Abstract
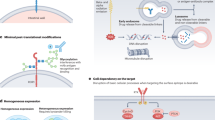
Improving the specificity of prostate cancer treatment requires ligands that bind selectively and with ultra-high affinity to tumour-associated targets absent from healthy tissues. Prostatic acid phosphatase has emerged as an alternative target to prostate-specific membrane antigen, as it is expressed in a broader subset of prostate cancers and is not detected in healthy organs such as the salivary glands and kidneys. Here, to discover selective binders to prostatic acid phosphatase, we constructed two DNA-encoded chemical libraries comprising over 6.7 million small molecules based on proline and phenylalanine scaffolds. Screening against the purified human prostatic acid phosphatase yielded OncoACP3, a small organic ligand with picomolar binding affinity. When radiolabelled with lutetium-177, OncoACP3 selectively accumulated in enzyme-expressing tumours with a long residence time (biological half-life greater than 72 h) and a high tumour-to-blood ratio (>148 at 2 h after administration). Lutetium-177-labelled OncoACP3 cured tumours in mice at low, well-tolerated doses. Its conjugation to the cytotoxic agent monomethyl auristatin E facilitated tumour-selective payload deposition, resulting in potent anti-tumour activity. The modular structure of OncoACP3 supports flexible payload delivery for the targeted treatment of metastatic prostate cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All data supporting the findings of this study are available within the Article and its Supplementary Information. Unprocessed images for ex vivo biodistribution studies connected to Fig. 6 and Extended Data Fig. 10, as well as raw data for PET–CT in vivo biodistribution work connected to Extended Data Fig. 9, are available via figshare at https://doi.org/10.6084/m9.figshare.28668596 (ref. 55). Source data are provided with this paper.
References
Van Der Veldt, A. A. M. et al. Biodistribution and radiation dosimetry of 11C-labelled docetaxel in cancer patients. Eur. J. Nucl. Med. Mol. Imaging 37, 1950–1958 (2010).
Van Der Veldt, A. A. M. et al. Toward prediction of efficacy of chemotherapy: a proof of concept study in lung cancer patients using [11C]docetaxel and positron emission tomography. Clin. Cancer Res. 19, 4163–4173 (2013).
Van Cutsem, E. et al. Phase III study of docetaxel and cisplatin plus fluorouracil compared with cisplatin and fluorouracil as first-line therapy for advanced gastric cancer: a report of the V25 study group. J. Clin. Oncol. 24, 4991–4997 (2006).
Lyu, X. et al. The global landscape of approved antibody therapies. Antibody Ther. 5, 233–257 (2022).
Dennis, M. S. et al. Imaging tumors with an albumin-binding Fab, a novel tumor-targeting agent. Cancer Res. 67, 254–261 (2007).
Cazzamalli, S., Dal Corso, A., Widmayer, F. & Neri, D. Chemically defined antibody- and small molecule-drug conjugates for in vivo tumor targeting applications: a comparative analysis. J. Am. Chem. Soc. 140, 1617–1621 (2018).
Lu, G. et al. Co-administered antibody improves penetration of antibody–dye conjugate into human cancers with implications for antibody–drug conjugates. Nat. Commun. 11, 5667 (2020).
Thurber, G. M., Schmidt, M. M. & Wittrup, K. D. Antibody tumor penetration: transport opposed by systemic and antigen-mediated clearance. Adv. Drug Deliv. Rev. 60, 1421–1434 (2008).
Cilliers, C., Guo, H., Liao, J., Christodolu, N. & Thurber, G. M. Multiscale modeling of antibody–drug conjugates: connecting tissue and cellular distribution to whole animal pharmacokinetics and potential implications for efficacy. AAPS J. 18, 1117–1130 (2016).
Zana, A. et al. A comparative analysis of fibroblast activation protein-targeted small molecule–drug, antibody–drug, and peptide–drug conjugates. Bioconjugate Chem. 34, 1205–1211 (2023).
Yuan, F. et al. Vascular permeability in a human tumor xenograft: molecular size dependence and cutoff size. Cancer Res. 55, 3752–3756 (1995).
Strosberg, J. et al. Phase 3 trial of 177 Lu-dotatate for midgut neuroendocrine tumors. N. Engl. J. Med. 376, 125–135 (2017).
