Abstract
Climate change is rapidly driving environmental shifts, posing an increasing threat to global biodiversity. Interspecific introgression—in which genetic material is transferred from one species to another following hybridization—may facilitate climate adaptation by introducing new genetic variation, which could mitigate species’ vulnerability to changing conditions. Here, using population and ecological genomic approaches and genetic offset modelling for future climates, we show that hybrid mountainous birds showed reduced vulnerability to climate change compared with non-hybrid counterparts. While geographic isolation and ecological heterogeneity promoted species divergence and distinct climatic niche requirements, gene flow persists at contact zones between these species. Maintaining current gene flow rates is projected to buffer against climate change risks over the next 40 generations. These findings demonstrate the role of interspecific introgression in enhancing climate resilience and future survival, and emphasize the conservation importance of preserving gene flow among species with narrow environmental tolerances.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Resequencing data generated in this study have been deposited in the National Genomics Data Centre (https://db.cngb.org/) under the accession number CNP0006945. Datasets used in this study have been deposited to GitHub at https://github.com/willright28/Project-for-three-Fulvetta-species and Zenodo at https://doi.org/10.5281/zenodo.17271739 (ref. 104). Source data are provided with this paper.
Code availability
Analysis scripts can be found at GitHub at https://github.com/willright28/Project-for-three-Fulvetta-species and Zenodo at https://doi.org/10.5281/zenodo.17271739 (ref. 104).
References
Scheffers, B. R. et al. The broad footprint of climate change from genes to biomes to people. Science 354, aaf7671 (2016).
Aitken, S. N. & Whitlock, M. C. Assisted gene flow to facilitate local adaptation to climate change. Annu. Rev. Ecol. Evol. Syst. 44, 367–388 (2013).
Browne, L. et al. Adaptational lag to temperature in valley oak (Quercus lobata) can be mitigated by genome-informed assisted gene flow. Proc. Natl Acad. Sci. USA 116, 25179–25185 (2019).
De-Kayne, R. et al. Genomic architecture of adaptive radiation and hybridization in Alpine whitefish. Nat. Commun. 13, 4479 (2022).
Kagawa, K. & Takimoto, G. Hybridization can promote adaptive radiation by means of transgressive segregation. Ecol. Lett. 21, 264–274 (2018).
Brauer, C. J. et al. Natural hybridization reduces vulnerability to climate change. Nat. Clim. Chang 13, 282–289 (2023).
Chen, Y. et al. The combination of genomic offset and niche modelling provides insights into climate change-driven vulnerability. Nat. Commun. 13, 4821 (2022).
Rahbek, C. et al. Building mountain biodiversity: geological and evolutionary processes. Science 365, 1114–1119 (2019).
Fjeldså, J., Bowie, R. C. K. & Rahbek, C. The role of mountain ranges in the diversification of birds. Annu. Rev. Ecol. Evol. Syst. 43, 249–265 (2012).
Wu, Y. J. et al. Explaining the species richness of birds along a subtropical elevational gradient in the Hengduan Mountains. J. Biogeogr. 40, 2310–2323 (2013).
Qu, Y. et al. Long-term isolation and stability explain high genetic diversity in the Eastern Himalaya. Mol. Ecol. 23, 705–720 (2014).
Qu, Y. et al. Lineage diversification and historical demography of a montane bird Garrulax elliotii—implications for the Pleistocene evolutionary history of the eastern Himalayas. BMC Evol. Biol. 11, 174 (2011).
Chen, Y. et al. 2023. Alpine burrow-sharing mammals and birds show similar population-level climate change risks. Nat. Clim. Change 13, 990–996 (2023).
Ruegg, K. et al. Ecological genomics predicts climate vulnerability in an endangered southwestern songbird. Ecol. Lett. 21, 1085–1096 (2018).
Bay, R. A. et al. Genomic signals of selection predict climate driven population declines in a migratory bird. Science 359, 83–86 (2018).
Li, C. et al. Two Antarctic penguin genomes reveal insights into their evolutionary history and molecular changes related to the Antarctic environment. Gigascience 3, 27 (2014).
Pirri, F. et al. Selection-driven adaptation to the extreme Antarctic environment in Emperor penguin. Heredity 129, 317–326 (2022).
Wang, L. C. et al. Involvement of the Arabidopsis HIT1/AtVPS53 tethering protein homologue the acclimation of the plasma membrane to heat stress. J. Exp. Bot. 62, 3609–3620 (2011).
