Abstract
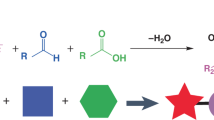
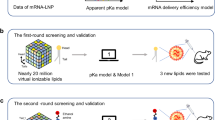
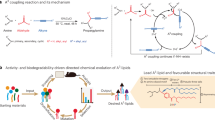
Efficient mRNA delivery to specific tissues requires optimized ionizable lipids, yet the role of lipid spatial conformation in organ targeting and endosomal escape remains underexplored. Here we developed a library of lipids with diverse amino heads, degradable linkers and hydrophobic tails, generating distinct three-dimensional conformations. Molecular dynamics simulations revealed the dynamic conformations of these lipids during organic–aqueous phase transitions, and experimental validation confirmed that head and tail arrangements are key determinants of delivery efficiency and organ specificity. To accelerate lipid discovery, dynamic conformation data were converted into 2D density images to train machine learning models for lipid selection. AI-guided candidates, notably lipid P1, adopted stable three-tail cone-shaped conformations that promoted IgM protein corona formation and enabled spleen-targeted mRNA delivery. In preclinical models, P1-based mRNA vaccines triggered strong antibody and T-cell responses, leading to marked tumour suppression. These results highlight the pivotal role of lipid spatial conformation and the potential of AI-driven strategies to optimize lipid nanoparticles for organ-specific mRNA delivery.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All relevant data supporting the findings of this study are available within the Article, its Supplementary Information files or Source Data files. Source data are provided with this paper.
Code availability
The custom code for constructing the 2D density maps of lipids in ML is available in Supplementary Information files.
References
Su, L.-J. et al. Delivery of mRNA for cancer therapy: progress and prospects. Nano Today 53, 102013 (2023).
Huang, X. G. et al. The landscape of mRNA nanomedicine. Nat. Med. 28, 2273–2287 (2022).
Baden, L. R. et al. Efficacy and safety of the mRNA-1273 SARS-CoV-2 vaccine. N. Engl. J. Med. 384, 403–416 (2021).
Polack, F. P. et al. Safety and efficacy of the BNT162b2 mRNA Covid-19 vaccine. N. Engl. J. Med. 383, 2603–2615 (2020).
Zhang, Y., Sun, C., Wang, C., Jankovic, K. E. & Dong, Y. Lipids and lipid derivatives for RNA delivery. Chem. Rev. 121, 12181–12277 (2021).
Sabnis, S. et al. A novel amino lipid series for mRNA delivery: improved endosomal escape and sustained pharmacology and safety in non-human primates. Mol. Ther. 26, 1509–1519 (2018).
Akinc, A. et al. The Onpattro story and the clinical translation of nanomedicines containing nucleic acid-based drugs. Nat. Nanotechnol. 14, 1084–1087 (2019).
Zhang, Y. et al. Close the cancer–immunity cycle by integrating lipid nanoparticle–mRNA formulations and dendritic cell therapy. Nat. Nanotechnol. 18, 1364–1374 (2023).
Lee, S. M. et al. A systematic study of unsaturation in lipid nanoparticles leads to improved mRNA transfection in vivo. Angew. Chem. Int. Ed. 60, 5848–5853 (2021).
Hajj, K. A. et al. Branched-tail lipid nanoparticles potently deliver mRNA in vivo due to enhanced ionization at endosomal pH. Small 15, e1805097 (2019).
Hou, X. et al. Vitamin lipid nanoparticles enable adoptive macrophage transfer for the treatment of multidrug-resistant bacterial sepsis. Nat. Nanotechnol. 15, 41–46 (2020).
Zhao, X. et al. Imidazole-based synthetic lipidoids for in vivo mRNA delivery into primary T lymphocytes. Angew. Chem. Int. Ed. 59, 20083–20089 (2020).
Qiu, M. et al. Lung-selective mRNA delivery of synthetic lipid nanoparticles for the treatment of pulmonary lymphangioleiomyomatosis. Proc. Natl Acad. Sci. USA 119, e2116271119 (2022).
Cheng, Q. et al. Selective organ targeting (SORT) nanoparticles for tissue-specific mRNA delivery and CRISPR-Cas gene editing. Nat. Nanotechnol. 15, 313–320 (2020).
Liu, S. et al. Membrane-destabilizing ionizable phospholipids for organ-selective mRNA delivery and CRISPR-Cas gene editing. Nat. Mater. 20, 701–710 (2021).
Hatit, M. Z. C. et al. Nanoparticle stereochemistry-dependent endocytic processing improves in vivo mRNA delivery. Nat. Chem. 15, 508–515 (2023).