Sartor, O. et al. Lutetium-177–PSMA-617 for metastatic castration-resistant prostate cancer. N. Engl. J. Med. 385, 1091–1103 (2021).
Hughes, J. P., Rees, S. S., Kalindjian, S. B. & Philpott, K. L. Principles of early drug discovery. Br. J. Pharmacol. 162, 1239–1249 (2011).
MacArron, R. et al. Impact of high-throughput screening in biomedical research. Nat. Rev. Drug Discov. 10, 188–195 (2011).
Goodnow, R. A., Dumelin, C. E. & Keefe, A. D. DNA-encoded chemistry: enabling the deeper sampling of chemical space. Nat. Rev. Drug Discov. 16, 131–147 (2017).
Neri, D. & Lerner, R. A. DNA-encoded chemical libraries: a selection system based on endowing organic compounds with amplifiable information. Annu. Rev. Biochem. 8, 479–502 (2018).
Gura, T. DNA helps build molecular libraries for drug testing. Science 350, 1139–1140 (2015).
Decurtins, W. et al. Automated screening for small organic ligands using DNA-encoded chemical libraries. Nat. Protoc. 11, 764–780 (2016).
Mannocci, L. et al. High-throughput sequencing allows the identification of binding molecules isolated from DNA-encoded chemical libraries. Proc. Natl Acad. Sci. USA 105, 17670–17675 (2008).
Leamon, C. P. et al. Folate targeting enables durable and specific antitumor responses from a therapeutically null tubulysin B analogue. Cancer Res. 68, 9839–9844 (2008).
Cazzamalli, S. et al. Enhanced therapeutic activity of non-internalizing small-molecule–drug conjugates targeting carbonic anhydrase IX in combination with targeted interleukin-2. Clin. Cancer Res. 24, 3656–3667 (2018).
Millul, J. et al. An ultra-high-affinity small organic ligand of fibroblast activation protein for tumor-targeting applications. Proc. Natl Acad. Sci. USA 118, 1–10 (2021).
Krutzek, F., Donat, C. K., Ullrich, M. & Stadlbauer, S. Design, synthesis, and biological evaluation of small-molecule-based radioligands with improved pharmacokinetic properties for imaging of programmed death ligand 1. J. Med. Chem. 66, 15894–15915 (2023).
Ruigrok, E. A. M. et al. In vitro dose effect relationships of actinium-225- and lutetium-177-labeled PSMA-I&T. Eur. J. Nucl. Med. Mol. Imaging 49, 3627–3638 (2022).
Thurin, N. H. et al. Epidemiology of metastatic castration-resistant prostate cancer: a first estimate of incidence and prevalence using the French nationwide healthcare database. Cancer Epidemiol. 69, 1–6 (2020).
Cancer in men: prostate cancer is #1 for 118 countries globally. American Cancer Society https://www.cancer.org/research/acs-research-news/prostate-cancer-is-number-1-for-118-countries-worldwide.html (2024).
Satapathy, S. et al. [177Lu]Lu-PSMA-617 versus docetaxel in chemotherapy-naïve metastatic castration-resistant prostate cancer: final survival analysis of a phase 2 randomized, controlled trial. J. Nucl. Med. 64, 1726–1729 (2023).
Mahajan, S., Grewal, R. K., Friedman, K. P., Schöder, H. & Pandit-Taskar, N. Assessment of salivary gland function after 177Lu-PSMA radioligand therapy: current concepts in imaging and management. Transl. Oncol. 21, 1–8 (2022).
Heynickx, N., Herrmann, K., Vermeulen, K., Baatout, S. & Aerts, A. The salivary glands as a dose limiting organ of PSMA- targeted radionuclide therapy: a review of the lessons learnt so far. Nucl. Med. Biol. 98–99, 30–39 (2021).
Expression of ACP3 in tumors and healthy organs (IHC). The Human Protein Atlas https://www.proteinatlas.org/ENSG00000014257-ACP3 (2024).
Muniyan, S. et al. Human prostatic acid phosphatase: structure, function and regulation. Int. J. Mol. Sci. 14, 10438–10464 (2013).
Quintero, I. B. et al. Prostatic acid phosphatase is not a prostate specific target. Cancer Res. 67, 6549–6554 (2007).