Piñol, R. A. et al. Preoptic BRS3 neurons increase body temperature and heart rate via multiple pathways. Cell Metab. 33, 1389–1403 (2021).
Guilherme, A. et al. Neuronal modulation of brown adipose activity through perturbation of white adipocyte lipogenesis. Mol. Metab. 16, 116–125 (2018).
Graham, A. M. et al. Adaptive introgression of the beta-globin cluster in two Andean waterfowl. Heredity 127, 107–123 (2021).
Pizzagalli, M. D., Bensimon, A. & Superti-Furga, G. A guide to plasma membrane solute carrier proteins. FEBS J. 288, 2784–2835 (2020).
Zhang, Y., Guo, W., Zhang, Y., Zhang, H. & Wu, C. Insights into hypoxic adaptation in Tibetan chicken embryos from comparative proteomics. Comp. Biochem. Physiol. Part D. 31, 100602 (2019).
Qu, Y. et al. Rapid phenotypic evolution with shallow genomic differentiation during early stages of high elevation adaptation in Eurasian tree sparrows. Natl Sci. Rev. 7, 113–127 (2020).
Huerta-Sánchez, E. et al. Altitude adaptation in Tibetans caused by introgression of Denisovan-like DNA. Nature 512, 194–197 (2014).
Marchese, C. Biodiversity hotspots: a shortcut for a more complicated concept. Glob. Ecol. Conserv. 3, 297–309 (2015).
He, J. K., Lin, S. L., Li, J. T., Yu, J. H. & Jiang, H. S. Evolutionary history of zoogeographical regions surrounding the Tibetan Plateau. Commun. Biol. 3, 415 (2020).
Freeman, B. G., Scholer, M. N., Ruiz-Gutierrez, V. & Fitzpatrick, J. W. Climate change causes upslope shifts and mountaintop extirpations in a tropical bird community. Proc. Natl Acad. Sci. USA 115, 11982–11987 (2018).
Qu et al. The influence of geological events on the endemism of East Asian birds studied through comparative phylogeography. J. Biogeogr. 42, 179–192 (2015).
Song, G. et al. Phylogeography of the Alcippe morrisonia (Aves: Timaliidae): long population history beyond late Pleistocene glaciations. BMC Evol. Biol. 9, 143 (2009).
Excoffier, L., Dupanloup, I., Huerta-Sanchez, E., Sousa, V. C. & Foll, M. Robust demographic inference from genomic and SNP data. PLoS Genet. 9, e1003905 (2013).
Li, H. & Durbin, R. Inference of human population history from individual whole-genome sequences. Nature 475, 493–496 (2011).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Bailey, R. I. Bayesian hybrid index and genomic cline estimation with the R package gghybrid. Mol. Ecol. Resour. 24, e13910 (2024).
Blischak, P. D. et al. HyDe: a python package for genome-scale hybridization detection. Syst. Biol. 67, 821–829 (2018).
Schrider, D. R., Ayroles, J., Matute, D. R. & Kern, A. D. Supervised machine learning reveals introgressed loci in the genomes of Drosophila simulans and D. sechellia. PLoS Genet. 14, e1007341 (2018).
Atwater, D. Z., Ervine, C. J. & Barney, N. Climatic niche shifts are common in introduced plants. Nat. Ecol. Evol. 2, 34–43 (2018).
Petitpierre, P. et al. Climatic niche shifts are rare among terrestrial plant invaders. Science 335, 1344–1348 (2012).
Liu, C., Wolter, C., Xian, W. & Jeschke, J. M. Most invasive species largely conserve their climatic niche. Proc. Natl Acad. Sci. USA 38, 23643–23651 (2020).
Brown, J. L. & Carnaval, A. C. A tale of two niches: methods, concepts, and evolution. Front. Biogeogr. 11, e44158 (2019).
Thuiller, W., Lafourcade, B., Engler, R. & Araújo, M. B. BIOMOD—a platform for ensemble forecasting of species distributions. Ecography 32, 369–373 (2009).
Guo, Y., Li, X., Zhao, Z. & Nawaz, Z. Predicting the impacts of climate change, soils and vegetation types on the geographic distribution of Polyporus umbellatus in China. Sci. Total Environ. 648, 1–11 (2019).
Fournier-Level, A. et al. A map of local adaptation in Arabidopsis thaliana. Science 34, 86–80 (2011).
