Abstract
In most eukaryotic cells, euchromatin is localized in the nuclear interior, whereas heterochromatin is enriched at the nuclear envelope (NE). This conventional chromatin organization is established by heterochromatin tethering to the NE; however, its importance for cellular homeostasis is largely unexplored. One tether is constituted by the lamin B receptor (LBR) in mammals, but the enigmatic nature of other redundant tethers has hampered functional analyses. Here we demonstrate that downregulation of abundant, ubiquitous NE proteins can induce the global detachment of heterochromatin from the NE and its repositioning to the nuclear interior. We identify LBR and lamina-associated polypeptide 2 (LAP2) as key factors for peripheral heterochromatin positioning in differentiated and pluripotent mammalian cells. Their long-term loss leads to changes in three-dimensional chromatin organization and a reduction in repressive epigenetic marks, especially H3K27me3. These changes are associated with massive deregulation of gene expression, activation of antiviral innate immunity, and defects in cell fate determination.
Similar content being viewed by others
Main
Heterochromatin is positioned at the nuclear periphery and around nucleoli, whereas euchromatin resides in the nuclear interior and in the vicinity of nuclear pore complexes1,2,3,4,5,6,7. This ‘conventional’ nuclear architecture is an almost universal feature of eukaryotic nuclei and is achieved by tethering of heterochromatin to the nuclear envelope (NE)8,9. In metazoan cells, peripheral heterochromatin is tethered to the nuclear lamina, a network formed by proteins of the inner nuclear membrane (INM) and nuclear lamins10,11,12, comprising the A-type lamins, A and C, and the B-type lamins, B1 and B2, in mammals. Genomic regions at the nuclear lamina are referred to as lamina-associated domains (LADs). They are up to 10 Mb in size, gene-poor, enriched in epigenetic heterochromatic marks, and correspond by-and-large to the B compartments defined by Hi-C, especially the B.2 subcompartment13,14,15. Up to 40% of genomic regions are assigned as LADs, but many LADs dynamically reshuffle into nucleolus-associated domains (NADs) upon cell division5,14,15,16,17,18,19.
An outstanding question in the field concerns the mechanisms that govern the peripheral positioning of mammalian heterochromatin. The prevailing hypothesis posits that heterochromatin attachment to the NE of mammals is accomplished by two molecular assemblies, referred to as A- and B-type tethers8. The A-type tether is organized by A-type lamins and functionalized by unknown proteins of the INM, whereas the B-type tether is constituted by the INM protein lamin B receptor (LBR)8, which binds lamin B120. The presence of either the A-type or B-type tether is sufficient to maintain heterochromatin at the periphery, with the dominance of the respective tethers differing according to the differentiation stage of the cells8.
Peripheral positioning of heterochromatin has been implicated in the regulation of gene expression and thereby cell fate specification during differentiation and development7,12,14,21,22,23,24. Artificial tethering of genomic loci to the nuclear lamina can lead to their repression25,26, indicating that peripheral localization can promote gene silencing. LADs impede transcriptional activity and store silent genomic loci in the absence of their activating factors27,28, although certain peripherally tethered loci can escape repression29,30,31. Conversely, disruption of heterochromatin, local chromatin decondensation or transcriptional activation of genes can cause the relocation of genes into the nuclear interior32,33,34, suggesting that chromatin structure influences peripheral gene localization.
The anchorage of heterochromatin at the NE seems to be a fundamental feature of metazoan genome organization. However, a full understanding of the mechanisms governing heterochromatin localization to the INM as well as of the functional relevance of peripheral heterochromatin sequestration are currently lacking, because of the innate redundancy of the tethering mechanism. Besides its potential role in repressing genes found in LADs, it is not clear if other aspects of genome homeostasis and nuclear function are jeopardized when heterochromatin is detached from the nuclear periphery.
We set out to identify which mammalian INM proteins contribute to bulk heterochromatin tethering to the NE. We show that the downregulation of a suite of ubiquitously expressed NE proteins leads to detachment of heterochromatin from the nuclear periphery, and we identify major contributions of LBR and lamina-associated polypeptide 2 (LAP2) to peripheral heterochromatin localization. Loss of these tethers causes changes in gene expression, chromatin organization, histone methylation, transposon repression and cell differentiation, highlighting the functional importance of heterochromatin attachment to the NE.
Results
Depletion of abundant NE proteins causes detachment of constitutive heterochromatin
The attachment of constitutive heterochromatin to the NE in mammalian cells is assumed to be redundantly governed by LBR and undefined INM proteins8. To identify NE–heterochromatin tethers and considering redundancy, we employed a reverse genetics approach and collectively depleted 12 abundant, ubiquitously expressed NE factors using a mixture of siPOOLs35. The targeted factors comprised the A-type lamins (lamin A and lamin C), the B-type lamins (lamin B1 and lamin B2) and eight INM proteins (LBR, the four LEM (LAP2, emerin, MAN1)-domain proteins LAP2β, emerin, MAN1 and LEMD2, the SUN domain proteins SUN1 and SUN2, and LAP1; Fig. 1a). Only 11 siPOOLs (si-11) were needed for depletion of the 12 factors, because lamin A and lamin C can be depleted by the same siPOOL.
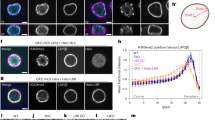
a, The selected set of 12 NE factors and their corresponding categories (lamins: lamin A, lamin C, lamin B1 and lamin B2; INM proteins: LBR, LAP2, emerin, LEMD2, MAN1, SUN1, SUN2 and LAP1) targeted by a combination of siPOOLs (si-11, including one siPOOL that targets both lamin A and lamin C). Each individual siPOOL is composed of ~30 different siRNAs. b, Representative confocal images of HCT116 and RPE-1 cells either treated with control siRNA (si-Ctr) or a combination of 11 siPOOLs for 72 h, fixed and subjected to immunofluorescence using an anti-H3K9me3 antibody. DNA was stained with Hoechst. N = 3 experiments. Scale bars, 5 μm. c, Immunoblots confirming the downregulation of select NE proteins upon si-11 depletion. The asterisk indicates an unspecific band in the HCT116 lamin B2 blot. d, Quantification procedure of the fraction of H3K9me3-positive heterochromatin at the nuclear rim. H3K9me3 intensity was quantified in an ~0.4-μm-wide ring at the nuclear periphery and expressed as the ratio of the average integrated fluorescence intensities at the nuclear rim over the entire nuclear area (nuclear rim + nuclear interior). e, Quantification of accumulation of the H3K9me3 signal at the NE of HCT116 cells (total number of cells, n ≥ 227; P <1.0 × 10−15) and RPE-1 cells (total number of cells, n ≥ 86, P <1.0 × 10−15) treated as in b (****P ≤ 0.0001, two-sided Welch’s t-test, mean ± s.d., N = 3 experiments). Numerical and statistics source data as well as unprocessed blots are available as source data.
To examine effects on heterochromatin positioning, we initially analysed HCT116 cells, a near-diploid human carcinoma cell line (Fig. 1b). Heterochromatin was visualized by immunofluorescence using an antibody against H3K9me3 that marks constitutive heterochromatin and is enriched in LADs14,36,37. Immunoblotting of a subset of the targeted factors confirmed their downregulation (Fig. 1c), which was further verified by RNA sequencing (RNA-seq; Extended Data Fig. 1a and Supplementary Table 1). HCT116 cells treated with control small interfering RNA (si-Ctr) displayed the typical peripheral enrichment of H3K9me3 and a few intranuclear heterochromatin foci (Fig. 1b). In contrast, cells treated with the si-11 pool showed a strong decrease in peripheral H3K9me3 staining and a striking accumulation of intranuclear heterochromatin clusters, reminiscent of chromocentres (Fig. 1b). Comparison with Hoechst and co-staining with a nucleolar marker indicated that a fraction of H3K9me3-positive foci was in the vicinity of nucleoli, indicating that some LADs may be converted into NADs (Extended Data Fig. 1b).
To quantify the loss of peripheral heterochromatin, we delineated the nuclear contours based on the Hoechst signal and measured the mean integrated density of H3K9me3 in an ~0.4-μm-wide ring at the nuclear periphery relative to the entire nucleus (Fig. 1d). These quantifications showed a significant decrease in H3K9me3 at the NE upon depletion of the 12 targeted NE proteins in both HCT116 and human retinal pigment epithelial cells (RPE-1 cells) (Fig. 1e). Analysis of H4K20me3 and H3K27me3, which decorate constitutive and facultative heterochromatin, respectively, supported the global loss of heterochromatin from the NE (Extended Data Fig. 1c,d). The repositioning of H3K9me3-positive heterochromatin into the nuclear interior was confirmed in human RPE-1 cells (Fig. 1b) as well as in mouse NIH/3T3 fibroblasts and embryonic stem cells (mESCs) (Extended Data Fig. 1e,f). Taken together, depletion of these 12 factors is sufficient to induce the detachment of heterochromatin from the NE in a variety of mammalian cell lines.
Depletion of LBR and lamin A/C is insufficient to globally release constitutive heterochromatin from the NE
Landmark studies in mouse rod cells have demonstrated that the absence of LBR and lamin A/C is sufficient to cause nuclear chromatin inversion8. We therefore examined whether downregulation of LBR and lamin A/C was also sufficient for heterochromatin detachment from the NE in HCT116 cells. Since siRNA-mediated downregulation of A-type lamins was inefficient (Fig. 1c), we generated LMNA knockout (KO) cells (Extended Data Fig. 2a). We then compared the effect of LBR depletion on peripheral heterochromatin positioning in HCT116 wild-type (WT) and LMNA KO cell lines (Extended Data Fig. 2b–d). The individual loss of either LBR (siP-LBR) or lamin A/C (LMNA KO cells) slightly reduced enrichment of H3K9me3-positive chromatin at the NE and caused the formation of small intranuclear heterochromatin foci. Their combined loss clearly exacerbated these defects. However, only the combined depletion of all other ten NE factors in LMNA KO cells led to a gross reduction of peripheral heterochromatin and the formation of large intranuclear chromatin clusters (LMNA KO, si-10). This shows that the absence of LBR and lamin A/C is insufficient to release heterochromatin from the NE to the same extent as the loss of all 12 proteins, indicating that factors other than LBR and lamin A/C contribute to bulk heterochromatin tethering in HCT116 cells. Even in the presence of lamin A/C (HCT116 WT), loss of the other ten proteins had a stronger effect than loss of LBR alone, supporting a contribution of other factors (Extended Data Fig. 2b–d).
Identification of mammalian INM proteins that anchor heterochromatin
To identify the factors that promote NE tethering of constitutive heterochromatin, we used H3K9me3 staining and the localization of centromeres as a complementary readout. Many centromeres are localized at the nuclear periphery throughout the cell cycle, and the remaining portion is found adjacent to nucleoli38,39,40. Consistent with recent analyses in HeLa cells40, we observed that ~60% of centromeres were localized close to the NE in HCT116 WT cells (Extended Data Fig. 3a,b). A similar fraction of centromeres was located near the NE in LMNA KO cells, whereas RNA interference (RNAi)-mediated depletion of the other ten selected NE factors (si-10) diminished peripheral centromere localization to less than 10%.
To deconvolute which of these factors promote NE–heterochromatin tethering, we used HCT116 LMNA KO cells. We grouped the ten remaining factors into five bins corresponding to their respective protein families (B-type lamins, LBR, LEMs, SUNs and LAP1; Figs. 1a and 2a). We first downregulated each group individually, using si-10 as positive control. As expected, we observed a significant decrease in peripheral H3K9me3-marked heterochromatin upon depletion of LBR (step 1 in Fig. 2a; Extended Data Fig. 4a,b,g), but a large fraction of heterochromatin remained at the NE. Depletion of the other factors did not lead to discernible effects on peripheral H3K9me3 staining. Consistently, knockdown of LBR also strongly reduced the fraction of peripheral centromeres from 60% to 25% (Extended Data Fig. 3c). Centromere localization was also affected by the depletion of LEM proteins and slightly by LAP1.
a, Schematic depiction of the deconvolution approach performed in HCT116 LMNA KO cells. The proteins targeted by si-10 were initially distributed into five groups: LBR, B-type lamins (lamin B: lamin B1 and lamin B2), LEM family (LEMs: LAP2, LEMD2, MAN1 and emerin), SUN proteins (SUNs: SUN1 and SUN2) and LAP1. The corresponding sets of siPOOLs were first individually tested (step 1), identifying LBR. In the following steps, the contributions of additional factors were evaluated as outlined. Arrows indicate the major additional factors affecting peripheral heterochromatin identified in each step. b, Representative confocal images of H3K9me3-positive heterochromatin in HCT116 LMNA KO cells treated with si-Ctr or the indicated combination of siPOOLs for 72 h (step 3 of deconvolution), identifying LAP2. Scale bars, 5 μm. c, Quantification of the accumulation of H3K9me3 at the NE for b (total number of cells n ≥ 133, NS P > 0.05, ****P ≤ 0.0001, one-way analysis of variance (ANOVA), mean ± s.d., N = 3 experiments). d, Analysis of centromere positioning at the NE (also Extended Data Fig. 3). Representative confocal images of CREST, CLIMP-63 and Hoechst (DNA) throughout five microscopy slices in HCT116 LMNA KO cells treated with si-Ctr or the indicated combination of siPOOLs for 72 h (step 3 of deconvolution). Scale bar, 5 μm. e, Quantification of the proportion of centromeres at the nuclear periphery in cells treated with si-Ctr or the indicated combinations of siPOOLs for 72 h (total number of cells n ≥ 28, NS P > 0.05, *P ≤ 0.05, **P ≤ 0.01, ****P ≤ 0.0001, one-way ANOVA, mean ± s.d., N = 3 experiments). f, Immunoblotting of LBR, LAP2β and select other NE proteins to determine the efficiency of siRNA-induced protein depletion. Numerical and statistics source data as well as unprocessed blots are available as source data.
In the second deconvolution step, we depleted each of the four remaining protein groups together with LBR in the LMNA KO cells (step 2 in Fig. 2a). This revealed that the LEM protein family had the strongest impact on both readouts (Extended Data Figs. 4c,d,g and 3d). Only co-depletion of LAP1 and the B-type lamins with LBR caused a small reduction in the fraction of peripheral centromeres (Extended Data Fig. 3d).
To pinpoint which LEM proteins contribute to peripheral heterochromatin positioning, we individually downregulated LAP2, emerin, LEMD2 or MAN1 together with LBR (Fig. 2a, step 3). This led to a significant effect of LAP2 depletion for both readouts. Cells co-depleted of LBR and LAP2 contained large chromocentre-like heterochromatin clusters and displayed a reduction in peripheral H3K9me3-positive heterochromatin (Fig. 2b,c,f). Co-downregulation of LBR and LAP2 also strongly affected peripheral centromere positioning, reducing the fraction of centromeres at the nuclear periphery to ~10%, akin to co-depletion of LBR and all LEMs (Fig. 2d,e). Co-depletion of none of the other LEM proteins with LBR caused significant changes, indicating that LAP2 and LBR are chiefly responsible for peripheral centromere positioning. Notably, downregulation of LBR and LAP2 was not sufficient to fully recapitulate the si-10 phenotype (Fig. 2c,e), suggesting that additional factors may contribute.
Testing the remaining three groups (B-type lamins, SUNs and LAP1) in conjunction with LBR and LAP2 co-depletion (step 4; Extended Data Figs. 3e and 4e–g) did not show any further enhancement of the phenotype. Collectively, these experiments identify the INM proteins LBR and LAP2 as the major players in tethering of constitutive heterochromatin to the NE in HCT116 LMNA KO cells.
The membrane-anchored isoforms of LAP2 promote heterochromatin attachment to the NE
Human cells express four isoforms of LAP2 (α, β, γ and δ; Fig. 3a), of which LAP2α and LAP2β are the most abundant41. All isoforms can interact with chromatin via their N-terminal LEM-like and LEM domains. LAP2β, γ and δ are embedded in the INM (Fig. 3a), whereas LAP2α localizes to the nucleoplasm. We next wished to examine whether it is indeed the membrane-bound isoforms that synergize with LBR in promoting heterochromatin localization to the NE, especially in cells that still contain A-type lamins. To ensure the equal absence of LBR in all cells, we first generated HCT116 LBR KO cells (Extended Data Fig. 5a). H3K9me3 localization in these cells was slightly perturbed, evident by more prominent intranuclear heterochromatin foci and an uneven H3K9me3 distribution at the NE (Fig. 3b, si-Ctr). Additional downregulation of the other NE proteins including LAP2 caused a global detachment of heterochromatin from the NE, as expected (Fig. 3b,c, si-10b). The individual depletions of either LAP2 or lamin A/C in the HCT116 LBR KO background caused similar effects on peripheral heterochromatin localization, leading to a reduced accumulation of the H3K9me3 signal at the NE and more accentuated intranuclear heterochromatin clusters.
