Abstract
Cell-free enzymatic reaction networks (ERNs) enable the production of value-added compounds in a single reaction. However, allosteric interactions, product inhibition, reversibility and competition for shared cofactors only become apparent once the ERN is assembled. These emergent dynamics create kinetic barriers that limit the overall yield. Here we introduce a model-guided optimal design strategy to generate time-dependent ‘recipes’ for batch reactions, in which every component can be added repeatedly at specified amounts, at any time. We apply the method to two ERNs: the pentose phosphate pathway and a branched nucleotide salvage pathway. In the pentose phosphate pathway, optimized inputs increased AMP production up to 5.7-fold and raised glucose-to-product conversion from ~12% in the control to ~48% using a time-dependent input. In the salvage pathway, time-dependent dosing balanced competing branches and increased UTP yield ~21-fold relative to composition-matched all-at-once dosing. Timed batch inputs provide a generally applicable route to optimizing a complex reaction sequence.

Similar content being viewed by others
Main
Cell-free enzymatic reaction networks (ERNs) are assemblies of extracted enzymes that partially reconstitute metabolic pathways outside of living systems. This in vitro approach offers a route to scale the production of valuable compounds in a single reaction1,2,3,4,5,6,7,8,9,10 or build functional systems11,12,13,14. Compared with traditional in vivo metabolic engineering, the use of cell-free ERNs avoids difficult product purification steps, improves yields, enables compounds that are toxic to cells to be produced and negates the sequestration of input substrates by other metabolic pathways15,16,17,18,19. Although purified enzymes and added cofactors are costly, they offer greater flexibility and control over the construction of optimized multi-enzymatic cascades20.
Leveraging these advantages, several groups have recently reported major breakthroughs in the development of complex ERNs, starting from simple and cheap materials such as glucose or fixing CO2 to produce valuable compounds such as monoterpenes, polyhydroxybutyrate, cannabinoids, malate or 6-deoxyerythronolide B21,22,23,24,25,26. It should be noted that complex networks can also be constructed using crude cell lysates for production of n-butanol, mevalonate or limonene27,28,29. Despite these advances, translating such successes into consistently high-yielding, cost-effective ERNs remains challenging.
Optimizing the initial ERN reaction mixture for optimal yields can be laborious; it is often impossible to identify a single set of initial concentrations that represents a global optimum30,31,32,33,34,35,36. The underlying challenge arises because enzyme cascades are frequently partially reversible, leading to suboptimal equilibria, and because large ERNs exhibit emergent activation and inhibition processes that affect enzyme efficiency37. These emergent kinetic barriers include allosteric regulation, product inhibition and competition for shared cofactors and only become apparent once all enzymes and cofactors are added to the reactor38,39.
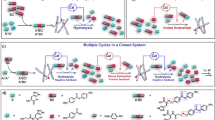
To overcome these barriers, we introduce a model-guided strategy to improve ERN yields by designing a time-dependent ‘recipe’ for each network component (Fig. 1a). This approach expands the accessible design space exponentially (Fig. 1b), as each reagent can be added multiple times with an independently varied timing and amount. Within this enlarged space lie trajectories that can bypass kinetic bottlenecks and drive the cascade towards higher substrate conversion.
a, A kinetic model is constructed utilizing an active learning cycle which maps ERN behaviour; this model is then used to design a sequence of time-dependent inputs for the batch reaction. Batch reactions with time-dependent inputs (optimized) are then compared with standard batch experiments where all reagents are added at t = 0 (control). b, A visualization of the combinatorial explosion of the potential inputs space λ if time-dependent inputs are allowed. c, An overview of the model ERN used in this study, the PPP. AMP and PRPP are the final products. The concentrations of enzymes, glucose and cofactors can be controlled and are added to the reactor. Enzymes used in the network are HK, G6PDH, 6PGDH, PRI and PRPPS; cofactors and their products are ATP, ADP, AMP, NADP and reduced NADP (NADPH). Four observed species were quantified using HPLC.
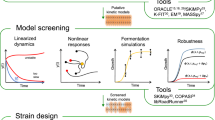
Yet, experimentally identifying such optimal trajectories is infeasible without an accurate, predictive map of ERN dynamics. Therefore, we first parameterize a model with an active learning workflow40,41,42,43,44,45,46,47 that designs maximally informative experiments about ERN behaviour48,49,50,51,52,53 (see Fig. 2 for a detailed overview). As shown previously, this workflow performs best in systems that can sustain continuous perturbations; therefore, we conduct these experiments in a flow reactor. The resulting time-series data are used to train and test a small set of mechanistic model candidates, each incorporating slightly different regulatory assumptions. The model with the highest predictive accuracy is then selected to design batch mode experiments featuring optimized, time-dependent input profiles.
(1) The OED algorithm designs a sequence of pulsed inputs for each species that flow into a temperature-controlled CSTR (volume ~121 µl). (2) These species include five enzymes: HK, G6PDH, 6PGDH, PRI and PRPPS and ATP, NADP and glucose. Eight individual syringes control the unique inflow profile of each species. The outflow of the CSTR reactor is collected, quenched and measured offline by HPLC. We observe AMP, ADP, ATP and NADP. (3) The first experiment is added to a database and used to train a model, and from thereon, every new experiment added to the database serves as test data as the pipeline uses the other experiments as training data. (4) When the model has sufficient predictive power to control the catalytic processes inside the reactor, the active learning cycle stops.