Han, X. et al. An ionizable lipid toolbox for RNA delivery. Nat. Commun. 12, 7233 (2021).
Dong, W. et al. Multicomponent synthesis of imidazole-based ionizable lipids for highly efficient and spleen-selective messenger RNA delivery. J. Am. Chem. Soc. 146, 15085–15095 (2024).
Chen, I. J. & Foloppe, N. Is conformational sampling of drug-like molecules a solved problem? Drug Dev. Res. 72, 85–94 (2010).
Xu, Y. et al. AGILE platform: a deep learning powered approach to accelerate LNP development for mRNA delivery. Nat. Commun. 15, 6305 (2024).
Li, B. et al. Accelerating ionizable lipid discovery for mRNA delivery using machine learning and combinatorial chemistry. Nat. Mater. 23, 1002–1008 (2024).
Witten, J. et al. Artificial intelligence-guided design of lipid nanoparticles for pulmonary gene therapy. Nat. Biotechnol. 43, 1790–1799 (2025).
Philipp, J. et al. pH-dependent structural transitions in cationic ionizable lipid mesophases are critical for lipid nanoparticle function. Proc. Natl Acad. Sci. USA 120, e2310491120 (2023).
Yu, H. et al. Real-time pH-dependent self-assembly of ionisable lipids from COVID-19 vaccines and in situ nucleic acid complexation. Angew. Chem. Int. Ed. 62, e202304977 (2023).
Akinc, A. et al. A combinatorial library of lipid-like materials for delivery of RNAi therapeutics. Nat. Biotechnol. 26, 561–569 (2008).
Patel, P., Ibrahim, N. M. & Cheng, K. The importance of apparent pKa in the development of nanoparticles encapsulating siRNA and mRNA. Trends Pharmacol. Sci. 42, 448–460 (2021).
Jayaraman, M. et al. Maximizing the potency of siRNA lipid nanoparticles for hepatic gene silencing in vivo. Angew. Chem. Int. Ed. 51, 8529–8533 (2012).
Zong, Y., Lin, Y., Wei, T. & Cheng, Q. Lipid nanoparticle (LNP) enables mRNA delivery for cancer therapy. Adv. Mater. 35, e2303261 (2023).
Ouyang, R., Curtarolo, S., Ahmetcik, E., Scheffler, M. & Ghiringhelli, L. M. SISSO: a compressed-sensing method for identifying the best low-dimensional descriptor in an immensity of offered candidates. Phys. Rev. Mater. 2, 083802 (2018).
Ouyang, R., Yoo, S.-H. & Jang, W. Small dataset machine-learning approach for efficient design space exploration: engineering ZnTe-based high-entropy alloys for water splitting. npj Comput. Mater. 10, 166 (2024).
Liu, X. et al. Finding predictive models for singlet fission by machine learning. npj Comput. Mater. 8, 70 (2022).
Wang, T. et al. Nature of metal–support interaction for metal catalysts on oxide supports. Science 386, 915–920 (2024).
Chen, V. et al. Applying interpretable machine learning in computational biology—pitfalls, recommendations and opportunities for new developments. Nat. Methods 21, 1454–1461 (2024).
Nel, A. E. et al. Understanding biophysicochemical interactions at the nano–bio interface. Nat. Mater. 8, 543–557 (2009).
Dilliard, S. A., Cheng, Q. & Siegwart, D. J. On the mechanism of tissue-specific mRNA delivery by selective organ targeting nanoparticles. Proc. Natl Acad. Sci. USA 118, e2109256118 (2021).
Schvartz, I., Seger, D. & Shaltiel, S. Vitronectin. Int. J. Biochem. Cell Biol. 31, 539–544 (1999).
Dormond, O., Foletti, A., Paroz, C. & Rüegg, C. NSAIDs inhibit alpha V beta 3 integrin-mediated and Cdc42/Rac-dependent endothelial-cell spreading, migration and angiogenesis. Nat. Med. 7, 1041–1047 (2001).
Ehrenstein, M. R. & Notley, C. A. The importance of natural IgM: scavenger, protector and regulator. Nat. Rev. Immunol. 10, 778–786 (2010).
Li, Y. et al. Immunoglobulin M perception by FcμR. Nature 615, 907–912 (2023).
Helmink, B. A. et al. B cells and tertiary lymphoid structures promote immunotherapy response. Nature 577, 549–555 (2020).
Hu, X. et al. Landscape of B cell immunity and related immune evasion in human cancers. Nat. Genet. 51, 560–567 (2019).
Zhu, Y. et al. Screening for lipid nanoparticles that modulate the immune activity of helper T cells towards enhanced antitumour activity. Nat. Biomed. Eng. 8, 544–560 (2024).