Vihko, P. et al. Immunoscintigraphic evaluation of lymph node involvement in prostatic carcinoma. Prostate 11, 51–57 (1987).
Babaian, R. J. & Lamki, L. M. Radioimmunoscintigraphy of prostate cancer. Semin. Nucl. Med. XIX, 309–321 (1989).
Waight, A. B. et al. Structural basis of microtubule destabilization by potent auristatin anti-mitotics. PLoS ONE 11, 1–14 (2016).
Dal Corso, A., Cazzamalli, S., Gébleux, R., Mattarella, M. & Neri, D. Protease-cleavable linkers modulate the anticancer activity of noninternalizing antibody–drug conjugates. Bioconjugate Chem. 28, 1826–1833 (2017).
Doronina, S. O. et al. Development of potent monoclonal antibody auristatin conjugates for cancer therapy. Nat. Biotechnol. 21, 778–784 (2003).
Oehler, S. et al. A DNA-encoded chemical library based on chiral 4-amino-proline enables stereospecific isozyme-selective protein recognition. Nat. Chem. 15, 1431–1443 (2023).
Favalli, N. et al. Stereo- and regiodefined DNA-encoded chemical libraries enable efficient tumour-targeting applications. Nat. Chem. 13, 540–548 (2021).
Beers, S. A. Phosphatase inhibitors-lll. Benzylamino phosphonicacids as potent inhibitors of human prostatic acid phosphatase. Bioorg. Med. Chem. 4, 1693–1701 (1996).
Bao, G. et al. Targeting CXCR4/CXCL12 axis via [177Lu]Lu-DOTAGA.(SA.FAPi)2 with CXCR4 antagonist in triple-negative breast cancer. Eur. J. Nucl. Med. Mol. Imaging 51, 2744–2757 (2024).
Backhaus, P. et al. Translational imaging of the fibroblast activation protein (FAP) using the new ligand [68Ga]Ga-OncoFAP-DOTAGA. Eur. J. Nucl. Med. Mol. Imaging 49, 1822–1832 (2022).
Srinivasarao, M., Galliford, C. V. & Low, P. S. Principles in the design of ligand-targeted cancer therapeutics and imaging agents. Nat. Rev. Drug Discov. 14, 203–219 (2015).
Kratochwil, C. et al. PSMA-targeted radionuclide therapy of metastatic castration-resistant prostate cancer with 177Lu-labeled PSMA-617. J. Nucl. Med. 57, 1170–1176 (2016).
Roberts, M. S., Magnusson, B. M., Burczynski, F. J. & Weiss, M. Enterohepatic circulation. Clin. Pharmacokinet. 41, 751–790 (2002).
Fleck, C. & Bräunlich, H. Factors determining the relationship between renal and hepatic excretion of xenobiotics. Arzneimittelforschung 40, 942–946 (1990).
Hemmingsson, J. et al. Specific uptake in the bone marrow causes high absorbed red marrow doses during [177Lu]Lu-DOTATATE treatment. J. Nucl. Med. 64, 1456–1462 (2023).
Zana, A. et al. Fibroblast activation protein triggers release of drug payload from non-internalizing small molecule drug conjugates in solid tumors. Clin. Cancer Res. 28, 5440–5454 (2022).
Wichert, M. et al. Dual-display of small molecules enables the discovery of ligand pairs and facilitates affinity maturation. Nat. Chem. 7, 241–249 (2015).
Puglioli, S. et al. Selective tumor targeting enabled by picomolar fibroblast activation protein inhibitors isolated from a DNA-encoded affinity maturation library. Chem 9, 411–429 (2023).
Ceci, F. et al. 68Ga-PSMA-11 PET/CT in recurrent prostate cancer: efficacy in different clinical stages of PSA failure after radical therapy. Eur. J. Nucl. Med. Mol. Imaging 46, 31–39 (2019).
Violet, J. et al. Dosimetry of 177Lu-PSMA-617 in metastatic castration-resistant prostate cancer: correlations between pretherapeutic imaging and whole-body tumor dosimetry with treatment outcomes. J. Nucl. Med. 60, 517–523 (2019).