We applied the workflow to two model ERNs in vitro, the pentose phosphate pathway (PPP; Fig. 1c) and a branched nucleotide salvage pathway (NSP; Extended Data Fig. 1) and achieved yield improvements of more than 5- to 21-fold, respectively, compared with control reactions, where all components were added simultaneously at t = 0. This work demonstrates that an understanding of emergent ERN dynamics enables the design of non-intuitive, time-dependent input profiles, which can overcome kinetic barriers and drive cascades towards higher product formation in batch mode.
Results and discussion
Learning ERN dynamics with optimally designed training data
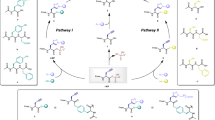
Figure 1c shows one of the ERN cascades used in this study. The system contains five enzymes (hexokinase (HK), glucose-6-phosphate dehydrogenase (G6PDH), 6-phosphogluconic dehydrogenase (6PGDH), phosphoriboisomerase (PRI) and phosphoribosyl pyrophosphate synthetase (PRPPS)), two cofactors (nicotinamide adenine dinucleotide phosphate oxidized (NADP) and adenosine triphosphate (ATP)) and one substrate (glucose (GLU)). At the start, glucose is converted to glucose-6-phospate (G6P) by HK using ATP as a cofactor. G6PDH using NADP as a cofactor converts G6P to 6-phosphoglucono-δ-lactone (6PGL), which undergoes spontaneous hydrolysis to 6-phosphogluconate (6PGA). Subsequently, 6PGA is converted to ribulose-5-phosphate (Ru5P) by 6PGDH using NADP as a cofactor, which is reversibly isomerized to ribose-5-phosphate (Ro5P) by PRI. In the last step, Ro5P is converted to phosphoribosyl pyrophosphate (PRPP) by PRPPS using ATP as a cofactor (PRPPS and PRI were purified according to Arthur et al.54; for more information see Supplementary Section 2.3). The final product PRPP is the precursor for the nucleotide synthesis pathway ERN40 (described in Supplementary Section 4.1), as well as a building block for DNA and RNA. The observed intermediates were quantified using high-performance liquid chromatography (HPLC). Supplementary Section 2.1 (Supplementary Table 2 and Supplementary Figs. 2 and 3) shows the concentrations used for all reagents, as well as indicative yields for the final product.
We first validated that the PPP functions as a cell-free ERN in batch mode (Supplementary Fig. 2). Under static conditions, only ~12% of glucose was converted to final products, illustrating the limitations of fixed-input operation described above. The cascade involves cofactors that induce conformational changes necessary for substrate binding through ordered sequential bi–bi mechanisms55,56,57,58,59,60. As these cofactors are consumed and regenerated, their concentrations fluctuate, rendering reaction rates inherently time-dependent. Several steps are potentially reversible within the concentration ranges tested, further constraining overall flux. In addition, literature reports extensive allosteric regulation in this pathway, notably the inhibition of PRPPS by adenosine diphosphate (ADP), a feature conserved across organisms. Together, these factors create dynamic kinetic barriers that cannot be mitigated by static dosing at t = 0. Overcoming them requires a parameterized model capable of capturing the emergent, time-dependent behaviour of the network61 (Supplementary Section 3).
To obtain such a model, we applied an active learning workflow that integrates optimal experimental design (OED) with flow chemistry to systematically probe a wide range of reaction conditions. In this approach, a continuous stirred tank reactor (CSTR; Supplementary Figs. 4 and 5) is subjected to a programmed sequence of out-of-equilibrium perturbations. Enzymes, substrates and cofactors are continuously supplied through separate inlets while the mixed reaction medium is withdrawn at the same rate, ensuring steady operation. This set-up produces kinetic data that are maximally informative for model parameterization (Supplementary Section 3.1). As shown in Fig. 2, the process is iterative. The most recent experiment serves as test data, and the previous runs are used for training. The cycle concludes once a model can predict the test data.
An in silico analysis indicated that a 6-h optimally designed flow experiment was sufficient to parameterize a model (Supplementary Section 5.1 and Supplementary Figs. 15 and 16). Thus, we implemented the workflow experimentally, performing two >6-h flow runs as the first train–test cycle. In total, eight syringe pumps, each delivering one component—HK, G6PDH, 6PGDH, PRI, PRPPS, glucose, ATP or NADP—were connected to a 121-µl CSTR with a single outlet line (Supplementary Section 2.2 and Supplementary Figs. 4 and 5). Samples were collected sequentially, quenched and analysed offline by HPLC (Supplementary Section 1.3 and Supplementary Fig. 1).
The results of two flow experiments are summarized in Fig. 3, where Fig. 3a shows the training of the model and Fig. 3b shows a flow experiment designed to test the obtained model. Each panel represents one iteration and includes both the optimally designed inflow profiles (Fig. 3a,b, left) and the corresponding experimental time-series data (Fig. 3a,b, right). The training experiment included a 250-min manually designed segment as a control (Supplementary Fig. 7). Total flow rates ranged from 97–1,670 µl h−1 in the first iteration and 297–989 µl h−1 in the second, corresponding to residence times of 4–75 min and 7–24 min, respectively. To expand the explored input space and generate sufficiently complex test data, the stock concentrations of enzymes and substrates in each syringe were varied between the two iterations (Supplementary Section 4.1 and Supplementary Fig. 8, enzyme concentrations inside the reactor).
a, Training the model. Left: the inflow profile of the first iteration with the flow rates of each enzyme and substrate; the dashed black line represents the total flow rate. Right: data as measured using HPLC (black triangles): AMP, ADP, ATP and NADP. The convergence of the training process to this data is given by the mean (thick coloured line); the uncertainty of the fit is visualized by shaded area (s.d. of 2) utilizing the 25 best fits. For the flow per species, the actual concentrations of enzyme species are shown in Supplementary Fig. 8. b, Testing the model. Left: the 25 best fits from iteration one are used to predict the OED experiment performed for iteration two. Right: predictions of the time series data are visualized in an identical manner to a.