Zhao, S. et al. Acid-degradable lipid nanoparticles enhance the delivery of mRNA. Nat. Nanotechnol. 19, 1702–1711 (2024).
Chen, J. et al. Combinatorial design of ionizable lipid nanoparticles for muscle-selective mRNA delivery with minimized off-target effects. Proc. Natl Acad. Sci. USA 120, e2309472120 (2023).
Chaudhary, N. et al. Amine headgroups in ionizable lipids drive immune responses to lipid nanoparticles by binding to the receptors TLR4 and CD1d. Nat. Biomed. Eng. 8, 1483–1498 (2024).
Miao, L. et al. Delivery of mRNA vaccines with heterocyclic lipids increases anti-tumor efficacy by STING-mediated immune cell activation. Nat. Biotechnol. 37, 1174–1185 (2019).
He, Z. et al. A multidimensional approach to modulating ionizable lipids for high-performing and organ-selective mRNA delivery. Angew. Chem. Int. Ed. 62, e202310401 (2023).
Xue, L. et al. Combinatorial design of siloxane-incorporated lipid nanoparticles augments intracellular processing for tissue-specific mRNA therapeutic delivery. Nat. Nanotechnol. 20, 132–143 (2025).
Han, X. et al. Fast and facile synthesis of amidine-incorporated degradable lipids for versatile mRNA delivery in vivo. Nat. Chem. 16, 1687–1697 (2024).
Van Der Spoel, D. et al. GROMACS: fast, flexible, and free. J. Comput. Chem. 26, 1701–1718 (2005).
Lee, J. et al. CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. J. Chem. Theory Comput. 12, 405–413 (2016).
Huang, J. & MacKerell, A. D. Jr. CHARMM36 all-atom additive protein force field: validation based on comparison to NMR data. J. Comput. Chem. 34, 2135–2145 (2013).
Vanommeslaeghe, K. et al. CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010).
Yu, W., He, X., Vanommeslaeghe, K. & MacKerell, A. D. Jr. Extension of the CHARMM General Force Field to sulfonyl-containing compounds and its utility in biomolecular simulations. J. Comput. Chem. 33, 2451–2468 (2012).
Sarzynska, J., Popenda, M., Antczak, M. & Szachniuk, M. RNA tertiary structure prediction using RNAComposer in CASP15. Proteins 91, 1790–1799 (2023).
Popenda, M. et al. Automated 3D structure composition for large RNAs. Nucleic Acids Res. 40, e112 (2012).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865 (2008).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 27–28 (1996).
Acknowledgements
This work was supported by the National Key R&D Program of China (2022YFA1205700 to Y.-X.L. and H.W., 2024YFD2101600 to Y.-X.L., 2024YFA1509600 and 2022YFA1203200 to Y.G.), the National Natural Science Foundation of China (32371458 to Y.-X.L., 22273014 to Y.G.), the Beijing Natural Science Foundation (L242036 to Y.-X.L.), the Basic Research Cooperation Special Foundation of Beijing–Tianjin–Hebei (22J00017 to Y.-X.L.) and the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB1030000 to Y.G.). Y.-X. L. acknowledges the start-up funding from the National Center for Nanoscience and Technology and the Chinese Academy of Sciences. The funders had no roles in the study design, data collection and analysis, decision to publish, and preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Y.-X.L. and L.-J.S. conceived the idea and designed the experiments. Y.W. and Y.-X.L. directed this project. L.-J.S., Z.-H.J., M.-X.X., M.-Z.Y., C.L. and J.Z. performed the synthesis experiments of ionizable lipids. L.-J.S., C.Y., Q.C. and Z.-H.J. performed the in vitro and in vivo screening studies. L.-J.S. collected and analysed the cryo-transmission electron microscopy data under the supervision of K.F.; R.L. conducted the MD experiments and machine learning under the supervision of Y.G.; L.-J.S. and Z.-H.J. collected and analysed the proteomics data. N.-N.W., L.-J.S. and Z.-H.J. performed the organ-targeting experiments. N.-N.W. and H.G. performed the mRNA vaccine experiments under the supervision of Y.W. and Y.-X.L.; L.M., Y.Z. and H.W. provided reagents and conceptual advice. Z.-H.J., L.-J.S., R.L. and N.-N.W. prepared figures and the first draft of the paper. Y.W., Y.-X.L., Y.G. and H.W. revised the paper. All authors discussed the results and computational methods and commented on the paper.