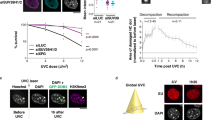
a, Schematic depiction of the four isoforms of LAP2 expressed in human cells. Note that the presence of LAP2δ at the protein level is still contested. b, Representative confocal images of H3K9me3 localization in HCT116 LBR KO cells treated with si-Ctr or the indicated combinations of siPOOLs for 72 h. si-10b refers to the siPOOL mixture including siP-lamin A/C but not siP-LBR. Scale bars, 5 μm. c, Quantification of the accumulation of H3K9me3 at the NE in b (total number of cells n ≥ 137, ****P ≤ 0.0001, one-way ANOVA, mean ± s.d., N = 3 experiments). d, Quantification of the proportion of centromeres at the nuclear periphery in cells treated as in b (total number of cells n ≥ 27, NS P > 0.05, ****P ≤ 0.0001, one-way ANOVA, mean ± s.d., N = 3 experiments). e, Immunoblotting of LAP2 (detecting LAP2α and LAP2β, asterisks), lamin A/C and emerin to determine the efficiency of siRNA-induced protein depletion. f, Representative confocal images of H3K9me3 localization in HCT116 WT and LBR/TMPO/LMNA TKO cells as well as of TKO cells expressing LAP2β-GFP after 48 h of transient transfection. Scale bars, 5 μm. g, Quantification of the accumulation of H3K9me3 at the NE in f (total number of cells n ≥ 35, ****P ≤ 0.0001, one-way ANOVA, mean ± s.d., N = 3 experiments). Numerical and statistics source data as well as unprocessed blots are available as source data.
For group-specific depletion of LAP2 isoforms, we exploited siPOOLs targeting either LAP2α alone or all membrane-bound isoforms together. These isoform-specific depletions worked, as shown in LAP2 immunofluorescence and immunoblotting analyses (Fig. 3b,e). Downregulation of the membrane-bound isoforms (siP-LAP2β, γ and δ) but not of LAP2α mirrored the effect of downregulation of all LAP2 variants (siP-LAP2). This was confirmed by an analysis of centromere positioning (Fig. 3d). We conclude that the INM-bound isoforms of LAP2 promote heterochromatin tethering to the NE. Notably, co-depletion of lamin A/C together with the membrane-bound or all LAP2 isoforms affected H3K9me3 localization more strongly, again indicating an A-type lamin-dependent contribution to heterochromatin tethering that remains to be defined.
The role of LAP2 peripheral heterochromatin localization was confirmed by a rescue experiment exploiting HCT116 cells lacking LBR, LAP2 and LMNA/C (LBR/TMPO/LMNA triple KO (TKO); Extended Data Figs. 5f and 6i). Re-expression of LAP2β-GFP partially rescued the enrichment of H3K9me3-positive chromatin at the NE (Fig. 3f,g). In the rescued cells, we also noticed a still more prominent nucleolar association of H3K9me3 than in WT cells.
Loss of LBR and LAP2 causes widespread changes in gene expression
Most differentiated eukaryotic cells position their heterochromatin underneath the NE; however, the biological consequences of a global loss of peripheral heterochromatin have not been explored. Having a system that allowed the detachment of heterochromatin at will, we set out to determine the consequences on gene expression and genome organization.
The initial RNA-seq analysis had shown that the short-term depletion of NE proteins in WT HCT116 cells by RNAi (si-11, 72 h) affected gene expression very mildly (Extended Data Fig. 1a). Only a small number of genes became deregulated more than two-fold (41). The 28 upregulated genes included genes from inter-LAD (19) and LAD (9) regions (Extended Data Fig. 6a; LADs previously identified by DamID19), whereas the 13 downregulated genes mainly comprised genes targeted by si-11. Furthermore, ATAC-seq (Assay for Transposase-Accessible Chromatin with sequencing), which measures the accessibility of genomic regions42, demonstrated that only a small fraction of the genome (~0.9%, |log2-fold change| ≥ 1) exhibited altered accessibility following short-term depletion of NE tethers (Extended Data Fig. 6b,c). Thus, the short-term effects of RNAi-induced heterochromatin release from the NE on gene expression are minor, even though chromatin forms large intranuclear clusters. We also conducted the experiment for six days, which stimulated more pronounced changes in gene expression (Extended Data Fig. 6d,e), indicating that the duration of downregulation may matter.
To analyse the effects of persistent detachment of heterochromatin from the NE, we generated a panel of HCT116 KO cell lines that carry single, double and triple deletions of the genes for LBR, LMNA/C and LAP2 (TMPO gene) (Extended Data Fig. 5a–f), focusing mostly on the LBR/TMPO double KO (DKO) and LBR/TMPO/LMNA TKO cell lines. These cells proliferated nearly like WT cells (Extended Data Fig. 5g,h). We then examined how the loss of LBR, LMNA/C and/or LAP2 affected the localization of peripheral heterochromatin using antibodies for H3K9me3 and H3K27me3 (Fig. 4a,b). Consistent with RNAi, we observed a decrease in H3K9me3 at the NE in the DKO and TKO backgrounds, accompanied by the appearance of intranuclear clusters of heterochromatin. Similarly, H3K27me3-positive heterochromatin was detached from the NE, showing that facultative heterochromatin is also affected.
a, Representative confocal images of HCT116 WT and the indicated KO cell lines, fixed and subjected to immunofluorescence using antibodies against H3K9me3 or H3K27me3. DNA was stained using Hoechst. Scale bar, 5 μm. b, Quantification of the accumulation of H3K9me3 at the NE relative to its total nuclear signal (total number of cells n ≥ 91, NS P > 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001, one-way ANOVA, mean ± s.d., N = 3 experiments) and H3K27me3 signal (total number of cells n ≥ 92, *P ≤ 0.05, **P ≤ 0.01, **P ≤ 0.001, ****P ≤ 0.0001, one-way ANOVA, mean ± s.d., N = 3 experiments) at the nuclear rim in the various KO backgrounds (a). c, Volcano plots showing differentially expressed genes identified by RNA-seq (HCT116 LBR/TMPO DKO and LBR/TMPO/LMNA TKO versus HCT116 WT cells, N = 3 experiments). Numbers of genes with adjusted P ≤ 0.05 and |log2(fold change)| ≥ 2 are indicated in the volcano plots (two-tailed Wald test and Benjamini–Hochberg procedure). Left: table summarizing two- and four-fold changes in gene expression (|log2-fold change| (LFC)). Right: Venn diagrams indicating the overlap in up- and downregulated genes between LBR/TMPO DKO and LBR/TMPO/LMNA TKO cells. d, Stacked bar plots showing the proportion of differentially expressed genes from c assigned to LAD and inter-LAD regions. A one-sided Fisher’s exact test was used to evaluate the enrichment of upregulated genes in LADs compared to inter-LADs (P values: DKO 1.69 × 10−25; TKO 1.99 × 10−31). All loci with a |log2(fold change)| ≥ 1 and adjusted P ≤ 0.05 were considered significant. e, GO analysis of upregulated genes, showing the ‘Biological process’ category. Genes from c with adjusted P ≤ 0.05 and log2(fold change) ≥ 2 in HCT116 LBR/TMPO DKO and LBR/TMPO/LMNA TKO cells were considered upregulated. A one-sided Fisher’s exact test was used to determine terms with significant enrichment. The x axis shows the –log10(Padj) values for the respective terms. f, Heatmap of innate immunity genes upregulated in the KO backgrounds and part of the identified GO terms ‘negative regulation of viral genome’ and ‘defence response to virus’ from e. g, Volcano plots of RNA expressed from transposable elements identified by TEtranscripts for LBR/TMPO DKO and LBR/TMPO/LMNA TKO cells versus HCT116 WT cells (N = 3, dashed lines indicate changes with P ≤ 0.05 and |log2(fold change)| ≥ 0.585; two-tailed Wald test and Benjamini–Hochberg procedure). DNA, DNA transposon; LINE, long interspersed nuclear elements; LTR, long terminal repeat retrotransposon; satellite, satellite repeats; SINE, small interspersed nuclear elements. Numerical and statistics source data as well as unprocessed blots are available as source data.
Bulk RNA-seq revealed that the deletion of heterochromatin tethers caused widespread changes to the transcriptome (Fig. 4c, Extended Data Fig. 6f and Supplementary Table 1). Hundreds of genes were deregulated by more than four-fold (1,659 in LBR/TMPO DKO genes; 1,360 in TKO), among them transcription factors, which may amplify the effect. About 80% of deregulated genes were upregulated, with a slight preference for genomic regions originally assigned to LADs (Fig. 4d). When comparing the different genotypes, we noted that deletion of LBR alone already caused considerable changes (Extended Data Fig. 6f). LBR and TMPO co-deletion was synergistic when considering downregulated genes, consistent with the notion that the combined loss of LBR and LAP2 leads to a greater extent of heterochromatin collapse, potentially burying genes and diminishing gene expression.
To discern functionally linked clusters among the upregulated genes that might be part of a cellular response to the inflicted changes, we performed Gene Ontology (GO) analysis (Fig. 4e). This identified GO terms pertaining to antiviral innate immunity, chemokine signalling and extracellular matrix organization. STRING analysis of the top upregulated protein-coding genes (Extended Data Fig. 6g) identified a gene cluster of cytokine genes (for example, IFN1B) and innate immunity factors (Fig. 4f), including cytosolic dsRNA sensors43 (RIG-1, MDA-5/IFIH1), oligoadenylate synthases44 (for example, OAS2) and APOBEC cytidine deaminases45 that are potent inhibitors of retroviral replication. However, an induction of innate immunity can be caused by diverse cellular problems, including a dysregulation of epigenetic gene silencing and thereby expression of retrotransposons such as endogenous retroviruses (ERVs)46,47,48, defects in pre-mRNA splicing49, or loss of R-loop resolution causing the occurrence of cytoplasmic RNA–DNA hybrids50. To deduce whether the expression of transposable elements (TEs) was altered following loss of NE–heterochromatin tethers, we analysed our bulk RNA-seq data. This indeed revealed a deregulation of TEs in both the LBR/TMPO DKO and LBR/TMPO/LMNA TKO cell lines, especially of LTR retrotransposons (Fig. 4g). In addition, we also observed a dysregulation of splicing, including changes in intron retention (Extended Data Fig. 6h). It is currently unclear whether these changes in TE expression and splicing or other defects govern innate immune activation.
Persistent loss of heterochromatin tethers causes defects in the maintenance of repressive epigenetic marks
To examine how altered gene expression relates to changes in genome organization, we first analysed alterations in genome accessibility. ATAC-seq revealed that both DKO and TKO cells showed ~14% of detected genomic regions changing accessibility more than two-fold (Fig. 5a and Supplementary Table 2). Increased and decreased accessibility were almost balanced (Fig. 5a). Genes with increasing accessibility largely overlapped between DKO and TKO cells (59%, Fig. 5b), indicating that changes are not random. Among the more accessible genomic regions, genes originally positioned in LADs were slightly overrepresented (Fig. 5c), indicating that LAD genes are preferentially affected by the long-term loss of peripheral heterochromatin localization. Further analysis showed a weak correlation between the expression of protein-coding genes and accessibility changes in promoter regions (Fig. 5d, R = 0.2 for DKO and TKO), suggesting that other effects contribute.
a, Stacked bar plot showing the proportion of differentially accessible ATAC-seq peaks in nucleosome-free regions (Nfr) for LBR/TMPO DKO and LBR/TMPO/LMNA TKO cells (N = 3). All loci (no selection for genes) with |log2(fold change)| ≥ 1 and adjusted P ≤ 0.05 were considered significantly changed (two-tailed Wald test and Benjamini–Hochberg procedure). b, Venn diagrams indicating the overlap in changes in genome accessibility between LBR/TMPO DKO and LBR/TMPO/LMNA TKO, related to a. c, Stacked bar plots showing proportions of differentially accessible ATAC-seq peaks (|log2(fold change)| ≥ 1 and adjusted P ≤ 0.05) from a assigned to LAD and inter-LAD regions. A one-sided Fisher’s exact test was used to evaluate the enrichment of peaks with increased accessibility in LADs compared to inter-LADs. P values: DKO 2.36 × 10−238; TKO 1.14 × 10−204. d, Spearman correlation between changes in gene expression (RNA-seq) and promoter accessibility (ATAC-seq) for protein-coding genes. The promoter was defined as the region 3,000 bp upstream and 500 bp downstream of the transcription start site (TSS). Significantly affected genes (|log2(fold change)| ≥ 1 and adjusted P ≤ 0.05) in RNA-seq and ATAC-seq datasets (two-tailed Wald test and Benjamini–Hochberg procedure) are highlighted in magenta and unchanged genes are coloured in grey. Number of genes, n = 13,768 for both comparisons; for both DKO versus WT and TKO versus WT: R = 0.20 and P < 2.2 × 10−16. e, Representative immunoblot of H3, H3K9me2, H3K9me3, H3K27me3 and H4K20me3 relative to β-actin in LBR/TMPO DKO and LBR/TMPO/LMNA TKO cells. N = 3. See also Extended Data Fig. 6i,j. f, Quantification of protein levels (normalized to β-actin) for histone modifications in LBR/TMPO DKO and LBR/TMPO/LMNA TKO cells (NS P > 0.05 *P ≤ 0.05, **P ≤ 0.01, ****P ≤ 0.0001, one-way ANOVA, mean ± s.e.m., N = 3 experiments). Numerical and statistics source data as well as unprocessed blots are available as source data.
Considering the extensive upregulation of gene expression, we wondered whether there were any defects in the maintenance of repressive histone marks and therefore examined their levels by quantitative immunoblotting. Strikingly, although H3K9me3 and H4K20me3 remained nearly unaffected, H3K27me3 was strongly reduced in the LBR/TMPO DKO and LBR/TMPO/LMNA TKO cells (Fig. 5e,f and Extended Data Fig. 6i,j). We also observed a reduction in H3K9me2. Notably, there were no major changes in mRNA levels of polycomb repressive complex (PRC) components that would directly explain the decrease in H3K27me3 (Extended Data Fig. 6k and Supplementary Table 4). Only the mRNA levels of CBX2 (PRC1 component) dropped, whereas CBX7 was upregulated. CBX7 is normally expressed in pluripotent cells and replaced by CBX2 during lineage commitment51, indicating a potential confusion in cell state-specific gene expression.
The reduction in H3K27me3 levels suggested that de-repression of H3K27me3-marked genes might contribute to the upregulation of gene expression. H3K27me3 chromatin immunoprecipitation (ChIP) indeed revealed that the mean H3K27me3 peak intensity strongly dropped in the DKO and TKO cells (Fig. 6a), consistent with the reduction in overall H3K27me3 levels (Fig. 5f). Most peaks were strongly reduced, and only a small fraction of regions showed an increased H3K27me3 signal. When specifically looking at changes in the promoter region of protein-coding genes, about one-third of upregulated genes showed a decrease in H3K27me3 levels (Fig. 6b) and genes originally positioned in LADs were preferentially affected (Fig. 6c; an example is shown in Fig. 6f).
a, Metagene profiles (top) and heatmap (bottom) of the RPGC-normalized H3K27me3 ChIP-seq signal for HCT116 WT, LBR/TMPO DKO and LBR/TMPO/LMNA TKO cells, ranked according to H3K27me3 enrichment in HCT116 WT cells (N = 3 experiments). RPGC, reads per genomic content (1× normalization). b, Stacked bar plots showing the proportion of genes with the indicated status of H3K27me3 (two-sided quasi-likelihood F-test) among the downregulated, unchanged and upregulated protein-coding genes (related to a and Fig. 4c). c, Stacked bar plots showing the proportion of regions with differential H3K27me3 signal (two-sided quasi-likelihood F-test) from a assigned to LAD and inter-LAD regions (|log2(fold change)| ≥ 1 and adjusted P ≤ 0.05) assigned to LAD and inter-LAD regions. A one-sided Fisher’s exact test was used to evaluate the enrichment of peaks with decreased H3K27me3 signal in LADs compared to inter-LADs. P values: DKO 8.94 × 10−118; TKO 9.25 × 10−70. d, Spearman correlation between changes in gene expression (RNA-seq) and H3K27me3 ChIP-seq signal in the promoters (−3,000 to +500 bp of TSS) of protein-coding genes. Significantly affected genes (|log2(fold change)| ≥ 1 and adjusted P ≤ 0.05) in RNA-seq (two-tailed Wald test and Benjamini–Hochberg procedure) and H3K27me3 ChIP-seq datasets (two-sided quasi-likelihood F-test) are highlighted in magenta and unchanged genes are coloured in grey. Number of genes, n = 2,954; DKO versus WT R = −0.20 and P < 2.2 × 10−16; TKO versus WT, R = −0.22 and P < 2.2 × 10−16. e, Metagene profiles of RPGC-normalized H3K27me3 ChIP-seq signal for the top 100 up- and downregulated genes from Fig. 4c for HCT116 LBR/TMPO DKO and LBR/TMPO/LMNA TKO cells relative to WT. Start and End represent gene start and end. RPGC, reads per genomic content (1× normalization). f, RNA-seq, ATAC-seq and H3K27me3 ChIP-seq profiles for an example upregulated gene (ADAM28), originally positioned in a LAD region, comparing HCT116 WT, DKO and TKO cells. The region overlaid in grey highlights the start of the gene. Gene upregulation coincides with increased accessibility (ATAC-seq) in its promoter and a drop in H3K27me3 throughout its gene body (ChIP-seq). Statistics source data are available as source data.