Because the relevant mechanistic features of the ERN were not known a priori, we used time-series data from the first experiment to train four ordinary differential equation (ODE) models, each incorporating slightly different regulatory assumptions. These models were then tested against data from the second optimally designed experiment. The best-performing model accurately captured both training and test data (R2 = 0.892) (Supplementary Figs. 9 and 10), indicating sufficient flexibility without overparameterization. During model selection, we confirmed that ADP inhibits PRPPS, as models lacking this interaction failed to converge as well (Supplementary Fig. 11). Adding additional activation or inhibition terms, or using Hill-type kinetics, did not improve the model (Supplementary Section 5.2 and Supplementary Figs. 18 and 19).
Trained models unlock time-dependent batch inputs
With an appropriately parameterized model, we switched to batch mode and tested the central hypothesis, namely, that that time-dependent reagent additions can overcome kinetic barriers and substantially improve ERN yields. To maintain favourable conditions throughout the reaction, the optimization was unconstrained in both timing and frequency of inputs, allowing new enzyme, cofactor or substrate additions every 5 min.
Figure 4a shows three optimized input sequences, each sampling different stock concentrations to expand the explored design space (Supplementary Section 4.2 and Supplementary Table 4). The y axis indicates the added volume and the x axis the time in minutes; each colour corresponds to a specific enzyme or substrate (HK, G6PDH, 6PGDH, PRI, PRPPS, glucose, ATP or NADP). These non-intuitive dosing schedules involve repeated additions of multiple components at varying times and amounts. Figure 4b compares model predictions (orange) with control reactions (green), where all reagents were added at t = 0. Both the simulations and experiments show substantial increases in final product concentrations. The shaded regions indicate prediction uncertainty (s.d. of 2). Figure 4c shows measured adenosine monophosphate (AMP) concentrations after 120 min, demonstrating up to a 5.6-fold improvement relative to the control (experiment 3). Finally, Fig. 4d reports overall yields, expressed as the percentage of glucose converted to AMP. The largest gain occurred in experiment 2, where yield rose from ~12% in the control to ~48% under optimized, time-dependent dosing.
a, Three time series showing each addition of input species within the ERN: HK, G6PDH, 6PGDH, PRI, PRPPS, ATP, NADP and GLU. The y axis shows the volume of the additions to the reactor in µl and the x axis the time of additions. The algorithm allows an input to be given every 5 min, and each experiment (1–3) samples from different stock solutions. b, The time series plots show the prediction of the model for the time-dependent batch experiment (orange) and its corresponding control (green), (n = 10, 95% confidence intervals). c, A bar plot of the concentrations in μM of one of the final products AMP as measured on the HPLC. The time-dependent batch experiments (orange) have substantially higher final AMP concentrations compared with the control (green) as predicted in b and final concentration with a 5.6-fold increase compared with the control for experiment 3 (number above orange bar). Additional control experiments show that the spontaneous ATP hydrolysis does not affect outcomes (Supplementary Fig. 13). d, The yields indicate that a notably larger percentage of the initial substrate, glucose, was converted to AMP in all experiments compared with the control (yield = 100 n(AMP)final/n(glucose)added). Notably, in experiment 2, nearly half of the input substrate was transformed into the product.
Optimally timed input sequences lower costs
In Fig. 5, we demonstrate flexibility by showing we can target cost in silico by optimizing input schemes that minimize the cost per unit of the other, more valuable output molecule: PRPP (AMP serves as an experimentally observable proxy). Figure 5a shows an in silico comparison between optimized time-dependent batch runs and equivalent controls by static dosing at t = 0. Each orange circle’s size reflects the fold change between the control and the optimized reaction. The y axis is the cost per milligram produced; the x axis is the final product concentration. On average, time-dependent dosing results in substantially lower costs (~17-fold, based on 440 samples of the input space; Fig. 5b). When we take the best-performing control versus the best-performing time-dependent experiment, we obtained a ~16-fold lower cost (Fig. 5c). We used Sigma-Aldrich price listings as a reference for this investigation (Supplementary Section 5.3).
a, A scatterplot of 440 in silico experiments: in orange, the optimized time-dependent inputs, and in green, the equivalent static dose control at t = 0. The size of the optimized dots represents the ratio difference between the control and the time sensitive dosing. The y axis represents the log-scaled cost per milligram unit in euros; the x axis shows the final concentration of the product AMP or PRPP once the reaction is at steady state. b, The average cost per milligram for the 440 time-dependent experiments and 440 equivalent controls. Error bars signify the standard error of the mean for optimizations performed for both the optimized and control experiments (n = 440). c, The lowest-cost experiment found in the best-performing time-dependent and best-performing control experiment.
