Abstract
Triple-negative breast cancer (TNBC) is characterized by a high degree of immune infiltrate in the tumour microenvironment, which may influence the fate of TNBC cells. We reveal that loss of the tumour suppressive transcription factor Elf5 in TNBC cells activates intrinsic interferon-γ (IFN-γ) signalling, promoting tumour progression and metastasis. Mechanistically, we find that loss of the Elf5-regulated ubiquitin ligase FBXW7 ensures stabilization of its putative protein substrate IFN-γ receptor 1 (IFNGR1) at the protein level in TNBC. Elf5low tumours show enhanced IFN-γ signalling accompanied by an increase of immunosuppressive neutrophils within the tumour microenvironment and increased programmed death ligand 1 expression. Inactivation of either programmed death ligand 1 or IFNGR1 elicited a robust anti-tumour and/or anti-metastatic effect. A positive correlation between ELF5 and FBXW7 expression and a negative correlation between ELF5, FBXW7 and IFNGR1 expression in the tumours of patients with TNBC strongly suggest that this signalling axis could be exploited for patient stratification and immunotherapeutic treatment strategies for Elf5low patients with TNBC.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The GSEA data can be accessed using the accession numbers GSE122090 and GSE122180. Previously published MDA-MB-231 GSEA data can be accessed at GSE32144. The data and/or reagents that support the findings of this study are available from the corresponding author upon reasonable request. Source data for Figs. 1–8 and Extended Data Figs. 1–5 and 7–10 are provided online. Each experiment was repeated independently with similar results.
Change history
04 August 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41556-021-00733-7
References
Kwa, M. J. & Adams, S. Checkpoint inhibitors in triple-negative breast cancer (TNBC): where to go from here. Cancer 124, 2086–2103 (2018).
Liu, Z., Li, M., Jiang, Z. & Wang, X. A comprehensive immunologic portrait of triple-negative breast cancer. Transl. Oncol. 11, 311–329 (2018).
Garcia-Teijido, P., Cabal, M. L., Fernandez, I. P. & Perez, Y. F. Tumor-infiltrating lymphocytes in triple negative breast cancer: the future of immune targeting. Clin. Med. Insights Oncol. 10, 31–39 (2016).
Kumar, S. et al. DeltaNp63-driven recruitment of myeloid-derived suppressor cells promotes metastasis in triple-negative breast cancer. J. Clin. Invest. 128, 5095–5109 (2018).
Quail, D. F. & Joyce, J. A. Microenvironmental regulation of tumor progression and metastasis. Nat. Med. 19, 1423–1437 (2013).
Markowitz, J., Wesolowski, R., Papenfuss, T., Brooks, T. R. & Carson, W. E. 3rd Myeloid-derived suppressor cells in breast cancer. Breast Cancer Res. Treat. 140, 13–21 (2013).
Hanahan, D. & Coussens, L. M. Accessories to the crime: functions of cells recruited to the tumor microenvironment. Cancer Cell 21, 309–322 (2012).
Zhou, J. et al. Cancer-associated fibroblasts correlate with tumor-associated macrophages infiltration and lymphatic metastasis in triple negative breast cancer patients. J. Cancer 9, 4635–4641 (2018).
Yu, T. & Di, G. Role of tumor microenvironment in triple-negative breast cancer and its prognostic significance. Chin. J. Cancer Res. 29, 237–252 (2017).
Deng, L. et al. Immune profiles of tumor microenvironment and clinical prognosis among women with triple-negative breast cancer. Cancer Epidemiol. Biomark. Prev. 28, 1977–1985 (2019).
Yuan, Z. Y., Luo, R. Z., Peng, R. J., Wang, S. S. & Xue, C. High infiltration of tumor-associated macrophages in triple-negative breast cancer is associated with a higher risk of distant metastasis. Onco. Targets Ther. 7, 1475–1480 (2014).
Choi, Y. S., Chakrabarti, R., Escamilla-Hernandez, R. & Sinha, S. Elf5 conditional knockout mice reveal its role as a master regulator in mammary alveolar development: failure of Stat5 activation and functional differentiation in the absence of Elf5. Dev. Biol. 329, 227–241 (2009).
Harris, J. et al. Socs2 and elf5 mediate prolactin-induced mammary gland development. Mol. Endocrinol. 20, 1177–1187 (2006).
