Abstract
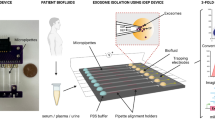
Circulating extracellular vesicles can be used for tumour diagnostics. However, current isolation methods are time consuming, require manual handling and are prone to contamination. Here we report on SpinEx (separation-processing integration for extracellular vesicles), a compact disc device for automatic isolation and multiplex immunolabelling of whole-blood samples. SpinEx integrates on-disc chromatography, centripetal liquid transfer and bead-based vesicle capture with antibody labelling. The system processes 150 µl of whole blood, enriching and labelling vesicles for 16 protein targets in under 75 minutes. Detection is performed by measuring dual fluorescence signals from labelled extracellular vesicles captured on microbeads. In a pilot clinical study, SpinEx was used to process 221 plasma samples for multiplex profiling of 30 vesicle-associated proteins. Using fluorescence flow cytometry to analyse cancer-specific biomarker expression, we found that vesicles processed by SpinEx distinguished cancer from non-cancer samples with 90% accuracy and 97% specificity, and classified 5 tumour types with 96% accuracy. SpinEx enables automated and multiplex processing of extracellular vesicles from blood, which may support the development of clinically viable assays for cancer detection and classification.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The raw patient datasets generated and analysed during the study are available from the corresponding authors, subject to approval from the Institutional Review Boards of MGH, Kyungpook National University Medical Center and Chonnam National University Hwasun Hospital. Source data are provided with this paper.
Code availability
Source codes for the machine learning are available at https://github.com/kylie0914/SpinEx.
References
Chi, K. R. The tumour trail left in blood. Nature 532, 269–271 (2016).
Siravegna, G., Marsoni, S., Siena, S. & Bardelli, A. Integrating liquid biopsies into the management of cancer. Nat. Rev. Clin. Oncol. 14, 531–548 (2017).
Heitzer, E., Haque, I. S., Roberts, C. E. S. & Speicher, M. R. Current and future perspectives of liquid biopsies in genomics-driven oncology. Nat. Rev. Genet. 20, 71–88 (2019).
Ignatiadis, M., Sledge, G. W. & Jeffrey, S. S. Liquid biopsy enters the clinic—implementation issues and future challenges. Nat. Rev. Clin. Oncol. 18, 297–312 (2021).
Théry, C., Ostrowski, M. & Segura, E. Membrane vesicles as conveyors of immune responses. Nat. Rev. Immunol. 9, 581–593 (2009).
Xu, R. et al. Extracellular vesicles in cancer—implications for future improvements in cancer care. Nat. Rev. Clin. Oncol. 15, 617–638 (2018).
van Niel, G., D’Angelo, G. & Raposo, G. Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell Biol. 19, 213–228 (2018).
Buzas, E. I. The roles of extracellular vesicles in the immune system. Nat. Rev. Immunol. 23, 236–250 (2023).
Im, H. et al. Label-free detection and molecular profiling of exosomes with a nano-plasmonic sensor. Nat. Biotechnol. 32, 490–495 (2014).
Vinik, Y. et al. Proteomic analysis of circulating extracellular vesicles identifies potential markers of breast cancer progression, recurrence, and response. Sci. Adv. 6, eaba5714 (2020).
Skog, J. et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat. Cell Biol. 10, 1470–1476 (2008).
Blandin, A. et al. Lipidomic analysis of adipose-derived extracellular vesicles reveals specific EV lipid sorting informative of the obesity metabolic state. Cell Rep. 42, 112169 (2023).
Ferguson, S. et al. Single-EV analysis (sEVA) of mutated proteins allows detection of stage 1 pancreatic cancer. Sci. Adv. 8, eabm3453 (2022).
Jo, A. et al. Inaugurating high-throughput profiling of extracellular vesicles for earlier ovarian cancer detection. Adv. Sci. 10, e2301930 (2023).
Liu, C. et al. Low-cost thermophoretic profiling of extracellular-vesicle surface proteins for the early detection and classification of cancers. Nat. Biomed. Eng. 3, 183–193 (2019).
Hinestrosa, J. P. et al. Early-stage multi-cancer detection using an extracellular vesicle protein-based blood test. Commun. Med. 2, 29 (2022).
Shin, H. et al. Single test-based diagnosis of multiple cancer types using exosome-SERS-AI for early stage cancers. Nat. Commun. 14, 1644 (2023).
Shao, H. et al. Protein typing of circulating microvesicles allows real-time monitoring of glioblastoma therapy. Nat. Med. 18, 1835–1840 (2012).