Correlation analysis confirmed a weak negative correlation between changes in H3K27me3 and gene expression (R = −0.20 DKO; R = −0.22 TKO; Fig. 6d), suggesting that altered H3K27me3 levels may explain deregulation for a fraction of genes. This notion was supported by comparing the mean H3K27me3 intensity profiles for the top 100 up- and downregulated genes between WT and KO cells, which showed a reduction of the H3K27me3 mean signal strength for upregulated genes, and an increase for downregulated genes (Fig. 6b,f). Similarly, we also analysed changes in H3K27me3 for upregulated TEs (Fig. 4g), and noticed a drop in the mean H3K27me3 signal for upregulated TE copies originally positioned in LADs in both the DKO and TKO cells (Extended Data Fig. 6l and Supplementary Table 3). For comparison, we also performed H3K9me3 ChIP, which did not reveal similar changes in the profiles.
Loss of peripheral heterochromatin tethering causes changes in 3D genome organization
To analyse how the release of heterochromatin from the NE impacts genome organization, we mapped the 3D genome topology using Pico-C52,53,54. We generated datasets for HCT116 WT, DKO and TKO cells, obtaining maps at low kilobase (kb) resolution (Fig. 7a and Extended Data Fig. 7a–c). These revealed extensive new interactions both within and between chromosomes in the KO genotypes (exemplified for chromosomes 4 and 5 in Fig. 7a, rectangular areas; Extended Data Fig. 7d). Analysis of trans compartment interactions showed that novel contacts are preferentially established between B.2 compartments, the compartment type housing LADs (Fig. 7b). This might indicate that, upon release of LADs from the NE, the collapse of heterochromatin into intranuclear clusters yields novel B.2–B.2 contacts, which is especially prominent in the TKO background, in line with the strength of chromocentre formation (Figs. 7b and 4a).
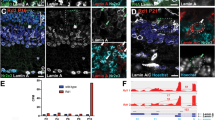
a, Pico-C contact maps of chromosomes 4 and 5 in HCT116 WT, DKO and TKO cells, revealing increased long-range cis and trans interactions between B.2 compartments (black boxes) that are more pronounced in TKO. log2 ratio maps (mutant/control) are shown (middle (DKO) and right panel (TKO)), highlighting interaction gains. Compartment annotations (A.1, A.2, B.1, B.2) are shown below. (N = 3 experiments). b, Aggregate trans-compartment contact maps show a specific increase in B.2–B.2 interactions in DKO and TKO, compared to control. c, Genome-wide compartment trends. Top: compartment distribution across the genome. Bottom: fraction of loci undergoing compartment changes, predominantly gains or losses of B.2 identity. d, Neighbourhood analysis of loci switching from B.1 to B.2 (right) or to A compartments (left) in TKO cells shows that most loci gain the identity of their local neighbourhood. e, DamID profiles for LMNB1 and NAD at lost, stable or gained B.2 loci. Lost B.2 regions are enriched for LMNB1, whereas stable B.2 loci show higher NAD association. f, De novo motif analysis reveals enrichment of CTCF and AP-1 complex motifs (for example, FOSL1) at sites undergoing compartment switching. g, Deeply sequenced Pico-C maps (8-kb resolution) illustrate local structural changes at B.2 regions in DKO and TKO, accompanied by changes in chromatin accessibility (ATAC-seq), histone modifications (H3K9me3 and H3K27me3) and transcription (RNA-seq). LMNB1 and NAD data are from ref. 94, HP1α data from ref. 95, and CTCF binding from ENCODE (UCSC Genome Browser, TFBS Clusters v3). Numerical source data are available as source data.
In addition, a substantial fraction of the genome (29.3% DKO; 32.5% TKO) changed compartment identity (Fig. 7c). Shifts to more active compartments (from B to A) prevailed, whereas transitions from A to B were less prominent. Most widespread were transitions from B.1, which is enriched in H3K27me3-marked facultative heterochromatin, into the transcriptionally more active A category (Fig. 7c, lower graphs). Notably, the genomic environment seems to impose a bias to the direction of compartment switches: loci switching from B.1 to A and from B.1 to B.2 are preferentially surrounded by A and B.2 compartments, respectively (Fig. 7d and Extended Data Fig. 7e,f). Compared to the rest of the genome, genes encompassed by the 469 regions switching from B.1 to A in both the DKO and TKO cells are overrepresented for upregulated genes (Extended Data Fig. 7g).
Interestingly, the compartment most affected in its dimensions is the B.2 compartment, which has undergone a net reduction in size. When the affected genomic B.2 regions are aligned with LADs assigned by DamID, the fraction of changing B.2 regions show a high LMNB1 DAM signal, indicating that they originate from LADs, whereas stable B.2 regions are enriched for NADs (Fig. 7e). Thus, compartment changes are probably caused by LAD untethering from the NE. When we zoom into such B.2 regions that undergo switches, their changes seem to be influenced by their surrounding genomic neighbourhood and changes in histone modifications. For example, shrinking of a B.2 compartment is accompanied by a reduction in the broad H3K9me3 mountain across the B.2 domain and a drop in the H3K27me3 ChIP signal in the neighbouring B.1 regions (Extended Data Fig. 7f,h). We also observed a gain of pronounced topologically associating domain (TAD) substructure in compartments changing from B.2 to B.1 and A. These regions showed a marked increase in accessibility by ATAC-seq, probably explained by de novo CTCF loading, consistent with an enrichment of CTCF binding sites in lost B.2 compartments (Fig. 7f). Conversely, compartments that adopt a B.2 state lose their TAD substructure (Extended Data Fig. 7i).
Loss of LBR and LAP2 affects the differentiation of mESCs
LADs are established shortly after fertilization55, with cumulative changes accompanying lineage commitment and cell fate determination56,57,58,59,60. Chromatin organization in mESCs is globally affected by the deletion of lamins61, pointing at an important function of the nuclear periphery in genome organization in pluripotent cells.
To investigate the role of the identified NE–heterochromatin tethers in mESCs, we generated Lbr/Lmna DKO, Lbr/Tmpo DKO and Lbr/Tmpo/Lmna TKO in the blastocyst-derived mESC line E14TG2a (Extended Data Fig. 8). We maintained these cells in a naïve pluripotent ground state in the presence of LIF/2i62. Immunofluorescence analysis of H3K9me3, H4K20me3 and H3K27me3 revealed that both Lbr/Tmpo DKO and Lbr/Tmpo/Lmna TKO cells lost the enrichment of heterochromatin at the nuclear periphery and showed an accumulation of chromocentre-like heterochromatin clusters in the nuclear interior (Fig. 8a–c). In contrast, Lbr/Lmna DKO cells maintained a large fraction of heterochromatin at the nuclear rim, while also showing intranuclear clusters.
a, Representative confocal images of WT, Lbr/Lmna DKO, Lbr/Tmpo DKO and Lbr/Tmpo/Lmna TKO E14TG2a mESCs, fixed and subjected to immunofluorescence using anti-H3K9me3, anti-H4K20me3 and anti-H3K27me3 antibodies. DNA was stained using Hoechst. Note that mouse cells possess chromocentres, which are clusters of centromeric heterochromatin typically found attached to the NE in WT cells. Scale bar, 5 μm. b, Schematic depiction of the quantification method. The fraction of antibody-stained heterochromatin at the nuclear rim was quantified using ten line scans across the NE per analysed cell nucleus, with each line reaching 1 μm into the nucleus and the cytoplasm. c, Quantification of fluorescence signal at the nuclear rim in a, showing the mean normalized fluorescence intensity at each point of the line scan (top) and the highest peak mean intensity (bar plots). Lbr/Lmna DKO, Lbr/Tmpo DKO and Lbr/Tmpo/Lmna TKO mESCs (total number of cells n = 12, **P ≤ 0.01, ****P ≤ 0.0001, Welch’s t-test, N = 3 experiments, mean ± s.d.). d, Volcano plots showing differentially expressed genes identified by RNA-seq (Lbr/Tmpo DKO and Lbr/Tmpo/Lmna TKO versus WT cells, N = 3). Genes with an adjusted P ≤ 0.05 (two-tailed Wald test and Benjamini–Hochberg procedure) are marked as up- or downregulated if |log2(fold change)| ≥ 1. e, GO analysis showing the ‘Biological process’ category. Genes from d with P ≤ 0.05 and log2(fold change)| ≥ 1 in Lbr/Tmpo DKO and Lbr/Tmpo/Lmna TKO cells were considered upregulated. A one-sided Fisher’s exact test was used to determine terms with significant enrichment. The x axis shows the –log10(Padj) values for the respective terms. f, Heatmap of piRNA pathway genes upregulated in Lbr/Tmpo DKO and Lbr/Tmpo/Lmna TKO cells. g, Representative phase contrast images of E14TG2a WT, Lbr/Tmpo DKO, Lbr/Lmna DKO and Lbr/Tmpo/Lmna TKO cells after ten days of embryoid body differentiation. Scale bar, 200 μm. h, Quantification of cell numbers after ten days of embryoid body differentiation (****P ≤ 0.0001, one-way ANOVA, mean ± s.e.m., N = 3 experiments). i, Representative immunoblot of H3K27me3 and H3 relative to β-actin in Lbr/Tmpo DKO and Lbr/Tmpo/Lmna TKO cells. N = 3 experiments. j, Quantification of H3K27me3 (normalized to β-actin) in Lbr/Tmpo DKO and Lbr/Tmpo/Lmna TKO cells (****P ≤ 0.0001, one-way ANOVA, N = 3 experiments, mean ± s.e.m.). k, Heatmap displaying the expression of marker genes for the three germ layers, primed epiSCs and pluripotency in Lbr/Tmpo DKO and Lbr/Tmpo/Lmna TKO cells as identified by RNA-seq after ten days of embryoid body differentiation. Numerical and statistics source data as well as unprocessed blots are available as source data.
To examine the effects of loss of heterochromatin tethering to the NE on genome organization and gene expression, we performed ATAC-seq and RNA-seq, focusing on Lbr/Tmpo DKO and Lbr/Tmpo/Lmna TKO cells. Compared to what we had observed in differentiated HCT116 cells (Figs. 4 and 5), ATAC-seq showed fewer changes in chromatin accessibility in mESCs (Extended Data Fig. 9a,b and Supplementary Table 2), with a larger fraction of genomic regions becoming less accessible, probably because chromatin in stem cells is generally more accessible at the outset. Consistently, RNA-seq identified prominent changes in gene expression (Fig. 8d), with downregulation of genes prevailing in both DKO and TKO cells (Extended Data Fig. 9c).
GO analysis highlighted the upregulation of genes related to retrotransposon silencing, interferon response, piRNA processing, and, for the TKO, adaptive immunity (Fig. 8e). Upregulated genes related to ‘retrotransposon silencing and piRNA processing’ comprised piRNA pathway genes, including Ddx4/Vasa, Piwil2, Piwil4, Maelstrom (Mael), Tdrd1 and Tdrd563 (Fig. 8f), but we did not observe a significant upregulation of TEs (Extended Data Fig. 9d). As both the DKO and TKO cell lines maintain expression of pluripotency markers (Pou5f1/Oct4, Nanog) and do not show signs of precocious differentiation when maintained in LIF/2i (Extended Data Fig. 9e), it will be interesting to investigate whether the upregulation of piRNA biogenesis genes reflects an induced innate immune response to the activation of retrotransposons64 or the emergence of cells that have adopted a different cell state65.
Finally, we used embryoid body differentiation (EB) to examine whether loss of heterochromatin tethering affects differentiation. By withdrawal of LIF/2i, cells were induced to differentiate into the germ layers endoderm, mesoderm and ectoderm, and allowed to cluster into EBs by growth in suspension for six days. After EB reattachment to gelatin-coated plates, we observed a marked reduction in both the size of the EBs and the total number of cells recovered after ten days of differentiation, both for the DKO and TKO cells. This suggests that a large proportion of cells are unable to successfully transition from pluripotency to a differentiated state (Fig. 8g,h and Extended Data Fig. 9f). At the same time, global H3K27me3 levels were drastically reduced in differentiated DKO and TKO cells (Fig. 8i,j).
RNA-seq revealed that cells contained in the developed EBs showed changes in the relative abundance of germ-layer markers (Fig. 8k, Extended Data Fig. 9f and Supplementary Table 1). The expression of markers for endoderm (Foxa1, Dab2) and mesoderm (Foxf1, Hand2 and Hand1) was reduced in the KO cells, whereas ectoderm, especially neuroectoderm, markers (Sox1, Nestin) were significantly increased, indicating that the induced changes in chromatin organization impair cell fate determination. The ectoderm differentiation pathway is considered the ‘default’ differentiation pathway, as ectoderm enhancers are already accessible in E4.5 pluripotent epiblast cells, whereas endo- and mesoderm enhancers are not66. We particularly noticed a strong upregulation of genes expressed in neuroectoderm, in particular factors involved in GABAergic synaptic transmission, indicating that some cells might have further differentiated precociously (Extended Data Fig. 9g). In conclusion, mESCs that have lost NE–heterochromatin tethers change their gene-expression profile, have severe defects in forming EBs, and are compromised in the fidelity of cell fate determination.
Discussion
We have demonstrated that the collective downregulation of a set of ubiquitously expressed NE proteins leads to the detachment of heterochromatin from the nuclear periphery. Among these factors, we identified LBR and the membrane-bound isoforms of LAP2 as the major NE tethers for bulk H3K9me3-marked heterochromatin, including centromeres. Their persistent loss causes changes in gene expression, chromatin organization and the maintenance of repressive histone marks, especially of H3K27me3. mESCs lacking these tethers have defects in the transition from pluripotency to differentiation and cell fate determination.
The current model of nuclear organization posits that the conventional radial positioning of chromatin is driven by the active tethering of heterochromatin to the nuclear lamina67. This view is based on groundbreaking studies of the inverted chromatin architecture in rod cell nuclei of nocturnal animals8 and on modelling approaches exploiting polymer simulations9. In support of this model, we find that downregulation of the identified tethers leads to a detachment of bulk heterochromatin from the NE, providing direct experimental evidence for the importance of active tethering to peripheral heterochromatin positioning. Following loss of tethering, the detached heterochromatin accumulated prominently around nucleoli, suggesting that some of the affected genomic regions are converted into NADs68,69, consistent with the known reshuffling of genes between LADs and NADs even in unperturbed, dividing cells16,17. Because LADs and NADs both provide a mostly repressive environment for gene expression, this repositioning is not expected to cause immediate effects on gene activity. In addition, some of the detached heterochromatin formed dense nucleoplasmic foci reminiscent of chromocentres, supporting a model proposing a tug of war between adhesive forces executed by NE–heterochromatin tethers and cohesive forces generated by heterochromatin bundling factors70.
Our systematic deconvolution experiments revealed a major role of LBR and LAP2 in the peripheral positioning of constitutive heterochromatin. LAP2 was originally identified as a lamin B1 interactor71, which would place it in the B-type tether category, but a functional interplay with A-type lamins is also possible. To tether heterochromatin, LBR and LAP2 must decode specific molecular cues of heterochromatin such as epigenetic histone marks, heterochromatin-resident proteins or DNA methylation. LBR can recognize several such heterochromatin features, including HP1 (which binds H3K9me372,73), the methyl CpG binding protein MeCP274 and H4K20me275. In contrast, it is unclear how LAP2 promotes NE attachment of constitutive heterochromatin. LAP2 possesses an N-terminal DNA-binding LEM-like domain of unknown specificity that may contribute to tethering76. The adjacent barrier-to-autointegration factor (BAF)-binding LEM domain77 is unlikely to confer heterochromatin specificity, because BAF binds chromatin irrespective of its epigenetic status78. In addition, LAP2β possesses an internal region that interacts with HDAC379 to support NE anchorage of so-called lamina-associated sequences (LASs)80. These DNA elements are characterized by extended GAGA motifs found prominently in polycomb response elements81. GAGA motifs associate with the transcriptional repressor cKrox, which serves as a bridge to HDAC3 and LAP2β80. Our data suggest that LAP2’s role in tethering heterochromatin goes beyond LASs, because its downregulation also affected constitutive heterochromatin, including centromeres. Thus, it remains to be defined how the membrane-bound isoforms of LAP2 function as tethers. Interestingly, de(p)letion of A-type lamins exacerbated the effect of LBR and LAP2 loss on peripheral heterochromatin localization, indicating that A-type lamin-associated factors might also contribute. These may include LAP1, the LEM protein emerin (which delocalizes to the endoplasmic reticulum upon deletion of A-type lamins82), or nucleoplasmic factors such as PRR1483,84.