Zhou, J. et al. Elf5 is essential for early embryogenesis and mammary gland development during pregnancy and lactation. EMBO J. 24, 635–644 (2005).
Chakrabarti, R. et al. Elf5 regulates mammary gland stem/progenitor cell fate by influencing notch signaling. Stem Cells 30, 1496–1508 (2012).
Chakrabarti, R. et al. Elf5 inhibits the epithelial–mesenchymal transition in mammary gland development and breast cancer metastasis by transcriptionally repressing Snail2. Nat. Cell Biol. 14, 1212–1222 (2012).
Mojic, M. & Takeda, K. & Hayakawa, Y.The dark side of IFN-γ: its role in promoting cancer immunoevasion. Int. J. Mol. Sci. 19, E89 (2017).
Schreiber, R. D., Old, L. J. & Smyth, M. J. Cancer immunoediting: integrating immunity’s roles in cancer suppression and promotion. Science 331, 1565–1570 (2011).
Mimura, K. et al. PD-L1 expression is mainly regulated by interferon gamma associated with JAK-STAT pathway in gastric cancer. Cancer Sci. 109, 43–53 (2018).
Zaidi, M. R. et al. Interferon-γ links ultraviolet radiation to melanomagenesis in mice. Nature 469, 548–553 (2011).
Yang, J. & Weinberg, R. A. Epithelial–mesenchymal transition: at the crossroads of development and tumor metastasis. Dev. Cell 14, 818–829 (2008).
Thiery, J. P. Epithelial–mesenchymal transitions in tumour progression. Nat. Rev. Cancer 2, 442–454 (2002).
Polyak, K. & Weinberg, R. A. Transitions between epithelial and mesenchymal states: acquisition of malignant and stem cell traits. Nat. Rev. Cancer 9, 265–273 (2009).
Thiery, J. P., Acloque, H., Huang, R. Y. & Nieto, M. A. Epithelial–mesenchymal transitions in development and disease. Cell 139, 871–890 (2009).
Lin, N. U. et al. Clinicopathologic features, patterns of recurrence, and survival among women with triple-negative breast cancer in the National Comprehensive Cancer Network. Cancer 118, 5463–5472 (2012).
De Ruijter, T. C., Veeck, J., de Hoon, J. P., van Engeland, M. & Tjan-Heijnen, V. C. Characteristics of triple-negative breast cancer. J. Cancer Res. Clin. Oncol. 137, 183–192 (2011).
Dent, R. et al. Triple-negative breast cancer: clinical features and patterns of recurrence. Clin. Cancer Res. 13, 4429–4434 (2007).
Sorlie, T. et al. Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc. Natl Acad. Sci. USA 100, 8418–8423 (2003).
Echeverria, G. V. et al. High-resolution clonal mapping of multi-organ metastasis in triple negative breast cancer. Nat. Commun. 9, 5079 (2018).
Aprelikova, O. et al. Development and preclinical application of an immunocompetent transplant model of basal breast cancer with lung, liver and brain metastases. PLoS ONE 11, e0155262 (2016).
Katayama, H. et al. An autoimmune response signature associated with the development of triple-negative breast cancer reflects disease pathogenesis. Cancer Res. 75, 3246–3254 (2015).
Jezequel, P. et al. Gene-expression molecular subtyping of triple-negative breast cancer tumours: importance of immune response. Breast Cancer Res. 17, 43 (2015).
Choi, Y. S., Cheng, J., Segre, J. & Sinha, S. Generation and analysis of Elf5-LacZ mouse: unique and dynamic expression of Elf5 (ESE-2) in the inner root sheath of cycling hair follicles. Histochem. Cell Biol. 129, 85–94 (2008).
Ayers, M. et al. IFN-γ-related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Invest. 127, 2930–2940 (2017).
Liu, X. et al. Activation of the JAK/STAT-1 signaling pathway by IFN-γ can down-regulate functional expression of the MHC class I-related neonatal Fc receptor for IgG. J. Immunol. 181, 449–463 (2008).
Villarino, A. V., Kanno, Y., Ferdinand, J. R. & O'Shea, J. J. Mechanisms of Jak/STAT signaling in immunity and disease. J. Immunol. 194, 21–27 (2015).
Shackleton, M. et al. Generation of a functional mammary gland from a single stem cell. Nature 439, 84–88 (2006).
Yeh, C. H., Bellon, M. & Nicot, C. FBXW7: a critical tumor suppressor of human cancers. Mol. Cancer 17, 115 (2018).