Chen, G. et al. Exosomal PD-L1 contributes to immunosuppression and is associated with anti-PD-1 response. Nature 560, 382–386 (2018).
Zhou, E. et al. Circulating extracellular vesicles are effective biomarkers for predicting response to cancer therapy. eBioMedicine 67, 103365 (2021).
Ghodasara, A., Raza, A., Wolfram, J., Salomon, C. & Popat, A. Clinical translation of extracellular vesicles. Adv. Healthc. Mater. 12, e2301010 (2023).
Sunkara, V., Woo, H. K. & Cho, Y. K. in Extracellular Vesicles: Applications to Regenerative Medicine, Therapeutics and Diagnostics (eds Chrzanowski, W. et al.) 22–44 (The Royal Society of Chemistry, 2021).
Simonsen, J. B. What are we looking at? Extracellular vesicles, lipoproteins, or both. Circ. Res. 121, 920–922 (2017).
Vergauwen, G. et al. Confounding factors of ultrafiltration and protein analysis in extracellular vesicle research. Sci Rep. 7, 2704 (2017).
Johnsen, K. B., Gudbergsson, J. M., Andresen, T. L. & Simonsen, J. B. What is the blood concentration of extracellular vesicles? Implications for the use of extracellular vesicles as blood-borne biomarkers of cancer. Biochim. Biophys. Acta Rev. Cancer 1871, 109–116 (2019).
Onódi, Z. et al. Isolation of high-purity extracellular vesicles by the combination of iodixanol density gradient ultracentrifugation and bind-elute chromatography from blood plasma. Front. Physiol. 9, 1479 (2018).
Zhang, X., Borg, E. G. F., Liaci, A. M., Vos, H. R. & Stoorvogel, W. A novel three step protocol to isolate extracellular vesicles from plasma or cell culture medium with both high yield and purity. J. Extracell. Vesicles 9, 1791450 (2020).
Havers, M., Broman, A., Lenshof, A. & Laurell, T. Advancement and obstacles in microfluidics-based isolation of extracellular vesicles. Anal. Bioanal. Chem. 415, 1265–1285 (2023).
Madou, M. et al. Lab on a CD. Annu. Rev. Biomed. Eng. 8, 601–628 (2006).
Gorkin, R. et al. Centrifugal microfluidics for biomedical applications. Lab Chip 10, 1758–1773 (2010).
Kim, T. H. et al. A lab-on-a-disc with reversible and thermally stable diaphragm valves. Lab Chip 16, 3741–3749 (2016).
Grumann, M., Geipel, A., Riegger, L., Zengerle, R. & Ducrée, J. Batch-mode mixing on centrifugal microfluidic platforms. Lab Chip 5, 560–565 (2005).
Watts, A. S. et al. Centrifugal microfluidics with integrated sensing microdome optodes for multiion detection. Anal. Chem. 79, 8046–8054 (2007).
Woo, H. K. et al. Urine-based liquid biopsy: non-invasive and sensitive AR-V7 detection in urinary EVs from patients with prostate cancer. Lab Chip 19, 87–97 (2018).
Klatt, J. N. et al. Miniaturization, parallelization, and automation of endotoxin detection by centrifugal microfluidics. Anal. Chem. 93, 8508–8516 (2021).
Park, J., Sunkara, V., Kim, T. H., Hwang, H. & Cho, Y. K. Lab-on-a-disc for fully integrated multiplex immunoassays. Anal. Chem. 84, 2133–2140 (2012).
Kong, L. X., Perebikovsky, A., Moebius, J., Kulinsky, L. & Madou, M. Lab-on-a-CD: a fully integrated molecular diagnostic system. J. Lab. Autom. 21, 323–355 (2016).
Kim, T. H. et al. FAST: size-selective, clog-free isolation of rare cancer cells from whole blood at a liquid–liquid interface. Anal. Chem. 89, 1155–1162 (2017).
Woo, H. K. et al. Exodisc for rapid, size-selective, and efficient isolation and analysis of nanoscale extracellular vesicles from biological samples. ACS Nano 11, 1360–1370 (2017).
Sunkara, V. et al. Fully automated, label-free isolation of extracellular vesicles from whole blood for cancer diagnosis and monitoring. Theranostics 9, 1851–1863 (2019).
Dong, L. et al. Comprehensive evaluation of methods for small extracellular vesicles separation from human plasma, urine and cell culture medium. J. Extracell. Vesicles 10, e12044 (2020).
Hill, W. D., Rothfus, R. R. & Li, K. Boundary-enhanced sedimentation due to settling convection. Int. J. Multiphase Flow 3, 561–583 (1977).