Short-term depletion of NE factors by RNAi caused the global release of heterochromatin from the nuclear periphery, but had little immediate consequences on chromatin accessibility and gene expression. In contrast, deletion of the genes encoding for LBR and LAP2 caused massive changes in the transcriptome, indicating that their persistent absence has more severe consequences. In HCT116 DKO and TKO cells, we observed a significant general decrease in the levels of silencing heterochromatin marks, especially in H3K27me3 and H3K9me2. About one-third of the upregulated protein-coding genes displayed a reduction in H3K27me3, linking the decrease in maintenance of this epigenetic mark to the deregulation of the genome. The drop in H3K27me3 suggests that facultative heterochromatin might be especially vulnerable upon release from the NE. Micro-C revealed that a considerable fraction of B.1 compartments, which are usually enriched in H3K27me3-marked chromatin15,85, change their identity to either A or B.2. Although changes to B.2 will be neutral for gene expression, shifts to A bring genes into a different, permissive chromatin environment, making them more susceptible to activation. The question arising is whether the reduction in H3K27me3 drives the compartment changes or whether, in reverse, altered chromatin positioning disrupts the maintenance of epigenetic marks. We favour a model of heterochromatin release from the NE being causative for changes in 3D genome organization, with subsequent defects in epigenetic maintenance in cycling cells (Extended Data Fig. 10). The restoration rate of H3K27me3 after replication is known to correlate with dense chromatin86. Thus, changes in 3D genome organization upon release of peripheral heterochromatin may perturb the nucleation and spread of polycomb domains that occur every cell cycle87, leading to the erasure of H3K27me3-marked nucleosomes over time. We propose that the peripheral positioning of polycomb domains in close vicinity to constitutive heterochromatin at the nuclear lamina may support the maintenance of their epigenetic state.
Among the factors that were upregulated in HCT116 KO cells, we identified a cluster of type I interferon-regulated innate immunity genes. Currently, we do not know what the cause for activation of innate immunity is, but this response coincided with deregulation of LTR retrotransposons. Interestingly, we observed a drop in H3K27me3 at upregulated TEs originally positioned in LADs. Recently, H3K27me3 has been suggested to contribute to silencing of TEs in diverse eukaryotes such as plants, ciliates and fungi88,89. Although H3K9 and DNA methylation prevail in TE repression in mammals, it is possible that H3K27me3, perhaps together with a decrease in H3K9me2, may also contribute to the de-repression of TEs, in particular of LTRs. We also noted changes in splicing, potentially caused by the spatial repositioning of genomic loci relative to nuclear speckles90.
In summary, the massive deregulation of gene expression in KO cells might be the cumulative effect of different classes of change. We propose that these include (1) immediate deregulation of a minor fraction of genes upon release of heterochromatin from the nuclear periphery, (2) deregulation of genes by inflicted changes in 3D genome organization with a long-term impact on compartment identity and epigenetic control, (3) indirect consequences of the former, for example, by effects on the expression levels of transcription factors, and (4) a cellular response to the consequences of the inflicted changes in gene expression and genome organization (innate immunity activation).
A feature of pluripotent cells is an open state of chromatin organization, enabling plasticity for reorganization during cell fate determination and differentiation91. mESCs lacking the heterochromatin tethers already displayed changes in gene expression in the pluripotent state. Downregulation of genes prevailed, probably because the open state of chromatin in ESCs was affected by the formation of intranuclear heterochromatin clusters. When ESCs exit pluripotency, the correct establishment and maintenance of different heterochromatin types accompany cell lineage determination and differentiation56,92. Consistent with this notion, we observed defects in cell fate determination and differentiation when NE–heterochromatin tethers were deleted, extending recent work ablating lamin and LBR expression93. Upon induction of differentiation, a large fraction of cells were unable to transition out of pluripotency. At the same time, proliferating cells showed defects in cell fate determination, suggesting an important contribution of peripheral heterochromatin localization during a developmental period that involves substantial alterations in genome organization. The confusion in cell fate manifested in a signature indicating precocious differentiation into neurectoderm, the default differentiation pathway of mESCs, suggesting that cells were unable to properly integrate the upstream differentiation signals.
We suspect that the peripheral attachment of heterochromatin is important also for other cellular features, whether it be replication timing and DNA repair or the mechanical rigidity of cell nuclei, especially in cell types that experience deformation or mechanical strain. The conventional nuclear architecture with a sheet of heterochromatin spread out underneath the NE seems to have evolved to prevent the undesired collapse and clustering of heterochromatin, which may impair cellular plasticity, probably with fatal consequences for development and differentiation.
Methods
Antibodies
The following primary antibodies were used in this study: rabbit anti-H3K9me3 (Abcam 8898, 1:500), mouse anti-H3K9me3 (Active Motif AB_2935892, 1:500), mouse anti-H3K9me2 (Abcam 1220, 1:500), rabbit anti-H4K20me3 (Abcam 9053), rabbit anti-H3K27me3 (Millipore 07-449, 1:500, and Active Motif AB_2561020, 1:500 (ChIP)), rabbit anti-H3 (Abcam 1791, 1:1,000), H2Av antibody (Active Motif 61686), rabbit anti-LBR(hu) (Abcam 32535, 1:500), rabbit anti-lamin A/C (Proteintech, 10298-1-AP, 1:1,000), mouse anti-lamin A/C (Kerafast, LASS2D9, 1:500), rabbit anti-lamin B1 (Abcam 16048, 1:500), rabbit anti-lamin B2 (Abcam 155319, 1:500), rabbit anti-LAP1 (Abcam 86307, 1:500), mouse anti-LAP2 (BD Transduction Laboratories 611000, 1:1,000), rabbit anti-Lap2 (Proteintech 1465-1-AP, 1:1,000), rabbit anti-emerin (Abcam 40688, 1:1,000), rabbit anti-LEMD2 (Sigma HPA017340, 1:200), rabbit anti-SUN196, rabbit anti-SUN2 (Sigma HPA001209, 1:500), mouse anti-beta-actin (SantaCruz sc-47778, 1:1,000), mouse anti-CLIMP-63 (Enzo Life Sciences ALX804-604-C100, 1:200), human anti-CREST (ImmunoVision HCT0100, 1:1,000), rabbit anti-Lbr(mouse) (Abnova PAB15583, 1:2,000) and rabbit anti-emerin(mouse) (Novus Biologicals NBP1-87692, 1:1,000). The rabbit anti-RRP12 antibody has been described previously97. The specificity of the H3K9me2 and H3K9me3 antibodies used in this study were validated previously98 and confirmed using siPOOLs targeting their cognate histone methyltransferases.
The secondary antibodies used included mouse Alexa Fluor 488 (ThermoFisher Scientific, A-11001), mouse Alexa Fluor 594 (ThermoFisher Scientific, A-11005), rabbit Alexa Fluor 488 (ThermoFisher Scientific, A-11034), rabbit Alexa Fluor 594 (ThermoFisher Scientific, A-11012), human Alexa Fluor 647 (ThermoFisher Scientific, A-21445) used for immunofluorescence at 1:300, and mouse Alexa Fluor Plus 680 (ThermoFisher Scientific, A32729) and rabbit Alexa Fluor Plus 800 (ThermoFisher Scientific, A32735) used for blotting at 1:10,000.
Cell lines and cell culture
HCT116 (CCL-247; ATCC), hTERT-RPE-1 (kind gift from J. Mansfeld, Institute of Cancer Research, London) and NIH/3T3 (kind gift from V. Vogel, ETHZ) cells were grown in Dulbecco’s modified Eagle medium (DMEM; Gibco), supplemented with 10% fetal calf serum (FCS; Eurobio Scientific) and 100 µg ml−1 penicillin/streptomycin (Corning) at 37 °C, 5% CO2. ES-E14TG2a mESCs (CRL-1821; ATCC) were grown in DMEM (Sigma) supplemented with 13% FCS (Sigma), 0.32% 2-mercaptoethanol (Gibco) and 100 µg ml−1 penicillin/streptomycin (Corning). Media were freshly supplemented with 1,000 units ml−1 recombinant mouse LIF protein (ESGRO), 1 µM MEK/ERK inhibitor PD0325901 (Sigma) and 3 µM GSK inhibitor CHIR99021 (Sigma, 2i). The cells were plated at a density of 2.65 × 105 cells per cm2 on plates coated with 0.2% gelatin from porcine skin (Sigma), then grown at 37 °C, 8% CO2, and passaged every 48 h using 0.05% trypsin-EDTA for detachment (Gibco). Cells were frozen in FCS (Sigma) supplemented with 10% dimethyl sulfoxide (DMSO; Gibco).
Cell proliferation and viability assays
To examine cell proliferation in HCT116 KO cell lines, cells were seeded at a density of 4 × 104 per well of a six-well plate, allowed to settle for 4 h, and counted at 0 h, 20 h and 44 h in a counting chamber (Neubauer). To measure the cell viability of E14TG2a mESCs during differentiation, cells were seeded into 96-well plates coated with 0.2% gelatin from porcine skin (Sigma) at a density of 1,000 cells per well using an EL406 microplate washer dispenser (BioTek) and grown in the absence of LIF/2i. Cell viability was measured 24, 48 and 72 h after seeding by the addition of 10 µg µl−1 resazaurin (Sigma). After 4 h of incubation, fluorescence was measured using a Tecan Infinite 200 plate reader with an excitation wavelength of 544 nm and emission wavelength of 590 nm.
Flow cytometry
To perform cell-cycle analysis of HCT116 cells, cells were fixed in 70% ethanol at −20 °C overnight. Staining was performed with 300 μM propidium iodide (Sigma) in 1 ml of staining buffer (50 mM HEPES pH 7.5, 700 mM NaCl, 12 mM CaCl2). Cells were incubated for 15 min at room temperature (r.t.) before analysis on a BD LSRFortessa flow cytometer (BD Biosciences) utilizing a 488-nm laser and detected with a 610/20 filter at the Flow Cytometry Core Facility of ETH Zürich. The flow cytometry results were analysed using FlowJo v10.8 software (BD Life Sciences) utilizing univariate Watson (Pragmatic) modelling.
Molecular cloning
The coding region of full-length LAP2β was amplified from pEGFP-N3-LAP2β99 and cloned into KpnI and NotI sites of pCDNA5/FRT/TO (Invitrogen), which had been modified by inserting the enhanced green fluorescent protein (EGFP) coding sequence into the XhoI and ApaI restriction sites, allowing for the expression LAP2β-EGFP.
Transient transfection of cells
Transfection of HCT116 cells with plasmid vectors was carried out using jetPRIME (Polypus). In each well of a six-well plate, 100 µl of jetPRIME buffer, 3 µl of jetPRIME and 1 µg of plasmid DNA were mixed and incubated for 15 min. This mixture was added to cells, and the medium was exchanged the following day.
RNAi
RNAi-mediated depletion of target proteins was accomplished using siPOOLs (siTOOLs Biotech) at a working concentration of 2 nM to deplete the mRNAs encoding for the following proteins: human LBR, LMNA, TOR1AIP1 (LAP1), EMD, TMPO (LAP2), LEMD2, LEMD3 (MAN1), SUN1, SUN2, LMNB1, LMNB2, TMPO isoform alpha, TMPO isoforms beta, delta and gamma, and mouse Lbr, Lmna, Tor1aip1, Emd, Tmpo, Lemd2, Lemd3, Sun1, Sun2, Lmnb1 and Lmnb2. The siPOOL negative control (siTOOLs Biotech) was used for all cell lines except for HCT116 cells, for which the AllStars negative control (Qiagen), a single non-targeting siRNA, was used instead (because high concentrations of the siPOOL control affected the HCT116 cell nuclear morphology, although the other cell lines were seemingly unaffected). In each well of a six-well plate, for every 1 nM siPOOL, 20 µl of Opti-MEM (Gibco) and 0.1 µl of RNAiMAX (ThermoFisher Scientific) were mixed, incubated for 15 min at r.t. and added to cells. For mouse cells, a double pulse of RNAi was carried out, re-transfecting cells 48 h after the initial treatment. Samples were collected after a total of 96 h.
CRISPR-Cas9-mediated generation of KO cell lines
CRISPR RNA guides were designed using the webtools ‘E-CRISP’ and ‘CHOPCHOP’. The guides 5′-GTGAAGTGGTAAGAGGTCGA-3′ and 5′-CAATGTGACGCTGCCGGCCG-3′ were used to target the first exons of the LBR and TMPO genes, respectively. The DNA templates of the guide RNA along with their reverse complement strands were ordered as single-stranded DNA oligonucleotides (Merck) and annealed, yielding double-stranded DNA inserts with overhangs complementary to the vector pC2P-Cas9100. The inserts were ligated into the linearized vector. To target the LMNA gene, the previously described pC2P-gLMNA/C vector was used101. The resulting plasmids (1 µg) were transfected into HCT116 cells using jetPrime (Polyplus). After 24 h, the cells were detached and transferred to a 10-cm plate and selected with 10 µg ml−1 of puromycin (Sigma) for 48 h. The puromycin was washed away, and clones resistant to puromycin were picked, expanded and characterized. Clones containing frameshift mutations were identified using polymerase chain reaction (PCR) and Sanger sequencing, with indels being identified using the TIDE web tool102 and through manual inspection of the sequencing chromatograms. To screen the LBR KO clones via PCR, the primers 5′-GATCACGGTAGGTGGGTAACA-3′ and 5′-ATCCTCTGCCTTCAAACCGA-3′ were used. LBR KO clone #2 was used for the creation of the double and triple KO cell lines and in the corresponding RNA-seq experiments. To screen the KO clones for edits in the TMPO gene, the primers 5′-TTTGTGTCCGCGAGTTTTTGTTCCG-3′ and 5′-GCCAAACGCCCGCCTTTGTAG-3′ were utilized. Finally, edits made in the LMNA gene were uncovered using the primers 5′-TTTCTCCCCGACCCTTTTGC-3′ and 5′- GGCAAAGTTATCGGCCTCCA-3′.
CRISPR-Cas9-mediated gene KO of Lbr, Tmpo and Lmna in ES-E14TG2a cells was done as previously described103. In brief, paired CRISPR/Cas9 sgRNAs targeting the start and stop codons of the gene of interest were designed. The guide pair 5′-GTTGAGGGTGAAGTGGTCAG-3′ and 5′-GTAGCACGGCATGATCAGAC-3′ was used to target Lbr, 5′-CGGGCACGGCACAGAACTCG-3′ and 5′-CCAAGGGAGCATACTTCGGGACG-3′ to target Tmpo, and 5′-GCGGCGCGCCACCCGCAGTG-3′ and 5′-CTCGGCCTCGTGTAGTGCCG-3′ to target Lmna. The respective sgRNAs targeting the start codon were cloned into pLentiCRISPR-EGFP (Addgene plasmid ID 75159), and those targeting the stop codons were cloned into pLentiCRISPR-mCherry (Addgene plasmid ID 75161), whereas for Lbr, both guides were cloned into pX458-GFP (Addgene plasmid ID 48138). A total of 160,000 cells were seeded into a well of a six-well plate. For co-transfection of both vectors the next day, 2 μg of each vector was added to 250 μl of reduced-serum medium (Opti-MEM; Gibco). In parallel, 250 μl of Opti-MEM (Gibco) was mixed with 10 μl of Lipofectamine 2000 (L2000; Invitrogen). The two solutions were combined and incubated at r.t. for 20 min, then 500 μl of the mixture was added to the cells to yield a final volume of 1.5 ml of medium. Media were exchanged after 6 h. At 48 h after transfection, the cells were sorted (BDFACSAria, Flow Cytometry Core Facility, ETHZ) to isolate single GFP and mCherry double-positive cells, or in the case of Lbr, GFP-positive cells. These were plated in 96 gelatin-coated wells and grown to colonies. Clones were characterized by PCR using primer pairs (PPs) flanking the sgRNAs designed with Primer-BLAST104, immunoblotting and by sequencing across the gene deletion site. The following PPs flanking the sgRNAs binding sites were used: (1) Lbr KO: PP1-F 5′-AGTTGTCCAGTCTGATGCCTG-3′, PP1-R 5′-TGGGCCTATACCTCACTCCC-3′; PP2-F 5′-ACAGTGTTTTTGCTACTGTTTAGC-3′, PP2-R 5′-ACAAAGCTTGCCAAACATCAAAT-3′; (2) Tmpo KO: PP1-F 5′-TATCAGCGTCCGAGGGGAA-3′, PP1-R 5′-TGCAGGTAGAGCTGCACGTA-3′, PP2-F 5′-GTGATACGTCAATCCCCTGCT-3′, PP2-R 5′-TGAGATCCGTGTAAGCATCCTG-3′; (3) Lmna KO, PP1-F 5′-AAGGGGTGTTCTGCAGAGTG-3′, PP1-R 5′-AGAATCAAGGGTCTTGCGGG-3′, PP2-F 5′-CTTGCAGTCCCTCTTGCATC-3′, PP2-R 5′-AATCGCCGCACCTCTAGACT-3′. The E14TG2a Lbr KO cell line was used to generate the Lbr/Tmpo and Lbr/Lmna DKO cell lines. The Lbr/Lmna DKO was used as a background for generation of the Lbr/Tmpo/Lmna TKO cell line.