Gossage, L., Eisen, T. & Maher, E. R. VHL, the story of a tumour suppressor gene. Nat. Rev. Cancer 15, 55–64 (2015).
Shin, H. Y. et al. Hierarchy within the mammary STAT5-driven Wap super-enhancer. Nat. Genet. 48, 904–911 (2016).
Marchetti, M. et al. Stat-mediated signaling induced by type I and type II interferons (IFNs) is differentially controlled through lipid microdomain association and clathrin-dependent endocytosis of IFN receptors. Mol. Biol. Cell 17, 2896–2909 (2006).
Takeuchi, Y. & Nishikawa, H. Roles of regulatory T cells in cancer immunity. Int. Immunol. 28, 401–409 (2016).
Vinay, D. S. et al. Immune evasion in cancer: mechanistic basis and therapeutic strategies. Semin. Cancer Biol. 35, S185–S198 (2015).
Facciabene, A., Motz, G. T. & Coukos, G. T-regulatory cells: key players in tumor immune escape and angiogenesis. Cancer Res. 72, 2162–2171 (2012).
Zhou, Q., Facciponte, J., Jin, M., Shen, Q. & Lin, Q. Humanized NOD-SCID IL2rg−/− mice as a preclinical model for cancer research and its potential use for individualized cancer therapies. Cancer Lett. 344, 13–19 (2014).
Andre, M. C. et al. Long-term human CD34+ stem cell-engrafted nonobese diabetic/SCID/IL-2Rγnull mice show impaired CD8+ T cell maintenance and a functional arrest of immature NK cells. J. Immunol. 185, 2710–2720 (2010).
DeRose, Y. S. et al. Tumor grafts derived from women with breast cancer authentically reflect tumor pathology, growth, metastasis and disease outcomes. Nat. Med. 17, 1514–1520 (2011).
Gradishar, W. J. et al. NCCN Guidelines Insights: Breast Cancer, Version 1.2017. J. Natl Compr. Cancer Netw. 15, 433–451 (2017).
Blows, F. M. et al. Subtyping of breast cancer by immunohistochemistry to investigate a relationship between subtype and short and long term survival: a collaborative analysis of data for 10,159 cases from 12 studies. PLoS Med. 7, e1000279 (2010).
Lu, W. & Kang, Y. Epithelial–mesenchymal plasticity in cancer progression and metastasis. Dev. Cell 49, 361–374 (2019).
Soundararajan, R. et al. Targeting the interplay between epithelial-to-mesenchymal-transition and the immune system for effective immunotherapy. Cancers (Basel) 11, E714 (2019).
McNutt, M. Cancer immunotherapy. Science 342, 1417 (2013).
Vikas, P., Borcherding, N. & Zhang, W. The clinical promise of immunotherapy in triple-negative breast cancer. Cancer Manag. Res. 10, 6823–6833 (2018).
Dirix, L. Y. et al. Avelumab, an anti-PD-L1 antibody, in patients with locally advanced or metastatic breast cancer: a phase 1b JAVELIN Solid Tumor study. Breast Cancer Res. Treat. 167, 671–686 (2018).
Nanda, R. et al. Pembrolizumab in patients with advanced triple-negative breast cancer: phase Ib KEYNOTE-012 study. J. Clin. Oncol. 34, 2460–2467 (2016).
Li, C. H., Karantza, V., Aktan, G. & Lala, M. Current treatment landscape for patients with locally recurrent inoperable or metastatic triple-negative breast cancer: a systematic literature review. Breast Cancer Res. 21, 143 (2019).
Gyorffy, B. et al. An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res. Treat. 123, 725–731 (2010).
Busino, L. et al. Fbxw7α- and GSK3-mediated degradation of p100 is a pro-survival mechanism in multiple myeloma. Nat. Cell Biol. 14, 375–385 (2012).
Bayne, L. J. & Vonderheide, R. H. Multicolor flow cytometric analysis of immune cell subsets in tumor-bearing mice. Cold Spring Harb. Protoc. 2013, 955–960 (2013).
Pearton, D. J., Broadhurst, R., Donnison, M. & Pfeffer, P. L. Elf5 regulation in the trophectoderm. Dev. Biol. 360, 343–350 (2011).