Woo, H. K. et al. Characterization and modulation of surface charges to enhance extracellular vesicle isolation in plasma. Theranostics 12, 1988–1998 (2022).
Van Deun, J. et al. Integrated dual-mode chromatography to enrich extracellular vesicles from plasma. Adv. Biosyst. 4, e1900310 (2020).
Latour, R. A. Fundamental principles of the thermodynamics and kinetics of protein adsorption to material surfaces. Colloids Surf. B 191, 110992 (2020).
Jonsson, A. L., Roberts, M. A. J., Kiappes, J. L. & Scott, K. A. Essential chemistry for biochemists. Essays Biochem. 61, 401–427 (2017).
Lyklema, J. Proteins at solid–liquid interfaces a colloid-chemical review. Colloids Surf. 10, 33–42 (1984).
Lee, K. et al. Multiplexed profiling of single extracellular vesicles. ACS Nano 12, 494–503 (2018).
Spitzberg, J. D. et al. Multiplexed analysis of EV reveals specific biomarker composition with diagnostic impact. Nat. Commun. 14, 1239 (2023).
Park, J. et al. An integrated magneto-electrochemical device for the rapid profiling of tumour extracellular vesicles from blood plasma. Nat. Biomed. Eng. 5, 678–689 (2021).
Jeong, S. et al. Integrated magneto-electrochemical sensor for exosome analysis. ACS Nano 10, 1802–1809 (2016).
Hallal, S., Tűzesi, Á, Grau, G. E., Buckland, M. E. & Alexander, K. L. Understanding the extracellular vesicle surface for clinical molecular biology. J. Extracell. Vesicles 11, e12260 (2022).
Shao, H. et al. Chip-based analysis of exosomal mRNA mediating drug resistance in glioblastoma. Nat. Commun. 6, 6999 (2015).
Hong, S., Park, K. S., Weissleder, R., Castro, C. M. & Lee, H. Facile silicification of plastic surface for bioassays. Chem. Commun. 53, 2134–2137 (2017).
Yang, K. S. et al. Extracellular vesicle analysis allows for identification of invasive IPMN. Gastroenterology 160, 1345–1358.e11 (2021).
Lee, Y., Cho, W., Sung, J., Kim, E. & Park, S. B. Monochromophoric design strategy for tetrazine-based colorful bioorthogonal probes with a single fluorescent core skeleton. J. Am. Chem. Soc. 140, 974–983 (2018).
Roederer, M. Compensation in flow cytometry. Curr. Protoc. Cytom. 2, 1.14.1–1.14.20 (2022).
Leys, C., Ley, C., Klein, O., Bernard, P. & Licata, L. Detecting outliers: do not use standard deviation around the mean, use absolute deviation around the median. J. Exp. Soc. Psychol. 49, 764–766 (2013).
Osher, S. & Fedkiw, R. P. Level set methods: an overview and some recent results. J. Comput. Phys. 169, 463–502 (2001).
Han, L. S. Hydrodynamic entrance lengths for incompressible laminar flow in rectangular ducts. J. Appl. Mech. 27, 403–409 (1960).
Granger, R. A. Fluid Mechanics (Dover Publications, 1995).
Cortes, C. & Vapnik, V. Support-vector networks. Mach. Learn. 20, 273–297 (1995).
Song, J. et al. Amplifying mutational profiling of extracellular vesicle mRNA with scope. Nat. Biotechnol. 43, 1485–1495 (2025).
Acknowledgements
We thank D. G. You (MGH) and H. Lee (MGH) for their assistance in bead imaging. This work was supported in part by NIH 1U01CA279858 (C.M.C. and H.L.), U01CA284982 (H.L. and C.M.C.), R01CA229777 (H.L.), R01CA239078 (H.L.), R01HL163513 (H.L.), R01CA237500 (H.L.), R21CA267222 (H.L.), R01CA264363 (C.M.C. and H.L.) and R61CA297878 (H.L.); the Korea Health Industry Development Institute grant HR22C1832 (J.S.P.); Samsung Research Funding and Incubation Center of Samsung Electronics SRFC-IT1901-51 (S.C.); and a National Research Foundation grant funded by the Korean government 2022M3A9B6018217 (S.C.).