Embryoid body differentiation assay
Embryoid body (EB) differentiation assays were performed as described in ref. 105. In brief, at day zero, 1 × 106 cells were seeded in 10-cm Petri dishes using media lacking LIF and 2i and carefully spread evenly across the plate by swirling. On day three, the EBs were plated onto two 10-cm dishes. At day six, EBs from each dish were transferred into gelatin-coated T75 flasks and grown until day ten. At days zero and ten of differentiation, the cells were analysed by phase contrast microscopy with a Nikon Ti-E with an air objective Plan Fluor ×10 NA 0.30 Ph1 DL WD 15.2 mm objective. Images were acquired with Nikon NIS-Elements software, and samples were taken for immunoblotting and RNA extraction.
Immunoblot analysis
Whole-cell extracts were generated by taking up cells in sodium dodecyl sulfate (SDS) sample buffer (75 mM Tris pH 7.8, 20% (vol/vol) glycerol, 4% SDS, 50 mM dithiothreitol (DTT) and 0.1% bromophenol blue) and separated by SDS polyacrylamide gel electrophoresis (SDS–PAGE). Proteins were transferred onto a nitrocellulose membrane (Amersham Protran, 10600015) by semi-dry blotting. The membranes were incubated three times in blocking solution (5% (wt/vol) milk powder, 0.1% Tween in 1× phosphate buffered saline (PBS)) for 5 min. The membrane was incubated with the primary antibody diluted in blocking solution at 4 °C overnight, washed with blocking solution three times for 5 min, and then exposed to fluorescently labelled secondary antibodies diluted in blocking solution for 35 min at r.t. Finally, the membrane was washed three times with 0.1% Tween in 1× PBS for 5 min. Signals were detected using an Odyssey (LI-COR) imaging system.
Immunofluorescence
For immunofluorescence, cells on coverslips were fixed with 2% paraformaldehyde (Sigma-Aldrich) in PBS for 10 min at r.t. The cells were washed twice for 3 min using 1× PBS, permeabilized with 0.2% Triton-X100 (Sigma) or 0.2% Triton-X100 and 0.02% SDS in PBS for 12 min at r.t., then washed three times with 1× PBS for 5 min. Blocking was performed in blocking solution (2% bovine serum albumin, Pan-Biotech) and 0.1% Tween in 1× PBS) for at least 45 min at r.t. Primary antibodies were incubated overnight at 4 °C in blocking solution at different concentrations (anti-H3K9me3 1:500 (rb)/1:200 (ms), anti-H3K27me3 1:500, anti-H4K20me3 1:500, anti-RRP12 1:2,000, anti-LAP2 1:2,000, anti-LBR 1:1,000, anti-lamin A/C 1:1,000, anti-CREST 1:1,000, anti-CLIMP-63 1:200). The following day, the coverslips were washed three times for 3 min using washing buffer (0.1% Tween in 1× PBS). Secondary antibodies were diluted 1:300 in blocking solution, added to the coverslips, and incubated for 35 min at r.t. The coverslips were then washed three times using washing buffer, before a final wash with 1× PBS for 3 min. Finally, the cells were stained with 1 µg ml−1 Hoechst (Sigma-Aldrich, 63493) for 10 min at r.t. and washed with 1× PBS. The coverslips were mounted onto glass slides using 3 µl of VECTASHIELD (Vector Laboratories) and sealed with nail polish. The samples were imaged with a Zeiss LSM 880 or a Zeiss LSM 780 confocal laser scanning microscope, using a ×63, 1.4-NA oil plan-apochromat objective. Details on microscopy are summarized in Supplementary Table 5.
Quantification of fluorescently labelled chromatin at the nuclear periphery
To quantify the fraction of antibody-labelled heterochromatin at the nuclear periphery in differentiated cells, the cells were subjected to immunofluorescence with the indicated antibodies. The images were first split into separate channels. Segmentation of cell nuclei was performed using the Hoechst channel with the nucleAIzer model106. To compute the nuclear rim, mathematical morphology was used to erode the mask image with a rectangular structuring element. The size of the structuring element was set to equal 8 pixels of erosion (~0.42 µm). The difference between the nuclear mask and its eroded version determined the pixels that belong to the nuclear rim. If a few pixels of random shift were noted in the images, this was fixed by calculating the phase cross-correlation between the channels and transforming the heterochromatin channel. The rim mask was then applied to the channel used for heterochromatin labelling and the underlying pixel intensities were summed and divided by the area of the rim mask. The same was done with the entire nucleus, and the average nuclear intensity was determined. The ratio of the average rim and nuclear intensities was used for further statistics. The segmented images were individually inspected, and cells for which the segmentation was poor or faulty, for instance two cells being counted as one, were excluded. The data were further analysed using GraphPad Prism 10 (GraphPad), differences between samples were tested for significance (Welch’s t-test or one-way ANOVA) and graphs plotted.
To quantify the fraction of heterochromatin at the nuclear periphery of ES-E14TG2a cells, the above method did not produce robust results due to the variable size and presence of chromocentres at the nuclear periphery. Therefore, for these cells, quantification of the signal intensity of histone marks at the nuclear periphery was performed using Fiji ImageJ107. For each cell nucleus, ten line scans of 2 μm distributed along the entire nuclear perimeter were taken, each extending 1 μm into the cytoplasm and 1 μm into the nuclear interior. The intensity of each point of the line scan was normalized against the total nuclear signal and plotted using GraphPad. An ordinary one-way ANOVA was performed on normalized values corresponding to the peak signal at the NE using GraphPad Prism 10.
Quantification of centromere positioning
To measure the fraction of centromeres at the nuclear periphery, fixed HCT116 cells were subjected to immunofluorescence using anti-CREST and anti-CLIMP-63 antibodies, stained with Hoechst, and images were taken on a Zeiss LSM 880 confocal laser scanning microscope, using a ×63, 1.4-NA oil plan-apochromat objective, taking 33 z-stacks of 0.39-µm step size. These images were processed in Fiji, and a composite stack of images containing Hoechst, a nuclear envelope marker (CLIMP-63), and centromeres (CREST) was generated. Using the measure function of Fiji, the number of centromeres positioned within a 0.5-µm-wide ring at the nuclear periphery was determined in 2D throughout stacked z-planes, thereby examining the entire 3D nuclear space. Each centromere focus was only counted once, in the plane where it was brightest. The distance of 0.5 µm was considered as within the range of a LAD based on previous work16,60. For the distance measurements, the measuring pointer was placed at the centre of each focus, and the shortest distance to the nuclear periphery was determined. If the centromere foci were too close to each other, not allowing resolution of single centromeres, these foci were counted as one centromere. After the peripheral centromeres had been counted and measured, the number of internal centromeres was determined to calculate the fraction of peripheral centromeres over all centromeres. GraphPad Prism 10 was used to perform statistical tests and plot graphs.
Total RNA extraction and RNA-seq
To prepare samples for RNA-seq, three individual batches of cells were plated on the same day and further processed in parallel. Freshly collected cells were lysed with 500 µl of QIAzol lysis reagent (Qiagen #79306) and mixed by gentle pipetting. Subsequently, 100 µl of chloroform (Sigma) was added, and the samples were vortexed for 10 s and spun at 12,000g for 15 s at 4 °C. The upper phase was transferred into a new tube, 600 μl of isopropanol (Sigma) added, and the samples were incubated overnight at −20 °C. After centrifugation at 16,000g for 10 min at 4 °C, the RNA pellet was washed in 75% ethanol and resuspended in RNase-free water (Invitrogen). DNase digestion of RNA was performed using the RNase-Free DNase Set from Qiagen, following the manufacturer’s instructions. RNA was then precipitated by adding sodium acetate (Sigma) to a final concentration of 0.3 M, and adding three volumes of 100% ethanol overnight at −20 °C. RNA was recovered by centrifugation at 16,000g for 10 min and washed twice with 70% ethanol. The RNA was dissolved in RNase-free water (Invitrogen).
RNA-seq was performed at the Functional Genomic Center Zurich. The quality of the isolated RNA was determined using a fragment analyser (Agilent). Only samples with a 260 nm/280 nm ratio between 1.8–2.1 and a 28S/18S ratio within 1.5–2 were further used for library preparation. An Illumina Ribo Zero Plus system (Illumina) was used in the succeeding steps. Briefly, total RNA samples (25–1,000 ng) were depleted of ribosomal RNA and then reverse-transcribed into double-stranded complementary DNA (cDNA). The cDNA samples were fragmented, end-repaired, and adenylated before ligation of an anchor. Fragments containing the anchor on both ends were selectively enriched by PCR at the same time adding the index with unique dual indices. The quality and quantity of the enriched libraries were validated using the fragment analyser (Agilent). The libraries were normalized to 10 nM in Tris-HCl 10 mM, pH 8.5 with 0.1% Tween20. The Novaseq X (Illumina) was used for cluster generation and sequencing according to a standard protocol. Sequencing experiments were paired-end at 2 × 150 bp.
ATAC-seq
To prepare samples, three individual batches of cells were plated on the same day in six wells (for si-11) or 10-cm plates (for KO cells). Cell batches were then further processed in parallel, during all subsequent steps. For collection of cells, the cells were washed twice with 1× PBS and detached using 0.5 mM EDTA in 1× PBS. The cells were counted and 80,000 cells were pelleted. Pellets were resuspended in 200 µl of freezing medium (10% DMSO in FCS) and slowly frozen at −80 °C. ATAC-seq was performed at QuickBiology (Monrovia, California). The cells were washed and lysed to generate a crude preparation of nuclei. This crude extract was then treated with Tn5 transposase. Tn5 transposase tagmentation simultaneously fragments the genome and tags the resulting DNA with Illumina sequencing adapters. DNA fragments were PCR-amplified and subsequently purified using the MinElute PCR purification kit (Qiagen). The ATAC-seq library was prepared as described in the protocol in ref. 108. After quality control using an Agilent TapeStation 4200 (Agilent Technologies) and Life Technologies Qubit 3.0 Fluorometer (Life Technologies), the libraries were sequenced (150-bp paired-end reads) on an Illumina HiSeq sequencer.
ChIP
ChIP was carried out as previously described109,110,111. In brief, 1% formaldehyde in DMEM was added to HCT116 WT, DKO or TKO cells for 10 min, then quenched with 0.1 M glycine. Isolated nuclei were lysed and sonicated using a Bioruptor Pico ultrasonic cell disruptor (Diagenode) to shear genomic DNA to an average fragment size of 200 bp. Chromatin (20 μg) was diluted to a total volume of 500 μl with ChIP buffer (16.7 mM Tris pH 8.1, 167 mM NaCl, 1.2 mM EDTA, 0.01% SDS, 1.1% Triton X100) and pre-cleared with 10 μl of packed sepharose beads for 2 h at 4 °C (ref. 111). Pre-cleared chromatin was incubated with ChIP-grade antibodies against H3K9me3 or H3K27me3 overnight, then 20 ng of Drosophila melanogaster S2 chromatin and 0.5 μg of H2Av antibody were added as spike-in to each sample. Dynabeads Protein A (Invitrogen, 10002D) or G (Invitrogen, 10004D) was added111. After 4 h at 4 °C, beads were pelleted and washed, and bound chromatin was eluted with 1% SDS and 100 mM NaHCO3. After digestion with Proteinase K (Thermo Scientific) at 50 °C for 3 h and reversion of crosslinking (65 °C, overnight), DNA was purified with phenol/chloroform, ethanol-precipitated and quantified by quantitative PCR. For ChIP-seq analyses, the quantity of the isolated DNA was determined with a Qubit 4 fluorometer (Life Technologies). Libraries were prepared using NEBNext Ultra II DNA Library Prep for Illumina (New England Biolabs, E7645S and E7645L), following the manufacturer’s protocol. Briefly, ChIP and input samples (15 ng) were end-repaired and polyadenylated. The adapters containing the index for multiplexing were then ligated. The quality and quantity of the enriched libraries were validated using a 4200 TapeStation system (Agilent). Sequencing was performed on an Illumina NovaSeq6000 machine with single-end 100-bp reads.
Micro-C (Pico-C)
Pico-C maps were obtained as described in refs. 54,112,113. Briefly, HCT116 cells were fixed, and digestion tests were carried out to determine the optimal amount of MNase. MNase reactions were stopped by the addition of ethylene glycol tetraacetic acid (EGTA), and the samples were washed with MB2 (50 mM NaCl, 10 mM Tris-HCl pH 7.5 and 10 mM MgCl2). After this, the supernatant was removed, and 112.5 µl of extraction buffer (50 mM Tris-HCl pH 8, 50 mM NaCl, 1 mM EDTA, 1% SDS) and 12.5 µl of 2 mg ml−1 Proteinase K were added. The samples were maintained at 65 °C overnight to reverse crosslinking. The following day, phenol:chloroform:isoamylalcohol DNA extraction was carried out followed by RNA removal via RNase A treatment. DNA concentrations were quantified using Qubit and run on a 3.5% Tris, acetate and EDTA (TAE) gel or Bioanalyzer run. MNase concentrations that yielded ~60–70% mono-nucleosomal and 40–30% di-nucleosomal DNA were used in subsequent experiments. Once the optimal MNase concentration was determined, libraries were generated using ~2 million cells per individual library, as described in ref. 54.
Analysis of multi-omics data
Reference genomes and annotations
The reference genomes for human (GRCh38; annotation version 108) and mouse (GRCm39; annotation version 112) were obtained from ENSEMBL114.
Bulk RNA-seq
The paired-end reads from bulk RNA-seq datasets were processed to remove the Illumina universal adapter (AGATCGGAAGAGCACACGTCTGAACTCCAGTCA) using Cutadapt (v4.4) with the parameters ‘-q 25 -m 25’115. The resulting filtered reads were aligned to their respective reference genome using STAR aligner (v2.7.10b) with default parameters116. Coordinate-sorted alignment files were indexed using SAMtools (v1.17)117. Exon mapped reads were counted using featureCounts from the subread package (v2.0.6)118. Subsequent data analysis was performed with custom scripts in R (R Core Team, 2024). Data wrangling and visualization were performed using the tidyverse package (v2.0.0)119. Differential expression analysis was conducted using DESeq2 (v1.44)120. Differentially expressed genes were defined as having an absolute log(fold change) > 1 and adjusted P < 0.05. Differentially expressed genes were further subjected to GO analysis using TopGO (Bioconductor.org)121.
To evaluate the expression of TEs, the filtered reads from above were aligned to their respective reference genome using STAR aligner (v2.7.10b) with the parameters ‘--winAnchorMultimapNmax 200 --outFilterMultimapNmax 100’. The resulting alignment files were used to quantify the read counts for genes and TEs using TETranscripts (v2.2.3)122. Differential expression analysis was performed using DESeq2 (v1.44) and differentially expressed TEs were defined as having an absolute log(fold change) > 0.585 and adjusted P < 0.05.
ATAC-seq
ATAC-seq datasets were processed to remove the Nextera adapter (CTGTCTCTTATACACATCT) using Cutadapt (v4.4) with the parameters ‘-q 25 -m 25’115. The resulting filtered reads were aligned to the reference human genome using bowtie2 aligner (v2.5.1) with the parameter ‘--very-sensitive’ (Bowtie2123). The alignment file was sorted by coordinate, indexed, and filtered for mitochondrial reads using SAMtools (v1.17). The filtered alignment file was further processed to keep only uniquely mapped, non-duplicated reads using sambamba (v1.0.1)124. Reads were further processed using alignmentSieve from deepTools (v3.5.5) to account for transposon dimerization before insertion. Positive and negative strand alignments were shifted by +4 bp and −5 bp, respectively. Additionally, fragments of 100 bp, corresponding to nucleosome-free regions, were retained with the parameters ‘--minMappingQuality 25 --maxFragmentLength 100 --ATACshift --BED’125. Peaks were called on the resulting BEDPE files using MACS3 (v3.0.1)126 with the parameters ‘--call-summits --bdg --nomodel --nolambda --gsize hs --keep-dup all --format BEDPE --qvalue 0.05’. The predicted peaks were filtered for ENCODE project blacklisted regions using BEDTools (v2.31.0) intersect (ENCODE blacklist127; BEDTools128).