Deome, K. B., Faulkin, L. J. Jr, Bern, H. A. & Blair, P. B. Development of mammary tumors from hyperplastic alveolar nodules transplanted into gland-free mammary fat pads of female C3H mice. Cancer Res. 19, 515–520 (1959).
Chakrabarti, R. & Kang, Y. Transplantable mouse tumor models of breast cancer metastasis. Methods Mol. Biol. 1267, 367–380 (2015).
Welte, T. et al. Oncogenic mTOR signalling recruits myeloid-derived suppressor cells to promote tumour initiation. Nat. Cell Biol. 18, 632–644 (2016).
Chakrabarti, R. et al. ΔNp63 promotes stem cell activity in mammary gland development and basal-like breast cancer by enhancing Fzd7 expression and Wnt signalling. Nat. Cell Biol. 16, 1004–1015 (2014).
Acknowledgements
We thank L. King (University of Pennsylvania) for critical reading of the manuscript and helpful discussions. We thank A. Minn (University of Pennsylvania) for helpful discussions. We thank the Penn Vet Comparative Pathology Core for assistance with embedding, sectioning, consultation and slide evaluation. The Penn Vet Comparative Pathology Core is supported by the Abramson Cancer Center Support Grant (P30 CA016520). We thank Dr. Andres J.P. Klein-Szanto (Fox Chase Cancer Center) for helpful discussions and consultation on primary and metastatic tumour samples. We thank Y. Kang (Princeton University) for the HEK293T, EpRas, 4T1, LM2 and BT549 cell lines and lentiviral vector pLEX-hFL2iG. We thank S. Ran (Southern Illinois University) for the HCC1806 cell line. We thank the Eastern Division of the Cooperative Human Tissue Network at The University of Pennsylvania for providing human breast cancer fixed tissues from patients. We thank A. Welm (University of Utah) for the PDX tumour tissues. We thank the members of the Flow Cytometry Core at the Children’s Hospital of Philadelphia and University of Pennsylvania. We thank the Penn Vet Imaging Core for confocal microscopy. This work was supported by grants from the American Cancer Society, an NCI-K22 grant to R.C. (K22CA193661-01) and an NCI-R01 (R01 CA237243-01A1) grant to R.C.
Author information
Authors and Affiliations
Contributions
S.Singh and R.C. designed all of the experiments. S.Singh, S.Kumar, R.K.S., A.N., H.M., G.T., S.Kim and M.B. performed all of the experiments. M.A.B. performed all of the clinical and statistical analyses using human breast cancer samples and datasets. J.T. performed the RNA-Seq analysis and generated the GSEA datasets. R.S. and L.B. provided experimental support in the FBXW7-related experiments. S.Sinha provided the Elf5+/− and Elf5–GFP mice. R.M.Z. and S.Y.F. provided the reagents and valuable insights into the work. S.Singh and R.C. wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 C3-T+; Elf5+/- tumor cells show increased tumorigenesis in REAR mice.
a, mRNA expression of Elf5 in tumor cells from C3-T+; Elf5+/- (n = 10) and C3-T+; Elf5+/+ (n = 3) individual tumors. qPCR values were normalized to Gapdh. Experiments were performed thrice with qPCR in technical duplicate. b, H&E staining of tumors shows more mesenchymal cells in C3-T+; Elf5+/- tumors. Images are representative of two independent experiments. c, Quantification of mitotic index in tumor sections stained with H&E (right panel), n= 3, C3-T+; Elf5+/- and n=4, C3-T+; Elf5+/+ individual tumors were used. d, Schematic showing representation of experiment in REAR recipient mice. 225,000 cells/MFP of sorted tumor cells from C3-T+; Elf5+/+ and C3-T+; Elf5+/- tumors were injected into contralateral mammary fat pad of REAR mice REAR mice. C3-T+; Elf5+/- tumor cells formed e, tumors earlier, n=6 tumors/group f, g, with rapid growth and volume at sacrifice, n=5 tumors for C3-T+; Elf5+/- and n=4 tumors for C3-T+; Elf5+/+ group. Log-rank test was used for KM plots to calculate p-value, (C3-T+; Elf5+/+, n=6 tumors; C3-T+; Elf5+/-, n=6 tumors in (e). f, Two-way ANOVA test was used with Bonferroni post hoc test to calculate statistical significance. Data presented as mean ± SEM. h, i, Metastatic nodules visible in lungs of REAR mice injected with C3-T+; Elf5+/- tumor cells, n=6 mice/genotype, p=0.049. Experiment performed once. j, k, H&E images showed increased mesenchymal features and invasive structures in tumors formed from C3-T+; Elf5+/- tumor cells. Images are representative to two independent experiments. (a, c, i) Two-tailed student’s t test was used to calculate statistical significance. Data presented as mean ± SEM. Scale bars, (b) 40 µm, (h), 40 µm, (j) 60 µm, (k) 200 µm. **p < 0.01.