We demonstrate the generalizability of our workflow by applying it to a second ERN: a branched NSP converting PRPP with adenine or uracil to ATP and UTP via five enzymes (APRT, UPRT, AK, PK and UMPK) and ten control inputs (Extended Data Fig. 1 and Supplementary Section 4.1). We first mapped the kinetics of this network (Extended Data Fig. 2) following the method outlined above. The trained model was used to generate a time-dependent protocols for batch mode experiments, balancing substrate inputs and cofactor availability. Experimental validation achieved a ~21-fold increase in UTP yield and PRPP conversion compared with the composition-matched all-at-once control used as an internal reference (Extended Data Fig. 3 and Supplementary Section 4.2). A negative-control experiment validated the model’s predictive power. This test case demonstrates that this workflow effectively transfers to branched, competing networks and that time-dependent control can guide these ERNs through different productive states.
Conclusion
Optimizing the performance of complex ERNs remains challenging because hidden interactions and unfavourable kinetic barriers often emerge once enzymes operate together in a network. By introducing time as an explicit dimension in the optimization process, we expand the accessible design space by many orders of magnitude beyond traditional static approaches that only vary initial conditions. For example, assuming eight components each dosed between one and ten arbitrary levels yields 108 possible experiments. When time-dependent dosing is introduced by allowing 24 sequential additions (one every 5 min, as in Fig. 4a), this space grows to 10192 possible input sequences. Within this vast space, non-intuitive dosing trajectories exist that can substantially increase product yield, yet exhaustive experimental exploration is infeasible, particularly when enzymes and cofactors are costly or available in limited amounts.
To access this space efficiently, a predictive model is essential. We therefore used an active learning workflow that designs maximally informative pulse experiments, enabling rapid model parameterization within a single OED iteration. Although a flow reactor provides an ideal platform for these experiments owing to its ability to support continuous perturbations, the approach can, in principle, be adapted to other reactor formats if sufficient dynamic and multi-species time series data can be acquired. This means both the kinetic characterization and production could be done in batch mode.
The resulting model captures the key sensitivities governing ERN dynamics and allows systematic exploration of time-dependent input strategies. Using this model, we designed batch mode addition schedules that minimize inhibitory interactions and shift reversible reactions towards higher conversion. This strategy improved product yields for two ERNs between 5- and 21-fold, respectively, relative to conventional static dosing, which served as an internal benchmark. Although our experiments focused on maximizing total yield, we also demonstrated in silico that the same framework can accommodate multiple objectives by identifying a route that would yield products at 16-fold lower cost.
Overall, our results demonstrate that model-guided, time-dependent input design can overcome kinetic bottlenecks inherent to complex biochemical networks. Because our workflow is data-driven and mechanistic rather than pathway-specific, it is in principle network-agnostic. As predictive modelling and automated experimentation continue to advance, such time-dependent ‘recipes’ are poised to unlock most efficient and cost-effective routes to value-added compounds.
Methods
Microfluidic reactor and control unit
A CSTR (volume of 121 µl for the first and second iteration of PPP experiments and 158 µl for the NSP experiment) made of poly(methyl methacrylate) equipped with four or five inlets, which were doubled to eight or ten using Y connectors. The outlet tubes differed between experiments and had their own volume (109 µl and 69 µl for the first and second PPP experiments and 69 µl for NSP experiment, respectively) (Supplementary Figs. 4 and 5). Enzyme and substrate input flow rates were controlled by Cetoni neMESYS syringe pumps and Hamilton syringes and changed every 12 or 15 min. The ranges of the lowest and highest flow rates for the first and second PPP flow experiments and NSP experiment were 97–1,670, 297–988 and 97–1,670 µl h−1, respectively. The ranges of residence time for the first and second PPP flow experiments and NSP experiment were 4–75, 7–24 and 6–98 min, respectively. Reactor content was sampled by collecting droplets at the outlet (39 µl and 29 µl for the first and second PPP flow experiments and 27 µl for the NSP experiment, respectively) using a fraction collector, and the time of each drop was recorded using an infrared (IR) detector (Supplementary Fig. 5). The IR sensor was fitted to the fraction collector head to precisely monitor the time drops were collected; the control software can be found at ref. 62. PPP samples from the CSTR outlet tube were dropped directly into a guanidinium chloride solution (8 M, 117 µl and 87 µl for the first and second PPP flow experiments, respectively), where the sample-to-guanidinium-chloride ratio was 1:3, and quenched immediately. The NSP experiment samples were dropped directly into a formic acid solution (30 V%, 3 µl), where the sample-to-formic-acid ratio was 9:1, and quenched immediately. Quenched samples were centrifuged for 5 min at 6,708g, the supernatant (30 µl for PPP samples, 20 µl for NSP samples) was transferred to the micro insert of HPLC vial and measured by HPLC. PPP samples were measured by HPLC method 1, whereas NSP samples were measured by method 2. See Supplementary Section 2.2 for more information about the flow set-up, whereas stock concentrations for flow experiments can be found in Supplementary Section 4.1.
HPLC measurements
HPLC analyses were performed on Shimadzu Nexera X3 instrument. The conditions were: Inertsil ODS-4 C18 column (3-μm pore size, 150 mm × 4.6 mm; GL Science) and a guard column (3-μm pore size; 10 mm × 4.6 mm) at 40 °C. Analysis was based off of ref. 55.
