greene, J., Segaran, A. & Lord, S. Targeting OXPHOS and the electron transport chain in cancer; molecular and therapeutic implications. Semin Cancer Biol. 86, 851–859 (2022).
Zhang, E. et al. Identification of subgroups along the glycolysis-cholesterol synthesis axis and the development of an associated prognostic risk model. Hum. Genom. 15, 53 (2021).
Jiang, M., Gu, X., Xu, Y. & Wang, J. Metabolism-associated molecular classification and prognosis signature of head and neck squamous cell carcinoma. Heliyon 10, e27587 (2024).
Li, L. et al. Cuproptosis/OXPHOS tendency prediction of prognosis and immune microenvironment of esophageal squamous cell carcinoma: bioinformatics analysis and experimental validation. Gene 902, 148156 (2024).
Galber, C., Acosta, M. J., Minervini, G. & Giorgio, V. The role of mitochondrial ATP synthase in cancer. Biol. Chem. 401, 1199–1214 (2020).
Wang, T., Ma, F. & Qian, H. L. Defueling the cancer: ATP synthase as an emerging target in cancer therapy. Mol. Ther. Oncolytics. 23, 82–95 (2021).
Cheng, Y. et al. Downregulation of ATP5F1D inhibits mtROS/NLRP3/caspase-1/GSDMD axis to suppress pyroptosis-mediated malignant progression of endometrial cancer. Int Immunopharmacol. 139, 112808 (2024).
Poliakov, E., Managadze, D. & Rogozin, I. B. Generalized portrait of cancer metabolic pathways inferred from a list of genes overexpressed in cancer. Genet. Res. Int. 2014, 646193 (2014).
Wang, Z. et al. High throughput proteomic and metabolic profiling identified target correction of metabolic abnormalities as a novel therapeutic approach in head and neck paraganglioma. Transl. Oncol. 14, 101146 (2021).
Yi, J., Gao, W., Wu, C., Yang, R. & Yu, C. A multidimensional pan-cancer analysis of NDUFA4L2 and verification of the oncogenic value in colon cancer. FASEB J. 39, e70300 (2025).
Shang, B. et al. ALDOC promotes non-small cell lung cancer through affecting MYC-mediated UBE2N transcription and regulating Wnt/beta-catenin pathway. Aging 15, 9614–9632 (2023).
Pes, G. M., Errigo, A., Soro, S., Longo, N. P. & Dore, M. P. Glucose-6-phosphate dehydrogenase deficiency reduces susceptibility to cancer of endodermal origin. Acta Oncol. 58, 1205–1211 (2019).
Lan, T. et al. Glucose-6-phosphate dehydrogenase maintains redox homeostasis and biosynthesis in LKB1-deficient KRAS-driven lung cancer. Nat. Commun. 15, 5857 (2024).
Kim, J. W. & Dang, C. V. Multifaceted roles of glycolytic enzymes. Trends Biochem. Sci. 30, 142–150 (2005).
de Padua, M. C. et al. Disrupting glucose-6-phosphate isomerase fully suppresses the “Warburg effect” and activates OXPHOS with minimal impact on tumor growth except in hypoxia. Oncotarget 8, 87623–87637 (2017).
Yang, H. D. et al. T-cell immune regulator 1 enhances metastasis in hepatocellular carcinoma. Exp. Mol. Med. 50, e420 (2018).
Gao, W. et al. The role of S-nitrosylation of PFKM in regulation of glycolysis in ovarian cancer cells. Cell Death Dis. 12, 408 (2021).
Shen, X. et al. PFKM promotes the progression of gastric cancer by up-regulating CNTN1 expression through H3K18la modification. Appl. Biochem. Biotechnol. 197, 5885–5901 (2025).
Cunningham, J. T., Moreno, M. V., Lodi, A., Ronen, S. M. & Ruggero, D. Protein and nucleotide biosynthesis are coupled by a single rate-limiting enzyme, PRPS2, to drive cancer. Cell 157, 1088–1103 (2014).