Author information
Authors and Affiliations
Contributions
H.-K.W., C.K., C.M.C., J.S.P. and H.L. designed the study, prepared the figures and wrote the paper. H.-K.W., Y.K.C. and H.K. conducted the experiments. J.J. conducted a Western blot. H.-K.W. designed the device, and H.K. and S.C. assisted in its implementation. H.K. and D.-H.J. performed the fluidic simulation. H.-K.W., Y.C., L.-N.D., M.A., I.P. and H.L. developed the machine learning models. C.K., S.Y.P., C.M.C. and J.S.P. analysed clinical data. C.K., S.Y.P. and J.S.P. acquired clinical samples. All authors contributed to writing the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors (H.-K.W., Y.C. and H.L.) are inventors on an invention disclosure related to components of the SpinEx technology, which was filed and assigned to MGH. This intellectual property is related to the methods and devices described in this paper. The other authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks Lingxin Chen, Mei He and Takao Yasui for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 SpinEx operation steps.
1 A whole-blood (150 µL) sample is loaded. 2 Plasma is separated from the whole blood. 3 Plasma is transferred to an on-disc chromatography column, and an elution buffer is introduced. 4 A dense oil pushes the enriched EVs toward the inner side for further processing. 5 EVs are transferred to an assay chamber containing polystyrene beads. EVs bind to the bead surface via physisorption. 6 EV-bead complexes are washed by introducing a buffer. 7 TFP-biotin reagent is introduced to the assay chamber, and EVs are biotinylated. TFP, 2,3,5,6-tetrafluorophenyl. 8 Excess biotinylation reagent is removed via washing. 9 A block solution (10% Superblock) is introduced to the assay chamber. 10 The blocking solution is removed with another washing step. 11 EV-bead complexes are dispensed into eight labeling compartments, each containing a primary antibody (1° Ab) against a target protein. 12 Dye-conjugated streptavidin and secondary antibody (2° Ab) are introduced to the compartments. The labelled EV-bead complexes are ready for fluorescent measurements (for example, microscopy, flow cytometry).
Extended Data Fig. 2 Workflow for machine learning analyses.
(a) Constructing a model for cancer diagnosis. The LASSO regression determined the most informative marker sets from the training set (n = 154; left). The selected markers were used to train an SVM model for binary classification (cancer versus non-cancer). This trained model was evaluated using an independent test set (n = 67; right). (b) Model for five-cancer classification. Data from all cancer patients (n = 157) were used. The model performed a one-versus-one classification to differentiate between five tumor types.
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods, Figs. 1–25 and Tables 1–3.
Supplementary Video 1 (download MP4 )
SpinEx disc fluidic workflow. This video illustrates the overall sequence of assay steps performed within a single device, including on-disc plasma separation, dual-mode chromatography–based EV enrichment, bead-based EV capture and biotinylation, and multiplexed fluorescent labeling across eight downstream chambers.
Supplementary Video 2 (download MP4 )
Density-based centripetal fluid transport. This video shows the simulation of density-driven fluidic exchange during the centripetal transfer step. The high-density oil phase displaces the aqueous EV solution toward the disc center under rotation (1800 rpm), enabling rapid (<2 s) transfer between adjacent assay chambers.
Supplementary Video 3 (download MP4 )
Fluidic simulations of a cascading elution. This video shows the simulated flow pattern of the on-disc chromatography module during disc rotation. The eluate first fills the chamber closest to the column outlet (F3) and then sequentially transfers to the adjacent chambers (F2 and F1), producing an inverted elution order compared with conventional column devices. This cascading liquid transfer explains why early fractions are collected in F1 and late fractions, including potential (V)LDL-rich eluates, are collected in F3, enabling optimized selection of the EV-enriched fraction in F2.
Source data
Source Data Fig. 2 (download XLSX )
Raw numerical data used to generate the plots in Fig. 2.
Source Data Fig. 3 (download ZIP )
Uncropped transmission electron microscopy micrograph and uncropped single-EV fluorescence microscopy images used in Fig. 3.
Source Data Fig. 3 (download XLSX )
Raw numerical data used to generate the plots in Fig. 3.
Source Data Fig. 4 (download XLSX )
Raw numerical data used to generate the plots in Fig. 4.
Source Data Fig. 5 (download ZIP )
Uncropped scanning electron microscopy micrograph and uncropped bead fluorescence microscopy images used in Fig. 5.
Source Data Fig. 5 (download XLSX )
Raw numerical data used to generate the plots in Fig. 5.
Source Data Fig. 6 (download XLSX )
Raw numerical data used to generate the plots in Fig. 6.
Source Data Fig. 7 (download XLSX )
Raw numerical data used to generate the plots in Fig. 7.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Woo, HK., Kim, C., Choi, Y. et al. Automated disc device for multiplexed extracellular vesicle isolation and labelling from liquid biopsies in cancer diagnostics. Nat. Biomed. Eng (2026). https://doi.org/10.1038/s41551-025-01601-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41551-025-01601-7