Cunha, L., Baete, K., Leijen, C. & Jamar, F. Main challenges in radiation protection with emerging radionuclide therapies. Quart. J. Nucl. Med. Mol. Imaging 67, 14–28 (2023).
Georgiev, T. et al. Discovery of high-affinity ligands for prostatic acid phosphatase via DNA-encoded library screening enables targeted cancer therapy. figshare https://doi.org/10.6084/m9.figshare.28668596 (2025).
Sharma, S. K. et al. A rapid bead-based radioligand binding assay for the determination of target-binding fraction and quality control of radiopharmaceuticals. Nucl. Med. Biol. 71, 32–38 (2019).
Galbiati, A. et al. A dimeric FAP-targeting small-molecule radioconjugate with high and prolonged tumor uptake. J. Nucl. Med. 63, 1852–1858 (2022).
Acknowledgements
We thank C. Pellegrino (University Hospital Zurich) and R. De Luca (Philochem AG) for their support with the generation of hACP3+ tumour cell lines. C. Comacchio (Philochem AG) is acknowledged for the production of human recombinant TNAP. G. Neri (Philogen S.p.A.) is acknowledged for the thorough revision of the manuscript. R. Konyarska is acknowledged for preparing cover art designs and M. Satama for proofreading the manuscript to ensure clarity and accessibility for a broader audience. G. Rotta is acknowledged for providing the rendered mice images for Figs. 1 and 8. We received no specific funding for this work.
Author information
Authors and Affiliations
Contributions
All authors have contributed to the preparation of this manuscript. T.G., F.M., E.P., S.O., S.C. and D.N. designed the experiments. F.M., M.M. and S.O. synthesized the chemical libraries with support from G.B. and N.F. F.M., S.O. and G.B. performed the selections. F.M., T.G., S.O., G.B. and N.F. analysed the high-throughput screening data. Hits were synthesized and validated by T.G., F.M., A.C. and S.O. I.B. and F.M. expressed and biotinylated the proteins. In vitro and in vivo experiments were performed by T.G., F.M., M.M., S.O., Y.P. and S.C. L.P. and E.G. performed mass spectrometry experiments, including ex vivo biodistribution studies with SMDCs. P.F. and F.B. performed, analysed and interpreted ACP3 and PSMA immunohistochemistry staining on human healthy tissues and prostate cancer lesions.
Corresponding authors
Ethics declarations
Competing interests
D.N. is the co-founder, CEO, CSO and president of the Scientific Advisory Board of Philogen. T.G., F.M., I.B., M.M., Y.S.P.K., L.P., E.G., N.F., E.P., S.O. and S.C. are employed by Philochem AG, the research and development unit of the Philogen group. Some of the findings included in this manuscript have been patented. T.G., F.M., S.O., S.C., D.N. and Y.S.P.K. are listed in the patent application PCT/EP2024/080350 (filed on 25 October 2024). The other authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks Iain J. McEwan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Additional ACP3 and PSMA IHC staining results on Prostate Cancer tissue sections (primary and metastatic lesions).
Gleason and H-scores are presented along with the specific patient ID from which the samples were obtained (University of Pisa’s biobank, Department of Surgical, Medical, Molecular Pathology, and Critical Area; approval from the Ethical Committee of the University of Pisa, PROTOCOL N. 9989 - 20th of February 2019).
Extended Data Fig. 2 Additional selection fingerprints after screening of SOP-DEL.
Selection fingerprints after screening of SOP-DEL against ACP3 (a), tissue non-specific alkaline phosphatase (TNAP, negative control, b), streptavidin-coated beads (DynabeadsTM MyOneTM Streptavidin C1) in PBS buffer (no protein, negative control, c) or TNAP buffer (no protein, negative control, d). DEL selections were performed in duplicates. Dot colour and size correspond to the normalized sequence counts. (a, left) Total Counts (TCs) = 1,461,935, Average Count (AC) = 1.02, Cut-off = 8; (a, right) TCs = 1,502,438, AC = 1.05, Cut-off = 8; (b, left) TCs = 1,556,461, AC = 1.09, Cut-off = 25; (b, right) TCs = 1,640,950, AC = 1.14, Cut-off = 25; (c) TCs = 1,050,742, AC = 0.73, Cut-off = 15; (d) TCs = 1,580,160, AC = 1.10, Cut-off = 35.