Frichot, E. & François, O. L. E. A. An R package for landscape and ecological association studies. Methods Ecol. Evol. 6, 925–929 (2015).
Forester, B. R. et al. Detecting spatial genetic signatures of local adaptation in heterogeneous landscapes. Mol. Ecol. 25, 104–120 (2016).
Dikmen, S. et al. Genome-wide association mapping for identification of quantitative trait loci for rectal temperature during heat stress in Holstein cattle. PLoS ONE 8, e69202 (2013).
Kim, T. W. et al. A critical role for IRAK4 kinase activity in Toll-like receptor–mediated innate immunity. J. Exp. Med. 204, 1025–1036 (2007).
Weldenegodguad, M. et al. Adipose gene expression profiles reveal insights into the adaptation of northern Eurasian semi-domestic reindeer (Rangifer tarandus). Commun. Biol. 4, 1170 (2021).
Heinrich, E. C. et al. Increased levels of interleukin-6 (IL-6) in Andean males with chronic mountain sickness and sea-level participants after one day at high altitude may reflect differences in IL-6 regulation. FASEB J. 32, lb479 (2018).
Ferrer-Admetlla, A., Liang, M., Korneliussen, T. & Nielsen, R. On detecting incomplete soft or hard selective sweeps using haplotype structure. Mol. Biol. Evol. 31, 1275–1291 (2014).
Voight, B. F., Kudaravalli, S., Wen, X. & Pritchard, J. K. A map of recent positive selection in the human genome. PLoS Biol. 4, e72 (2006).
Ellis, N., Smith, S. J. & Pitcher, C. R. Gradient forests: calculating importance gradients on physical predictors. Ecology 93, 156–168 (2012).
Haller, B. C. & Messer, P. W. SLiM 4: multispecies eco-evolutionary modeling. Am. Nat. 201, E127 (2023).
Matz, M. V., Treml, E. A., Aglyamova, G. V. & Bay, L. K. Potential and limits for rapid genetic adaptation to warming in a Great Barrier Reef coral. PLoS Genet. 14, e1007220 (2018).
Matz, M. V., Treml, E. A. & Haller, B. C. Estimating the potential for coral adaptation to global warming across the Indo-West Pacific. Glob. Change Biol. 26, 3473–3481 (2020).
Dong, F. et al. Potential millennial-scale avian declines by humans in southern China. Glob. Change Biol. 28, 5505–5513 (2022).
Bird, J. P. et al. Generation lengths of the world’s birds and their implications for extinction risk. Conserv. Biol. 34, 1252–1261 (2020).
Clarke, R. T. et al. Confidence limits for regression relationships between distance matrices: estimating gene flow with distance. J. Agric. Biol. Environ. Stat. 7, 361–372 (2002).
Rieseberg, L. H., Archer, M. A. & Wayne, R. K. Transgressive segregation, adaptation and speciation. Heredity 83, 363–372 (1999).
Seehausen, O. Hybridization and adaptive radiation. Trends Ecol. Evol. 19, 198–207 (2004).
Rieseberg, L. H. Hybrid speciation in wild sunflowers. Ann. Mo. Bot. Gard. 93, 34–48 (2006).
Rieseberg, L. H. et al. Major ecological transitions in wild sunflowers facilitated by hybridization. Science 301, 1211–1216 (2003).
Ungerer, M. C., Baird, S. J. E., Pan, J. & Rieseberg, L. H. Rapid hybrid speciation in wild sunflowers. Proc. Natl Acad. Sci. USA 95, 11757–11762 (1998).
Stelkens, R. B., Brockhurst, M. A., Hurst, G. D. D. & Greig, D. Hybridization facilitates evolutionary rescue. Evol. Appl. 7, 1209–1217 (2014).
Stelkens, R. B., Brockhurst, M. A., Hurst, G. D. D., Miller, E. L. & Greig, D. The effect of hybrid transgression on environmental tolerance in experimental yeast crosses. J. Evol. Biol. 27, 2507–2519 (2014).
Zhang, D. C., Zhang, Y. H., Boufford, D. E. & Sun, H. Elevational patterns of species richness and endemism for some important taxa in the Hengduan Mountains, southwestern China. Biodivers. Conserv. 18, 699–716 (2009).
Zhang, R. Z., Zheng, D., Yang, Q. Y. & Liu, Y. H. Physical Geography of Hengduan Mountains (Science Press, 1997).