Peaks from the three replicates of a condition were intersected using BEDTools (v2.31.0) with the parameter ‘-f 0.50 -r’128. Peaks detected in at least two of three replicates of a condition were considered consensus regions corresponding to open chromatin. Differentially accessible regions were identified using DESeq2 (v1.44) and annotated using the Chipseeker package (v1.40)129.
To compare ATAC-seq with RNA-seq data, reads within peaks that overlapped with protein-coding genes and within 3,000 bp upstream and 500 bp downstream of the TSS were summed to obtain reads within promoters. Spearman correlation was calculated using the ggpubr R package (https://rpkgs.datanovia.com/ggpubr/).
ChIP-seq
ChIP-seq datasets were processed to remove the Illumina adapters using Cutadapt (v4.4) with the parameters ‘--poly-a -q 25 -m 25 -a AGATCGGAAGAGCACACGTCTGAACTCCAGTCA -A AGATCGGAAGAGCGTCGTGTAGGGAAAGAGTGT’115. The resulting filtered reads were aligned to the reference human genome using bowtie2 aligner (v2.5.1) with the parameter ‘--very-sensitive’ (Bowtie2123). The alignment file was sorted by coordinate, indexed, and filtered for mitochondrial reads using SAMtools (v1.17). The filtered alignment file was further processed to keep only uniquely mapped, non-duplicated reads using sambamba (v1.0.1)124.
For differential H3K27me3 binding analysis, the alignment files were further filtered for ENCODE project blacklisted regions using BEDTools (v2.31.0) intersect (ENCODE blacklist127; BEDTools128). The resulting alignment files were analysed using the csaw R package (v1.42)130. Binary alignment map (BAM) files were read using windowCounts with the parameters ‘width = 2000, spacing = 500, ext = 150’. The global background width was set to 10,000 bp, and only windows with more than two-fold enrichment over the background level were considered for analysis. After normalization for compositional bias and statistical modelling, windows between KO and control samples were compared using glmQLFTest. Windows fewer than 1,000 bp apart were consolidated and subjected to an improved Bonferroni procedure to compute combined P values131. Regions with |log2-fold change| ≥ 1 and adjusted P < 0.05 were considered differentially affected.
To compare ChIP-seq with RNA-seq data, H3K27me3 windows from above were annotated using the Chipseeker package (v1.40)129. The windows that belonged to protein-coding genes and within 3,000 bp upstream to 500 bp downstream of TSS were considered for further analysis. For genes with isoforms, the ensemble canonical isoform was considered. If an ensemble canonical isoform was not present, then the transcript with H3K27me3 signal closest to the TSS was considered. If two transcripts of a gene had an H3K27me3 signal equidistant from the TSS, then the transcript with the longest length was chosen as a representative of that gene. Spearman correlation was calculated using the ggpubr R package (https://rpkgs.datanovia.com/ggpubr/).
For visualization, the filtered BAM files from above were converted into normalized bigWig signal tracks using the bamCoverage module of the deepTools package (v3.5.5)125. The parameters used were ‘--effectiveGenomeSize 2913022398 --binSize 1 --normalizeUsing RPGC --extendReads 150 --outFileFormat bigwig’. For each condition, the signal mean was calculated using the bigwigAverage module.
Peak calling was performed using SICER2 with the parameters ‘-w 1000 -f 150 -g 2000 -fdr 0.001’ for H3K27me3 and ‘-w 20000 -f 150 -g 40000 -fdr 0.001’ for H3K9me3. Regions from the three replicates of a condition were intersected using BEDTools (v 2.28.0) using default parameters. Regions detected in all three replicates of a condition were considered for further analysis.
Heatmaps and metagene profiles were created using the computeMatrix and plotHeatmap modules from the deepTools package (v 3.5.6). The parameters for computeMatrix were ‘--referencePoint center -b 10000 -a 10000 -bs 50 --sortRegions descend --smartLabels --missingDataAsZero -p max’, and for the heatmaps were ‘--sortRegions keep --zMax 7.0 --yAxisLabel Mean coverage --xAxisLabel distance from center --averageTypeSummaryPlot mean --refPointLabel center --legendLocation none’.
Pico-C data analysis
Briefly, Pico-C reads were mapped to the hg38 genome using BWA-MEM (v0.7.17). Aligned reads were then processed with Pairtools (v1.0.3132) using the parse2 function to rescue complex walks. The resulting pair files were sorted and converted to the FAN-C pairs format (v0.9.18133). Duplicates were removed, and quality control metrics were extracted. We then generated FAN-C .hic files. Principal component analysis was performed on these merged.hic files using FAN-C, with a sample size of 100,000 interactions, focusing on distances between 100 kb and 1 Mb to assess variation and clustering across the samples. Because biological replicates clustered well across all stages, the .hic files of replicates were merged. The merged files were binned at different resolutions and normalized using iterative correction. We then used a custom Python script to determine the bin size where 80% of bins had over 1,000 reads, as done previously15, to get an idea of the optimal resolution of each stage. Finally, the merged FAN-C .hic files were converted to balanced cooler (v0.8.6134) .mcool files using cooler zoomify.
Compartment analysis
Compartment analysis was carried out using Calder2135 with 20-kb bin size, which displayed good correlation (r >0.5) with the RNA-seq and ATAC-seq signals. Compartment switches were identified by extracting bins with different compartments between WT and each of the mutant samples using GenomicRanges136. Common switches across the mutant samples were identified using BEDTools128 intersect. These shared switches were kept for subsequent analysis. To quantify compartment enrichment around genomic features, we used a custom R function that normalizes compartment annotations using GenomicRanges (GRanges objects) to a binned matrix centred on target regions (±1 Mb by default). Each bin was annotated based on compartment state (A.1, A.2, B.1, B.2) and used to calculate the fraction of each compartment across all bins. Input regions were imported from BED files and converted to GRanges, and compartment annotations were simplified for consistent labelling. The resulting data frame includes per-bin percentages for downstream visualization. GGplot2 was then used for plotting137.
Statistics and reproducibility
No statistical method was used to predetermine sample size. The experiments were not randomized, and the investigators were not blinded to allocation during experiments and outcome assessment. Information on the statistical tests used for each analysis and reproducibility is included in the relevant sections, and described in the methods and figure legends, and relevant code provided in the GitHub repository138.
Reporting Summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this Article.
Data availability
Sequencing data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE281919. Data supporting the findings of this study are available from the corresponding author on reasonable request. Source data are provided with this paper.
Code availability
All relevant code used for the analysis of high-throughput sequencing data is available at GitHub (https://github.com/ps-puneetsharma/lewis_et_al_2025).
References
Heitz, E. Das Heterochromatin der Moose. I. Jahr. Wiss. Bot. 69, 762–818 (1928).
Cremer, T. et al. The 4D nucleome: evidence for a dynamic nuclear landscape based on co-aligned active and inactive nuclear compartments. FEBS Lett. 589, 2931–2943 (2015).
Joffe, B., Leonhardt, H. & Solovei, I. Differentiation and large scale spatial organization of the genome. Curr. Opin. Genet. Dev. 20, 562–569 (2010).
Solovei, I., Thanisch, K. & Feodorova, Y. How to rule the nucleus: divide et impera. Curr. Opin. Cell Biol. 40, 47–59 (2016).
van Koningsbruggen, S. et al. High-resolution whole-genome sequencing reveals that specific chromatin domains from most human chromosomes associate with nucleoli. Mol. Biol. Cell 21, 3735–3748 (2010).
Harr, J. C., Gonzalez-Sandoval, A. & Gasser, S. M. Histones and histone modifications in perinuclear chromatin anchoring: from yeast to man. EMBO Rep. 17, 139–155 (2016).
Buchwalter, A., Kaneshiro, J. M. & Hetzer, M. W. Coaching from the sidelines: the nuclear periphery in genome regulation. Nat. Rev. Genet. 20, 39–50 (2019).
Solovei, I. et al. LBR and lamin A/C sequentially tether peripheral heterochromatin and inversely regulate differentiation. Cell 152, 584–598 (2013).
Falk, M. et al. Heterochromatin drives compartmentalization of inverted and conventional nuclei. Nature 570, 395–399 (2019).
Gruenbaum, Y., Margalit, A., Goldman, R. D., Shumaker, D. K. & Wilson, K. L. The nuclear lamina comes of age. Nat. Rev. Mol. Cell Biol. 6, 21–31 (2005).
Burke, B. & Stewart, C. L. The nuclear lamins: flexibility in function. Nat. Rev. Mol. Cell Biol. 14, 13–24 (2013).
Wong, X., Melendez-Perez, A. J. & Reddy, K. L. The nuclear lamina. Cold Spring Harb. Perspect. Biol. 14, a040113 (2022).
Harr, J. C. et al. Directed targeting of chromatin to the nuclear lamina is mediated by chromatin state and A-type lamins. J. Cell Biol. 208, 33–52 (2015).
van Steensel, B. & Belmont, A. S. Lamina-associated domains: links with chromosome architecture, heterochromatin, and gene repression. Cell 169, 780–791 (2017).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Kind, J. et al. Single-cell dynamics of genome–nuclear lamina interactions. Cell 153, 178–192 (2013).
Bersaglieri, C. et al. Genome-wide maps of nucleolus interactions reveal distinct layers of repressive chromatin domains. Nat. Commun. 13, 1483 (2022).
Bizhanova, A., Yan, A., Yu, J., Zhu, L. J. & Kaufman, P. D. Distinct features of nucleolus-associated domains in mouse embryonic stem cells. Chromosoma 129, 121–139 (2020).
van Schaik, T., Vos, M., Peric-Hupkes, D., Hn Celie, P. & van Steensel, B. Cell cycle dynamics of lamina-associated DNA. EMBO Rep. 21, e50636 (2020).
Worman, H. J., Yuan, J., Blobel, G. & Georgatos, S. D. A lamin B receptor in the nuclear envelope. Proc. Natl Acad. Sci. USA 85, 8531–8534 (1988).
Cabianca, D. S. & Gasser, S. M. Spatial segregation of heterochromatin: uncovering functionality in a multicellular organism. Nucleus 7, 301–307 (2016).
Lemaitre, C. & Bickmore, W. A. Chromatin at the nuclear periphery and the regulation of genome functions. Histochem. Cell Biol. 144, 111–122 (2015).
Briand, N. & Collas, P. Lamina-associated domains: peripheral matters and internal affairs. Genome Biol. 21, 85 (2020).
Mattout, A., Cabianca, D. S. & Gasser, S. M. Chromatin states and nuclear organization in development—a view from the nuclear lamina. Genome Biol. 16, 174 (2015).
Reddy, K. L., Zullo, J. M., Bertolino, E. & Singh, H. Transcriptional repression mediated by repositioning of genes to the nuclear lamina. Nature 452, 243–247 (2008).
Finlan, L. E. et al. Recruitment to the nuclear periphery can alter expression of genes in human cells. PLoS Genet. 4, e1000039 (2008).
Akhtar, W. et al. Chromatin position effects assayed by thousands of reporters integrated in parallel. Cell 154, 914–927 (2013).
Leemans, C. et al. Promoter-intrinsic and local chromatin features determine gene repression in LADs. Cell 177, 852–864.e14 (2019).
Guelen, L. et al. Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature 453, 948–951 (2008).
Kumaran, R. I. & Spector, D. L. A genetic locus targeted to the nuclear periphery in living cells maintains its transcriptional competence. J. Cell Biol. 180, 51–65 (2008).
Wu, F. & Yao, J. Identifying novel transcriptional and epigenetic features of nuclear lamina-associated genes. Sci. Rep. 7, 100 (2017).
Brueckner, L. et al. Local rewiring of genome-nuclear lamina interactions by transcription. EMBO J. 39, e103159 (2020).
Therizols, P. et al. Chromatin decondensation is sufficient to alter nuclear organization in embryonic stem cells. Science 346, 1238–1242 (2014).
Cabianca, D. S. et al. Active chromatin marks drive spatial sequestration of heterochromatin in C. elegans nuclei. Nature 569, 734–739 (2019).
Hannus, M. et al. siPools: highly complex but accurately defined siRNA pools eliminate off-target effects. Nucleic Acids Res. 42, 8049–8061 (2014).
Padeken, J. & Heun, P. Nucleolus and nuclear periphery: velcro for heterochromatin. Curr. Opin. Cell Biol. 28, 54–60 (2014).
Martens, J. H. et al. The profile of repeat-associated histone lysine methylation states in the mouse epigenome. EMBO J. 24, 800–812 (2005).
Weierich, C. et al. Three-dimensional arrangements of centromeres and telomeres in nuclei of human and murine lymphocytes. Chromosome Res. 11, 485–502 (2003).
Solovei, I. et al. Differences in centromere positioning of cycling and postmitotic human cell types. Chromosoma 112, 410–423 (2004).
Keller, D. et al. Non-random spatial organization of telomeres varies during the cell cycle and requires LAP2 and BAF. iScience 27, 109343 (2024).
Dechat, T., Vlcek, S. & Foisner, R. Review: Lamina-associated polypeptide 2 isoforms and related proteins in cell cycle-dependent nuclear structure dynamics. J. Struct. Biol. 129, 335–345 (2000).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21.29.1–21.29.9 (2015).
Rehwinkel, J. & Gack, M. U. RIG-I-like receptors: their regulation and roles in RNA sensing. Nat. Rev. Immunol. 20, 537–551 (2020).
Schwartz, S. L. & Conn, G. L. RNA regulation of the antiviral protein 2′-5′-oligoadenylate synthetase. Wiley Interdiscip. Rev. RNA 10, e1534 (2019).
Bishop, K. N. et al. Cytidine deamination of retroviral DNA by diverse APOBEC proteins. Curr. Biol. 14, 1392–1396 (2004).
Chiappinelli, K. B. et al. Inhibiting DNA methylation causes an interferon response in cancer via dsRNA including endogenous retroviruses. Cell 162, 974–986 (2015).
Chen, Y. G. & Hur, S. Cellular origins of dsRNA, their recognition and consequences. Nat. Rev. Mol. Cell Biol. 23, 286–301 (2022).
Gazquez-Gutierrez, A., Witteveldt, J., Heras, S. R. & Macias, S. Sensing of transposable elements by the antiviral innate immune system. RNA 27, 735–752 (2021).
Bowling, E. A. et al. Spliceosome-targeted therapies trigger an antiviral immune response in triple-negative breast cancer. Cell 184, 384–403.e321 (2021).
Crossley, M. P. et al. R-loop-derived cytoplasmic RNA–DNA hybrids activate an immune response. Nature 613, 187–194 (2023).
Morey, L. et al. Nonoverlapping functions of the Polycomb group Cbx family of proteins in embryonic stem cells. Cell Stem Cell 10, 47–62 (2012).
Slobodyanyuk, E., Cattoglio, C. & Hsieh, T. S. Mapping mammalian 3D genomes by Micro-C. Methods Mol. Biol. 2532, 51–71 (2022).
Jerkovic, I. & Cavalli, G. Understanding 3D genome organization by multidisciplinary methods. Nat. Rev. Mol. Cell Biol. 22, 511–528 (2021).
Maziak, N. et al. Three-dimensional genome reorganization foreshadows zygotic genome activation in Drosophila. Nat. Genet. https://doi.org/10.1038/s41588-026-02503-3 (2026).
Borsos, M. et al. Genome-lamina interactions are established de novo in the early mouse embryo. Nature 569, 729–733 (2019).
Padeken, J., Methot, S. P. & Gasser, S. M. Establishment of H3K9-methylated heterochromatin and its functions in tissue differentiation and maintenance. Nat. Rev. Mol. Cell Biol. 23, 623–640 (2022).
Madsen-Osterbye, J., Abdelhalim, M., Baudement, M. O. & Collas, P. Local euchromatin enrichment in lamina-associated domains anticipates their repositioning in the adipogenic lineage. Genome Biol. 23, 91 (2022).
Robson, M. I. et al. Constrained release of lamina-associated enhancers and genes from the nuclear envelope during T-cell activation facilitates their association in chromosome compartments. Genome Res. 27, 1126–1138 (2017).
Shah, P. P. et al. An atlas of lamina-associated chromatin across twelve human cell types reveals an intermediate chromatin subtype. Genome Biol. 24, 16 (2023).
Kind, J. et al. Genome-wide maps of nuclear lamina interactions in single human cells. Cell 163, 134–147 (2015).
Zheng, X. et al. Lamins organize the global three-dimensional genome from the nuclear periphery. Mol. Cell 71, 802–815 e807 (2018).
Silva, J. et al. Promotion of reprogramming to ground state pluripotency by signal inhibition. PLoS Biol. 6, e253 (2008).