Method 1 for PPP samples: we prepared elution solution A as 50 mM aqueous potassium phosphate (pH 6.0) buffer filtered over a 0.22-µm membrane and solution B as 100% MeOH. The method 1 elution gradient was as follows: 100% solution A for 2 min, 0–12.5% linear gradient of solution B for 8 min, 12.5% solution B for 2 min, 12.5–40% linear gradient of solution B for 6 min, 40% solution B for 2 min, 40–0% linear gradient of solution B for 3 min and 100% solution A for 7 min. The flow rate was maintained at 1 ml min−1; the injection volume was 10 µl, 15 µl and 20 µl for the batch, first and second flow experiments, respectively. The ultraviolet detection was at 254 nm.
Method 2 for NSP samples: we prepared elution solution A as 100 mM aqueous potassium phosphate buffer (pH 6.4) with 8 mM ion-pair reagent tetrabutylammonium bisulfate filtered over a 0.22-µm membrane and solution B as 70% solution A with 30% acetonitrile. The method 2 elution gradient was as follows: 100% solution A for 13 min, 0–77% linear gradient of solution B for 22 min, 77–100% solution B for 1 min, 100% solution B for 15 min and 100% solution A for 4 min. The flow rate was maintained at 1 ml min−1; the injection volume was 3 µl and 1 µl for the batch and flow experiments, respectively. The ultraviolet detection was at 254 nm.
See Supplementary Section 1.3 for more information on the HPLC and detected compounds.
Plasmid preparation
-
(1)
PRI
The gene for PRI was amplified from Escherichia coli K12-derived XL-1 blue competent cells using primers (bold and underline are the restriction sites used for cloning into the pET28a expression vector):
RpiA-Fw: 5′TTAGTCAGTCATATGACGCAGGATGAATTGAAAAAAGCAGTAGG3′
RpiA-Rv: 5′ATAGATAGCCTCGAGTCATTTCACAATGGTTTTGACACCGTCAG3′
DNA oligonucleotides were from Integrated DNA Technologies, standard desalted.
-
(2)
PRPPS
The plasmid for PRPPS originates from ref. 63.
-
(3)
UMPK
The gene for UMPK was amplified from Escherichia coli K12-derived XL-1 blue competent cells using primers:
UMPK-Fw: 5′GCATGACGTAGTACATATGGCTACCAATGCAAAACCCGTCTATAAACGC3′
UMPK-Rv: 5′ACTGGTACGCCTCGAGTTATTCCGTGATTAAAGTCCCTTCTTTTTCACCC3′
DNA oligonucleotides were from Integrated DNA Technologies, standard desalted.
-
(4)
APRT and UPRT
The plasmids for enzymes UPRT and APRT were a kind gift from Arthur and Dayie’s lab and originate from ref. 54.
Materials can be obtained via contact with the corresponding authors.
See Supplementary Section 2.3 for more information on the plasmid preparation.
Preliminary batch reactions
In a batch experiment, all five PPP network enzymes (HK, G6PDH, 6PGDH, PRI and PRPPS) together with substrate glucose and cofactors ATP and NADP were mixed in a buffer (100 mM HEPES, 10 mM MgCl2 and 1 mM CaCl2, pH 7.8) and left to react for 120 min at 30 °C while shaking. At every 0, 5, 10, 20, 40, 60 and 120 min reaction timepoint a sample (20 µl) was taken from each reaction tube, quenched using guanidinium chloride (8 M, 60 µl), where the sample-to-guanidinium-chloride ratio was 1:3, and vortexed. After this, all samples were centrifuged for 5 min at 6,708g, and the supernatant (30 µl) was analysed offline by HPLC. Conditions and results of preliminary batch reactions can be found in Supplementary Section 2.1.
Time-dependent batch experiments
Optimized time-dependent input experiments and control experiments were performed in batch conditions, where all inputs of control experiments were added at t = 0, whereas in optimized experiments, inputs were added at specific times and volumes, as shown in Fig. 4a for PPP and Extended Data Fig. 3 for NSP. Enzyme and substrate stock concentrations are presented in Supplementary Table 4. At 120 and 150 min (PPP experiments 1–3), 370 min (NSP experiment 1) and 355 min (NSP experiment 2) reaction timepoints samples from control and optimized reaction tubes were taken, quenched and measured as described in Supplementary Section 1.3. Conditions for time-dependent batch experiments as well as enzyme and substrate stock concentrations can be found in Supplementary Section 1.2.
See Supplementary Section 3.1 for a comprehensive description of the optimization procedure. See Supplementary Sections 3.2 and 3.3 for more information about the algorithm and software. See Supplementary Section 5 for an additional analysis to gauge the utility of the optimal design.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Raw data and notebooks used to generate the figures are available at the Huckgroup GitHub Data subfolder via GitHub at http://GitHub.com/huckgroup/TimedBatchReactions/. The estimated parameters for each of the modelling steps, the input flow profiles and raw data can be found in the extended data file Excel sheet available in the GitHub repository. Source data are provided with this paper.
Code availability
The code is available in the Software subfolder via GitHub at http://GitHub.com/huckgroup/TimedBatchReactions/. The folder contains demonstrations and explanations. For more information on the experimental software, please contact bob.vansluijs@gmail.com. Several elements of the software are present in previous work64. ODE solvers utilize AMICI, a C++ compiler for ODE solvers65. The custom software was written in Python 3.7.
References
Ghosh, S. et al. Exploring emergent properties in enzymatic reaction networks: design and control of dynamic functional systems. Chem. Rev. 124, 2553–2582 (2024).
Burgener, S., Luo, S., McLean, R., Miller, T. E. & Erb, T. J. A roadmap towards integrated catalytic systems of the future. Nat. Catal. 3, 186–192 (2020).