Extended Data Fig. 2 Hyperplastic mammary glands from C3-T+; Elf5+/- tumors show increased P-STAT1 and P-STAT3 protein levels and Elf5 is predominantly expressed in tumor cells.
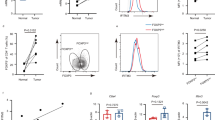
a, b, IHC staining indicates increased staining of IFNGR1 and P-STAT1 protein in hyperplastic mammary glands harvested from C3-T+; Elf5+/- compared to C3-T+; Elf5+/+ mice. One representative image is shown from multiple experiments (n=5). Arrows in (b) indicate P-STAT1+ cells. c, d, IHC staining indicates increased staining of P-STAT1 (Ser727) and P-STAT3 (Tyr705) proteins in tumors harvested from C3-T+; Elf5+/- compared to C3-T+; Elf5+/+ mice. n=5 individual tumors were taken in each group and quantification was done using 16 random fields. Two-tailed Mann-Whitney U test was used to calculate statistical significance and data is represented as mean ± SEM in (c) and mean ± SD in (d). e, Immunofluorescence image showing expression of GFP (Elf5) in tumor cells and their absence in stromal cells in tumor tissues obtained from C3-T+; Elf5-GFP. Absence of GFP in stromal cells of C3-T+ was confirmed by immunofluorescence in n=5 tumors. f, Flow cytometry graph showing presence of Elf5 predominantly in epithelial cells in tumors obtained from C3-T+; Elf5-GFP, one representative image shown from multiple experiments (n=5). FOV; Field of view. Scale bar in (a, c, d, e) 40 µm, (b) 20 µm.
Extended Data Fig. 3 Elf5 negatively correlates with IFN-γ signaling in C3-T+ tumors.
a, Schematic diagram showing generation of C3-T+; Elf5-GFP mice. b, Immunofluorescence images showing Elf5 loss during progression from mammary gland (N) to hyperplasia (H) to tumors. K14 antibody was used to mark basal epithelial cells. Experiment was repeated twice with similar results. c, Histogram showing %GFP+ cells in epithelial enriched tumor cells in C3-T+; Elf5-GFP mice. Tumor cell enriched population was sorted using CD31, CD45 and Tert119 markers following published protocol37. Red line denotes WT tumor cells as control. Experiment was repeated thrice with similar results. d, qPCR analysis of Elf5 expression in GFP+ and GFP- tumor cells sorted from C3-T+; Elf5-GFP mice, n=3. qPCR values were normalized to the housekeeping gene Gapdh. Experiments were performed three times, each with qPCR in technical duplicate, and data presented as the mean ± SD. Two- tailed Student’s t test was used to calculate statistical significance. e-i, GSEA graphs showing high EMT signatures, (e); increased invasive signatures, (f); high expression of signatures associated with stemness, (g); and increased IFN-γ signaling in GFP- (Elf5-) tumor cells, (h-i). n=2 GFP+ and n=4 GFP- samples were used for RNA sequencing and GSEA analysis. Statistical significance was assessed by comparing the ES to enrichment results generated from 1000 random permutations of the gene set to obtain p values (nominal p value) j, Western blot showing high P-STAT1 and STAT1 in GFP- (Elf5-). Cell sorting was performed to isolate tumor cells and multiple independent tumors were pooled to make samples. This is a representative blot and experiment was repeated twice with n=9 tumors. k, Heat map showing increased ISG in GFP- (Elf5-) tumor cells. n=2 GFP+ and n=4 GFP- tumor cell population was used to generate heat map. Scale bar in (b), 20 µm.
Extended Data Fig. 4 FBXW7, a ubiquitin ligase is a direct target of ELF5.