Gao, F. et al. Identification of ubiquinol cytochrome c reductase hinge (UQCRH) as a potential diagnostic biomarker for lung adenocarcinoma. Open Biol. 6, 150256 (2016).
Bu, W., Cao, M., Wu, X. & Gao, Q. Prognosis prediction of head and neck squamous cell carcinoma through the basement membrane-related lncRNA risk model. Front Mol. Biosci. 11, 1421335 (2024).
Lai, J., Huang, R. & Huang, J. Predicting head and neck cancer response to radiotherapy with a chemokine-based model. Sci. Rep. 15, 28450 (2025).
Su, F. et al. Quantitative proteomics identified 3 oxidative phosphorylation genes with clinical prognostic significance in gastric cancer. J. Cell Mol. Med. 24, 10842–10854 (2020).
Chen, W., Yang, Z. & Chen, Y. A novel oxidative phosphorylation-associated gene signature for prognosis prediction in patients with hepatocellular carcinoma. Dis. Markers 2022, 3594901 (2022).
Zhan, Z., Lin, K. & Wang, T. Construction of oxidative phosphorylation-related prognostic risk score model in uveal melanoma. BMC Ophthalmol. 24, 204 (2024).
Lin, W. et al. A novel mitochondrial metabolism-related gene signature for predicting the prognosis of oesophageal squamous cell carcinoma. Aging 16, 9649–9679 (2024).
Liao, R. G. et al. A novel mitochondrial-related risk model for predicting prognosis and immune checkpoint blockade therapy response in uterine corpus endometrial carcinoma. Sci. Rep. 15, 1404 (2025).
Wu, C. S. et al. Integrated multi-omics analyses of oral squamous cell carcinoma reveal precision patient stratification and personalized treatment strategies. Cancer Lett. 614, 217482 (2025).
Huang, C. et al. Proteogenomic insights into the biology and treatment of HPV-negative head and neck squamous cell carcinoma. Cancer Cell. 39, 361–79.e16 (2021).
Zhou, Z. et al. Prognosis-related molecular subtyping in head and neck squamous cell carcinoma patients based on glycolytic/cholesterogenic gene data. Cancer Cell Int. 23, 37 (2023).
Kanehisa, M. Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Hanzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 14, 7 (2013).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. 102, 15545–15550 (2005).
Mootha, V. K. et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Thorsson, V. et al. The immune landscape of cancer. Immunity 48, 812–30.e14 (2018).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Simon, N., Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for Cox’s proportional hazards model via coordinate descent. J. Stat. Softw. 39, 1–13 (2011).
Tay, J. K., Narasimhan, B. & Hastie, T. Elastic net regularization paths for all generalized linear models. J. Stat. Softw. 106, 1–31 (2023).
Tang, D. et al. SRplot: a free online platform for data visualization and graphing. PLoS One 18, e0294236 (2023).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 47, D442–D450 (2019).
Acknowledgements
We thank TCGA project and PDC portal for the availability of the transcriptome and proteome data of HNSCC cohort. AG thanks CSIR India for fellowship. The authors are grateful to Prof. Saumitra Das (Department of Microbiology and Cell Biology, Indian Institute of Science, Bengaluru) for connecting us with clinicians. This research work was supported by BRIC-National Institute of Biomedical Genomic intramural funding (NIBMG-60116).
Author information
Authors and Affiliations
Contributions
Writing—Original draft preparation, S. Sau, A.G., S. Sinha, A.M.K.; Writing–Review and Editing, S.M.A.M., A.M.K.; Software and Methodology, S. Sau, A.M.K.; Investigation and conceptualization, S. Sau, A.M.K.; Supervision, A.M.K. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sau, S., Gupta, A., Sinha, S. et al. Metabolic gene expression-based stratification and prognostic risk predictive model of head and neck squamous cell carcinoma. npj Syst Biol Appl (2026). https://doi.org/10.1038/s41540-026-00689-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41540-026-00689-0