Li, Y., Zhang, Y. & Liu, M. Knockout gene-based evidence for PIWI-interacting RNA pathway in mammals. Front. Cell Dev. Biol. 9, 681188 (2021).
Yu, T. et al. The piRNA response to retroviral invasion of the koala genome. Cell 179, 632–643.e612 (2019).
Ramakrishna, N. B., Battistoni, G., Surani, M. A., Hannon, G. J. & Miska, E. A. Mouse primordial germ-cell-like cells lack piRNAs. Dev. Cell 57, 2661–2668.e2665 (2022).
Argelaguet, R. et al. Multi-omics profiling of mouse gastrulation at single-cell resolution. Nature 576, 487–491 (2019).
Feodorova, Y., Falk, M., Mirny, L. A. & Solovei, I. Viewing nuclear architecture through the eyes of nocturnal mammals. Trends Cell Biol. 30, 276–289 (2020).
Bersaglieri, C. & Santoro, R. Genome organization in and around the nucleolus. Cells 8, 579 (2019).
Cerqueira, A. V. & Lemos, B. Ribosomal DNA and the nucleolus as keystones of nuclear architecture, organization and function. Trends Genet. 35, 710–723 (2019).
Brandle, F., Fruhbauer, B. & Jagannathan, M. Principles and functions of pericentromeric satellite DNA clustering into chromocenters. Semin. Cell Dev. Biol. 128, 26–39 (2022).
Foisner, R. & Gerace, L. Integral membrane proteins of the nuclear envelope interact with lamins and chromosomes, and binding is modulated by mitotic phosphorylation. Cell 73, 1267–1279 (1993).
Ye, Q., Callebaut, I., Pezhman, A., Courvalin, J. C. & Worman, H. J. Domain-specific interactions of human HP1-type chromodomain proteins and inner nuclear membrane protein LBR. J. Biol. Chem. 272, 14983–14989 (1997).
Jenuwein, T. & Allis, C. D. Translating the histone code. Science 293, 1074–1080 (2001).
Guarda, A., Bolognese, F., Bonapace, I. M. & Badaracco, G. Interaction between the inner nuclear membrane lamin B receptor and the heterochromatic methyl binding protein, MeCP2. Exp. Cell. Res. 315, 1895–1903 (2009).
Hirano, Y. et al. Lamin B receptor recognizes specific modifications of histone H4 in heterochromatin formation. J. Biol. Chem. 287, 42654–42663 (2012).
Filipczak, D., Souchet, A., Georgiou, K., Foisner, R. & Naetar, N. Lamin chromatin binding is modulated by interactions of different LAP2α domains with lamins and chromatin. iScience 27, 110869 (2024).
Cai, M. et al. Solution structure of the constant region of nuclear envelope protein LAP2 reveals two LEM-domain structures: one binds BAF and the other binds DNA. EMBO J. 20, 4399–4407 (2001).
Zheng, R. et al. Barrier-to-autointegration factor (BAF) bridges DNA in a discrete, higher-order nucleoprotein complex. Proc. Natl Acad. Sci. USA 97, 8997–9002 (2000).
Somech, R. et al. The nuclear-envelope protein and transcriptional repressor LAP2β interacts with HDAC3 at the nuclear periphery, and induces histone H4 deacetylation. J. Cell Sci. 118, 4017–4025 (2005).
Zullo, J. M. DNA sequence-dependent compartmentalization and silencing of chromatin at the nuclear lamina. Cell 149, 1474–1487 (2012).
Chetverina, D., Erokhin, M. & Schedl, P. GAGA factor: a multifunctional pioneering chromatin protein. Cell. Mol. Life Sci. 78, 4125–4141 (2021).
Fong, L. G. et al. Prelamin A and lamin A appear to be dispensable in the nuclear lamina. J. Clin. Invest. 116, 743–752 (2006).
Poleshko, A. et al. The human protein PRR14 tethers heterochromatin to the nuclear lamina during interphase and mitotic exit. Cell Rep. 5, 292–301 (2013).
Kiseleva, A. A., Cheng, Y. C., Smith, C. L., Katz, R. A. & Poleshko, A. PRR14 organizes H3K9me3-modified heterochromatin at the nuclear lamina. Nucleus 14, 2165602 (2023).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Liu, C. et al. Histone H1 facilitates restoration of H3K27me3 during DNA replication by chromatin compaction. Nat. Commun. 14, 4081 (2023).
Veronezi, G. M. B. & Ramachandran, S. Nucleation and spreading maintain Polycomb domains every cell cycle. Cell Rep. 43, 114090 (2024).
Deleris, A., Berger, F. & Duharcourt, S. Role of Polycomb in the control of transposable elements. Trends Genet. 37, 882–889 (2021).
Hisanaga, T. et al. The Polycomb repressive complex 2 deposits H3K27me3 and represses transposable elements in a broad range of eukaryotes. Curr. Biol. 33, 4367–4380.e4369 (2023).
Gholamalamdari, O. et al. Major nuclear locales define nuclear genome organization and function beyond A and B compartments. eLife https://doi.org/10.7554/elife.99116 (2024).
Gaspar-Maia, A., Alajem, A., Meshorer, E. & Ramalho-Santos, M. Open chromatin in pluripotency and reprogramming. Nat. Rev. Mol. Cell Biol. 12, 36–47 (2011).
Kurimoto, K. et al. Quantitative dynamics of chromatin remodeling during germ cell specification from mouse embryonic stem cells. Cell Stem Cell 16, 517–532 (2015).
Marin, H. C. et al. The nuclear periphery confers repression on H3K9me2-marked genes and transposons to shape cell fate. Nat. Cell Biol. 27, 1311–1326 (2025).
Kumar, P. et al. Nucleolus and centromere tyramide signal amplification-seq reveals variable localization of heterochromatin in different cell types. Commun. Biol. 7, 1135 (2024).
Spracklin, G. et al. Diverse silent chromatin states modulate genome compartmentalization and loop extrusion barriers. Nat. Struct. Mol. Biol. 30, 38–51 (2023).
Sosa, B. A., Rothballer, A., Kutay, U. & Schwartz, T. U. LINC complexes form by binding of three KASH peptides to domain interfaces of trimeric SUN proteins. Cell 149, 1035–1047 (2012).
Wyler, E. et al. Tandem affinity purification combined with inducible shRNA expression as a tool to study the maturation of macromolecular assemblies. RNA 17, 189–200 (2011).
Poleshko, A. et al. H3K9me2 orchestrates inheritance of spatial positioning of peripheral heterochromatin through mitosis. eLife 8, e49278 (2019).
Ungricht, R., Klann, M., Horvath, P. & Kutay, U. Diffusion and retention are major determinants of protein targeting to the inner nuclear membrane. J. Cell Biol. 209, 687–703 (2015).
Welte, T. et al. The RNA hairpin binder TRIM71 modulates alternative splicing by repressing MBNL1. Genes Dev. 33, 1221–1235 (2019).
Luithle, N. et al. Torsin ATPases influence chromatin interaction of the Torsin regulator LAP1. eLife 9, e63614 (2020).
Brinkman, E. K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168 (2014).
Wettstein, R., Bodak, M. & Ciaudo, C. Generation of a knockout mouse embryonic stem cell line using a paired CRISPR/Cas9 genome engineering tool. Methods Mol. Biol. 1341, 321–343 (2016).
Ye, J. et al. Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinformatics 13, 134 (2012).
Bodak, M. & Ciaudo, C. Monitoring long interspersed nuclear element 1 expression during mouse embryonic stem cell differentiation. Methods Mol. Biol. 1400, 237–259 (2016).
Hollandi, R. et al. nucleAIzer: a parameter-free deep learning framework for nucleus segmentation using image style transfer. Cell Syst. 10, 453–458 e456 (2020).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Leone, S., Bar, D., Slabber, C. F., Dalcher, D. & Santoro, R. The RNA helicase DHX9 establishes nucleolar heterochromatin, and this activity is required for embryonic stem cell differentiation. EMBO Rep. 18, 1248–1262 (2017).
Santoro, R. Analysis of chromatin composition of repetitive sequences: the ChIP-Chop assay. Methods Mol. Biol. 1094, 319–328 (2014).
Lerra, L. et al. An RNA-dependent and phase-separated active subnuclear compartment safeguards repressive chromatin domains. Mol. Cell 84, 1667–1683.e1610 (2024).
Hsieh, T. S., Fudenberg, G., Goloborodko, A. & Rando, O. J. Micro-C XL: assaying chromosome conformation from the nucleosome to the entire genome. Nat. Methods 13, 1009–1011 (2016).
Ing-Simmons, E. et al. Independence of chromatin conformation and gene regulation during Drosophila dorsoventral patterning. Nat. Genet. 53, 487–499 (2021).
Harrison, P. W. et al. Ensembl 2024. Nucleic Acids Res. 52, D891–D899 (2024).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBNet.journal 17, 10–12 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Liao, Y., Smyth, G. K. & Shi, W. The Subread aligner: fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. 41, e108 (2013).
Wickham, H. et al. Welcome to the Tidyverse. J. Open Source Softw. 4, 1686 (2019).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Alexa, A. & Rahnenführer, J. topGO: Enrichment Analysis for Gene Ontology (Bioconductor, 2024).
Jin, Y., Tam, O. H., Paniagua, E. & Hammell, M. TEtranscripts: a package for including transposable elements in differential expression analysis of RNA-seq datasets. Bioinformatics 31, 3593–3599 (2015).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Tarasov, A., Vilella, A. J., Cuppen, E., Nijman, I. J. & Prins, P. Sambamba: fast processing of NGS alignment formats. Bioinformatics 31, 2032–2034 (2015).
Ramirez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Amemiya, H. M., Kundaje, A. & Boyle, A. P. The ENCODE blacklist: identification of problematic regions of the genome. Sci. Rep. 9, 9354 (2019).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Yu, G., Wang, L. G. & He, Q. Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Lun, A. T. & Smyth, G. K. csaw: a Bioconductor package for differential binding analysis of ChIP-seq data using sliding windows. Nucleic Acids Res. 44, e45 (2016).
SIMES, R. J. An improved Bonferroni procedure for multiple tests of significance. Biometrika 73, 751–754 (1986).
Open2C et al. Pairtools: from sequencing data to chromosome contacts. PLoS Comput. Biol. 20, e1012164 (2024).
Kruse, K., Hug, C. B. & Vaquerizas, J. M. FAN-C: a feature-rich framework for the analysis and visualisation of chromosome conformation capture data. Genome Biol. 21, 303 (2020).
Abdennur, N. & Mirny, L. A. Cooler: scalable storage for Hi-C data and other genomically labeled arrays. Bioinformatics 36, 311–316 (2020).
Liu, Y. Systematic inference of multi-scale chromatin sub-compartments using Calder2. Methods Mol. Biol. 2856, 213–221 (2025).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2016).
Sharma, P. Code for LBR and LAP2 mediate heterochromatin tethering to the nuclear periphery to preserve genome homeostasis (v1.0.0). GitHub https://github.com/ps-puneetsharma/lewis_et_al_2025 (2025).
Acknowledgements
We thank M. Jagannathan, T. Kleele, H. Baird and S. Pawar for critical comments on the paper, K. Mahan and other members of the Kutay laboratory for discussions, M. Jagannathan, Z. Smith, J. Veenvliet, C. Bücker, T. Aktas, R. Foisner and A. Wutz for helpful suggestions, and J. uit de Bos for advice on data analysis. Microscopy was performed on instruments of the Microscopy Center Scope M of ETHZ, RNA-seq was supported by C. Aquino and D. Rodrigues at the Functional Genomics Center Zurich, and the processing of genomics data was performed on the Euler high-performance computing cluster at ETH Zürich.
This work was supported by the Swiss National Science Foundation (SNSF) (310030_219203 and TMAG-3_209245 (U.K.)). The collaboration of U.K., R.S. and C.C. was supported by the Swiss NCCR ‘RNA and disease’ (51NF40-205601). P.H. and K.K. acknowledge support from a Lendület BIOMAG grant (2018–342), TKP2021-EGA09, Horizon-BIALYMPH, Horizon-SYMMETRY, Horizon-SWEEPICS, H2020-Fair-CHARM, CZI Deep Visual Proteomics, HAS-NAP3, an ELKH-Excellence grant from OTKA-SNN (139455/ARRS) and the Finnish Cancer Society. R.S. acknowledges support from the ERC (ERC-AdG-787074-NucleolusChromatin) and the Swiss Cancer Research Foundation (KFS-5488-02-2022). J.M.V. acknowledges support from the UK Medical Research Council (MC_UP_160510), the Academy of Medical Sciences and the Department of Business, Energy and Industrial Strategy (APR3\1017).
Funding
Open access funding provided by Swiss Federal Institute of Technology Zurich.
Author information
Authors and Affiliations
Contributions
R.L., V.S. and U.K. conceived the project. R.L., V.S., N.M., C.B., I.Z., J.M.V., R.S., P.S. and U.K. designed the experiments. R.L., V.S., N.M. and C.B. performed experiments. R.L. performed experiments in human cells, V.S. in mESCs, including all respective cell line generation, immunofluorescence, immunoblotting, RNA-seq and ATAC-seq experiments, guided by U.K. N.M. and R.L. performed Pico-C analysis, guided by J.M.V. C.B. and R.L. performed ChIP-seq experiments, guided by R.S. C.C. shared knowhow for work with mESCs. C.A. generated reagents. I.Z. carried out nucleolar stainings. K.K. quantified immunofluorescence on histone marks in HCT116 cells, guided by P.H. P.S. (with input from U.K., R.L. and V.S.) performed bioinformatics analyses of RNA-seq, ATAC-seq and ChIP-seq data, the latter supported by C.B. and R.S. N.M. analysed Micro-C data. R.L., V.S., P.S. and U.K. analysed all processed data. U.K. and R.L. wrote the paper. All authors discussed the results, and reviewed and edited the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks Susana Gonzalo and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Heterochromatin is detached from the nuclear periphery of human and mouse differentiated and pluripotent mouse cells upon depletion of eight INM proteins and the nuclear lamins.
a, Volcano plot of RNA-seq analysis of si-11-treated vs. si-Ctr treated HCT116 cells (N = 3 experiments). Genes with a |log2-fold change| ≥ 1 and an adjusted p-value ≤ 0.05 were considered significant (two-tailed Wald test and Benjamini–Hochberg procedure). b, Representative confocal images of HCT116 cells that were treated either with si-Ctr or si-11 for 72 h, fixed and subjected to immunofluorescence using anti-H3K9me3 and anti-RRP12 antibodies. DNA was stained using Hoechst. Scale bar, 5 μm. c, Representative confocal images of HCT116 cells that were treated either with si-Ctr or si-11 for 72 h, fixed and subjected to immunofluorescence using anti-H3K27me3 and anti-H4K20me3 antibodies. DNA was stained using Hoechst. Scale bars, 5 μm. d, Quantification of the accumulation of H3K27me3 and H4K20me3 at the nuclear rim in HCT116 cells that were either treated with si-Ctr or si-11 (total number of cells: n ≥ 52, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001, Welch’s t-test, mean ± SD, N = 3 experiments). e, Representative confocal images of NIH/3T3 cells and mESCs (E14TG2a) treated either with control siRNAs (si-Ctr) or a combination of 11 siPOOLs (si-11) for 72 h, fixed and subjected to immunofluorescence using an anti-H3K9me3 antibody. DNA was stained using Hoechst. Scale bars, 5 μm. f, Immunoblots confirming the downregulation of select mouse NE proteins. Numerical and statistics source data as well as unprocessed blots are available in source data.
Extended Data Fig. 2 H3K9me3-stained heterochromatin is detached from the nuclear periphery in HCT116 LMNA knockout cells upon the collective depletion of eight INM proteins and the B-type lamins.
a, Characterisation of the HCT116 LMNA KO clone. Immunofluorescence and immunoblotting analyses of lamin A/C in parental HCT116 WT and HCT116 LMNA KO using anti-lamin A/C antibodies. Manual decomposition of the sequence surrounding the PAM in the LMNA KO clone confirms frameshift mutations in both alleles. Genomic coordinates correspond to the location where the guide RNA binds. Scale bar, 5 µm. b, Representative confocal images of HCT116 WT and HCT116 LMNA KO cells that were either treated with si-Ctr, an siPOOL against LBR (siP-LBR) or a combination of 10 siPOOLs (si-10), directed against lamins B1 and B2 and the INM proteins LBR, LAP1, Emerin, LAP2, LEMD2, MAN1, SUN1, and SUN2 (excluding the siPOOL for lamin A/C) for 72 h, fixed and subjected to immunofluorescence using an anti-H3K9me3 antibody. Scale bars, 5 μm. c, Quantification of the accumulation of H3K9me3 at the nuclear rim (total number of cells: n ≥ 199, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001, one-way ANOVA, mean ± SD, N = 3 experiments). d, Immunoblotting of lamin A/C, LBR, LAP2β and select other NE proteins to determine the efficiency of siRNA-induced protein depletion. Numerical and statistics source data as well as unprocessed blots are available in source data.