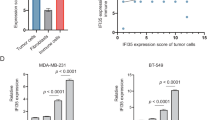
a, b, GSEA of GFP+ (Elf5+) and GFP- (Elf5-) tumor cells show upregulation of genes involved in proteasomal and ubiquitin mediated degradation in GFP+ (Elf5+) tumor cells as compared to GFP- (Elf5-) tumor cells, n=2 GFP+ and n=4 GFP- individual samples. c, List of ubiquitin ligases differentially expressed in GFP+ (Elf5+) and GFP- (Elf5-) tumor cells. d-g, qPCR analysis shows (d) increased ELF5 (e) increased FBXW7 (f) increased VHL and (g) no change in IFNGR1 mRNA in ELF5 expressing LM2 (lung metastatic derivative of MDA-MB-231) cells, n= independent 4 samples. h, i, Data mining for our published microarray data16 shows (h) decreased FBXW7 (i) and no change in VHL in cells upon transduction of ELF5 plasmid encoding mutation in DNA binding domain of ELF5 in MDA-MB-231 cells16 (n=3 independent biological samples/group). j, Western blot showing high ELF5, high FBXW7 and low IFNGR1 proteins upon re-expression of ELF5 in LM2 cells. k, Western blot images showing positive correlation between FBXW7 and ELF5, and their negative correlation with IFNGR1 in TNBC cells (MX1, LM2, HCC1806, BT549). MCF7 cells was used as positive control. l-n, qPCR of EpRas cells (n=6 independent biological samples/group) with re-expression of ELF5 shows (l) high Elf5 (m) high Fbxw7 (n) and no change in Ifngr1 mRNA levels. o, Expression of ELF5 in EpRas cells increases FBXW7 and decreases IFNGR1 protein levels as shown by western blotting. qPCR values were normalized to the housekeeping gene Gapdh. Experiments were performed three times, each with qPCR in technical duplicate, and data presented as the mean ± SD. Two tailed Student’s t test was used to calculate statistical significance. Please see uncropped WB images in Source data file.
Extended Data Fig. 5 ELF5 binds to multiple genomic loci of FBXW7 in normal mammary epithelial cells and ELF5 KD MX1 cells show increased expression of IFNGR1.
a, In silico ChIP-seq analysis showing multiple binding sites of ELF5 on genomic loci of FBXW7. Lac1 denotes mammary gland obtained from lactating mouse and P13 denotes mammary gland obtained from pregnant mouse at day 13. Black vertical lines show binding sites. b, FACS analysis showing increase in IFNGR1 protein expression upon stable knockdown of ELF5 in MX1 TNBC cells using multiple shRNAs. Data is representative of two independent experiments. c, d IHC images showing FoxP3 staining in periphery and core of tumor sections obtained from C3-T+; ELF5+/+ and C3-T+; ELF5+/- mice. n=3 samples were used. e, Contour plots and f, g, scatter plots showing increased Ly6G+ neutrophil population in tumors from C3-T+; Elf5+/- mice as compared to C3-T+; Elf5+/+ tumors, n=3, C3-T+; Elf5+/+ tumors and n=5, C3-T+; Elf5+/- individual tumors. h, Suppression of T-cell proliferation confirms that increased myeloid cells in tumors are Gr1+ myeloid cells. Tumor CD45+CD11b+Gr1+ neutrophils were used. Tumor neutrophils (Gr1+ myeloid cells) were enriched using Gr1 enrichment kit following manufacture’s protocol. Data is representative of two independent experiments. (f, g) Mann-Whitney U test was used to calculate statistical significance. Data presented as mean ± SEM. Scale bar, 200 μM.
Extended Data Fig. 6 Flow cytometry gates showing different immune populations.
All the gates were drawn according to published protocol from Dr. Vonderheide group59. (a, b) Different myeloid and lymphoid populations are shown in a and b respectively.
Extended Data Fig. 7 Alteration of Elf5 in tumor cells alters TME in REAR background.