Allendorf, F. W., Leary, R. F., Spruell, P. & Wenburg, J. K. The problems with hybrids: setting conservation guidelines. Trends Ecol. Evol. 16, 613–622 (2001).
Todesco, M. et al. Hybridization and extinction. Evol. Appl. 9, 892–908 (2016).
Hirashiki, C., Kareiva, P. & Marvier, M. Concern over hybridization risks should not preclude conservation interventions. Conserv. Sci. Pract. 3, e424 (2021).
Chen, Y. et al. Large-scale genome-wide reveals climate adaptive variability in a cosmopolitan pest. Nat. Commun. 12, 7206 (2021).
Urban, M. C. Escalator to extinction. Proc. Natl Acad. Sci. USA 115, 11871–11873 (2018).
Chen, S. Ultrafast one-pass FASTQ data preprocessing, quality control, and deduplication using fastp. iMeta 2, e107 (2023).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Van Der Auwera, G. & O’Connor, B. D. Genomics in the Cloud: Using Docker, GATK, and WDL in Terra (O’Reilly Media, 2020).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Lefort, V., Desper, R. & Gascuel, O. FastME 2.0: a comprehensive, accurate, and fast distance-based phylogeny inference program. Mol. Biol. Evol. 32, 2798–2800 (2015).
Wiens, B. J., DeCicco, L. H. & Colella, J. P. triangulaR: an R package for identifying AIMs and 825 building triangle plots using SNP data from hybrid zones. Heredity 134, 251–262 (2025).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. GigaScience 10, giab008 (2021).
Martin, S. H., Davey, J. W. & Jiggins, C. D. Evaluating the use of ABBA–BABA statistics to locate introgressed loci. Mol. Biol. Evol. 32, 244–257 (2015).
Cai et al. Near-complete phylogeny and taxonomic revision of the world’s babblers (Aves: Passeriformes). Mol. Phylogenet. Evol. 130, 346–356 (2019).
Ericson, P. G. & Irestedt, M. Comparative population genomics reveals glacial cycles to drive diversifications in tropical montane birds (Aves, Timaliidae). Avian Res. 13, 100063 (2022).
Browning, S. R. & Browning, B. L. Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am. J. Hum. Genet. 81, 1084–1097 (2007).
Xiao, H. et al. Adaptive and maladaptive introgression in grapevine domestication. Proc. Natl Acad. Sci. USA 120, e2222041120 (2023).
Lukicheva, S. & Mardulyn, P. Whole-genome sequencing reveals asymmetric introgression between two sister species of cold-resistant leaf beetles. Mol. Ecol. 30, 4077–4089 (2021).
Geneva, A. J., Muirhead, C. A., Kingan, S. B. & Garrigan, D. A new method to scan genomes for introgression in a secondary contact model. PLoS ONE 10, e0118621 (2015).
Hudson, R. R. A new statistic for detecting genetic differentiation. Genetics 155, 2011–2014 (2000).
Tian, J. et al. Projections of precipitation over China based on CMIP6 models. Stoch. Environ. Res. Risk Assess. 35, 831–848 (2021).
Lu, K. et al. Evaluating observed and future spatiotemporal changes in precipitation and temperature across China based on CMIP6-GCMs. Int. J. Climatol. 42, 7703–7729 (2022).
Boria, R. A., Olson, L. E., Goodman, S. M. & Anderson, R. P. Spatial filtering to reduce sampling bias can improve the performance of ecological niche models. Ecol. Model. 275, 73–77 (2014).
Liu, C. R., White, M. & Newell, G. Selecting thresholds for the prediction of species occurrence with presence-only data. J. Biogeogr. 40, 778–789 (2013).
Ge, S. X., Jung, D. & Yao, R. ShinyGO: a graphical gene-set enrichment tool for animals and plants. Bioinformatics 36, 2628–2629 (2020).
Szpiech, Z. A. & Hernandez, R. D. Selscan: an efficient multithreaded program to perform EHH-based scans for positive selection. Mol. Biol. Evol. 31, 2824–2827 (2014).
Meier, J. I., Marques, D. A., Wager, C. E., Excoffier, L. & Seehausen, O. Genomics of parallel ecological speciation in lake Victoria cichilids. Mol. Biol. Evol. 35, 1489–1506 (2018).