Extended Data Fig. 3 Additional selection fingerprints after screening of FMA-DEL.
Selection fingerprints after screening of FMA-DEL against prostatic acid phosphatase (ACP3, a), tissue non-specific alkaline phosphatase (TNAP, negative control, b), streptavidin-coated beads (DynabeadsTM MyOneTM Streptavidin C1) in PBS buffer (no protein, negative control, c) or TNAP buffer (no protein, negative control, d). DEL selections were performed in duplicates. Dot colour and size correspond to the normalized sequence counts. (a, left) TCs = 1,299,576, AC = 2.63, Cut-off = 10; a right) TCs = 1,258,947, AC = 2.55, Cut-off = 10; (b, left) TCs = 274,701, AC = 0.56, Cut-off = 10; (b, right) TCs = 253,457, AC = 0.51, Cut-off = 10; (c) TCs = 198,057, AC = 0.40, Cut-off = 10; (d) TCs = 286,279, AC = 0.58, Cut-off = 15.
Extended Data Fig. 4 Additional colorimetric ACP3 inhibition assays.
(a) Experiments performed with conjugates d9, d10, d11, and d12 from SOP-DEL. (b) Experiment performed with oligonucleotides bearing building block A2723 (d13 and d14) and (c) oligonucleotides bearing building block A2855 (d15 and d16) from FMA-DEL. Colorimetric ACP3 inhibition assay of B1326 (SOP-DEL), B125 (FMA-DEL), and their regioisomer derivatives were further performed (d, e). Possible exit-vector modifications of the benchmark scaffold were explored on the phenyl side (d) with compounds s13a-c, or benzyl side (e) with compounds s14a-c. ProXBM-DOTAGA, derived from compound s13c, was selected as the most potent conjugate, and used for further in vitro and in vivo studies. IC50 values are given as mean, with error bars indicating standard deviations of the replicates (n = 3).
Extended Data Fig. 5 Selectivity studies with DEL-derived hits by fluorescence polarization.
In comparison to (a) benchmark compound s56, DEL-derived-compounds (b) ProX2-(S)-Fluo (s27a), (c) ProX2-(R)-Fluo (s27b), (d) ProX1-(SS).Fluo (8), (e) ProX3-(S)-Fluo (9), (f) ProX3-(R)-Fluo (s33) presented >50-fold improvement of binding affinity. The molecules presented no-cross reactivity with alternative phosphatases (PLAP and TNAP) and bound to albumins (HSA and MSA) in the micromolar concentration range. Final compound concentration used corresponds to 5 nM for test items. Data is plotted as mean, with error bars indicating standard deviations of the replicates (n = 3, except for PLAP and TNAP in (d) where n = 2).
Extended Data Fig. 6 Radioligand bead- and cell-based assays.
(a) Radioligand bead-based assay with 177Lu-OncoACP3 (s44) and 177Lu-ProX3-(S)-DOTAGA (12). Magnetic Streptavidin-coated DynabeadsTM M-280 were functionalized with recombinant ACP3 and exposed to test compounds s44 and 12, without (full bars) or with (dashed bars) a 5,000-fold molar excess of blocking cold ACP3 ligands. Assay was performed in accordance with prior literature56. (b) Radioligand cell-based assay with 177Lu-OncoACP3 (s44) and 177Lu-ProX3-(S)-DOTAGA (12). Cells were exposed to test compounds s44 and 12, without (full bars) or with (dashed bars) a 1,000-fold molar excess of blocking cold ACP3 ligands. Assay was performed in accordance with prior literature57. Data is plotted as mean, with error bars indicating standard deviations of the replicates (n = 3).
Extended Data Fig. 7 Additional Autoradiography results.
Human prostate cancer tissue samples were exposed to ACP3-targeted 177Lu-labelled compounds s44 and 12. Patient IDs are presented on the left (University of Pisa’s tissue biobank).
Extended Data Fig. 8 Additional in vivo biodistribution studies with ACP3-targeted RLTs in tumour-bearing mice.