IHC analysis shows a, b, increased myeloid cells in C3-T+; Elf5+/- (n=8 FOV) tumors as compared to C3-T+; Elf5+/+ (n=6 FOV) c, d, decreased number of macrophages in C3-T+; Elf5+/- tumors (n=9 FOV) as compared to C3-T+; Elf5+/+ tumors (n=7 FOV), e, f, increased number of Foxp3+ cells in C3-T+; Elf5+/- tumors. (n=10 FOV for C3-T+; Elf5+/+ and n=11 FOV for C3-T+; Elf5+/-), n=3 individual tumors were used in a-f. g, j, increased Gr1+ myeloid cells (n=5 tumors/group) and Ly6G+ neutrophils (CD45+CD11b+Gr1+) (n=5 tumors/group) and decreased k, cytotoxic T-cells (CD45+CD8+) population in C3-T+; Elf5+/- tumors as compared to C3-T+; Elf5+/+ tumors (n=5 tumors/group) h, Macrophages (CD45+F4/80+) (n=5 C3-T+; Elf5+/+ tumors and n=6 C3-T+; Elf5+/- tumors) and i, Regulatory T-cells (CD45+CD3+FoxP3+) (n=3 tumors/group) were observed. IHC image showing increased FBXW7 (n=6 tumors for control and n=7 tumors for Elf5-OE from n=3 independent tumors). (l), decreased IFNGR1 (n=10 FOV for control and n=7 FOV for Elf5-OE from n=3 independent tumors) m, in Elf5-OE EpRas tumors. FACS plots showing no change in n, macrophages, o, CD4+ T-cells, p, increased Ly6C+ neutrophils in Elf5-OE tumors (n=6, control and n=4, Elf5-OE individual tumors). q FACS plots showing decreased number of Gr1+ myeloid cells in lungs of EpRas Elf5-OE tumor bearing mice (n=3 individual tumors/group). (a-j and n-q) Two-tailed student’s t test was used to compute p-value. (k, l, m) Mann-Whitney U test was used to calculate p value. Data are presented as the mean ± SEM. FOV; Field of view. (b, d, f, l, m) Boxplot data represent median, interquartile range, and spikes to upper and lower adjacent values. FOV; Field of view. (a, c, e, l, m) Images are representative of minimum three independent experiments, Scale bars, 40 µm.
Extended Data Fig. 8 IFN-γ signaling imparts more tumorigenic and metastatic potential.
IFN-γ treatment upregulates a, STAT1 and P-STAT1 (Data representative of minimum of three independent experiments) b, mRNA expression of IFN-γ target genes. qPCR values were normalized to Gapdh. Experiments were performed twice in technical duplicate, and data presented as mean ± SD. Two-sided Student’s t test used to calculate p-value. c, FACS analysis show upregulation of PD-L1 but not CTLA4 in IFN-γ-treated EpRas cells. Data representative of two independent experiments. d, e, IFN-γ treated EpRas cells form larger and more invasive tumors, n=5 tumors/group. Inset shows higher magnification images. Data represented as mean ± SD. f, g, Small metastatic nodules (black arrows) in lungs of mice injected with IFN-γ treated EpRas cells. g, GFP labelled IFN-γ treated EpRas cells show micro metastatic foci in lungs of mice (n=4), while control untreated cell injected mice show no micro metastatic foci (n=4). EpRas 250,000 cells/MFP were injected into mice for (g). Cells were labeled with GFP (lentiviral vector pLEX-hFL2iG was used) using standard method. White arrows show GFP+ micro metastatic foci in whole mount lung image. (e-g) Experiment was repeated twice with similar results. (h-l) IHC analysis shows high (h) P-STAT1+, (i) S100A8+ cells in IFN-γ treated EpRas tumors. (j) Decreased CD8+T-cells observed in IFN-γ treated tumors. (k, l) Increase in S100A8+ cells observed in IFN-γ treated EpRas lung sections (metastatic site) with quantification in the right panel. Boxplot data represent median, interquartile range, and spikes to upper and lower adjacent values. (h-l). Two-way ANOVA test was used with Bonferroni post hoc test to calculate p value in (d). Two-tailed Mann-Whitney U test was used to compute p-value for h, j, i, l and data is represented as mean ± SEM. n=10 random FOV were evaluated from 3 individual tumors (h, j) n=3 and 5 random FOV were evaluated (i), n=3 and 7 random FOV/group were used for quantification (l). (e-g) Representative images of minimum two independent experiments are presented. Scale bar, (e) 200µm, (f) 2mm, and (g) 500 μm and (h-k) 40 μm respectively. ***p < 0.001. FOV; Field of view.