Kim-Hellmuth, S. et al. Genetic regulatory effects modified by immune activation contribute to autoimmune disease association. Nat. Commun. 8, 266 (2017).
Blumstein, M. et al. Protocol for projecting allele frequency change under future climate change at adaptive-associated loci. STAR Protoc. 1, 100061 (2020).
Gain, C. et al. A quantitative theory for genomic offset statistics. Mol. Biol. Evol. 40, msad140 (2023).
Yair, S. & Coop, G. Population differentiation of polygenic score predictions under stabilizing selection. Philos. Trans. R. Soc. B 377, 20200416 (2022).
Peterman, W. E. ResistanceGA: an R package for the optimization of resistance surfaces using genetic algorithms. Methods Ecol. Evol. 9, 1638–1647 (2018).
Keenan, K., Mcginnity, P., Cross, T. F., Crozier, W. W. & Prodöhl, P. A. diveRsity: an R package for the estimation and exploration of population genetics parameters and their associated errors. Methods Ecol. Evol. 4, 782–788 (2013).
Bates, D., Mächler, M., Bolker, B. & Walker, S. Fitting linear mixed effects models using lme4. J. Stat. Softw. 67, 1–48 (2015).
Anderson, D. R. & Burnham, K. P. Avoiding pitfalls when using information-theoretic methods. J. Wildl. Manag. 66, 912–918 (2002).
Zhang, S. et al. Code and data for ‘Hybridisation mitigates climate change risk in mountainous birds’. Zenodo https://doi.org/10.5281/zenodo.17271739 (2025).
Danielson, J. J. & Gesch, D. B. Global Multi-resolution Terrain Elevation Data 2010 (GMTED2010) US Geological Survey Open-File Report 2011-1073, 26 p (2011).
Acknowledgements
This research was funded by the National Natural Science Foundation of China (NSFCU23A20162 and NSFC32401393). This research was funded by Young Elite Scientists Sponsorship Program by CAST (2023QNRC001), the Postdoctoral Fellowship Program of CPSF (GZC20232646), the China Postdoctoral Science Foundation (2023M743478) and Bingzhi Postdoctoral Fellowship Program of IOZ, CAS and Chinese Academy of Sciences President’s International Fellowship Initiative (2025PVA0187).
Author information
Authors and Affiliations
Contributions
Conceptualization: Y.Q. and P.G.P.E. Methodology: S.Z. and Y.C. Investigation: Y.Q. and Y.C. Visualization: S.Z., Y.C., W.Z. and X.W. Funding acquisition: Y.Q. and Y.C. Project administration: Y.Q. and F.L. Supervision: P.G.P.E., Y.Q. and W.Z. Writing – original draft: Y.Q. and S.Z. Writing – review and editing: Y.Q., P.G.P.E., S.Z. and Y.C.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Climate Change thanks Luciano Beheregaray, Junhu Su and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The historical demography of the three species inferred by pairwise sequentially Markovian coalescent (PSMC).
Following their divergence (Tdiv) approximately 216-221 thousand years ago, fratercula exhibited an initial population decline, hueti maintained a relatively stable population sizes, and davidi underwent continuous population growth. Notably, all species experienced synchronous population expansions beginning approximately 47 kya, a period that coincides with the estimated onset of interspecific gene flow (Tmig). However, all species decreased their effective population sizes (Ne) following the last glacial maximum (LGM).
Extended Data Fig. 2 Hybrid index estimations among davidi and hueti.
Pure reference populations are coded as S0 (blue, davidi) and S1 (red, hueti). The point estimates are based on the mode from the posterior distribution, and error bars indicate 95% credibility intervals based on an initial burn-in of 1,000 MCMC iterations followed by 5,000 iterations. Dashed blue and red lines define the 95% credible intervals for classifying a sample as parental or hybrid.
Extended Data Fig. 3 Triangle plots for hybrid index and interspecific heterozygosity.
Purple dots indicate pure davidi individuals. Green dots indicate pure hueti individuals. Hybrid individuals (red open circles) show reduced levels of interspecific heterozygosity, consistent with advanced-generation hybridization.
Extended Data Fig. 4 Principal component analysis of the 19 climatic variables selected the four climatic variable representing climate requirement of fulvettas.
Bio4, the temperature seasonality; bio10, the mean temperature of the warmest quarter; bio11, the mean temperature of the coldest quarter; bio17, the precipitation of the driest quarter.
Extended Data Fig. 5 Candidate genes showing signals of introgression.