(a) Comparison of 177Lu-ProX1-(SS)-DOTAGA (11) and 177Lu-ProX1-(RR)-DOTAGA (s39) at the 24 h time point and 62.5 nmol/kg dose. (b) Comparison of 177Lu-ProX2-DOTAGA enantiomers (s40 or s41) and 177Lu-ProX3-DOTAGA enantiomers (12 or s42) at the 2 h time point and 62.5 nmol/kg dose. (c) Comparison of 177Lu-ProX1-(SS)-DOTAGA (11), 177Lu-ProX3-(S)-DOTAGA (12), and 177Lu-ProXBM-DOTAGA (10) at the 2 h time point and 62.5 nmol/kg dose. Biodistribution studies were performed with BALB/c nu/nu female (a and b) and male (c and d) mice, at a molar activity ranging from 0.05 and 0.8 MBq/nmol (~1 MBq/mouse). Data for all biodistribution studies are presented as mean of %ID g−1 ± standard error of the mean (SEM) (n = 3 mice/group). Overall, differences in tumour uptake of ~10-20% ID/g at early time points were observed when repeating in vivo biodistribution experiments using the HT1080.hACP3 cell line (see also Fig. 5). This observation could be explained by differences in vascular permeability that naturally occur when implanting tumour cells in different animal batches. (d) In vivo biodistribution studies with 68Ga-OncoACP3 (s45, 62.5 nmol/kg) at the 1 h time point. Mice in the pre-blocking group were injected with cold OncoACP3 (50 nmol/mouse, ~2.5 µmol/kg – corresponding to a 40-fold molar excess as compared to the radioactive compound) 30 min before the administration of 68Ga-OncoACP3 (s45). Individual values are represented by circles for which bars display the average group %ID g−1 values. Error bars indicate the standard error of the mean (SEM) (n = 3 mice/group for the 68Ga-OncoACP3, n = 2 mice/group for the pre-blocking and ACP3-negative SK-RC-52.wt groups).
Extended Data Fig. 9 In vivo biodistribution and PET/CT imaging studies with 68Ga-OncoACP3 in tumour-bearing mice.
(a) In vivo biodistribution study with 68Ga-OncoACP3 (s45, 62.5 nmol/g, ~1 MBq/mouse) at 1 and 2 h post-injection in BALB/c nu/nu male mice (n = 3 mice/group). (b) Maximum intensity projection (MIP) obtained 1 h after injection (15 min exposure time). The arrows indicate the urinary bladder (U; highlighting renal excretion of the compound), the gallbladder (G; highlighting hepatobiliary excretion of the compound), and the tumour (T). (c) MicroPET images were obtained 1 h post-injection of 68Ga-OncoACP3 (s45) in the HT1080.hACP3 xenograft model. The subcutaneous tumour is highlighted by the cross section of the green (coronal), red (transversal) and yellow (para sagittal) planes. Dose: 62.5 nmol/kg (600 MBq/kg). A scale bar is included, displaying %ID g−1 (equal to 1/[g body weight]*SUV*100).
Extended Data Fig. 10 Additional ex vivo biodistribution studies with DEL-derived ACP3 ligands labelled with AF488.
Ex vivo biodistribution of ProX1-(SS)-AF488 (a, compound 14) and ProX3-(S)-AF488 (b, compound 15) in HT1080.hACP3 tumour-bearing mice, 2 h after intravenous administration. Compounds were administered at the 30 nmol/mouse dose (150 µL/animal, PBS solution). Green = compound 14 or 15, Blue = DAPI staining. Scale bar = 400 µm (full scale) or 100 µm (zoomed-in pictures).
Supplementary information
Supplementary Information (download PDF )
Detailed information on general procedures, Synthetic protocols, Compound characterization, Instrumentations and Supplemental results.
Source data
Source Data For Figs. 2–5, 7 and 8 and Extended Data Figs. 2–9a (download ZIP )
Source data: DEL selections (.txt files), enzymatic assays, SPR measurements, biodistribution studies, therapy experiments, FP studies, radioligand cell- and bead-based binding assays, flow cytometry analysis, unprocessed confocal microscopy and autoradiography images.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Georgiev, T., Migliorini, F., Ciamarone, A. et al. Discovery of high-affinity ligands for prostatic acid phosphatase via DNA-encoded library screening enables targeted cancer therapy. Nat. Biomed. Eng 10, 178–191 (2026). https://doi.org/10.1038/s41551-025-01432-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41551-025-01432-6