Corresponding authors
Ethics declarations
Competing interests
Y.-X.L., L.-J.S., Y.G. and R.L. have applied for patent applications (202510268720.4, China, 2025; 202411880079.1, China, 2024) related to this study. Y.-X.L. and J.Z. are founders and hold equity in Messerna BioTech. The other authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Feature extraction for ML training.
a, Twenty-two 3D spatial conformation features extracted from molecular density maps based on geometric properties (for example, angle, width and length). b, Six foundational 2D chemical structure features. c, Pearson correlation coefficient (PCC) matrix analysis of the 28 features. d, Feature importance analysis of the twenty-eight features, revealing their relative contribution to the model’s predictive performance. e, Prediction formula of the best-performing ML model (Model 1): ypre = c1D1 + c2D2 + c3D3 + c4D4 + c5D5 + c0, with coefficients and feature descriptions listed in the table.
Extended Data Fig. 2 ML model selection and testing.
a, Performance evaluation of the top four ML models based on accuracy, precision, recall, and F1-score, showing Model 1 as the best one. b, Comparison of ML-predicted and experimental measured delivery efficiencies for 15 test lipids, with MC3 served as a benchmark. c, Luciferase expression after treatment of HEK293T cells with LNPs (1 μg mL−1 mFluc, 24 h, n = 3 biologically independent samples). d, GFP expression after treatment of HEK293T cells with LNPs (1 μg mL−1 mGFP, 24 h, n = 3 independent biological samples). e, Representative confocal images of d. Scale bar, 50 μm. The data are presented as the mean ± s.d.
Extended Data Fig. 3 Changes in spatial conformations of lipid P1 in different media.
Conformational change of lipid P1 upon transition from ethanol to an acidic aqueous phase during LNP preparation. Part of this figure was created with Biorender.com.
Extended Data Fig. 4 Changes in spatial conformations of lipid P1 and lipid 9 during protonation.
Conformational change of lipids P1 (a) and 9 (b) in the medium transitioning from neutral pH to acidic pH, simulating the endosomal escape process.
Extended Data Fig. 5 An in-depth performance evaluation of lipid T2.
a, The chemical structure and spatial conformation of lipid T2. b, Representative bioluminescence images of the local injection alongside quantification of lipid T2 and ALC-0315 (positive control) after i.m. injection (0.75 mg kg−1 mFluc, 6 h, n = 3 biologically independent mice). c, Representative snapshots of MD simulations demonstrating the binding processes of lipids T2 and ALC-0315 (positive control). d, Calculated binding probability of lipid T2/ALC-0315 (positive control) to mRNA (n = 4 independent calculations). e, Snapshots of MD simulations showing the rapid binding of excess lipid T2/ALC-0315 (positive control) to mRNA at 10 ns and 30 ns. f, Binding ratio of lipid T2/ALC-0315 (positive control). g, Representative confocal images of the cellular uptake of lipids T2 and ALC-0315 (positive control) LNPs encapsulating Cy5-labelled mRNA in HEK293T cells at 2 h, 4 h, and 6 h, respectively. Scale bar, 10 μm. h, Quantification of cellular uptake in g (n = 6 biologically independent samples). i, Representative magnified confocal images and fluorescence colocalization analysis profiles of HEK293T cells after coincubation for 6 h, from three independent experiments. Scale bar, 5 μm. The data are all presented as the mean ± s.d. Statistical significance in d was analysed using a two-tailed unpaired Student’s t test. P values < 0.05 (*), P < 0.01 (**), P < 0.001 (***) and P < 0.0001 (****) were considered statistically significant. ns, not significant.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–32, Tables 1–5, Notes and Methods.
Supplementary Data 1 (download XLSX )
Source data for Supplementary Figs. 8–10, 13, 16, 18–21, 23, 25, 26, 28, 31, 32.
Supplementary Data 2 (download XLSX )
Source data for spatial conformation density maps in Figs. 2b, 4c, 5a, 5h, 6d; Extended Data Figs. 3, 4, 5a; Supplementary Figs. 2a, 3a, 4a, 5a, 6a, 11a, 14a, 15a, 17, 22, 24a.
Supplementary Data 3 (download ZIP )
Source data for machine learning models.
Supplementary Data 4 (download ZIP )
Source data for spatial conformation density maps preparation.
Supplementary Code 1 (download ZIP )
Source code for spatial conformation density maps data acquisition.
Source data
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Su, LJ., Wang, NN., Luo, R. et al. Artificial intelligence-guided design of LNPs for in vivo targeted mRNA delivery via analysis of the spatial conformation of ionizable lipids. Nat. Biomed. Eng (2026). https://doi.org/10.1038/s41551-026-01640-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41551-026-01640-8