Individual SNPs fixed for different alleles between parental A. davidi (purple) and hueti (blue) with heterozygous genotypes in hybrid individuals. (a) IRAK4, (b) KBTBD2, (c) IL6, (d) RAPGEF5. (a)-(d) Note the number of introgressed SNPs varies among hybrid individuals. (e) Introgressed climate-associated SNPs displayed clinal variation across the hybrid zone, with most cline centers coinciding with the primary contact zone.
Extended Data Fig. 6 The allele frequencies of the candidate genes showed cline variation with climatic variables.
Relationships between the allele frequency of SNPs in either of the KBTBD2, IRAK4, IL6 and RAPGEF5 and the mean temperature of the warmest quarter (bio10, left) or the precipitation of the driest quarter (bio17, right) were shown. Species distribution ranges produced by BirdLife International and Handbook of the Birds of the World from the IUCN Red List of Threatened Species (https://www.iucnredlist.org/species/22716644/94504173). Contemporary climate layers were sourced from the Climatologies at High Resolution for the Earth’s Land Surface Areas (CHELSA) database (https://chelsa-climate.org/).
Extended Data Fig. 7 Allele frequencies of the introgressed climate-associated SNPs in the hybrids and pure parental species.
(a) The introgressed climate-associated loci showed greater heterozygosity in hybrids than in pure parental species. Boxes show the median (centre line), interquartile range (box bounds: 25th and 75th percentiles), and minimum/maximum values (whiskers). Outliers are shown as individual points. P values were calculated using two-sided Wilcoxon rank-sum tests and adjusted for multiple comparisons using the Benjamini–Hochberg procedure. (b) Percentage of allele absence of the introgressed climate-associated SNPs from the hybrids and the parental species (relative to 2100 SSP5-8.5 climate scenario).
Extended Data Fig. 8 fd-detected introgressed climate-associated variants mitigates the climate change risk.
(a) Hybrid populations (that is, site-scale GO modeling, n = 7 sampling localities) exhibit lower genetic offset than non-hybrid populations (n = 13 sampling localities) under the 2070 and 2100 SSP5-8.5 climate scenarios. (b) Hybrid individuals (that is, range-based GO modeling, n = 3,925 grids) show the least genetic offset compared to each of the three species (fratercula, n = 12,521 grids, davidi, n = 9,477 grids, hueti, n = 11,427 grids). Box plots in (a) and (b) show the median (center line), interquartile range (box bounds: 25th and 75th percentiles), and minimum/maximum values (whiskers). Outliers are shown as individual points. P values were calculated using two-sided Wilcoxon rank-sum tests and adjusted for multiple comparisons using the Benjamini–Hochberg procedure. Species distribution ranges produced by BirdLife International and Handbook of the Birds of the World from the IUCN Red List of Threatened Species (https://www.iucnredlist.org/species/22716644/94504173).
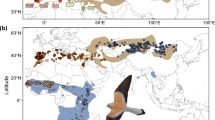
Extended Data Fig. 9 The parapatric distributions and contact zones of the fulvettas.
The ranges of fratercula, davidi, and hueti are delineated by black outlines. Putative contact zones, inferred from genetic admixture analyses, are highlighted in blue shade, with a putative hybrid zone marked in red outline. It is important to note that these inferred zones are dependent on the current sampling design and may represent underestimations of the true areas of sympatry. Species distribution ranges produced by BirdLife International and Handbook of the Birds of the World from the IUCN Red List of Threatened Species (https://www.iucnredlist.org/species/22716644/94504173). Map generated with ArcGIS v.10.6 (Esri) with elevation data from the Global Multi-resolution Terrain Elevation Data 2010 (GMTED2010), US Geological Survey105.
Extended Data Fig. 10 Slim simulations with varying σe, Esd, and σt yielded consistent results.
Each parameter combination was run 20 times to assess robustness. Results were consistent across simulations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–9 and Tables 1–8.
Supplementary Data 1 (download XLSX )
Sampling and sequencing information for the individuals used in this study.
Supplementary Data 2 (download XLSX )
Statistical source data for Supplementary Fig. 7.
Source data
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, S., Chen, Y., Zang, W. et al. Hybridization mitigates climate change risk in mountainous birds. Nat. Clim. Chang. 15, 1378–1387 (2025). https://doi.org/10.1038/s41558-025-02485-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41558-025-02485-w