Extended Data Fig. 9 IFN-γ signaling renders more metastatic properties to EpRas cells via PD-1/PD-L1 signaling.
a, b, Tail vein (TV) injection of control and IFN-γ treated EpRas cells, n=7 mice/group. c, d, Lung images and metastatic nodule numbers in NSG mice injected with EpRas cells treated with IFN-γ via TV (n=6 mice, control and n=7 mice, IFN-γ treated). e, f, TV injection of control and IFN-γ treated EpRas cells in mice treated with anti-IgG (n=5 mice), anti-PD-1 (n=5 mice) and anti-PD-L1 (n=6 mice) antibodies. g, h, IHC in tumors from control cells compared to IFNGR1-KD tumor cells, n=3 individual tumors were used and 7 random FOV were used. i, FBXW7-KD tumors cells grew faster than control, n=5 tumors/group. k, Western blot showed decreased FBXW7 in EpRas cells transduced with FBXW7 shRNAs compared to control. (l) IF analysis shows increased expression of IFNGR1 in FBXW7-KD cells compared to control. Experiment was repeated twice (k, l). j, m, TV injection of control and FBXW7-KD EpRas cells shows increased number of lung metastasis in FBXW7-KD mice compared to control mice, n=3 CTRL, n=4 KD1, and n=4 KD2 mice were used. Two-tailed Mann-Whitney U test was used to compute p-value in (b, d) and data represented as mean ± SEM (b) or ±S.D (d). Boxplot represent median, interquartile range, and spikes to upper and lower adjacent values (f, h). One-way ANOVA test was used with Tukey post hoc test to compute p-values of multiple comparison data. Data is represented as mean ± SEM (f, h, j). Two-way ANOVA test was used with Bonferroni post hoc test to calculate p values, data is represented as mean ± SD (i). Scale bar (a, c, e, m) 2 mm, (g) 40 µm (l) 100 µm. **p < 0.01, ***p < 0.001. FOV; Field of view.
Extended Data Fig. 10 High expression of ELF5/FBXW7 and low expression of IFNGR1 is protective in Mesenchymal, basal and immunomodulatory TNBC.
a–d, High expression of ELF5/FBXW7 and low expression of IFNGR1 is correlated with better Distant metastasis free survival (DMFS) in (a) Basal (n=24 for Elf5/FBXW7 and (n=32 for IFNGR1) (b) Immunomodulatory (n=47 patient samples for Elf5/FBXW7 and (n=36 patient samples for IFNGR1) (c) Mesenchymal TNBC subsets (n=57 for Elf5/FBXW7 and (n=57 patient samples for IFNGR1). (d) High expression of ELF5/FBXW7 and low expression of IFNGR1 is correlated with worse Distant metastasis free survival (DMFS) in Non-TNBC ER+ patients (n=437 patient samples for Elf5/FBXW7 and (n=1391 patient samples for IFNGR1). P-value was computed by Log Rank test. KM plotter database was used57. e, TNBC PDXs showing direct correlation between ELF5 and FBXW7, and indirect correlation with IFNGR1. 5 TNBC PDXs were used for this experiment. Two-tailed Mann-Whitney U test was used for statistical analysis and data is represented as mean ± SEM. (f-h) IHC images of TNBC PDXs showing direct correlation between ELF5 and FBXW7, and indirect correlation with IFNGR1. Scale bar, 40 µm.
Supplementary information
Supplementary Information (download XLSX )
Supplementary Tables 1–6.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download PDF )
Unprocessed western blots.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Rights and permissions
About this article
Cite this article
Singh, S., Kumar, S., Srivastava, R.K. et al. Loss of ELF5–FBXW7 stabilizes IFNGR1 to promote the growth and metastasis of triple-negative breast cancer through interferon-γ signalling. Nat Cell Biol 22, 591–602 (2020). https://doi.org/10.1038/s41556-020-0495-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41556-020-0495-y
This article is cited by
-
Unveiling personalised targets of PD-1 blockade hyperprogression in a cervical adenocarcinoma via longitudinal single-cell and spatial monitoring
npj Precision Oncology (2026)
-
Glycyrrhizin upregulates PTEN and suppresses oncogenic signaling in breast cancer
Medical Oncology (2026)
-
Interferon signaling pathways in health and disease
Molecular Biomedicine (2025)
-
Nuclear PD-L1 drives IFN-γ-promoted lung metastasis of triple-negative breast cancer via POLR2A-mediated transcriptional activation of LY6E
Breast Cancer Research (2025)
-
NAP1L1 degradation by FBXW7 reduces the deubiquitination of HDGF-p62 signaling to stimulate autophagy and induce primary cisplatin chemosensitivity in nasopharyngeal carcinoma
Molecular Cancer (2025)