Rasor, B. J. et al. Toward sustainable, cell-free biomanufacturing. Curr. Opin. Biotechnol. 69, 136–144 (2021).
Cai, T. et al. Cell-free chemoenzymatic starch synthesis from carbon dioxide. Science 373, 1523–1527 (2021).
Zhang, Y.-H., Sun, J. & Ma, Y. Biomanufacturing: history and perspective. J. Ind. Microbiol. Biotechnol. 44, 773–784 (2017).
Winkler, C. K. et al. Accelerated reaction engineering of photo (bio) catalytic reactions through parallelization with an open-source photoreactor. ChemPhotoChem 5, 957–965 (2021).
Vázquez-González, M., Wang, C. & Willner, I. Biocatalytic cascades operating on macromolecular scaffolds and in confined environments. Nat. Catal. 3, 256–273 (2020).
Benítez-Mateos, A. I., Padrosa, D. R. & Paradisi, F. Multistep enzyme cascades as a route towards green and sustainable pharmaceutical syntheses. Nat. Chem. 14, 489–499 (2022).
Markin, C. J. et al. Revealing enzyme functional architecture via high-throughput microfluidic enzyme kinetics. Science 373, eabf8761 (2021).
Jain, S., Ospina, F. & Hammer, S. C. A new age of biocatalysis enabled by generic activation modes. JACS Au 4, 2068–2080 (2024).
Helwig, B., van Sluijs, B., Pogodaev, A. A., Postma, J. & Huck, W. T. S. Bottom-up construction of an adaptive enzymatic reaction network. Angew. Chem. Int. Ed. Engl. 57, 14261–14265 (2018).
van Roekel, H. W. H. et al. Programmable chemical reaction networks: emulating regulatory functions in living cells using a bottom-up approach. Chem. Soc. Rev. 44, 7465–7483 (2015).
Yue, L., Wang, S., Zhou, Z. & Willner, I. Nucleic acid based constitutional dynamic networks: From basic principles to applications. J. Am. Chem. Soc. 142, 21577–21594 (2020).
Teders, M., Pogodaev, A., Bojanov, G. & Huck, W. T. S. Reversible photoswitchable inhibitors generate ultrasensitivity in out-of-equilibrium enzymatic reactions. J. Am. Chem. Soc. 143, 5709–5716 (2021).
Grubbe, W. S., Rasor, B. J., Krüger, A., Jewett, M. C. & Karim, A. S. Cell-free styrene biosynthesis at high titers. Metab. Eng. 61, 89–95 (2020).
Korman, T. P. et al. A synthetic biochemistry system for the in vitro production of isoprene from glycolysis intermediates. Protein Sci. 23, 576–585 (2014).
Ye, X. et al. Spontaneous high-yield production of hydrogen from cellulosic materials and water catalyzed by enzyme cocktails. ChemSusChem 2, 149–152 (2009).
Lin, B. & Yong, T. Whole-cell biocatalysts by design. Microb. Cell Fact. 16, 106 (2017).
Rollin, J. A., Tam, T. K. & Zhang, Y.-H. P. New biotechnology paradigm: cell-free biosystems for biomanufacturing. Green Chem. 15, 1708–1719 (2013).
Huffman, M. A. et al. Design of an in vitro biocatalytic cascade for the manufacture of islatravir. Science 366, 1255–1259 (2019).
Korman, T. P., Opgenorth, P. H. & Bowie, J. U. A synthetic biochemistry platform for cell free production of monoterpenes from glucose. Nat. Commun. 8, 15526 (2017).
Opgenorth, P. H., Korman, T. P. & Bowie, J. U. A synthetic biochemistry module for production of bio-based chemicals from glucose. Nat. Chem. Biol. 12, 393–395 (2016).
Valliere, M. A. et al. A cell-free platform for the prenylation of natural products and application to cannabinoid production. Nat. Commun. 10, 565 (2019).
Schwander, T., Schada von Borzyskowski, L., Burgener, S., Cortina, N. S. & Erb, T. J. A synthetic pathway for the fixation of carbon dioxide in vitro. Science 354, 900–904 (2016).
Diehl, C., Gerlinger, P. D., Paczia, N. & Erb, T. J. Synthetic anaplerotic modules for the direct synthesis of complex molecules from CO2. Nat. Chem. Biol. 19, 168–175 (2023).
Giaveri, S. et al. Integrated translation and metabolism in a partially self-synthesizing biochemical network. Science 385, 174–178 (2024).
Karim, A. S. & Jewett, M. C. A cell-free framework for rapid biosynthetic pathway prototyping and enzyme discovery. Metab. Eng. 36, 116–126 (2016).
Dudley, Q. M., Anderson, K. C. & Jewett, M. C. Cell-free mixing of Escherichia coli crude extracts to prototype and rationally engineer high-titer mevalonate synthesis. ACS Synth. Biol. 5, 1578–1588 (2016).
Dudley, Q. M., Nash, C. J. & Jewett, M. C. Cell-free biosynthesis of limonene using enzyme-enriched Escherichia coli lysates. Synth. Biol. 4, ysz003 (2019).
Morgado, G., Gerngross, D., Roberts, T. M. & Panke, S. Synthetic biology for cell-free biosynthesis: fundamentals of designing novel in vitro multi-enzyme reaction networks. Adv. Biochem. Eng. Biotechnol. 162, 117–146 (2018).
Hold, C., Billerbeck, S. & Panke, S. Forward design of a complex enzyme cascade reaction. Nat. Commun. 7, 12971 (2016).