Extended Data Fig. 3 The INM proteins LBR, LAP2 and LAP1 promote the peripheral positioning of centromeres.
a, Quantification of centromeres at the NE. Fixed cells were co-stained for centromeres (CREST antibody), CLIMP-63 (ER/NE network) and DNA (Hoechst). Z-stacks were acquired by confocal microscopy throughout the cell volume. All centromeres ≤ 0.5 μm from the nuclear periphery were considered to be positioned in LADs20. The fraction of peripheral centromeres was determined as the ratio between the number of peripheral centromeres and all identified centromeres. Scale Bar, 5 μm. b, Proof-of-concept experiment determining the proportion of centromeres within in 0.5 μm-wide ring at the nuclear periphery in si-Ctr and si-10-treated HCT116 LMNA KO cells as well as in si-Ctr-treated HCT116 WT cells (total number of cells: n ≥ 22, N = 2 experiments). c, Quantification of peripheral centromeres in HCT116 LMNA KO cells treated with si-Ctr or the indicated combination of siPOOLs for 72 h (Step 1 of deconvolution), identifying LBR (total number of cells: n ≥ 29, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001, one-way ANOVA, mean ± SD, N = 3 experiments). d, Quantification of peripheral centromeres in HCT116 LMNA KO cells treated with si-Ctr or the indicated combination of siPOOLs for 72 h (Step 2 of deconvolution), identifying the LEM-domain family of proteins (total number of cells: n ≥ 30, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001, one-way ANOVA, mean ± SD, N = 3 experiments). e Quantification of peripheral centromeres in HCT116 LMNA KO cells treated with si-Ctr or the indicated combination of siPOOLs for 72 h (Step 4 of deconvolution; total number of cells: n ≥ 25, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001, one-way ANOVA, mean ± SD, N = 3 experiments). Numerical and statistics source data are available in source data.
Extended Data Fig. 4 The INM proteins LBR and LAP2 promote the peripheral positioning of H3K9me3-positive heterochromatin.
a, Representative confocal images of H3K9me3-positive heterochromatin in HCT116 LMNA KO cells treated with si-Ctr or the indicated combinations of siPOOLs for 72 h (Step 1 of deconvolution), identifying LBR. Scale bars, 5 μm. b, Quantification of the accumulation of H3K9me3 at the nuclear rim (total number of cells: n ≥ 234, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001, one-way ANOVA, mean ± SD, N = 4 experiments). c, Representative confocal images of H3K9me3-positive heterochromatin in HCT116 LMNA KO cells treated with si-Ctr or the indicated combination of siPOOLs for 72 h (Step 2 of deconvolution), identifying LEM-domain family of proteins. Scale bars, 5 μm. d, Quantification of the accumulation of H3K9me3 at the nuclear rim (total number of cells: n ≥ 215, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001, one-way ANOVA, mean ± SD, N = 4 experiments). e, Representative confocal images of H3K9me3-positive heterochromatin in HCT116 LMNA KO cells treated with si-Ctr or the indicated combination of siPOOLs for 72 h (Step 4 of deconvolution). Scale bars, 5 μm. f, Quantification of the accumulation of H3K9me3 at the nuclear rim (total number of cells: n ≥ 142, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001, one-way ANOVA, mean ± SD, N = 3 experiments). g, Immunoblotting of LBR and select other NE proteins to determine the efficiency of siRNA-induced protein depletion. Numerical and statistics source data as well as unprocessed blots are available in source data.
Extended Data Fig. 5 Characterization of HCT116 knockout cell lines.
a, Characterization of the HCT116 LBR KO clone #1. Immunofluorescence and immunoblotting analyses of LBR in parental HCT116 WT and HCT116 LBR KO using anti-LBR antibodies. Manual decomposition of the sequence surrounding the PAM in the LBR KO clone #1 confirms frameshift mutations in both alleles. Genomic coordinates correspond to the location where the guide RNA binds. Used in Fig. 3. Scale Bar, 5 μm. b, Characterization of the HCT116 LBR KO clone #2. Manual decomposition of the sequence surrounding the PAM in the LBR KO clone #2 confirms frameshift mutations in both alleles. Used in Fig. 4 and Extended Data Fig. 6f, h, i. c, Characterization of the HCT116 TMPO KO clone, confirming frameshift mutations in both alleles. d, Characterization of the HCT116 LBR/TMPO DKO clone, confirming frameshift mutations in both TMPO alleles. Derived from LBR KO clone #2. e, Characterization of the HCT116 LBR/LMNA DKO clone, confirming frameshift mutations in both LMNA alleles. Derived from LBR KO clone #2. f, Characterization of the HCT116 LBR/TMPO/LMNA TKO clone. Manual decomposition of the sequence surrounding the PAM in the LBR/TMPO/LMNA TKO clone #1 confirms frameshift mutations in both LMNA alleles. Derived from LBR KO clone #2. g, FACS scatter plots demonstrating the gating strategy and histograms showing analysis of DNA content (propidium iodide (PI)) in HCT116 WT, LBR/TMPO DKO, and LBR/TMPO/LMNA TKO used to assess cell cycle distribution. Peaks corresponding to G1, S, and G2 phases are indicated. h, Bar plot indicating the percentage of cells with DNA content matching G1, S, and G2 phases in HCT116 WT, LBR/TMPO DKO, and LBR/TMPO/LMNA TKO as assessed from the flow cytometry histogram of DNA content; > G2 refers to polyploid cells (*p ≤ 0.05, **p ≤ 0.01, one-way ANOVA, mean ± SEM, N = 3 experiments). i, Growth curve of HCT116 WT, LBR/TMPO DKO, and LBR/TMPO/LMNA TKO. Growth rates were determined by cell counting. (Mean ± SEM, N = 3). Numerical and statistics source data as well as unprocessed blots are available in source data.
Extended Data Fig. 6 Changes in the transcriptome and chromatin organization upon loss of the heterochromatin tethers.
a, Stacked bar plots showing the proportion of differentially expressed genes (from Extended Data Fig. 1a) assigned to LAD and inter-LAD regions (one-sided Fisher’s exact test, p-value 0.19). b, Stacked bar plot showing the proportion of differentially accessible ATAC-seq peaks in nucleosome-free regions (Nfr) for si-11-treated HCT 116 cells after 72 h (N = 3). Loci with a |log2-fold change| ≥ 1, adjusted p-value ≤ 0.05 were considered as significantly changed (two-tailed Wald test and Benjamini–Hochberg procedure). c, Stacked bar plots showing the proportion of differentially accessible ATAC-seq peaks from (b) assigned to LAD and inter-LAD regions (one-sided Fisher’s exact, p-value 5.43e-160). d, Volcano plot showing differentially expressed genes (RNA-seq of si-Ctr or si-11-treated HCT116 cells after 6 d of RNAi (siRNA re-transfection at day 3) (N = 3 experiments); relates to Extended Data Fig. 1a. Number of genes with an adjusted p-value ≤ 0.05 and |log2-fold change| ≥ 1 (two-tailed Wald test and Benjamini–Hochberg procedure) are marked as up- or downregulated. e, Stacked bar plots showing the proportion of differentially expressed genes (from d) assigned to LAD and inter-LAD regions (one-sided Fisher’s exact test, p-value 0.997). f, Bar plots showing differentially expressed genes identified by RNA-seq in all KO genotypes (N = 3 experiments). Genes with an adjusted p-value ≤ 0.05 and |log2-fold change| ≥ 2 are considered as deregulated (two-tailed Wald test and Benjamini–Hochberg procedure). g, K-means clustering of the top 200 upregulated protein-coding genes in the LBR/TMPO DKO cells into a functional cluster. The genes within the cluster relating to the ‘defence response’ GO term are highlighted in green. Lines correspond to high confidence interactions based on STRING (interaction score > 0.7)115. h, Bar plots depicting the number of genes and transcripts identified in different deregulated splicing events. Events with IncLevelDifference > |0.15| and FDR < 0.05 were considered significant. i, Representative immunoblot of LBR, LAP2, lamin A/C, H3K27me3 and H3 relative to β-actin in the indicated HCT116 KO backgrounds (N = 3 experiments). j, Quantification of H3K27me3 levels (normalized to β-actin) in the different HCT116 KO backgrounds (ns p > 0.05, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001, one-way ANOVA, mean ± SEM, N = 3 experiments). k, Volcano plots showing transcriptional changes in histone methyltransferases (HMT), PRC1, PRC2 and HUSH complex members in HCT116 LBR/TMPO DKO and LBR/TMPO/LMNA TKO cell lines compared to WT. Genes with |log2-fold change| > 1 and padj < 0.05 (two-tailed Wald test and Benjamini–Hochberg procedure) were considered deregulated. Gene list: Supplementary Table 4. l, Metagene profiles of RPGC-normalized H3K9me3 and H3K27me3 ChIP-seq signal for two-fold up- and downregulated copies of TE classes (upregulated in Fig. 4c for HCT116 LBR/TMPO DKO and LBR/TMPO/LMNA TKO cells relative to WT). The copies were assigned to LAD and inter-LAD regions. Start and End represent TE copy start and end. RPGC: reads per genomic content (1x normalization). Numerical and statistics source data as well as unprocessed blots are available in source data.
Extended Data Fig. 7 The loss of LBR and LAP2 leads to reorganisation of 3D genome topology.
a, Principal component analysis of Pico-C libraries shows clear genotype separation across replicates, relates to Fig. 7. b, Pico-C reaches sub-5 kb resolution in all genotypes (cutoff of 80% bins with minimum 1k interactions, as done in ref. 15). c, Genome-wide Pico-C contact maps reveal no large-scale genomic aberrations in DKO or TKO HCT116 cell lines. d, Representative Micro-C maps from chr18 and chrX show genome-wide changes in compartmentalization between control and TKO. e, Neighbourhood analysis in DKO for the genomic regions commonly switching compartments mirrors the trends observed in TKO (main Fig. 7d), with loci switching from B.1 to A or B.2 adopting the identity of surrounding regions. The regions under consideration are those common to DKO and TKO cell lines. f, B.1 compartments undergoing switching exhibit accompanying changes in H3K9me3 and H3K27me3 enrichment. g, Stacked bar plots showing proportion of genes within the encompassed regions switching from B.1 to A (from e) with the indicated deregulation status compared to WT HCT116 cells. P-values (one-sided Fisher’s exact test) are depicted for upregulated genes in B.1 to A regions, compared to the rest of the genome (DKO 8.36e-16; TKO 7.16e-20). h, Gene density is higher around lost B.2 loci compared to stable or gained B.2 compartments. i, Deeply sequenced Pico-C maps and chromatin profiles (ATAC-seq, CTCF, RNA-seq, H3K9me3, H3K27me3) at a representative high-NAD region reveal a contrasting trend to main Fig. 7g, with the spread of repressive histone marks upon B.2 compartment gain. LMNB1 and NAD data94; HP1α95; CTCF binding from ENCODE (UCSC Genome Browser, TFBS Clusters v3). Statistics source data are available in source data.
Extended Data Fig. 8 Characterisation of mESCs knockout cell lines.
a, Characterisation of the E14TG2a Lbr KO. Genomic coordinates correspond to the locations that guides 1 and 2 bind. Manual decomposition of the sequence of the Lbr gene confirms deletion of the gene in both alleles. b, Characterisation of the E14TG2a Lmna KO. Manual decomposition of the sequence of the Lmna gene confirms deletion of the gene in both alleles. c, Characterisation of the E14TG2a Tmpo KO. Manual decomposition of the sequence of the Tmpo gene confirms deletion of the gene in both alleles. d, Characterisation of the E14TG2a Lbr/Tmpo DKO. Manual decomposition of the sequence of the Tmpo gene confirms deletion of the gene in both alleles. This KO was performed on the Lbr KO background. e, Characterisation of the E14TG2a Lbr/Lmna DKO. Manual decomposition of the sequence of the Lmna gene confirms deletion of the gene in both alleles. This KO was performed on the Lbr KO background. f, Characterisation of the E14TG2a Lbr/Tmpo/Lmna TKO. Manual decomposition of the sequence of the Tmpo gene confirms deletion of the gene in both alleles. This KO was performed on the Lbr/Lmna DKO background. g, Immunoblot of Lbr and Lap2 in the various knockout backgrounds to validate the loss of these proteins. Unprocessed blots are available in source data.
Extended Data Fig. 9 Changes in the transcriptome, chromatin organisation and cell differentiation upon loss of the heterochromatin tethers in mESCs.
a, Stacked bar plot showing the proportion of differentially accessible ATAC-seq peaks in nucleosome-free regions (Nfr) in Lbr/Tmpo DKO and Lbr/Tmpo/Lmna TKO mESCs (N = 3 experiments). Loci with a |log2-fold change| ≥ 1 and an adjusted p-value ≤ 0.05 (two-tailed Wald test and Benjamini–Hochberg procedure) were considered significant. b, Stacked bar plots showing the proportion of differentially accessible ATAC-seq peaks assigned to LAD and inter-LAD regions in regions originally designated as LADs in mESCs74 (one-sided Fisher’s exact test, decreased accessibility in LADs compared to inter-LADs, p-values: DKO 1.00e-16; TKO 8.73e-20). c, Stacked bar plots showing the proportion of differentially expressed genes (Fig. 8d) assigned to LAD and inter-LAD regions (one-sided Fisher’s exact test, downregulated genes in LAD compared to inter-LAD counterparts, p-values: DKO 0.522; TKO 3.64e-07). d, Volcano plots of transposable elements in E14TG2a mESCs Lbr/Tmpo DKO and Lbr/Tmpo/Lmna TKO vs. E14TG2a WT cells (N = 3 experiments). Genes with an adjusted p-value ≤ 0.5 are marked as up- or downregulated if |log2-fold change| ≥ 0.585 (two-tailed Wald test and Benjamini–Hochberg procedure). e, Cell viability of E14TG2a WT mESCs, Lbr/Tmpo DKO and Lbr/Tmpo/Lmna TKO cells during differentiation, calculated with resazaurin reduction-based assay, expressed as a percentage relative to the 24 h time point. (Mean ± SEM, N = 3 experiments). f, Bar plots of normalised read counts (RNA-seq) of marker genes for pluripotency, the three germ layers and primed epiSCs in E14TG2a WT, Lbr/Tmpo DKO and Lbr/Tmpo/Lmna TKO cells after 10 days of embryoid body differentiation (Mean ± SEM, N = 3 experiments). g, Bar plots of normalised read counts (RNA-seq) of select GABA receptor genes in E14TG2a WT, Lbr/Tmpo DKO and Lbr/Tmpo/Lmna TKO cells after 10 days of embryoid body differentiation (Mean ± SEM, N = 3 experiments). Numerical and statistics source data are available in source data.
Extended Data Fig. 10 Nuclear reorganization upon deletion of NE tethers in differentiated and pluripotent mammalian cells.
a, Nuclear envelope proteins observed to promote the peripheral positioning of different features of heterochromatin in mammalian cells. The INM proteins LBR and LAP2 were shown to contribute to the peripheral localisation of both H3K9me3 and centromeres in all cell-types examined, both in the presence and absence of lamin A/C. b, Model describing how the loss of LBR and LAP2 causes a global reduction of heterochromatin at the NE, accompanied by repositioning of heterochromatin into the vicinity of nucleoli and the formation of chromocenter-like foci within the nuclear interior together with the downstream consequences.
Supplementary information
Supplementary Table 1 (download XLSX )
Output of analysis performed on RNA-seq data for HCT116 cells and mESCs (Day 0 and Day 10).
Supplementary Table 2 (download XLSX )
Output of analysis performed on ATAC-seq data for HCT116 cells and mESCs (Day 0).
Supplementary Table 3 (download XLSX )
Changes in accessibility at regions within ±3 kb of TSSs for HCT116 cells and mESCs (Day 0).
Supplementary Table 4 (download XLSX )
Output of analysis for H3K27me3 ChIP for HCT116 cells relative to RNA-seq and list of histone methyltransferases (HMT), PRC1, PRC2 and HUSH complex members.
Supplementary Table 5 (download XLSX )
Light microscopy reporting Table.
Source data
Source Data Figs. and Extended Data Figs. (download PDF )
Unprocessed immunoblots for all figures and Extended Data figures.
Source Data Figs. and Extended Data Figs. (download XLSX )
Numerical source data.
Source Data Figs. and Extended Data Figs. (download XLSX )
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lewis, R., Sinigiani, V., Maziak, N. et al. LBR and LAP2 mediate heterochromatin tethering to the nuclear periphery to preserve genome homeostasis. Nat Cell Biol 28, 536–552 (2026). https://doi.org/10.1038/s41556-025-01822-7
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41556-025-01822-7