Taylor, C. J. et al. Flow chemistry for process optimisation using design of experiments. J. Flow. Chem. 11, 75–86 (2021).
Bujara, M., Schumperli, M., Pellaux, R., Heinemann, M. & Panke, S. Optimization of a blueprint for in vitro glycolysis by metabolic real-time analysis. Nat. Chem. Biol. 7, 271–277 (2011).
Shen, L. et al. A combined experimental and modelling approach for the Weimberg pathway optimisation. Nat. Commun. 11, 1098 (2020).
Pandi, A. et al. A versatile active learning workflow for optimization of genetic and metabolic networks. Nat. Commun. 13, 3876 (2022).
Borkowski, O. et al. Large scale active-learning-guided exploration for in vitro protein production optimization. Nat. Commun. 11, 1872 (2020).
Rondelez, Y. Competition for catalytic resources alters biological network dynamics. Phys. Rev. Lett. 108, 018102 (2012).
Baltussen, M. G., van de Wiel, J., Fernández Regueiro, C. L., Jakštaitė, M. & Huck, W. T. S. A Bayesian approach to extracting kinetic information from artificial enzymatic networks. Anal. Chem. 94, 7311–7318 (2022).
Taylor, C. J. et al. A brief introduction to chemical reaction optimization. Chem. Rev. 123, 3089–3126 (2023).
Duez, Q. et al. Quantitative online monitoring of an immobilized enzymatic network by ion mobility–mass spectrometry. J. Am. Chem. Soc. 146, 20778–20787 (2024).
van Sluijs, B. et al. Iterative design of training data to control intricate enzymatic reaction networks. Nat. Commun. 15, 1602 (2024).
van Sluijs, B., Maas, R. J., van der Linden, A. J., de Greef, T. F. & Huck, W. T. S. A microfluidic optimal experimental design platform for forward design of cell-free genetic networks. Nat. Commun. 13, 3626 (2022).
Cohn, D., Atlas, L. & Ladner, R. Improving generalization with active learning. Mach. Learn. 15, 201–221 (1994).
Angluin, D. Learning regular sets from queries and counterexamples. Inform. Comput. 75, 87–106 (1987).
Fedorov, V. V. Theory of Optimal Experiments (Elsevier, 2013).
Franceschini, G. & Macchietto, S. Model-based design of experiments for parameter precision: state of the art. Chem. Eng. Sci. 63, 4846–4872 (2008).
Kreutz, C. & Timmer, J. Systems biology: experimental design. FEBS J. 276, 923–942 (2009).
de Aguiar, P. F. et al. D-optimal designs. Chemom. Intell. Lab. Syst. 30, 199–210 (1995).
Sinkoe, A. & Hahn, J. Optimal experimental design for parameter estimation of an IL-6 signaling model. Processes 5, 49 (2011).
Raue, A. et al. Structural and practical identifiability analysis of partially observed dynamical models by exploiting the profile likelihood. Bioinformatics 25, 1923–1929 (2009).
Villaverde, A. F., Pathirana, D., Fröhlich, F., Hasenauer, J. & Banga, J. R. A protocol for dynamic model calibration. Brief. Bioinform. 23, bbab387 (2022).
Villaverde, A. F., Raimúndez, E., Hasenauer, J. & Banga, J. R. Assessment of prediction uncertainty quantification methods in systems biology. IEEE/ACM Trans. Comput. Biol. Bioinform. 20, 1725–1736 (2023).
Villaverde, A. F., Fröhlich, F., Weindl, D., Hasenauer, J. & Banga, J. R. Benchmarking optimization methods for parameter estimation in large kinetic models. Bioinformatics 35, 830–838 (2018).
Arthur, P. K., Luigi, J. A. & Dayie, T. K. Expression, purification and analysis of the activity of enzymes from the pentose phosphate pathway. Protein. Expr. Purif. 76, 229–237 (2011).
Nakajima, K. et al. Simultaneous determination of nucleotide sugars with ion-pair reversed-phase HPLC. Glycobiology 20, 865–871 (2010).
Cook, P. F. & Cleland, W. W. Enzyme Kinetics and Mechanism (Garland Science, 2007).
Mulcahy, P., O’Flaherty, M., Jennings, L. & Griffin, T. Application of kinetic-based biospecific affinity chromatographic systems to ATP-dependent enzymes: studies with yeast hexokinase. Anal. Biochem. 309, 279–292 (2002).
Velasco, P. et al. Purification, characterization and kinetic mechanism of glucose-6-phosphate dehydrogenase from mouse liver. Int. J. Biochem. 26, 195–200 (1994).
Hanau, S., Montin, K., Cervellati, C., Magnani, M. & Dallocchio, F. 6-Phosphogluconate dehydrogenase mechanism: evidence for allosteric modulation by substrate. J. Biol. Chem. 285, 21366–21371 (2010).
Switzer, R. L. in The Enzymes Vol. 10 (ed. Boyer, P. D.) 607–629 (Academic, 1974).
Willemoës, M., Hove-Jensen, B. & Larsen, S. Steady state kinetic model for the binding of substrates and allosteric effectors to Escherichia coli phosphoribosyl-diphosphate synthase. J. Biol. Chem. 275, 35408–35412 (2000).
Robinson, W. E. Labrador. GitHub https://GitHub.com/Will-Robin/labrador (2024).
Hove-Jensen, B., Harlow, K. W., King, C. J. & Switzer, R. L. Phosphoribosylpyrophosphate synthetase of Escherichia coli. Properties of the purified enzyme and primary structure of the prs gene. J. Biol. Chem. 261, 6765–6771 (1986).
Hu, X. et al. ARTseq-FISH reveals position-dependent differences in gene expression of micropatterned mESCs. Nat. Commun. 15, 3918 (2024).
Fröhlich, F. et al. AMICI: high-performance sensitivity analysis for large ordinary differential equation models. Bioinformatics 37, 3676–3677 (2021).
Acknowledgements
We thank W.E. Robinson (Radboud University, Nijmegen, the Netherlands) for building the IR drop detector. We thank the Radboud TechnoCentre, specifically A. de Kleine, for the design and manufacturing of the microreactors. This project is funded by the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (ERC Adv. Grant Life-Inspired, grant agreement no. 833466 and ERC PoC Grant OptiPlex, grant agreement no. 101069237 and grant agreement no. 862081 (CLASSY)). T.Z. acknowledges the Swiss National Science Foundation for financial support (P500PB_203166). B.v.S. acknowledges the Horizon Europe research and innovation programme under grant agreement no. 101120237 (ELIAS).
Author information
Authors and Affiliations
Contributions
B.v.S. conceived the study and designed the experiments. M.J. contributed to experimental work, microfluidics, HPLC and accompanying data analysis. B.v.S. contributed to software and modelling. T.Z supported preliminary studies and initial development of the HPLC approach. F.H.T.N. purified enzymes. M.J. and B.v.S. wrote the paper with input from W.T.S.H. All authors proofread the paper. W.T.S.H. and B.v.S. supervised the work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemistry thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 A conceptual overview of time-dependent experiments in batch for PRPP-fed salvage network.
A conceptual overview of time-dependent experiments in batch. a, A kinetic model is constructed utilizing an active learning cycle which maps enzymatic reaction network (ERN) behavior, this model is then used to design a sequence of time-dependent inputs for the batch reaction. Batch reactions with time-dependent inputs (optimized) are then compared to standard batch experiments where all reagents are added at t = 0 (control). b, Parallel phosphoribosyl pyrophosphate (PRPP)-fed salvage network. It converts adenine/uracil to adenosine triphosphate/ uridine triphosphate (ATP/UTP) via adenine phosphoribosyl transferase (APRT), uracil phosphoribosyl transferase (UPRT), adenylate kinase (AK), pyruvate kinase (PK), and uridine monophosphate kinase (UMPK). Inorganic pyrophosphate (PPi) is released after the first reactions with UPRT and APRT, while formed uridine monophosphate (UMP) and adenosine monophosphate (AMP) are further transformed to uridine diphosphate (UDP) and adenosine diphosphate (ADP) using UMPK, AK and ATP, and ultimately nucleoside diphosphates are converted to UTP and ATP using PK and phosphoenolpyruvate (PEP). The magnifying glasses indicate the species that are observed in the HPLC, the gears represent controlled inputs that is, species that are added to the reactor. There are 6 observables and 10 controlled reagents.
Extended Data Fig. 2 OED-in-flow pulse program and HPLC time-series.
a, Inflow program comprising a short manual validation segment followed by OED-designed pulses across 10 streams, the species are denoted by the color, the black dashed line represents the total flow rate, the first part of the experiment is manually designed (0-200 min). The in-flow species are: phosphoribosyl pyrophosphate (PRPP), uracil phosphoribosyl transferase (UPRT), phosphoenolpyruvate (PEP), uridine monophosphate kinase (UMPK), adenosine triphosphate (ATP), pyruvate kinase (PK), uracil, adenylate kinase (AK), adenine and adenine phosphoribosyl transferase (APRT). b, Representative time-series for adenine, ATP, uracil, UMP, UTP, and ADP used to train and select mechanistic ODE models. c, Demonstrates that the model fits to complex time-series data.
Extended Data Fig. 3 Overview of optimized nucleotide salvage pathway batch experiments.
Overview of optimized batch experiments. a, Optimized time-dependent batch schedules. The species are: phosphoribosyl pyrophosphate (PRPP), uracil phosphoribosyl transferase (UPRT), phosphoenolpyruvate (PEP), uridine monophosphate kinase (UMPK), adenosine triphosphate (ATP), pyruvate kinase (PK), uracil, adenylate kinase (AK), adenine and adenine phosphoribosyl transferase (APRT). b, Shows the trajectories of UTP production as predicted by the trained model. c) Shows the result of the batch experiment, specifically the UTP concentration for the optimized and control experiment. d, Shows the achieved yield, that is, the fraction of PRPP converted to UTP (yield =100 n(UTP)final/n(PRPP)added. e, Shows the ratio between the optimized and control experiment for both Experiment 1 and 2 by dividing the final measured concentration of the optimized batch experiment by its complementary control.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–19, Tables 1–8, materials and methods, preliminary experiments and experimental set-up, computational methods, additional experiments and additional computational analysis.
Source data
Source Data Fig. 3 (download XLSX )
Statistical source data file.
Source Data Fig. 4 (download XLSX )
Statistical source data file.
Source Data Fig. 5 (download XLSX )
Statistical source data file.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data file.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data file.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Jakštaitė, M., Zhou, T., Nelissen, F.H.T. et al. Timed batch inputs unlock substantially higher yields for enzymatic cascades. Nat. Chem. (2026). https://doi.org/10.1038/s41557-026-02138-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41557-026-02138-1